Abstract
Background
A growing number of studies have focused on the regulatory role of circular RNAs (circRNAs) in a variety of cancers. The purpose of this study was to investigate the effect of circRNA Keratin 14 (circKRT14) on the progression of esophageal cancer (EC).
Methods
The levels of circKRT14, miR-1256 and E2F transcription factor 3 (E2F3) were analyzed by real-time quantitative polymerase chain reaction (qRT-PCR) and western blot. The circular structure of circKRT14 was confirmed by RNase R digestion assay. Cell apoptosis, migration and invasion were detected by flow cytometry and transwell assay. The protein levels of related factors were determined by western blot. The relationship between miR-1256 and circKRT14 or E2F3 was verified by dual-luciferase reporter assay. The in vivo function of circKRT14 was studied by xenograft tumor assay.
Results
CircKRT14 was significantly increased in EC tissues and cells. CircKRT14 silencing inhibited EC cell proliferation, migration, and invasion, but promoted EC cell apoptosis in vitro. CircKRT1 acted as a sponge for miR-1256 in EC, and in-miR-1256 abolished the inhibitory effect of circKRT14 suppression on EC cell progression. E2F3 was a target of miR-1256 and functioned as an oncogene in EC cells. MiR-1256 curbed EC progression by downregulating E2F3. CircKRT14 could affect E2F3 expression by targeting miR-1256. CircKRT14 regulated EC progression in vivo through miR-1256/E2F3 axis.
Conclusions
These results uncovered that circKRT14 up-regulated the expression of E2F3 and promoted the malignant development of EC through sponging miR-1256.
Introduction
Esophageal cancer (EC) is a kind of upper gastrointestinal malignancy with high mortality, strong invasiveness and poor survival rate. 1 Surgery is the most important method for the treatment of EC, especially minimally invasive surgery represented by thoracoscopy, which is more effective and safer than traditional surgery and widely used in clinical practice. 2 With the continuous development of medical technology, remarkable progress has been made in the diagnosis and treatment of EC, but the prognosis of EC patients is still not satisfactory, and the pathogenesis of EC remains to be further clarified. 3 Therefore, in-depth study of the molecular mechanism of EC occurrence and development can provide new ideas for the treatment of EC.
Circular RNAs (circRNAs) are a special form of RNA that, in most cases, are formed by reverse splicing to form a continuous loop structure with covalently closing capability from the 5′ to 3′ ends, circRNAs are not affected by RNA exonuclease due to their structural specificity. 4 CircRNAs expressed stably in mammalian cells and have certain tissue, timing and disease specificity, 5 its function is to act as a miRNA sponge to relieve the inhibition of miRNA on its target genes and thus improve the expression level of target genes.6,7 CircRNAs have aroused interest in cancer research in the past few years. Researches have illustrated that many circRNAs acted great role in EC, 8 breast cancer, 9 lung cancer, 10 renal cell carcinoma, 11 cervical cancer 12 and other tumors. However, the effect of circKRT14 on EC progress has not been reported. Therefore, exploring the mechanism of EC progression may be of great significance for the development of new targeted drugs.
MiRNA are a sort of small nucleotide ncRNAs, mediating cell proliferation, apoptosis and migration and other life activities, and participating in tumor genesis and development. 13 MiR-1256 has been reported to act as a tumor suppressor in all kinds of malignant tumors, such as non-small cell lung cancer, 14 gastric cancer 15 and prostate cancer, 16 and is nearly relevant to the biological behavior of tumor malignancy. So far, the related studies of miR-1256 in EC have not been reported.
E2F3 (E2F transcription factor 3) belongs to E2F family and is widely taken part in cell proliferation, differentiation, apoptosis and other processes. 17 E2F3 has been revealed to play a vital role in the progression of bladder, 18 liver 19 and breast cancers. 20 In addition, Wang et al. 21 confirmed that E2F3 played a promoting role in esophageal squamous cell carcinoma . However, its mechanism of action in EC still needs further exploration.
The purpose of our work was to probe the functional and potential mechanisms of circKRT14 in EC and the effect of circKRT14 on EC progression via miR-1256/E2F3 axis.
Materials and methods
Source of organization
EC tissues (n = 63) and paracancerous tissues were obtained from Affiliated Hospital of Beihua University. All participating patients signed informed consent. This work was permitted by the Ethics Committee of Affiliated Hospital of Beihua University.
Cell culture and transfection
Esophageal cancer cell lines KYSE-150, KYSE-410, EC9706 and human esophageal epithelial cell line HET-1 were obtained from China Center for Type Culture Collection (Wuhan, China). All cells were cultured in RPMI1640 medium (Invitrogen, Carlsbad, CA, USA) with 10% fetal bovine serum (FBS, Invitrogen) in an incubator with 5% CO2 at 37°C.
Short hairpin RNA targeting circKRT14 (sh-circKRT14), miR-1256 mimics (miR-1256), miR-1256 inhibitor (in-miR-1256), E2F3 overexpression plasmids (E2F3) and their controls were purchased from Genepharma (Shanghai, China) and then transfected into KYSE-150 and EC9706 cells by using the LipofectamineTM3000 (Invitrogen) transfection reagent.
Quantitative real-time polymerase chain reaction (QRT-PCR)
Primers sequences used for PCR.

RNase R experiment
To determine the structure of circKRT14, RNA obtained from KYSE-150 and EC9706 cells was treated with RNase R (3 U/μg, TaKaRa) at 37°C. Then the level of circKRT14 was detected using qRT-PCR.
Clone formation experiment
After transfection, KYSE-150 and EC9706 cells of each group were distributed in 12-well plates with 5 × 102 cells/well, and cultivated in a cell incubator with 5% CO2 at 37°C. After 15 days, cells were stained with crystal violet (Beyotime, Shanghai, China) and observed under a microscope (Olympus, Tokyo, Japan).
5-Ethynyl-2′-deoxyuridine (EdU) assay
Transfected KYSE-150 and EC9706 cells were placed in 6-well plates with 2 × 105 cells/well. Cell proliferation was assessed using EdU kit (Beyotime). In brief, 50 μM EdU was appended to each well for 2 h, along with 4% paraformaldehyde for 30 min, then treated with 500 μL 0.5% Triton-X for 10 min. The cells were co-incubated with Apollo staining solution under dark conditions for 30 min. Finally, Hoechst 33,342 was added and incubated for 30 min. The results were observed by a microscope.
Flow cytometry
The transfected KYSE-150 and EC9706 cells were added into buffer solution and suspended again. Cells were stained with Annexin V solution (10 μL, Beyotime) and PI solution (10 μL, Beyotime) for 30 min. Finally, cell apoptosis was monitored by a flow cytometer (Agilent, Beijing, China).
Transwell assay
Transwell chamber was used for cell migration detection. In addition, transwell chamber was coated with Matrigel matrix (BD Biosciences, San Jose, CA, USA) for invasion analysis. Then, 100 μL cell suspension was pipetted into the upper chamber and added 500 μL RPMI-1640 medium was added to the lower chamber. Transwell chamber were incubated at 37°C for 24 h, and cells on the upper surface were gently wiped away. Cells passed into the lower chamber were fixed with paraformaldehyde (Beyotime) for 20 min, followed by staining of crystal violet (Beyotime) for 15 min. Finally, cells were observed under the microscope and counted in 5 randomly selected fields.
Western blot
RIPA buffer (Beyotime) was used to extract total proteins from cells and tissues. The protein concentration of samples was determined by BCA (Beyotime) method. The proteins were separated using sodium dodecyl sulfate-polyacrylamide gel electrophoresis, then the gels were transferred onto the PVDF (Millipore, Billerica, MA, USA) membrane and sealed with 5% milk for 1 h. PVDF membranes were incubated with the primary antibody against proliferating cell nuclear antigen (PCNA; ab18197, 1: 1000, Abcam), Bax (ab32503; 1:1000; Abcam), E-cadherin (Abcam, ab40772, 1:1000) and β-actin (bs-0061R; Bioss, Beijing, China) overnight at 4°C. Then, secondary antibody (bs-0295M-HRP; Bioss) were incubated to membranes for 1 h. ECL (Beyotime) hypersensitive luminescence solution was added to visualize the blots, and Image Lab was used to analyze the gray value of the blots
Dual-luciferase reporter gene assay
The circKRT14 or E2F3 3′UTR fragment containing wild type (WT) miR-1256 binding site or mutant type (MUT) miR-1256 binding site were cloned into the pGL3 vector (Promega, Madison, WI, USA) to obtain circKRT14 WT, circKRT14 MUT and E2F3 3′UTR WT, E2F3 3′UTR MUT plasmids, respectively. The plasmids were co-transfected with miR-NC or miR-1256 into KYSE-150 and EC9706 cells, and the dual-luciferase reporter gene kit (Promega) was prepared to examine the luciferase activity.
Xenograft mice model
Ten BALB/c nude mice were collected from Beijing Vital River Laboratory Animal Technology Co., Ltd (Beijing, China), and were randomly divided into sh-NC group and sh-circKRT14 group. KYSE-50 cells (5 × 107) were transfected into sh-NC or sh-circKRT14 and injected subcutaneously into nude mice. The tumor volume (volume = length × width2/2) was measured every week for five weeks. Thirty-five days later, the mice were sacrificed, and the tumor tissue was removed and weighed.
Immunohistochemical assay
Small pieces of fresh tumor tissue were taken and cut into paraffin sections. Antigen retrieval was performed in citrate buffer, and paraffin sections were blocked with 10% goat serum and cultured overnight with primary antibody, including ki-67 (ab92742; Abcam), PCNA (ab18197, Abcam), Bax (ab32503, Abcam), E-cadherin (Abcam, ab40772) at 4°C, and then incubated for 2 h with secondary antibody (ab205718, Abcam). DAB (Beyotime) was used to incubate with the slices, and hematoxylin (Beyotime) was employed for the re-staining of the slices. Finally, the slides were observed and photographed under a microscope.
Statistical analysis
The Student’s t-test and one-way ANOVA were employed for difference analysis. And a Pearson’s correlation coefficient was used to analyze the correlations. p < 0.05 (*p < 0.05, **p < 0.01, ***p < 0.001) was considered as statistically significant difference. Data were displayed as mean ± standard deviation (SD).
Results
CircKRT14 was increased in EC cells and tissues
The circBase (http://www.circbase.org/) revealed that circKRT14 was derived from Exons 2 and 3 of KRT14 gene (Figure 1A). the qRT-PCR results displayed that the level of circKRT14 in EC tissues was dramatically increased compared with normal tissues (Figure 1B). Also, circKRT14 was remarkably boosted in EC cells (KYSE-150, KYSE-410 and EC9706) compared with normal HET-1A cells (Figure 1C). KYSE-150 and EC9706 cells with more significant difference of circKRT14 were used as experimental cell lines. In addition, circKRT4 was resistant to RNase R, while the level of linear KRT14 was conspicuously reduced in KYSE-150 and EC9706 cells (Figure 1D and E). In conclusion, circKRT14 was a stable and upregulated circRNA in EC. Expression of circKRT14 in EC tissues and cell lines. (a) The schematic diagram of genomic location of circKRT14. (b) Relative expression of circKRT14 in EC tissues and normal tissues was measured by qRT-PCR. (c) Relative expression level of circKRT14 in EC cells and normal cells was detected by qRT-PCR. (d and e) The levels of circKRT14 and linear KRT14 were assessed by qRT-PCR assay in KYSE-150 and EC9706 cells treated with RNase R. **p < 0.01, ***p < 0.001.
CircKRT14 knockdown inhibited cell proliferation, migration, and invasion, but induced apoptosis in EC cells
To investigate the effect of circKRT14 on EC cell progression, we silenced circKRT14 expression in KYSE-150 and EC9706 cells by transfection of sh-circKRT14. The results indicated that sh-circKRT14 greatly reduced circFKRT14 level compared with the sh-NC group (Figure 2A). Colony formation and EdU assays demonstrated that circKRT14 silencing specially diminished colony formation number and cell proliferation capability compared with the control group (Figure 2B and C). Flow cytometry data uncovered that circKRT14 inhibition promoted cell apoptosis compared with the control group (Figure 2D). In addition, downregulation of circKRT14 suppressed the migration and invasion of KYSE-150 and EC9706 cells (Figure 2E and F). Moreover, circKRT14 knockdown apparently retarded the level of PCNA protein, and facilitated the protein levels of Bax and E-cadherin. These results suggested that knockdown of circRT14 curbed EC cell progression in vitro. Silencing of circKRT14 constrained EC cell progression in vitro. KYSE-150 and EC9706 cells were transfected with sh-NC or sh-circKRT14. (a) The content of circKRT14 in KYSE-150 and EC9706 cells was detected by qRT-PCR. (b) The number of colonies were measured through colony formation assay. (c) EdU analysis was used to assess the proliferation ability of KYSE-150 and EC9706 cells. (d) Flow cytometry assay was used to monitor cell apoptosis. (e and f) The invasion and migration of KYSE-150 and EC9706 cells was examined by transwell assay. (g and h) The protein levels of PCNA, Bax and E-cadherin were estimated through western blot. **p < 0.01, ***p < 0.001.
CircKRT14 was a sponge of miR-1256 in EC
To further explore the mechanism of circKRT14 on EC, we predicted the potential target miRNAs of circKRT14 by circBank and circinteractome database, and found that miR-223-3p, miR-657 and miR-1256 might be targeted by circKRT14 (Figure 3A). The qRT-PCR results manifested that the level of miR-1256 in sh-circKRT14 group was evidently increased compared with the control group, but there was no significant difference in the expression of miR-223-3p and miR-657 (Figure 3B and C). The binding sites between circKRT14 and miR-1256 were shown by circinteractome database (Figure 3D). We also discovered that miR-1256 overexpression obviously promoted miR-1256 levels in KYSE-150 and EC9706 cells compared with the control group (Figure 3E). As shown in Figure 3F and G, the luciferase activity was prominently reduced by the co-transfection of miR-1256 and circKRT14 WT, while there was no significant change in miR-1256 and circKRT14 MUT co-transfection group. The miR-1256 level was strikingly diminished in both EC tissues and cells compared with normal controls (Figure 3H and I). Moreover, miR-1256 level was negatively correlated with circKRT14 level in EC tissues (Figure 3J). In summary, circKRT14 directly targeted miR-1256 and regulated miR-1256 expression. CircKRT14 sponged miR-1256 to regulate miR-1256 expression. (A) We used circBank and circInteractome databases to screen out three miRNAs that may targeted by circKRT14. (b and c) The expression of miR-223-3p, miR-657 and miR-1256 in EC cells after knockdown of circKRT14 was assessed by qRT-PCR. (d) CircInteractome predicted the interaction sites between circKRT14 and miR-1256. (e) The abundance of miR-1256 in KYSE-150 and EC9706 cells transfected with miR-NC or miR-1256 was detected by qRT-PCR. (f and g) The interaction between circKRT14 and miR-1256 was verified by dual-luciferase reporter assay. (h and i) The content of miR-1256 in EC tissues and cells was determined by qRT-PCR assay. (j) The correlation between the levels of circKRT14 and miR-1256 in EC tissues was monitored. **p < 0.01, ***p < 0.001.
The inhibition effect of circKRT14 knockdown on EC cell progression was reversed by in-miR-1256
Next, we further explored the relationship between circKRT14 and miR-1256 through functional experiments. The data implicated that miR-1256 inhibitor conspicuously decreased miR-1256 level compared with the control group (Figure 4A). As shown in Figure 4B, circKRT14 inhibition substantially increased miR-1256 level, while miR-1256 inhibitor partially restored this change (Figure 4B). CircKRT14 silencing retarded the proliferation of KYSE-150 and EC9706 cells, while in-miR-1256 eliminated this inhibitory effect (Figure 4C and D). Similarly, miR-1256 inhibitor restored the promoting effect of circKRT14 knockdown on cell apoptosis (Figure 4E). In addition, the inhibitory effects of circKRT14 knockdown on cell migration and invasion were attenuated by miR-1256 inhibitor (Figure 4F and G). The regulatory changes of circKRT14 silencing on PCNA, Bax and E-cadherin protein levels were rescued by in-miR-1256 (Figure 4H and I). These results confirmed that circKRT14 mediated EC progression by sponging miR-1256 in part. MiR-1256 inhibitor mitigated circKRRT14 silencing-induced suppression on cell proliferation, migration and invasion in EC cells. (a) MiR-1256 level in KYSE-150 and EC9706 cells transfected with in-miR-NC or in-miR-1256 was detected through qRT-PCR. (b–i) KYSE-150 and EC9706 cells were transfected with sh-NC, sh-circKRT14, sh-circKRT14+in-miR-NC or sh-circKRT14+in-miR-1256. (b) The miR-1256 expression was analyzed using qRT-PCR. (b and d) Cell proliferation was examined using colony formation assay and EdU assay. (e) Cell apoptosis was tested by using flow cytometry assay. (f and g) Transwell assay was used to examine cell invasion and migration. (h and i) PCNA, Bax and E-cadherin protein levels were determined through western blot. *p < 0.05, **p < 0.01, ***p < 0.001.
E2F3 directly interacted with miR-1256
By using Targetscan analysis, E2F3 was found to contain the complementary binding sites of miR-1256 (Figure 5A). As shown in Figure 5B and C, luciferase activity was evidently reduced in KYSE-150 and EC9706 cells co-transfected with E2F3 3′UTR WT and miR-1256, while there was no significant difference in E2F3 3′UTR MUT and miR-1256 co-transfection group. In addition, the expression levels of E2F3 mRNA and protein in EC tissues and cells were strikingly higher than that of normal controls (Figure 5D-2013F). In addition, circKRT14 silencing effectively inhibited E2F3 level in KYSE-150 and EC9706 cells, while in-miR-1256 reversed this effect (Figure 5G and H). We also observed that E2F3 level was negatively correlated with miR-1256 level and positively correlated with circKRT14 level in EC tissues (Figure 5I and J). In summary, miR-1256 targeted E2F3 in EC cells and circKRT14 could regulate E2F3 expression by targeting miR-1256. E2F3 acted as a target of miR-1256. (a) The potential binding sites between E2F3 and miR-1256 were predicted by Targetscan. (b and c) Dual-luciferase reporter assay was used to analyze the luciferase activity of KYSE-150 and EC9706 cells. (d and e) The expression levels of E2F3 mRNA and protein in EC tissues were detected by using qRT-PCR and western blot. (f) E2F3 protein levels in EC cells were evaluated through western blot assay. (g and h) The levels of E2F3 mRNA and protein in KYSE-150 and EC9706 cells transfected with sh-NC, sh-circKRT14, sh-circKRT14+in-miR-NC or sh-circKRT14+in-miR-1256 were detected by qRT-PCR and western blot, respectively. (i and j) Pearson’s correlation coefficient was used for linear analysis between miR-1256 and E2F3 (i), as well as circKRT14 and E2F3 (j). **p < 0.01, ***p < 0.001.
E2F3 overexpression reversed the effect of miR-1256 overexpression on EC progression
The western blot results presented that E2F3 transfection obviously increased E2F3 level in KYSE-150 and EC9706 cells compared with the control group (Figure 6A). The miR-1256 overexpression strikingly reduced E2F3 level, while E2F3 promotion restored this change (Figure 6B). Cell cloning experiments demonstrated that miR-1256 overexpression prominently diminished the number of colonies in KYSE-150 and EC9706 cells, while E2F3 overexpression attenuated this effect (Figure 6C). Subsequently, EdU assay results displayed that upregulation of miR-1256 dramatically inhibited cell proliferation, while E2F3 overexpression rescued this inhibition effect (Figure 6D). Also, E2F3 overexpression eliminated the promoting influence of miR-1256 overexpression on cell apoptosis (Figure 6E). Transwell assay revealed that miR-1256 up-regulation markedly inhibited cell migration and invasion, while E2F3 overexpression weakened these effects (Figure 6F and G). Western blot assay showed that E2F3 overexpression in KYSE-150 and EC9706 cells saved the effects of miR-1256 overexpression on PCNA, Bax and E-cadherin protein levels (Figure 6H and I). These findings elucidated that miR-1256 played an inhibitory role in EC cells by partly inhibiting the expression of E2F3. E2F3 promotion partially relieved the function of miR-1256 overexpression. (a) The protein level of E2F3 was examined by western blot in KYSE-150 and EC9706 cells. (b–i) KYSE-150 and EC9706 cells were transfected with miR-NC, miR-1256, miR-1256 + pcDNA or miR-1256 + E2F3. (b) Relative E2F3 protein level was evaluated by western blot analysis. (c and d) Colony formation assay and EdU assay were used to estimated colony number and EdU positive cell rate. (e) The cell apoptosis was analyzed by flow cytometry. (f and g) Cell migration and invasion were assessed using Transwell assay. (h and i) Western blot was used for detecting PCNA, Bax and E-cadherin protein levels. *p < 0.05, **p < 0.01, ***p < 0.001.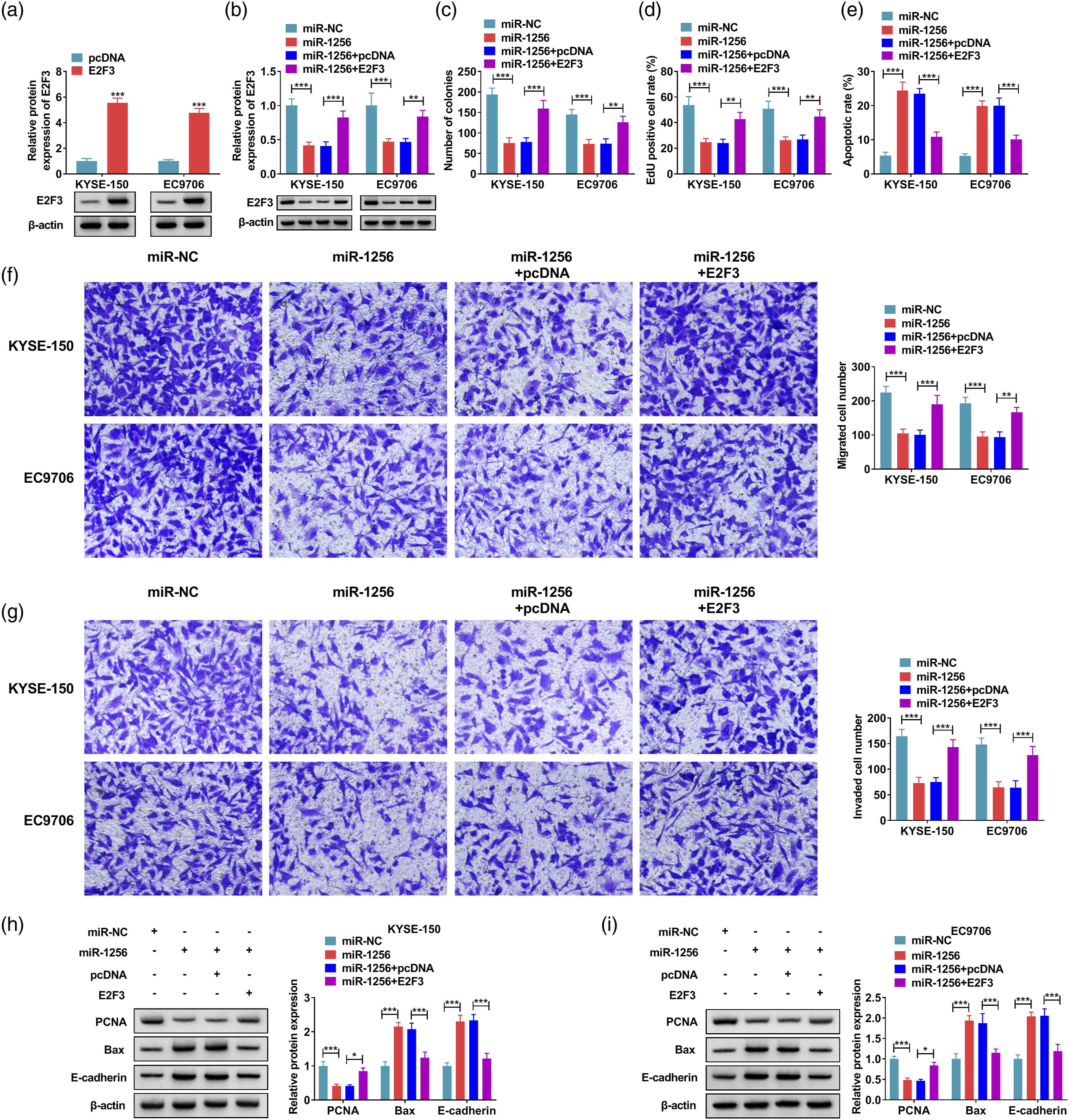
Silencing of circKRT14 curbed EC growth in vivo by regulating miR-1256/E2F3 axis
We used the xenograft mice model to study the effect of circKRT14 knockdown on tumor growth in vivo. CircKRT14 inhibition conspicuously constrained tumor volume and weight compared with the control group (Figure 7A-C). QRT-PCR results suggested that circKRT14 downregulation reduced the levels of circKRT14 and E2F3, and increased the level of miR-1256 in xenograft tumor tissues (Figure 7D). CircKRT14 silencing also obviously diminished the level of E2F3 protein in xenograft tumor tissues (Figure 7E). IHC assay indicated that circKRT14 knockdown downregulated protein levels of ki-67 and PCNA but upregulated protein levels of Bax E-cadherin in tumor tissues (Figure 7F). In conclusion, cirKRT14 downregulation reduced EC tumor growth and this function might be achieved by regulating miR-1256/E2F2 axis. Silencing of circKRT14 blocked tumor growth in vivo. (a and b) The tumor volume was estimated after being transfected with sh-NC or sh-circKRT14. (c) The tumor weight in sh-NC or sh-circKRT14 group was measured. (d) The levels of circKRT14, miR-1256 or E2F3 in tumor tissues were detected by qRT-PCR. (e) The western blot was used to assay the expression of E2F3 in tumor tissues. (f) IHC assay was performed to analyze the protein levels of ki-67, PCNA, Bax and E-cadherin. ***p < 0.001.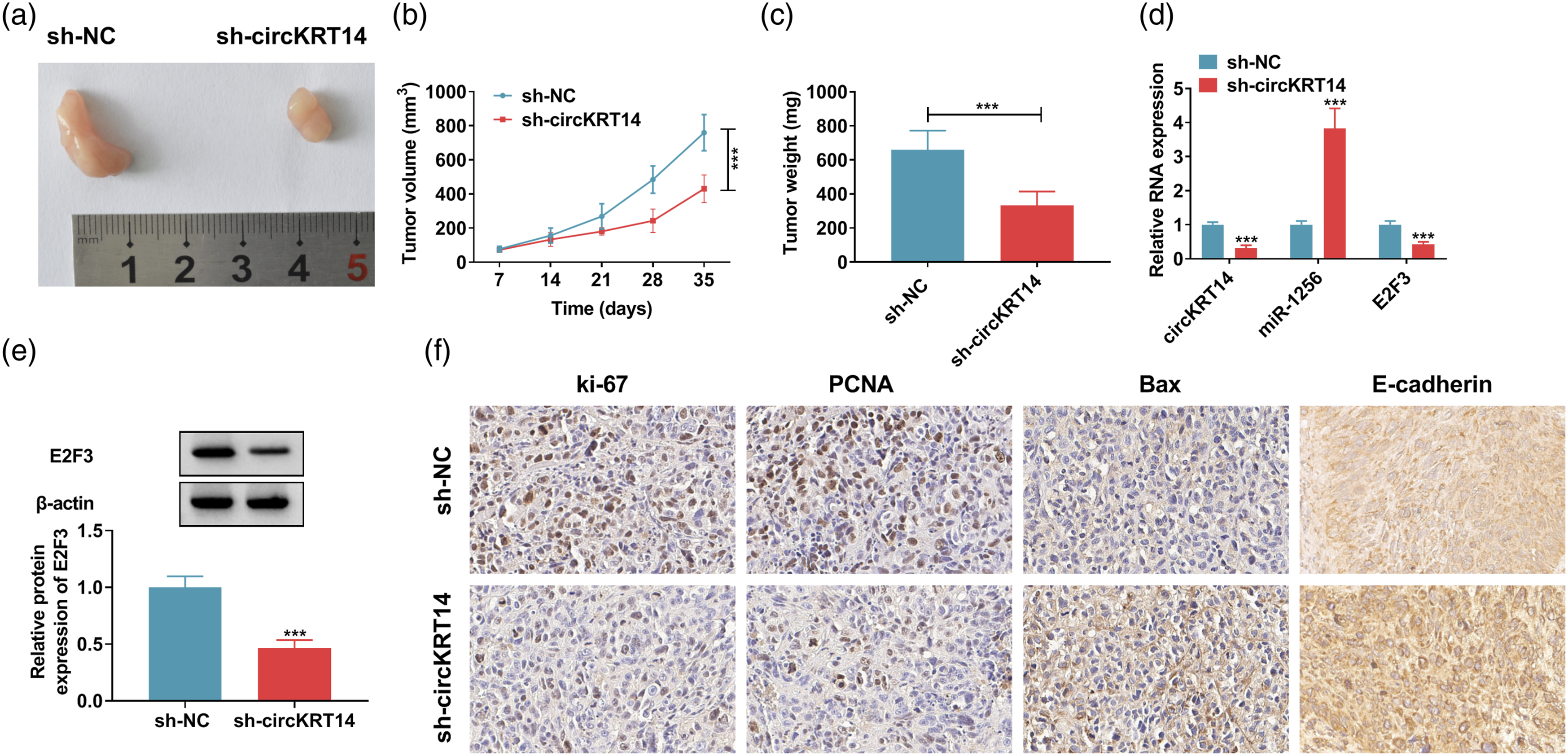
Discussion
EC is a common gastrointestinal malignancy and it has unclear pathogenic causes, including inflammation, chemical stimulation, trauma, genetic factors and lifestyle. 22 Due to the limitation of effective diagnosis and treatment, the clinical prognosis of EC is poor. 23 Therefore, in-depth understanding of the molecular mechanism of biological behavior of EC will provide important clues to boost the survival and quality of life of patients.
CircRNA is a hot field in tumor research because of its strong stability and circRNA the specificity of tissue timing. 24 A mass of literatures has indicated the abnormal expression levels and regulatory roles of various circRNAs in tumors. Recent studies have found that circ-HMGA2 was reinforced in lung adenocarcinoma, and circ-HMGA2 could promote lung adenocarcinoma metastasis via miR-1236-3p/ZEB1 axis. 25 Some scholars have confirmed that circ_0,044,516 played a role in GC progression by promoting HuR expression through sponging miR-149-5p. 26 Hou et al. showed that circ_0,025,033 promoted the evolution of ovarian cancer by targeting miR-184 to activate LSM4 expression. 27 In this experiment, the content of circKRT4 was significantly increased in EC cells and tissues. Further studies showed that circKRT14 silencing inhibited EC cell proliferation, migration and invasion but promoted apoptosis. Moreover, circKRT14 inhibition inhibited EC tumor growth in vivo. In conclusion, circKRT14 played as an oncogenic factor in EC.
A lot of researches have reported that miRNAs regulate a variety of cell pathophysiological behaviors and take part in the development of EC. 28 Song et al. uncovered that miR-140 inhibited the evolution of EC by targeting ZEB2 to regulate Wnt/β-catenin pathway. 29 Hu et al. 30 elucidated that miR-196a expedited the proliferation and migration of EC through the UHRF2/TET2 axis . MiR-133a-3p inhibited the malignant progression of EC by targeting CDCA8. 31 In addition, Wu et al. 32 found that miR-1256 inhibited cell evolution in papillary thyroid cancer. In this study, we discovered that miR-1256 was conspicuously confined in EC cells and tissues, and circKRT14 was predicted to have binding sites with miR-1256 by circInteractome. Besides, miR-1256 repression reversed the inhibitory impact of circKRT14 suppression on EC progression. These results suggested that miR-1256 inhibited EC development, and it acted as the target gene of circKRT14.
Next, target binding of miR-1256 to E2F3 was predicted by Targetscan and verified by dual-luciferase reporting experiment. The E2F family is a transcriptional regulatory family involved in many biological processes. 33 E2F3 acted as a member of the E2F family, which has carcinogenic properties. 34 It has been shown that E2F3 expression was regulated by miR-377-3p in non-small cell lung cancer. 35 Moreover, miR-217 inhibited pancreatic cancer development by targeting E2F3. 36 Herein, E2F3 was found as a target for miR-1256. E2F3 expression level was significantly increased in EC cells and tissues. Moreover, circKRT14 could regulate the expression of E2F3 via interacting with miR-1256. The inhibitory role of miR-1256 in EC progression was also achieved by downregulating E2F3 in part.
In conclusion, circKRT14 silencing inhibited EC progression in vitro and in vivo. Moreover, circKRT14 regulated EC cell progression via miR-1256/E2F3 axis, and it might be a potential diagnostic and therapeutic target for EC.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
