Abstract
Background
Circular RNAs (circRNAs) have been identified to participate in regulating multiple malignancies. Herein, this study aimed to explore the clinical significance, biological function, and regulatory mechanisms of circRNA lysophosphatidic acid receptor 3 (circLPAR3) in esophageal squamous cell carcinoma (ESCC) cell malignant phenotypes and Warburg effect.
Methods
The qRT-PCR and Western blot were used to detect the levels of genes and proteins. Glucose uptake and lactate production were detected to determine the Warburg effect. The effects of circLPAR3 on ESCC cell proliferation, apoptosis, and metastasis were evaluated by MTT, 5-ethynyl-2′-deoxyuridine (EdU), flow cytometry, wound healing, and transwell assays. The binding interaction between miR-873-5p and circLPAR3 or lactate dehydrogenase A (LDHA) was verified using dual-luciferase reporter and RIP assays. Xenograft mice models were established to conduct in vivo analysis.
Results
CircLPAR3 is a stable circRNA and was increased in ESCC tissues and cells. Functionally, circLPAR3 knockdown suppressed ESCC cell Warburg effect, proliferation, metastasis, and induced apoptosis in vitro, and impeded xenograft tumor growth and Warburg effect in ESCC mice models. Mechanistically, circLPAR3 served as a sponge for miR-873-5p, which targeted LDHA. Moreover, circLPAR3 could regulate LDHA expression by sponging miR-873-5p. Thereafter, rescue experiments suggested that miR-873-5p inhibition reversed the anticancer effects of circLPAR3 silencing on ESCC cells. Furthermore, miR-873-5p overexpression restrained ESCC cell Warburg effect and oncogenic phenotypes, which were abolished by LDHA up-regulation.
Conclusion
CircLPAR3 knockdown suppressed ESCC cell growth, metastasis, and Warburg effect by miR-873-5p/LDHA axis, implying a promising molecular target for ESCC therapy.
Keywords
Introduction
On a global scale, esophageal squamous cell carcinoma (ESCC) is the sixth leading cause of cancer-related mortality with approximately 330,000 new cases and over 270,000 deaths annually.1, 2 Although advances in comprehensive therapies, the outcome of ESCC remains poor due to the lack of early diagnosis, therapy resistance, metastasis, and frequent recurrence. 3 In order to better palliate symptoms and prolong survival, molecular-targeting agents have been investigated, while no effective approved molecular targeted therapeutics demonstrate the clinical utility in ESCC. 4 As a result, further investigations on the pathological mechanisms is essential for exploring novel avenues to prevent ESCC.
Circular RNAs (circRNAs) are closed circular molecules that lack 5′ and 3′ ends. 5 CircRNAs are considerable abundance across various species, and exhibit high specific, stability, and conservation relative to other types of RNAs. 6 Recent studies have reported that circRNAs are extensively involved in the regulation of a wide range of biological processes,7,8 and abnormal expression of circRNAs is identified in many types of cancer, which is closely associated with carcinogenesis, metastasis and cancer development.9-11 For example, circGSK3β facilitated ESCC cell invasion and migration by activating β-catenin signaling. 12 Circ_0,000,140 was demonstrated to act as a tumor suppressor to suppress the growth and metastasis of ESCC through augmenting Hippo pathway via miR-31/LATS2 axis. 13 Thus, circRNAs may be ideal biomarkers for clinical ESCC therapeutics. CircLPAR3 (ID: hsa_circ_0,004,390) is a circRNA derived from its parental gene lysophosphatidic acid receptor 3 (LPAR3). Shi et al. showed that circLPAR3 accelerated ESCC cell metastasis by up-regulating MET gene expression via miR-198. 14 Besides that, Cheng’s team suggested that circLPAR3 expedited ESCC cell proliferation, invasion and metastasis through the miR-375/miR-433-HMGB1 axis. 15 Warburg effect is the major cellular metabolic method of glucose metabolism in cancer cells in either anoxic or aerobic microenvironment, which is responsible for the sustaining growth of cancer cells.16,17 Currently, numerous evidence exhibits that circRNAs have role in regulating the Warburg effect in human cancers.18,19 However, the role of circLPAR3 in ESCC cell Warburg effect is still unclear.
Herein, we aimed to probe the action of circLPAR3 in ESCC cell malignant phenotypes and Warburg effect, moreover, the mechanism by which circLPAR3 regulates ESCC cell tumorigenesis was also explored.
Materials and methods
Clinical tissue specimens
The ESCC tissues and adjacent normal tissues were collected from 58 ESCC patients who had undergone esophagectomy at Second Affiliated Hospital of Xi’an Jiaotong University, and were then immediately stored at −80°C until use. All patients were newly diagnosed as ESCC through pathological examination, and did not received preoperative therapy. Written informed consent had been collected from all subjects before samples collection. This work was authorized by the Ethics Committee of Xi’an Jiaotong University.
Cell culture
ESCC cell lines (TE-1, ECA109, and KYSE150) and the normal esophageal cell line Het-1A, as well as HEK293 T cells were obtained from Cell Bank of Shanghai Institute of Cell Biology (Shanghai, China). All cell lines were authenticated and tested for mycoplasma contamination, and then cultivated in Dulbecco’s modified eagle’s medium (DMEM; Invitrogen, Carlsbad, CA, USA) containing 1% penicillin-streptomycin (Solarbio, Shanghai, China) and 10% fetal bovine serum (FBS, Invitrogen) at 37°C with 5% CO2.
Plasmid transfection
The circLPAR3-specific siRNAs (si1-circLPAR3 and si2-circLPAR3), pCD-ciR/circLPAR3 overexpression plasmid, pcDNA3.1/LDHA overexpression plasmid and their negative controls (si-NC, pCD-ciR or pcDNA) were synthesized by GeneChem (Shanghai, China). The miR-873-5p mimic, inhibitor, and negative controls (miR-NC or anti-miR-NC) were provided by Ribobio (Guangzhou, China). Then transient transfection in TE-1 and KYSE150 cells with 100 nM of siRNAs, 50 nM of miRNA mimic, or 100 ng of plasmids was implemented using Lipofectamine 2000 (Invitrogen).
Lentivirus vectors carrying short hairpin RNA (shRNA) targeting circLPAR3 (sh-circLPAR3) or sh-NC were constructed by Biowit Technology (Shenzhen, China). The stably transfected cells were selected by puromycin (2–5 μg/mL) to conduct animal experiments.
Quantitative real-time PCR (qRT-PCR)
Primers sequences used for qRT-PCR.

RNase R digestion and actinomycin D treatment
For RNase R treatment, total RNAs (3 μg) were interacted with 5 U/μg of RNase R (Epicentre Biotechnologies, Madison, WI, USA) or mock without enzymatic activity for 1.5 h at 37°C. Later, levels of circLPAR3 and linear LPAR3 were detected by qRT-PCR.
For Actinomycin D treatment, TE-1 and KYSE150 cells were mixed with 5 μg/mL Actinomycin D to block transcription, and the half-life of circLPAR3 and linear LPAR3 was evaluated using qRT-PCR at 0, 6, 12, and 18 h.
Glucose uptake and lactate production
TE-1 and KYSE150 cells, following indicated transfection, were incubated in a 60 mm plates with FBS-free medium for 24 h. Then glucose uptake and lactate production in the culture medium were detected using a glucose assay kit (Sigma-Aldrich, St Louis, MO, USA) and a Lactate Assay kit (BioVision, Mountain View, CA, USA) as per the manufacturer’s instruction as previously described. 21 The data were normalized to total cell proteins.
3-(4, 5-dimethylthiazol-2-yl)-2, 5-diphenyl-2H-tetrazolium bromide) (MTT) assay
Transfected TE-1 and KYSE150 cells were seeded into a 96-well plate and cultured for 1, 2, and 3 days. Then each well was incubated with 20 μL MTT regent (Sigma-Aldrich) for 4 h. At the end of incubation, the supernatants were removed and each well was mixed with 100 μL DMSO for 30 min. Later on, the absorbance at 570 nm was assessed using a microplate reader.
5-ethynyl-2′-deoxyuridine (EdU) assay
EdU assay with the Cell-Light™ EdU kit (Ribobio) was also utilized to assess cell proliferation as described before. 22 After transfection, TE-1 and KYSE150 cells were seeded onto a 96-well plate adding 90 μL of fresh culture medium and 10 μL of EdU reagent (RiboBio) and incubated for 2 h. Cells were fixed and permeabilized, followed by staining with Apollo reaction mixture for 30 min in the dark. Cell nucleus was stained with DAPI, and stained cells were determined by using a fluorescence microscope.
Flow cytometer
Transfected TE-1 and KYSE150 cells were harvested, resuspended in 500 μL 1×binding buffer to a concentration of 1 × 106/mL, and then stained with 5 μL FITC-conjugated Annexin V and PI (Beyotime) under darkness for 15 min. 23 Later on, apoptotic cells were detected by a flow cytometry (FACSCanto II, BD Biosciences, San Diego, CA, USA).
Wound healing assay
After transfection, TE-1 and KYSE150 cells were cultured in six-well plates. When reached to ∼100% confluence, cells were starved for 6 h and then scratched by using a sterile 200 μL pipette tip. After washing, the images were captured at 0 and 24 h culture, and the distance between the two edges of the wound was detected to calculate cell migration ability.
Transwell assay
Transwell inserts in 24-wells (Costar, Corning, Switzerland) with Matrigel-coated membrane (BD Biosciences) were used to determine cell invasion. Transfected TE-1 and KYSE150 cells (1 × 105) were suspended in 200 μL serum-free medium and inoculated into the upper chambers, and the lower chamber was filled with 500 μL of complete culture medium. 24 h later, cells invaded into the lower side of the membrane were fixed by ethanol and stained with 0.1% crystal violet, and then cells in five randomly selected fields were counted manually under the microscope.
Western blot
Total protein was extracted by using RIPA buffer containing phosphatase inhibitor cocktail and proteinase. After qualification by a BCA method, about 30 μg proteins were separated by 10% SDS-PAGE, and transferred onto the polyvinylidene fluoride (PVDF) membranes (Millipore, Darmstadt, Germany). Thereafter, primary antibodies included PCNA (1:5000, ab29), Vimentin (ab92547, 1:1000), Cleaved caspase 3 (1:2000, ab2302), Slug (1:1000, ab27568), lactate dehydrogenase (LDHA) (1:5000, ab52488) and β-actin (1:1000, ab6276) were used to incubate with the membranes overnight at 4°C, followed by the incubation with an HRP-conjugated second antibody (1: 1000, ab9482) for 2 h at 37°C. Protein bands were visualized by sensitive electrochemiluminescence (ECL) method (Beyotime). All antibodies were provided by Abcam (Cambridge, MA, USA).
Dual-Luciferase reporter assay
The sequences containing miR-873-5p binding sites in the circLPAR3 and LDHA 3′-UTR, and the site-directed mutagenesis in the binding sites generated by Genema were subcloned into the pmirGLO report luciferase vector (Invitrogen) to construct wild-type (wt) or mutated (mut) luciferase reporter vectors. Then vectors together with miR-873-5p mimic or the control (miR-NC) were co-transfected into HEK293T cells, and luciferase activity normalized to Renilla was analyzed after 48 h of transfection using the Dual Light System following the instruction.
RNA immunoprecipitation (RIP) assay
RIP assay was performed by using EZMagna RIP kit (Millipore) as per the manufacturer’s instruction. TE-1 and KYSE150 cells were transfected with miR-873-5p mimic or the control (miR-NC) for 48 h and then lysed by RIP lysis buffer. The lysates of cells were incubated with 5 μg of Ago2 antibody (Millipore) or rabbit IgG-coated beads overnight at 4°C. After treatment with proteinase K buffer, immunoprecipitated RNA was extracted and the enrichment of circLPAR3 was detected using qRT-PCR.
Tumor xenograft assay
Six-weeks-old BALB/c nude mice (male, n = 12) were purchased from Charles River Labs (Beijing, China) and maintained in pathogen-free conditions. TE-1 cells stably transfected with lentiviral-sh-circLPAR3 or lentiviral-sh-NC (5 × 106/0.2 mL PBS) were subcutaneously injected into the blindly randomized nude mice (n = 6/per group). Tumor size was measured every 5 days, and the volume of xenografts was assessed as the formula: volume = (length × width2)/2. At day 30, the mice were sacrificed, tumor masses were excised, weighed and collected for genes detection using qRT-PCR and Western blot, or fixed in formalin for IHC analysis as described previously. 24
Statistical analysis
Continuous variables are denoted as means ± SD. Statistical analyses were conducted with the t-test or paired t-test, or analysis of variance by GraphPad Prism 6 software (GraphPad, San Diego, CA, USA). Pearson’s correlation was applied for correlation analysis. p < 0.05 means significant differences.
Results
CircLPAR3 is highly expressed in ESCC tissues and cells
To evaluate the potential functions of circLPAR3 in ESCC, we initially investigated the expression profile of circLPAR3 in clinical ESCC samples. As shown in Figure 1(a), circLPAR3 expression was significantly higher in ESCC tissues than those in matched adjacent normal tissues. Furthermore, the relationship between circLPAR3 expression and clinical characteristics of ESCC patients were listed in Table 2. The expression of circLPAR3 is significantly correlated with differentiation (p = 0.004), TNM stage (p = 0.014), and lymph node metastasis (p = 0.002). Besides that, Kaplan-Meier survival curve suggested that high expression level of circLPAR3 was associated with shorter overall survival of patients with ESCC (Figure 2(b)). Thereafter, we observed a marked increase of circLPAR3 expression in ESCC cell lines (TE-1, ECA109, and KYSE150) compared with normal Het-1A cells (Figure 1(c)). CircLPAR3 is generated from exon 2 of its parental gene LPAR3 by back-splicing, it locates at chr1: 85,331,067–85,331,821 with the genomic length of 754 base pairs (bp) (Figure 1(d)). Thereafter, the stability of circLPAR3 was determined. The results of RNase R treatment indicated that circLPAR3 could resistant to the degradation by RNase R relative to the linear LPAR3 (Figure 1(e)). Moreover, Actinomycin D treatment suggested that the half-life of circLPAR3 exceeded 18 h, while that of linear LPAR3 mRNA was about 6 h (Figure 1(f) and (g)). Therefore, we confirmed that circLPAR3 was a stable circRNA and might be associated with ESCC progression. CircLPAR3 is highly expressed in ESCC tissues and cells. (a) qRT-PCR analysis of circLPAR3 expression in ESCC tissues and matched adjacent normal tissues. (b) Kaplan-Meier survival analysis of overall survival based on circLPAR3 expression in ESCC patients. (c) qRT-PCR analysis of circLPAR3 expression in ESCC cell lines (TE-1, ECA109, and KYSE150) and normal Het-1A cells. (d) Schematic diagram of the genomic location and formation of circLPAR3. (e) qRT-PCR analysis of circLPAR3 and linear LPAR3 expression in KYSE150 and TE-1 cells treated with RNase R or mock. (f) qRT-PCR analysis of circLPAR3 and linear LPAR3 expression in KYSE150 and TE-1 cells treated with Actinomycin D. **p <0.01, ***p <0.001. Clinicopathological factors and circLPAR3 expression in patients with ESCC. *p < 0.05, Chi-square test. CircLPAR3 knockdown suppresses ESCC cell Warburg effect and growth in vitro. (a–j) KYSE150 and TE-1 cells were transfected with si1-circLPAR3, si2-circLPAR3 or si-NC. (a, b) qRT-PCR analysis of circLPAR3 expression in cells after transfection. (c–f) Relative glucose uptake and lactate production in cells were detected by commercial kits. (g–i) Cell proliferation analysis by using MTT and EdU assays. (j) Flow cytometry for cell apoptosis analysis. *p <0.05, **p <0.01, ***p <0.001.


CircLPAR3 knockdown suppresses ESCC cell Warburg effect, growth and metastasis in vitro
Next, in order to investigate the biological roles of circLPAR3 in ESCC tumorigenesis, we constructed two siRNAs targeting circLPAR3 to knock down circLPAR3 in ESCC cells (KYSE150 and TE-1) (Figure 2(a) and (b)). Then the effect of circLPAR3 in Warburg effect was first determined. We proved that circLPAR3 silencing reduced glucose uptake and lactate production compared with the si-NC group in KYSE150 and TE-1 cells (Figure 2(c)–(f)). Thereafter, MTT and EdU assays manifested that knockdown of circLPAR3 inhibited cell viability and DNA synthesis activity, suggesting the repression of the proliferation in KYSE150 and TE-1 cells (Figure 2(g)–(i)). Besides, the results of flow cytometry showed that circLPAR3 knockdown led to apoptosis in KYSE150 and TE-1 cells (Figure 2(j)). Subsequently, the results of wound healing and transwell assays showed that both the migration and invasion abilities of KYSE150 and TE-1 cells were restrained after circLPAR3 knockdown (Figures 3(a) and (b)), Moreover, we also observed that circLPAR3 knockdown caused the decreases of PCNA, vimentin, and Slug protein levels as well as increase of cleaved caspase 3 protein level in KYSE150 and TE-1 cells (Figure 3(c)). Additionally, we also overexpressed circLPAR3 expression in ESCC cells to conduct gain-of-function assay. Following the transfection of circLPAR3 overexpression plasmids, circLPAR3 expression was significantly up-regulated (Fig. S1(A)). Then it was proved that circLPAR3 up-regulation promoted glucose uptake and lactate production (Fig. S1(B), (C)), enhanced cell proliferation (Fig. S1(D)-(F)), migration (Fig. S1(H)), invasion (Fig. S1(I)) and suppressed apoptosis (Fig. S1(G)) in KYSE150 and TE-1 cells. Taken together, circLPAR3 silencing suppressed ESCC cell Warburg effect and malignant phenotypes. CircLPAR3 knockdown suppresses ESCC cell metastasis in vitro. (a–c) KYSE150 and TE-1 cells were transfected with si1-circLPAR3, si2-circLPAR3 or si-NC. (a) Wound healing assay for cell migration. (b) Transwell assay for cell invasion. (c) Western blot analysis for the protein levels of PCNA, cleaved caspase 3, vimentin, and Slug in cells. **p <0.01, ***p <0.001.
MiR-873-5p is a target of circLPAR3
Previous researches have proposed that circRNA can function as competitive endogenous RNAs (ceRNAs) to sponge miRNAs to exert their functions.25,26 Thus, we hypothesized that circLPAR3 might also perform oncogenic effects through the ceRNA mechanism. Through the prediction of Circinteractome database, numerous miRNAs were found to have potential binding site on circLPAR3. According to the previous researches, the expression levels of seven predicted miRNAs were detected in ESCC with increased circLPAR3. Among them, miR-873-5p was found to be decreased after circLPAR3 up-regulation in KYSE150 and TE-1 cells (Figure 4(a) and (b)). Thus, we speculated that miR-873-5p might be a target of circLPAR3. Subsequently, in order to confirm this hypothesis, we mutated these response elements between miR-873-5p and circLPAR3 (Figure 4(c)) and cloned them into a luciferase reporter containing the 3′-UTR of circLPAR3. After the validation of the elevation efficiency of miR-873-5p mimic in HEK293T cells (Figure 4(d)), the dual-luciferase reporter assay was conducted, we proved that miR-873-5p mimic overtly reduced the luciferase activity of wild-type circLPAR3 luciferase reporter but not the mutated one in HEK293T cells (Figure 4(e)). Moreover, RIP assay suggested that circLPAR3 was significantly enriched in AGO2-containing beads samples compared to control IgG immunoprecipitates, suggesting the binding between circLPAR3 and miR-873-5p in an AGO2-dependent manner (Figure 4(f) and (g)). In addition, it was proved that miR-873-5p expression was elevated by circLPAR3 knockdown in KYSE150 and TE-1 cells (Figure 4(h)). In the meanwhile, qRT-PCR showed that miR-873-5p inhibitor significantly reduced the level of miR-873-5p in KYSE150 and TE-1 cells (Figure 4(i)), and then we discovered that miR-873-5p inhibition attenuated the increase of miR-873-5p in circLPAR3 down-regulated KYSE150 and TE-1 cells (Figure 4(j)). Next, miR-873-5p expression was found to be decreased in ESCC tissues compared with the normal tissues (Figure 4(k)), which was negatively correlated with circLPAR3 expression (Figure 4(l)). Similarly, a decreased miR-873-5p expression was also observed in ESCC cell lines (Figure 4(m)). Accordingly, circLPAR3 could function as a sponge for miR-873-5p and negatively regulated its expression in ESCC cells. MiR-873-5p is a target of circLPAR3. (a, b) The impacts of circLPAR3 on the expression levels of seven predicted miRNAs possessing the potential binding sites of circLPAR3. (c) The complementary binding sites of miR-873-5p on circLPAR3 predicted by Circinteractome database. (d) qRT-PCR analysis of miR-873-5p expression in HEK293T cells transfected with miR-873-5p mimic or miR-NC. (e) Dual-luciferase reporter assay for the luciferase activity of wild and mutated circLPAR3 reporter after miR-873-5p overexpression in HEK293 T cells. (f, g) RIP experiment was performed with Ago2 antibody and negative control IgG, followed by qRT-PCR to measure the levels of circLPAR3. (h) qRT-PCR analysis of miR-873-5p expression in KYSE150 and TE-1 cells transfected with si1-circLPAR3, si2-circLPAR3 or si-NC. (i) qRT-PCR analysis of miR-873-5p expression in KYSE150 and TE-1 cells transfected with anti-miR-873-5p or anti-miR-NC. (j) qRT-PCR analysis of miR-873-5p expression in KYSE150 and TE-1 cells transfected with si-NC, si1-circLPAR3, si1-circLPAR3 + anti-miR-NC, si1-circLPAR3 + anti-miR-873-5p. (k) qRT-PCR analysis of miR-873-5p expression in ESCC tissues and matched adjacent normal tissues. (l) MiR-873-5p expression was negatively correlated with circLPAR3 expression in ESCC tissues. (m) qRT-PCR analysis of miR-873-5p expression in ESCC cell lines (TE-1, ECA109, and KYSE150) and normal Het-1A cells. **p <0.01, ***p <0.001.
CircLPAR3 knockdown suppresses ESCC cell Warburg effect, growth, and metastasis via targeting miR-873-5p
In the following experiments, we explored the influence of circLPAR3/miR-873-5p axis on ESCC cell tumorigenesis. It was proved that miR-873-5p inhibition led to the elevation of glucose uptake and lactate production (Figure 5(a) and (b)), enhancement of cell proliferation (Figure 5(c)–(e)), suppression of cell apoptosis (Figure 5(f)), promotion of cell migration and invasion abilities (Figure 5(g) and (h)) in circLPAR3-decreased KYSE150 and TE-1 cells. Furthermore, Western blot analysis showed that the decreases of PCNA, vimentin, and Slug protein levels as well as increase of cleaved caspase 3 protein level in KYSE150 and TE-1 cells caused by circLPAR3 silencing were rescued by miR-873-5p inhibition (Figure 5(i) and (j)). Altogether, circLPAR3 regulated ESCC progression via miR-873-5p. CircLPAR3 knockdown suppresses ESCC cell Warburg effect, growth and metastasis via targeting miR-873-5p. (a, b) Relative glucose uptake and lactate production in cells were detected by commercial kits. (c–e) Cell proliferation analysis by using MTT and EdU assays. (f) Flow cytometry for cell apoptosis analysis. (g) Wound healing assay for cell migration. (h) Transwell assay for cell invasion. (i, j) Western blot analysis for the protein levels of PCNA, cleaved caspase 3, vimentin, and Slug in cells. *p <0.05, **p <0.01, ***p <0.001.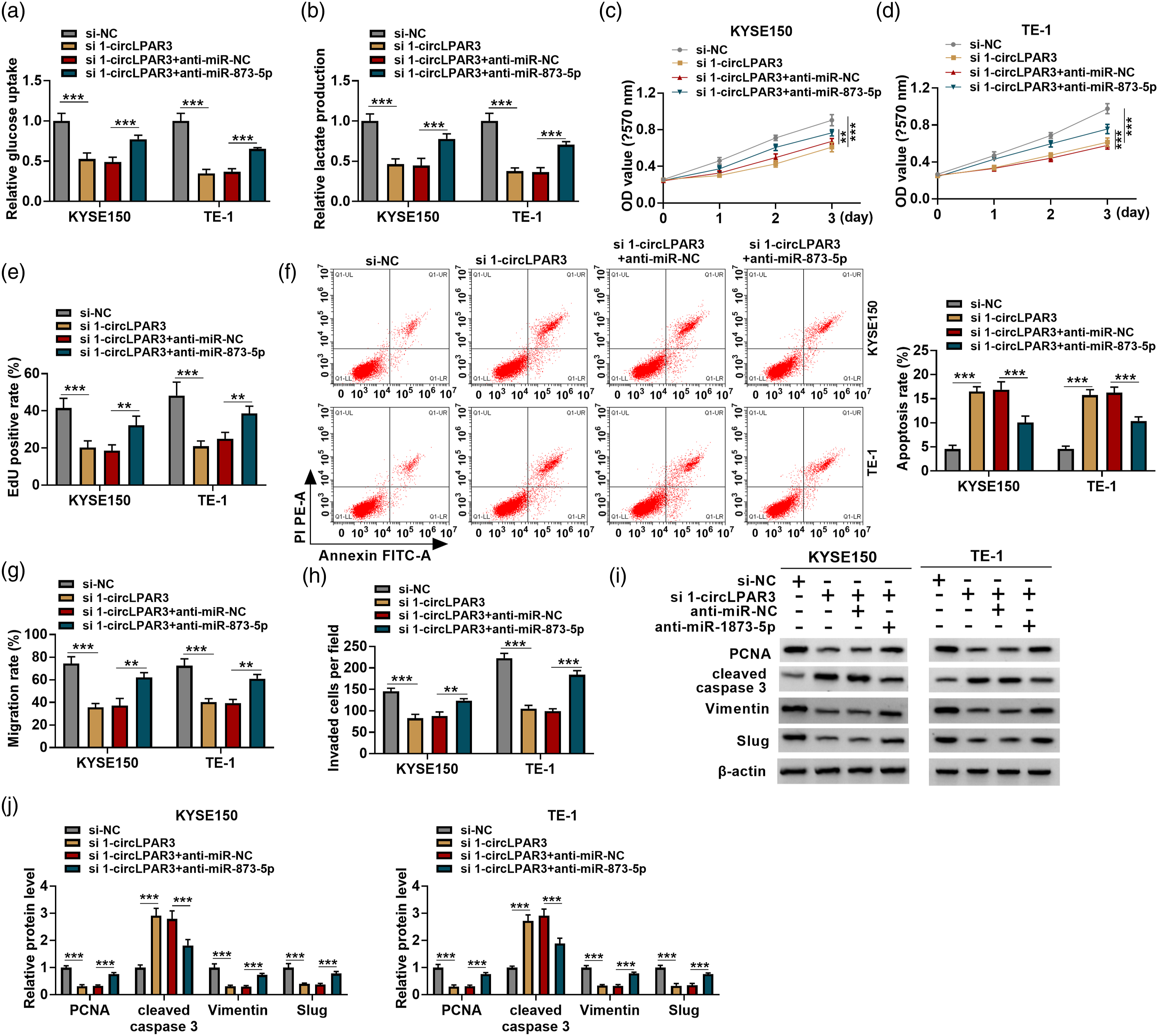
LDHA is a target of miR-873-5p, and circLPAR3 can regulate LDHA via sponging miR-873-5p
Based on the prediction of starbase database and previous researches, ten genes possessing putative binding sites on miR-873-5p were selected, among them, LDHA expression was discovered to be significantly reduced by miR-873-5p mimic in ESCC cells (Figure 6(a) and (b)). Thereafter, the response elements between miR-873-5p and LDHA were mutated to perform dual-luciferase reporter assay (Figure 6(c)), the results exhibited that miR-873-5p mimic reduced the luciferase activity of wild-type LDHA luciferase reporter, but failed to impact the luciferase activity of the mutated one in HEK293T cells (Figure 6(d)), implying the binding between miR-873-5p and LDHA. Subsequently, we confirmed the elevation efficiency of miR-873-5p mimic in KYSE150 and TE-1 cells (Figure 6(e)), then the results of Western blot showed that miR-873-5p mimic reduced LDHA expression, while miR-873-5p inhibitor elevated LDHA expression in KYSE150 and TE-1 cells (Figure 6(f)). Besides that, we also observed that circLPAR3 knockdown reduced the expression level of LDHA, which was rescued by miR-873-5p inhibition in KYSE150 and TE-1 cells (Figure 6(g)). After validating the elevation efficiency of LDHA vector (Figure 6(h)), it was demonstrated that LDHA overexpression reverted the decrease of LDHA expression induced by miR-873-5p mimic in KYSE150 and TE-1 cells (Figure 6(i)). Afterwards, the results of IHC staining and Western blot showed an increased protein level of LDHA in ESCC tissues (Figure 6(j)–(l)). Moreover, qRT-PCR suggested that LDHA mRNA was also increased in ESCC tissues (Figure 6(m)), and was negatively correlated with miR-873-5p expression (Figure 6(n)), and positively correlated with circLPAR3 expression (Figure 6(o)). Besides, LDHA expression was also elevated in ESCC cells (Figure 6(p)). In all, these results verified that miR-873-5p targeted LDHA and suppressed its expression, and circLPAR3/miR-873-5p/LDHA constituted a feedback loop in ESCC cells. LDHA is a target of miR-873-5p, and circLPAR3 can regulate LDHA via sponging miR-873-5p. (a, b) The impacts of miR-873-5p on the expression levels of ten predicted miRNAs possessing the potential binding sites of miR-873-5p. (c) The complementary binding sites of miR-873-5p on LDHA predicted by starbase database. (d) Dual-luciferase reporter assay for the luciferase activity of wild and mutated LDHA reporter after miR-873-5p overexpression in HEK293T cells. (e) qRT-PCR analysis of miR-873-5p expression in KYSE150 and TE-1 cells transfected with miR-873-5p mimic or miR-NC. (f) Western blot analysis for the protein level of LDHA in KYSE150 and TE-1 cells transfected with miR-873-5p mimic, inhibitor and the controls. (g) Western blot analysis for the levels of LDHA in KYSE150 and TE-1 cells transfected with si-NC, si1-circLPAR3, si1-circLPAR3 + anti-miR-NC, si1-circLPAR3 + anti-miR-873-5p. (h) The elevation efficiency of LDHA or pcDNA was validated using Western blot. (i) Western blot analysis for the levels of LDHA in KYSE150 and TE-1 cells transfected with miR-NC, miR-873-5p, miR-873-5p + pcDNA, or miR-873-5p + LDHA. (j) IHC staining for LDHA protein in ESCC tissues and adjacent normal tissues. (k–m) Western blot and qRT-PCR analysis for the protein and mRNA level of LDHA in ESCC tissues and adjacent normal tissues. (n) LDHA mRNA expression was negatively correlated with miR-873-5p expression. (o) LDHA mRNA expression was positively correlated with circLPAR3 expression. (p) Western blot analysis for LDHA protein level in ESCC cell lines (TE-1, ECA109, and KYSE150) and normal Het-1A cells. **p <0.01, ***p <0.001.
MiR-873-5p suppresses ESCC cell Warburg effect, growth and metastasis via targeting LDHA
Subsequently, the effects of miR-873-5p/LDHA axis on ESCC progression were explored. It was proved that miR-873-5p re-expression suppressed Warburg effect, evidenced by the decrease of glucose uptake and lactate production in KYSE150 and TE-1 cells, while LDHA up-regulation attenuated this condition (Figure 7(a) and (b)). Besides that, miR-873-5p re-expression caused the inhibition of cell proliferation (Figure 7(c)–(e)), enhancement of cell apoptosis (Figure 7(f)), as well as reduction of cell migration and invasion rates (Figure 7(g) and (h)) in KYSE150 and TE-1 cells, which were reversed by LDHA up-regulation (Figure 7(f)–(h)). Moreover, miR-873-5p mimic decreased the protein levels of PCNA, vimentin, and Slug, and increased the protein level of cleaved caspase 3 in KYSE150 and TE-1 cells, while co-transfection with LDHA abolished these changes (Figure 7(i) and (j)). Collectively, miR-873-5p/LDHA axis was responsible for ESCC progression. MiR-873-5p suppresses ESCC cell Warburg effect, growth and metastasis via targeting LDHA. (a–j) KYSE150 and TE-1 cells were transfected with miR-NC, miR-873-5p, miR-873-5p + pcDNA, or miR-873-5p + LDHA. (a, b) Relative glucose uptake and lactate production in cells were detected by commercial kits. (c–e) Cell proliferation analysis by using MTT and EdU assays. (f) Flow cytometry for cell apoptosis analysis. (g) Wound healing assay for cell migration. (h) Transwell assay for cell invasion. (i, j) Western blot analysis for the protein levels of PCNA, cleaved caspase 3, vimentin, and Slug in cells. *p <0.05, **p <0.01, ***p <0.001.
CircLPAR3 silencing impedes ESCC growth in vivo
We then also evaluated the effect of circLPAR3 on the growth of the xenograft tumors in nude mice. Silencing of circLPAR3 inhibited the growth of xenograft tumors in nude mice, manifested by the smaller tumor volume and weight in circLPAR3-decreased group (Figure 8(a)–(c)). Furthermore, IHC staining in xenograft tumors showed the protein of Ki-67, cleaved caspase 3, and LDHA was attenuated after circLPAR3 knockdown (Figure 8(d)–(f)). Besides that, molecular analysis suggested that the levels of circLPAR3 and LDHA were decreased while level of miR-873-5p was increased in xenograft tumors of circLPAR3-dereased group (Figure 8(g)–(i)). Moreover, we also showed that the protein levels of glucose transporter GLUT1 and the key glycolytic enzymes HK2 were decreased in xenograft tumors of sh-circLPAR3 group compared with the sh-NC group (Figure 8(j)). Collectively, circLPAR3 knockdown impeded ESCC growth in vivo. CircLPAR3 knockdown impedes the growth of Xenograft tumors in vivo. (a) The tumor volumes were examined every five days, and the growth curves were drawn. (b) The weight of xenograft tumors was analyzed, and the representative images in each group were showed (left panel). (d–f) IHC staining for Ki-67, leaved caspase 3, and LDHA in xenograft tumors. (g–i) qRT-PCR and Western blot analysis of circLPAR3, miR-873-5p and LDHA levels in xenograft tumors in each group. (j) Western blot analysis of GLUT1 and HK2 protein levels in xenograft tumors of each group. ***p <0.001.
Discussion
Previous studies have proposed that the progression of ESCC is a complicated process, involving the inactivation of anti-oncogenes and activation of oncogenes, and is often accompanied by epigenetic modifications.27–29 CircRNAs are important component of epigenetic regulation, which have been identified that can modulate the series of cancer phenotype, including proliferation, metastasis and Warburg effect.30,31 Warburg effect is one of hallmarks of cancers, increasing documents revealed that glycolysis is characterized by a vital metabolic event involved in promoting cancer cell survival, metastasis, stemness, and long-term maintenance. 32 In this study, we observed that circLPAR3 was increased in ESCC. Functionally, circLPAR3 silencing suppressed ESCC cell proliferation, invasion, metastasis, and induced apoptosis, accompanied with the decrease of PCNA, Vimentin, Slug and increase of cleaved caspase3 in cells in vitro. Besides that, we also observed the reductions of lactate generation and glucose uptake in ESCC cells after circLPAR3 silencing, suggesting the inhibition of Warburg effect. Moreover, subcutaneous xenograft model showed that circLPAR3 knockdown impeded tumor growth in vivo.
It has been revealed that circRNAs act as miRNA sponges to repress the level and function of miRNA.25,26 Besides that, circRNA-miRNA-mRNA regulatory mechanism has been identified to play vital roles in the development of ESCC. 33 Therefore, the miRNA-mRNA axis underlying circLPAR3 was investigated. We confirmed that circLPAR3 directly bound to miR-873-5p, which targeted LDHA. A previous study exhibited that miR-873 was decreased in esophageal cancer, and re-expression of miR-873 could repress cancer cell survival, invasion and migration. 34 Here, we hypothesized that miR-873-5p might also show tumor suppressing effects on ESCC. As expected, a decreased miR-873-5p expression was found in ESCC, importantly, restoration of miR-873-5p restrained ESCC cell Warburg effect, growth and metastasis. Furthermore, inhibition of miR-873-5p abolished the inhibitory action of circLPAR3 silencing on ESCC tumorigenesis. LDHA is a glycolytic enzyme that promotes glycolytic process by catalyzing the conversion of pyruvate into lactate, the up-regulation of LDHA is the basics of Warburg effect. 35 In human cancers, LDHA shows an aberrantly high expression, and is closely related to the malignant development of malignancies. 36 In ESCC, LDHA was demonstrated to promote cell growth, migration and tumorigenesis. 37 In the current work, we discovered that LDHA overexpression attenuated the anticancer functions of miR-873-5p in ESCC cells. Moreover, circLPAR3 could regulate LDHA through sponging miR-873-5p, indicating the circLPAR3/miR-873-5p/LDHA feedback loop in ESCC cells.
Currently, the potential value of RNA as a target for new small molecule therapeutics has have attracted great research interest for their diversity of impacts on the progression and recurrence of cancers. 38 The posttranscriptional gene regulatory mechanism known as RNA interference (RNAi) has been identified and more than 50 RNA-based drugs are currently under clinical testing. 39 This work implied that circLPAR3 specific shRNAs or siRNAs may be promising molecules used for ESCC clinical therapy by affecting the malignant biological behavior, indicating the pharmacological and toxicological applications in clinic.
In conclusion, we for the first time demonstrated that circLPAR3 silencing suppressed ESCC cell Warburg effect, growth and metastasis by targeting miR-873-5p/LDHA axis, revealing a potential therapeutic target for ESCC.
Supplemental Material
Supplemental Material - CircLPAR3 knockdown suppresses esophageal squamous cell carcinoma cell oncogenic phenotypes and Warburg effect through miR-873-5p/LDHA axis
Supplemental Material for CircLPAR3 knockdown suppresses esophageal squamous cell carcinoma cell oncogenic phenotypes and Warburg effect through miR-873-5p/LDHA axis by Yao Cheng, Zhenchuan Ma, Shiyuan Liu, Xiaoping Yang and Shaomin Li in Human & Experimental Toxicology
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the This work was supported by National Natural Scientific Foundation of Shaanxi (2020JQ-504; 2020JQ540).
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
