Abstract
Ergot alkaloids, a class of mycotoxins, induce vasoconstriction when consumed by animals and humans. Pregnant ewes (n = 16; 81.2 kg ± 7.7) were assigned fed endophyte-infected tall fescue seed (
Introduction
Mycotoxin contamination of our food and feed supply are predicted to increase with global warming and climate change. 1 Ergot alkaloids, a class of mycotoxins, are found in endophyte-infected (E+) tall fescue that is the primary cool-season, perennial grass grazed by ruminant animals. Ergovaline, the most prevalent ergot alkaloid in tall fescue seed, 2 causes potent and widespread vasoconstriction 3 and has been shown to constrict the umbilical and uterine arteries.2,4,5 The biological activity of ergot alkaloids originates from their similarity to biogenic amines such as norepinephrine, serotonin, and dopamine and they exert their effect by working as an agonist on three primary receptors including the D2 dopaminergic, α-2 adrenergic, and 5HT2a serotonergic.6–8 When consumed during gestation, ergot alkaloids induce placental insufficiency and intrauterine growth restriction (IUGR) most likely due to reduced uteroplacental blood flow.9–11 These effects appear most detrimental during late gestation when placental remodeling and exponential fetal growth occurs.5,12,13
In goats, ruminal infusion of E+ tall fescue seed extract induced vasoconstriction, as measured by luminal area of carotid artery using doppler ultrasound, by 2 d post dosing. 14 The carotid luminal area was reduced by 46% with E+ extract infusion compared to baseline values.
Vasoconstriction can lead to hypoxia and reduced nutrient supply to the fetus. Hypoxia can alter fetal development but depends on duration of hypoxia event and stage of the developing fetus. 15 In pregnant sheep, chronic hypoxia activates placental stress response and restricts fetal growth, which results in fetal brain-sparing. 16 Research has shown that both hypoxia and oxidative stress are involved in intrauterine growth restriction (IUGR) in twin bearing ewes. 17 However, little is known about how the duration of mycotoxin exposure during late gestation alters placental remodeling, nutrient transporters and fetal development. The pregnant sheep is an established model for human pregnancy and provides information on how mycotoxin exposure during pregnancy could impact placental and fetal development. 18 It may be that the immediate insult of mycotoxin exposure to promote vasoconstriction is most detrimental to the developing placenta/fetus or that chronic exposure to mycotoxins results in reduced nutrient transport and growth retardation. Therefore, the objective of this study was to examine the effect of feeding endophyte-infected tall fescue seed containing ergot alkaloids to ewes at specific stages of late gestation (gd86 to gd110 or gd133) on placental structure, fetal development, and cotyledon mRNA transcriptome.
Materials and methods
Experimental design
All animal experimental procedures were reviewed and approved by the Clemson University Institutional Animal Care and Use Committee (AUP 2018-055). Mature ewes (n = 21; 81.2 kg ± 7.7) were randomly divided into 3 groups (n = 9/group) for estrous synchronization. Each week, one group of ewes was synchronized using an intravaginal controlled internal drug release (CIDR) insert (Eazi-Breed CIDR, Zoetis Animal Health) for 7 d. Upon CIDR removal, ewes were given prostaglandin F2α (12.5 mg i.m; Lutalyse; Pfizer, New York, NY) and turned in with a purebred Texel ram (Geneseek, Myostatin Muscled) fitted with a marking harness. Ewes were examined for pregnancy and lamb number using a transabdominal ultrasound between d 45 to 60 of gestation (BCF Easi-Scan Portable Ultrasound Machine, BCF Technology Ltd, Rochester, MN). Ewes estimated to be carrying twins (n = 16) were randomly assigned to endophyte-infected tall fescue seed (E+; 4.14 μg ergovaline + ergovalinine/g seed) or a control total mixed ration (CON; 0 μg ergovaline + ergovalinine). Ewes were individually fed ergot alkaloid treatments (E+ or CON) during specific stages of gestation (gd86 to gd110 or gd86 to gd133) in a 2 × 2 factorial. Fescue seed contains the highest concentrations of ergovaline/ergovalinine compared to the leaf blades or stems with sheaths. 19 Fescue seed used in this study was analyzed for ergovaline and ergovalinine content. 20 All seed cultivars tested negative for ergotamine. Seed was fed to ewes at a total dose level of ergovaline/ergovalinine (E+ = 1.77 mg hd−1 d−1) per day starting on gd86. The dose level of ergovaline/ergovalinine fed in this study was based on previous research in which E+ seed was fed at a level of 0.8 mg/kg dry matter to induce vasocontriction. 20 To achieve this dose level, we individually fed ewes the same amount of dry matter intake each day containing the E+ seed at a quantity that equates to 0.8 mg/kg dry matter. 12 Nutrient analyses were performed on seed and TMR samples and rations were formulated to meet NRC requirements 21 for late gestating ewes. Two ewes had to be removed from the study (1 CON and 1 E+, gd133) due to reduced intake and body weight gain that required veterinary attention.
Prolactin
Blood samples were collected from ewes on the terminal day of gestation according to treatment. Whole blood samples were collected by jugular venipuncture into serum collection tubes. Whole blood was allowed to clot for 30 min at room temperature, stored on i.e. and then serum was separated by centrifugation at 537 x g for 20 min at 4°C. Serum was stored at −20°C for future analysis. Serum prolactin concentrations were measured using a Prolactin ELISA kit (K040-H1; Arbor Assays, Ann Arbor, MI) with an average intra-assay variance was 3.16% and inter-assay variance was 5.92%. Samples were spiked with prolactin standards and the recovery was 92.4%.
Harvest and placentome collection
Ewes underwent a terminal necropsy at gd110 (n = 4/treatment) or gd133 (n = 3/treatment). The uterus was exposed through a mid-ventral incision and the full uterus was tied off, excised proximal to the cervix and weighed. A small incision was made in the uterine wall for the collection of fluid and removal of each fetus. The fetuses were removed from the sac and the umbilical cord was clamped and cut. Each fetus was towel dried, weighed, and brain weight obtained. After the removal of the fetuses, any remaining uterine fluid was drained. Placentomes were removed from the uterine wall and classified by type according to Vatnick et al 22 . . For RNA sequencing, type B placentomes were removed, cotyledon tissue separated from caruncle, and cotyledon was immediately frozen in liquid nitrogen. A sample of each available placentome type was collected for histology. The placentome was halved and stored in formalin until paraffin embedding. After remaining placentomes were typed and counted, the height, maximum diameter, and minimum diameter measurements were taken on each placentome. Then the cotyledon and caruncle from each placentome were carefully separated and weighed individually. Once placentomes were removed from the uterine wall, all membranes were removed, drained of any remaining fluid, and weighed collectively. The remaining empty uterus was cleaned and weighed.
Histology
Placentome samples were paraffin embedded using a Tissue-Tek VIP 6 AI Vacuum Infiltration Processor (Sakura Finetek USA, Inc, Torrance, CA). Samples were sectioned at 8 μm thickness using a microtome and stained immediately following sectioning. The H&E staining procedure was utilized in this study to evaluate placental histomorphology. For consistency, type B placentomes were imaged in brightfield using a Leica Microsystems (Buffalo Grove, Illinois) M205FCA stereomicroscope coupled with a Leica DMC4500 color camera. Images were collected using a 1.0X (Numerical aperture= 0.05) objective with digital zoom, for a final magnification of ×20. Using the Navigator feature in the Leica LAS-X software, images of the entire placentome were collected and stitched together; a shading correction was applied to achieve an even field of illumination. Measurements, including height, width, and area, of regions of tissue representative of Wharton’s jelly according to Stegeman 23 were performed using the area tracing tool in the Leica LAS-X software. To demonstrate the appearance of the Wharton’s jelly regions in detail, additional high magnification images of a subset of samples were collected using the same microscope at ×200 magnification with a ×5.0 (numerical aperture = 0.5) objective.
RNA sequencing
For consistency, type B placentomes were selected for RNA sequencing analysis. Total cellular RNA was isolated from cotyledon samples using Trizol reagent and purified using a PureLink Mini RNA kit (Invitrogen, Thermo Fisher Scientific, Waltham, MA) according to manufacturer instructions. RNA quantity was estimated using a NanoDrop 1000 Spectrophotometer (ThermoFisher). RNA quality was assessed using an Agilent 2100 Bioanalyzer (Agilent, Santa Clara, CA). All samples utilized for RNAseq has a RIN value of 8 or greater. Samples were submitted to the Clemson University Genomics and Bioinformatics Facility for RNAseq analysis. Total RNA was quantified using the Broad Range Assay in the Qubit (ThermoFisher Scientific, Waltham, MA). Library preparation was conducted using the NEB Next Ultra II Library Prep Kit for Illumina according to manufacturer instructions. Samples were sequenced to an average sequence depth of 51M reads using an Illumina NextSeq 550 sequencing platform at 150 bp paired-end reads. Raw sequencing data was transferred to the Clemson University Palmetto Cluster for analysis. Downsampling was conducted to ensure uniform coverage using seqtk v1.3-r106 (https://github.com/lh3/seqtk). FastQC v.0.11.8 was utilized to check sample quality (http://www.bioinformatics.babraham.ac.uk/projects/fastqc2010). All low-quality bases and adaptor sequences were trimmed using TrimGalore (v0.38). 24 After trimming, 99.6% of the remaining reads were aligned to the ovine reference genome (ftp://ftp.ensembl.org/pub/release-96/fasta/ovis_aries/) using GSNAP v2018-07-04. 25 Subread’s feature Counts v2.0.0 was used to count uniquely mapped reads that aligned to known genes in the ovine genome. 26 Gene counts were analyzed using edgeR v3.22.5. 27 Genes were considered differentially expressed if their false discovery rate (FDR) was <0.05 and their |log2FC| was greater than or equal to 1.0.
Gene ontology (GO) and KEGG pathway analysis
Differentially expressed genes were subjected to Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analysis using clusterProfiler. 28
RT-qPCR validation
Gene expression analysis was conducted using quantitative real-time RT-PCR methods 29 to validate RNAseq results. Aliquots from the same RNA samples submitted for RNAseq were converted to cDNA using qScript cDNA SuperMix (Quanta Bio, Beverly, MA) according to manufacturer instructions. Real-time PCR was performed using a QuantStudio3 (Applied Biosystems, Thermo Fisher) and PowerUp (Applied Biosystems, Thermo Fisher) SYBR green according to the manufacturer’s directions. Genes for qPCR were selected based on having a published sequence in NCBI (https://www.ncbi.nlm.nih.gov/nucleotide/) and being differentially expressed by day and treatment. Primers for STARD (forward: GGGAACCTGTACAAAGGAGAAG; reverse: CATCCCACTGAGCTCGTAAAG) and GPR151 (forward: GATCTCAGCCAGGTGTTGATAG; reverse: GGCAACCTGCTCCAGTATT) were designed using PrimerQuest (Integrated DNA Technologies, IDT, Coralville, IA) and used to validate RNA-Seq results. The geometric mean of three housekeeping genes (glyceraldehyde 3-phosphate dehydrogenase, (GAPDH; forward: GGAGAAACCTGCCAAGTATGA; reverse: GCATCGAAGGTAGAAGAGTGAG), eukaryotic translation initiation factor 3 subunit K (EIF3K; forward: CTGACAGACAGCCAGCTAAA; reverse: CACGATGTTCTTGGGCTTAATG), and ubiquitously expressed prefoldin like chaperone (UXT; forward: CCACGGTTCTTACTAACACTTCT; reverse: TTGGCCAGCTCATATAC) was used for normalization. Normalized CT values (ΔCT = CT, gene − CT, HK) were calculated for each sample. 30 Relative gene expression was calculated using the ΔΔCT method and analyzed by ANOVA using GraphPad Prism (San Diego, CA).
Statistical analysis
Data were analyzed using a mixed procedure of SAS version 9.4 (SAS Institute Inc, Cary, NC) with fescue treatment (E+ vs CON), stage of gestation (gd110 or gd133) and two-way interaction in the model. Ewe was the experimental unit in this study. Statistical significance was declared at p < .05.
Results
Changes in ewe body weight, dry matter intake, and serum prolactin for ewes fed endophyte-infected (E+) tall fescue seed versus control (CON) at different stages of late gestation (gd110, gd133).

abMeans in the same row with different superscripts differ (p < .05) by gestational day.
cdMeans in the same row with different superscripts differ (p < .05) by fescue treatment.
Uterine components on a weight and percentage basis for ewes fed endophyte-infected (E+) tall fescue seed versus control (CON) at different stages of late gestation (gd110 or gd133).
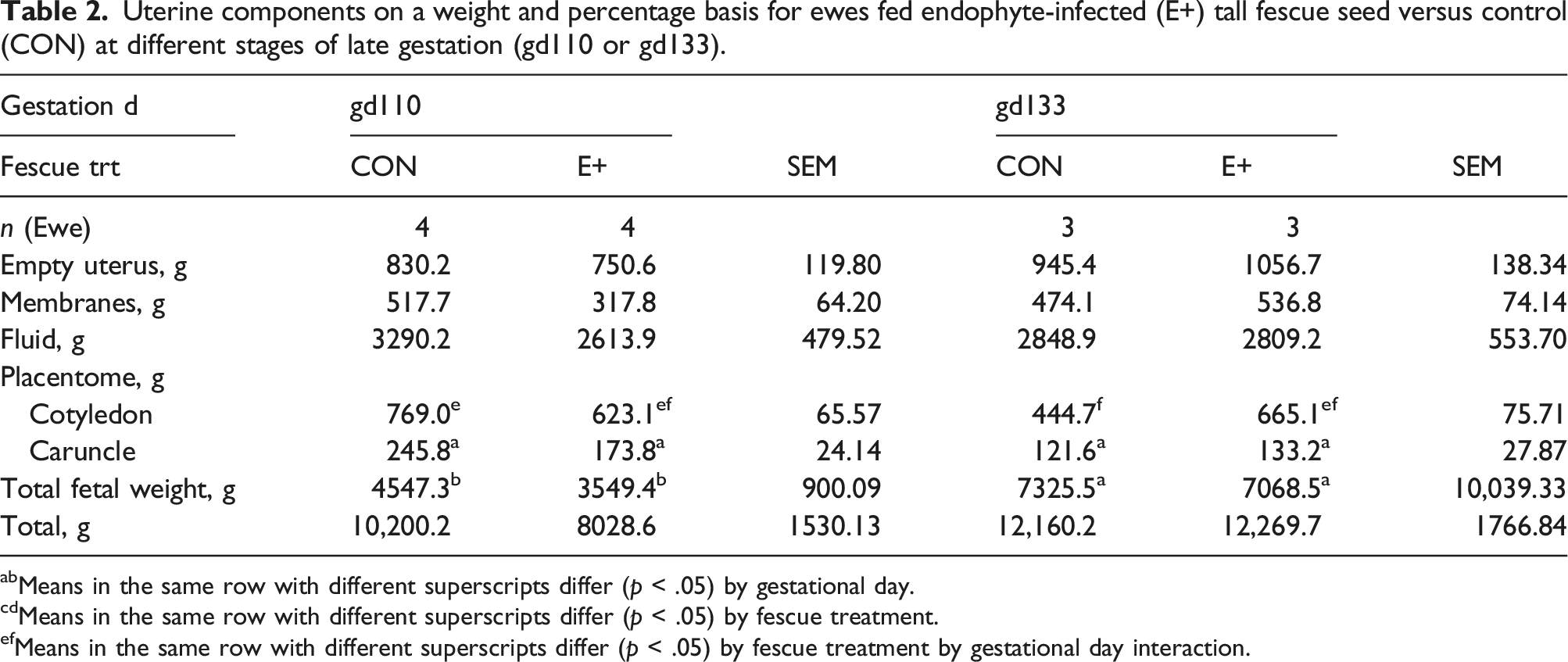
abMeans in the same row with different superscripts differ (p < .05) by gestational day.
cdMeans in the same row with different superscripts differ (p < .05) by fescue treatment.
efMeans in the same row with different superscripts differ (p < .05) by fescue treatment by gestational day interaction.
Placentome weight, number, and area by type for ewes fed endophyte-infected (E+) tall fescue seed versus control (CON) at different stages of late gestation (gd110, gd133).

abMeans in the same row with different superscripts differ (p < .05) by fescue treatment by gestational day interaction.
Changes in structure of the placentome were evaluated by histology and villi measurement were analyzed (Figure 1). Villi width was lower (p < .0001) at gd133 than gd110 but did not differ by ergot alkaloid treatment. The interaction between ergot alkaloid treatment and stage of gestation was significant (p < .05) for villi height and area of the placentome. Villi height and area were both greater (p < .05) for E+ than CON at gd110 but did not differ at gd133. Placental efficiency is calculated by relating the weight of the fetus to the cotyledon. Fetal to cotyledon weight was greater (p < .05) for CON than E+ at gd133 but did not differ between ergot alkaloid treatments at gd110 (Figure 2). Exposure to mycotoxins (E+) increased (p < 0.5) fetal brain to body weight ratio at both stages of gestation, which demonstrates asymmetrical growth and intrauterine growth restriction (IUGR) in lambs exposed to ergot alkaloids in utero. Placentome structure and fetal villi measurements for ewes fed endophyte-infected (E+) tall fescue seed compared to control diet (CON) at different stages of gestation (gd110 or gd133). The fetal to cotyledon weight (a) and brain to fetal body weight (b) ratios for ewes fed endophyte-infected (E+) tall fescue seed or control diet (CON).

Differentially expressed genes (FDR <0.05) in cotyledon tissue of ewes fed endophyte-infected (E+) tall fescue seed versus control (CON) by stage of gestation (gd86 to gd110 or gd86 to gd133).


Gene expression by qPCR of select genes for validation of sequencing project. (a). qPCR Gene Expression (b). StARD5 Pathway for Ovis aries (c). GPR151 Pathway for Ovis aries.
Differentially expressed genes (FDR <0.05; logFC >1.5) in cotyledon tissue of ewes from gd110 to gd133. Genes are displayed by level of significance.

Differentially expressed genes (FDR <0.05; logFC < −1.5) in cotyledon tissue of ewes from gd110 to gd133. Genes are displayed by level of significance.

Gene ontology enrichment of differentially expressed genes in cotyledon tissue of ewes by stage of gestation (gd110 to gd133).

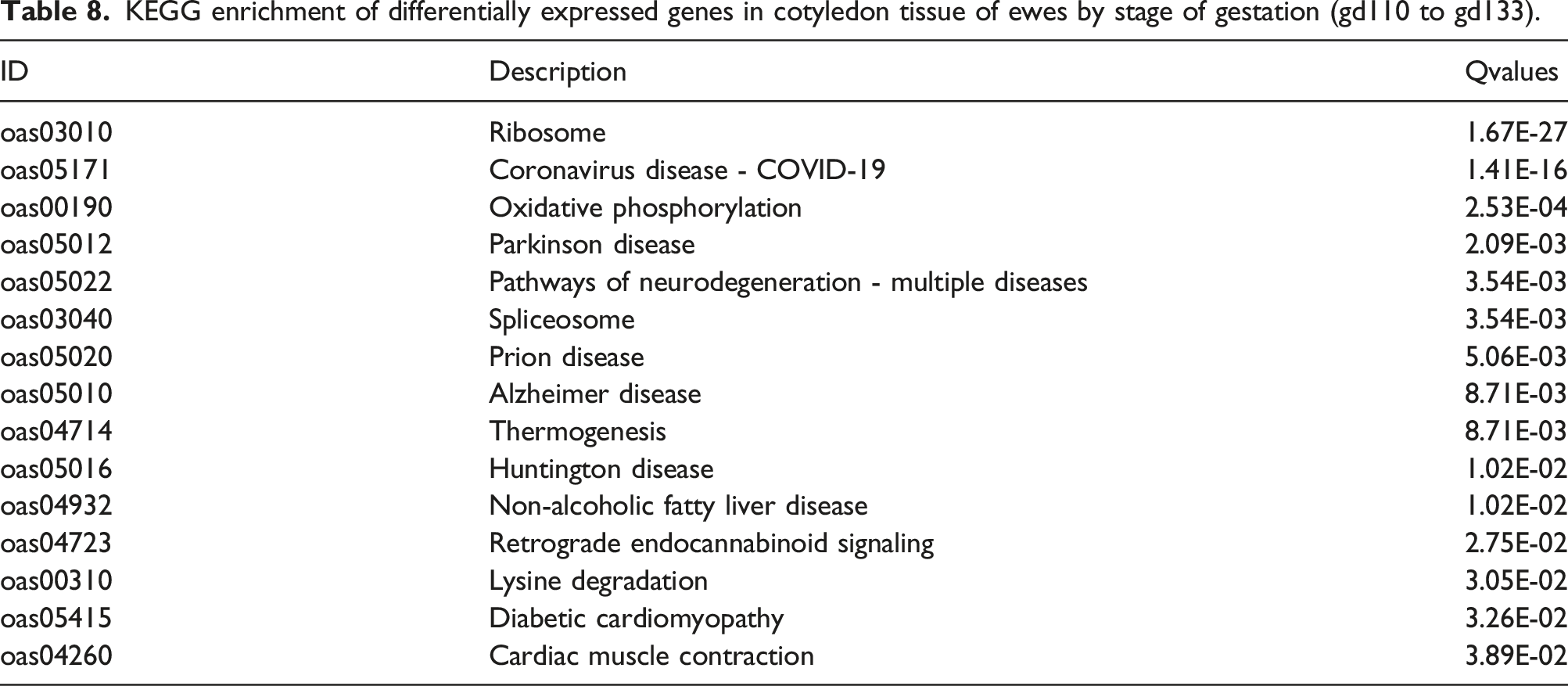
KEGG enrichment of differentially expressed genes in cotyledon tissue of ewes by stage of gestation (gd110 to gd133).

Discussion
Ergot alkaloids, a class of mycotoxins, are present in endophyte-infected tall fescue (E+) and consumption by ruminants (cattle, sheep, goats) induces vasoconstriction which reduces blood flow. 7 Limited research has examined how these mycotoxins alter placental and fetal development during gestation. Previous research found that exposure to ergot alkaloids during late gestation (gd86 to gd133) had the greatest impact on placental remodeling,13,29,31 fetal development 10 and subsequent postnatal growth 32 ; however, how these changes occur within this stage of gestation have not been examined. In this study, ergot alkaloid intake for E+ ewes was maintained by feeding tall fescue seed with known levels throughout late gestation. Feed intake was similar between treatments to reduce confounding of intake reduction by E+ exposure and its relationship to fetal growth. The diets, E+ and CON, were formulated to be isonitrogenous and isocaloric, and fed at amount to meet the requirements for pregnant ewes with twins during late gestation. Ergot alkaloid consumption (E+) reduced serum prolactin concentrations at gd110 by 29.6-fold and gd133 by 71.8-fold compared to controls. Reductions in serum prolactin levels are commonly reported in sheep grazing endophyte-infected tall fescue pastures33–35 or fed E+ tall fescue seed containing ergot alkaloids.11,31,36 The reductions in prolactin concentrations are caused by the structural similarity of the ergoline ring of ergot alkaloids to dopamine, which allows them to interact with D2 dopamine receptors. 7
During late gestation, placenta growth plateaus around gd80 37 to gd90 38 and then remodeling of the placenta occurs. 39 The ovine placenta experiences a period of rapid growth in which the fetal villi invade the maternal stroma of the caruncle .23,39,40 The fetal villi consist of a trophoblast layer that contains large regions ofamorphous connective tissue like Wharton’s jelly.23,39 As maternal vascular beds develop, the amorphous connective tissue disappears, and only thin strands remain. The decrease in weight of the cotyledon over time is related to this loss of connective tissue which contains a high amount of water. These changes were evident in control ewes were cotyledon and total placentome weights decreased at gd133 compared to gd110; however, cotyledon weight did not change by stage of gestation in E+ fed ewes. Placentome area was also lower at gd133 in control compared to gd110 but did not change in E+. Ewes fed E+ fescue seed also had greater placentome villi height and area than controls at gd110 or gd133 regardless of fescue treatment. These results suggest that E+ exposure slows the process of placental remodeling wherein maternal capillary beds replace large regions of amorphous connective tissue to facilitate nutrient exchange to the fetus. Britt et al. 13 identified greater amorphous connective tissue within the fetal villi from mycotoxin exposure that indicates underdeveloped maternal vascular beds that may hinder fetal growth.
The progression of placentome morphology has been extensively investigated but continues to draw mixed results. It is hypothesized that placentome morphology may shift towards more advanced type C and D placentomes during late gestation when the fetal demand for nutrients is at its highest; especially in cases of insufficient uterine environments such as undernutrition and hypoxia.41–44 Osgerby et al. 42 speculated that flattening of the placentome, which occurs in advancement, may be associated with attempts to increase nutrient delivery to the fetus. However, morphology is not always altered in cases of adverse intrauterine conditions. 45 In this study, ewes fed E+ fescue seed had a shift in placentome area to type C instead of type B at gd133. Placental efficiency, as defined by the ratio of fetal weight to placental weight, is indicative of how well the placenta adapts to meet the high nutrient demand for fetal growth. 46 In this study, ewes fed E+ seed had 30% less fetus per gram of cotyledonary tissue compared to controls at gd133. Prolonged exposure to E+ fescue seed appears to lower placental efficiency as indicated by a lower fetal:cotyledon weight ratio at gd133.
During late gestation, exponential fetal growth begins with over 80% of fetal weight obtained during this time.38,47 Previous research found that exposure to E+ fescue during late gestation reduced fetal BW at gd133 by 15% 11 and birth weight by 10.2%. 31 Brain to fetal body weight was increased with exposure to mycotoxins in this study, which is an indicator of asymmetrical growth and IUGR.15–17 This agrees with our earlier work that showed asymmetrical growth in lambs at gd133 10 after exposure to mycotoxins during late gestation that altered growth of the offspring and delayed puberty in females. 32 The asymmetrical growth was already established at gd110 and remained at gd133. These results suggest that ergot alkaloid exposure during this stage of gestation (gd86 to gd110) or that the immediate insult of ergot alkaloids (first 25 d on E+ treatment) is having the greatest impact on fetal growth and IUGR. Others have reported long-term consequences of offspring that were born to nutrient restricted dams.48–50
Exposure to ergot alkaloids differentially expressed 22 genes in cotelydon tissue at gd133 and 1 gene at gd133. Most the genes identified have an unknown role in placental and fetal development in the sheep but have been identified to be associated with hypertension and placental insufficiency in other species. Jerber et al. 51 discusses that CCDC151 is involved in cilia function but also may have additional roles in non-motile function including cell polarity and mitotic spindle orientation for vertebrates. Regulator of G protein signaling 5 (RGS5) gene regulates hemodynamic adaptation to pregnancy and loss of RGS5 causes gestational hypertension in the mouse. 52 Goyal et al. 53 reported that RGS5 was down regulated in a rat model of placental insufficiency. Other genes (SERPINA14 and STARD5) appear to be related to hormone and lipid metabolism. SERPINA14 plays a role as an immune regulator and mediates progesterone’s immunosuppressive response in the sheep. 54 The STARD5 gene is known to bind to cholesterol and 25-hydroxycholesterol to help with interchange of lipids between membranes. 55 In mice, STARD5 responds to stress and adjusts intracellular cholesterol homeostasis. 56 Fescue toxicosis has been shown to reduce plasma cholesterol levels in grazing cattle. 57 Edwards et al. 58 reported that nutrient restriction from gd35 to gd125 up-regulated GPR151 in placentomes from ewes that produced non-IUGR fetuses compared to IUGR fetuses. No differences in gene ontology or KEGG pathway analysis were noted for E+ compared to CON at gd110 or gd133. Previous research identified four genes (LECT2, SLC22A9, APOC3, MBL2) that were differentially expressed in the cotyledon from ewes fed E+ fescue from gd36 to gd133 and clusters associated with nutrient transport and cellular metabolism. 29 These differences in response to fescue may be related to the duration of exposure to ergot alkaloids with this short-term study having less impact on mRNA expression compared to long-term exposure.
From gd110 to gd133, 89 genes were differentially expressed (FDR <0.05; log2FoldChange < −1.5 or >1.5) in the cotyledon tissues. Many of these genes have associated roles with immune response and insulin action. CLEC3B was identified in bovine where it was up-regulated in pregnant versus non-pregnant heifers and appears to be involved in immune response to IFN-τ. 59 SFTPD has been shown to be an integral component of female reproductive tract and appears involved in immune response. 60 Others 42 have also observed decreased expression of IGFBP5 mRNA in the placentome of sheep from gd90 to 135. Jia et al. 61 proposed that IGFBP5 may be used as a marker for preeclampsia in humans because of its high expression in the preeclamptic placenta. RGS5 is reported to play a role in cardiovascular function during pregnancy 62 and serves as a marker for gestational hypertension in humans 63
Gene ontology analysis discovered enrichment of molecular function, cellular component, and biological process during gestation from gd110 to 133. KEGG pathway enrichment was found for pathways associated with ribosome, oxidative phosphorylation, splicesome, thermogenesis, and many disease states. Upon further investigation with KEGG pathways analysis using KEGG Mapper, oxidative phosphorylation is the primary driver between the overlap with disease states (Huntington’s disease, Parkinson’s disease, Alzheimer’s disease, and NAFLD) as well as several cancer pathways. In all mammals, placental energy demand is met primarily by oxidative phosphorylation. 64 This shows a comprehensive up-regulation of genes associated with ribosomal function. Research suggests the rate of rRNA transcription is tightly linked to ribosome biogenesis, which regulates cell growth and proliferation. 65
Exposure to E+ fescue at specific stages during late gestation show reduced serum prolactin concentration and slowed placental remodeling which resulted in placental inefficiency and asymmetrical growth of the fetus that was evident by gd110 and continued to gd133. Transcriptomic changes in the cotyledon tissue of E+ were identified but most of the genes identified have an unknown role and further research is needed to determine their involvement in placental remodeling and mycotoxin exposure. These results suggest that the response to mycotoxin exposure in the pregnant sheep model has an immediate impact on placental remodeling and fetal development that persists throughout the duration of the exposure period.
Footnotes
Author contributions
SKD, JLK and JLB designed the research project and wrote the manuscript. JLB, MAG and SMJ conducted the animal experiment, sample collection and sample preparation. JLB and REN analyzed the RNA Seq data. RRP and TFB conducted the histology image analyses. All authors have read and approved the manuscript.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Technical Contribution No. 6801 of the Clemson University Experiment Station. This material is based upon work supported by NIFA/USDA, under project number 1700537. The Leica stereoscope used in this study is housed in the Clemson Light Imaging Facility and was purchased with funds from the Clemson University Division of Research.
Data availability statement
The datasets generated and analyzed during the current study are available in Gene Expression Omnibus (GEO) repository (wlalwaggtjyhlqj).
