Abstract
Background
Allergic rhinitis (AR) is a common immunoglobulin E-mediated immune response involved various cell types, while the role of nasal fibroblasts (NFs) in the pathogenesis of AR is less understood.
Purpose
The study aimed to uncover the gene expression profile of AR-derived NFs and the potential mechanism for the changed phenotype of AR-NFs.
Research Design
The primary NFs were isolated from 3 AR patients (AR-NFs) and 3 controls (Ctrl-NFs), and the proliferation, migration and interleukins production abilities of NFs were detected respectively. RNA-sequence was used to identify differentially expressed genes (DEGs) in AR-NFs. Transcription factor (TF) regulatory network and bioinformatic analyses were both conducted to clarify the biological roles of DEGs including the TFs. The DEG with the highest validated |fold change (FC)| value, detected by qPCR, was selected for further confirmation.
Results
AR-NFs showed a higher proliferation and migration abilities as well as released higher levels of IL-33 and IL-6, compared to Ctrl-NFs. A total of 729 DEGs were screened out in AR-NFs. TF regulatory network indicated that BARX homeobox 1 (BARX1) and forkhead box L1 were the major node TFs. Bioinformatic analyses showed that a large number of DEGs including several target genes of BARX1 were both enriched cytokine-related GO terms, and immune- or inflammation-related pathways. BARX1 had the highest |FC| value, and silencing BARX1 in AR-NFs resulted in the significant downregulation of proliferation and migration abilities, and the production of interleukins.
Conclusions
Our study for the first time provided the gene expression profile of AR-derived NFs, and BARX1 could be developed as a potent target to alleviate the pathogenesis of AR.
Introduction
Allergic rhinitis (AR) is a common non-infectious inflammatory disease affecting 10–40% population worldwide. 1 AR is a specific immunoglobulin E (IgE)-mediated immune response against inhaled allergens, accompanied with classical symptoms including rhinorrhea, sneezing and nasal congestion, thus significantly affects quality of life and productivity at work or school. 2 Drug therapies such as anti-histamines, leukotriene receptor antagonists and nasal sprays, aiming to target mediators, cytokines, or nonspecific inflammation to attenuate allergic symptoms, are still current major treatment for AR patients.3,4 Recent studies indicate that other non-immune cells, such as epithelial cells and fibroblasts, also play important roles in the allergic response. It is reported that, epithelial cell-derived cytokines, including thymic stromal lympho-poietin (TSLP), interleukin (IL)-25 and -33, are able to regulate the function of dendritic cells (DCs), T cells, and mast cells, resulting in the production of proinflammatory cytokines and even the initiation of type-2 immune responses. 5 In addition, impairment of tight junctions molecules also plays causative roles in the pathogenesis of AR. 6
Nasal fibroblasts (NFs) are an important cell component of nasal mucosa, and also are another notable population of non-immune cells in the pathogenesis of AR. Several studies reveal that NFs are not only the effector cells against allergens but also the mediators response to various inflammatory cytokines. For instance, exposure of NFs to urban particulate matter and diesel exhaust particles causes the upregulation of IL-6 and IL-8,7,8 two important modulators of local inflammation. 9 IL-6 is also thought to amplify the allergic inflammatory response. 10 What is more, treatment of histamine, predominantly released from mast cells, also induces the release of IL-6 in NFs. 11 IL-17A and IL-21 are both involved in the pathogenesis of AR.12,13 It is reported that, stimulation of NFs from AR patients with IL-17A significantly up-regulates the production of TSLP. 14 Administration of IL-21 inhibits eotaxin release from NFs in murine AR models. 15 The important role of NFs in the development of AR indicates that, target regulating the function of NFs is a necessary and vital research direction in the multi-targets treatment of AR. The current studies have uncovered relative pathways activated in the effector NFs,8,16 but the deeper mechanism was not well understood.
In the present study, we aimed to investigate the phenotype and gene expression profile alterations of NFs isolated from AR patients (AR-NFs) compared to the control NFs (Ctrl-NFs). The biological roles of the differentially expressed genes (DEGs) in AR-NFs were then annotated by bioinformatic analyses, and transcription factors (TFs) regulatory network was also constructed. Finally, the role of interested DEG, BARX1 (BARX homeobox 1), playing important roles in tissue development and carcinogenesis,17,18 on phenotype of NFs was selected for further verification.
Materials and methods
Patients and specimens
Inferior turbinate mucosa tissue from six donors was collected for the isolation of NFs. Three donors (2 males, 1 female; mean age 32.67 ± 6.03) diagnosed with persistent AR complicated with turbinate hypertrophy were sorted as the AR group, while another three donors (2 females, 1 male; 28.67 ± 4.51) without AR and required a corrective septorhinoplasty was recruited as the Ctrl group. None of the patients had a history of asthma, or infections of the respiratory tract. For AR patients, only those without the use of corticosteroids, antibiotics, leukotriene modifiers and anti-histamines within a month before surgery were included. The fresh specimen was reserved in sterile phate-buffered saline (PBS) and transported to the laboratory for cell isolation within 3 h. This study was approved by the Ethics Committee of Shanghai Tenth Peoples’ Hospital, Tongji University, and written informed consent was obtained from all patients. All procedures conformed to the principles of the Declaration of Helsinki.
Nasal fibroblasts isolation and culture
The isolation of NFs was carried out following previous studies8,19 with some modifications. Fresh specimens were first washed with PBS supplemented with antibiotic-antimycotic solution (Gibco, Gaithersburg, MD) for three times to avoid contamination. Subsequently, the specimens were cut into small pieces (1–3 mm2) and digested in collagenase solution (500 U/mL, Sigma) at 37°C for 60 min. Next, the cells were collected by centrifugation at 1000 g for 5 min after the dissociated specimens were filtrated. The resulted NFs were cultured in DMEM medium containing 10% of fetal bovine serum (FBS, Invitrogen, Carlsbad, CA, USA), and 1% of penicillin/streptomycin (Invitrogen). The medium was changed every 3 days. All the cells were incubated at 37°C with 5% CO2.
Proliferation assay
Cells were collected and resuspended at a density of 3 × 104 cells/ml, and 100 µl of cells was then seeded into each well of the 96-well plate. Cell viability was detected daily for 7 days by MTT assay. Briefly, 10 µl of 5 mg/mL MTT solution (Sigma-Aldrich; Merck KGaA, Darmstadt, German) was added to each well and the cells were incubated at 37°C for 3 h. The absorbance of each well was detected at 490 nm after the formazan was totally resolved in the DMSO solution. Proliferation rate was calculated using the following formula: proliferation rate = A(day n)/A(day 1), n = 1–7.
Transwell assay
Briefly, cells at a density of 4 × 105 cells/mL was resuspended in the FBS-free medium. Then 100 µl of the cells were seeded into the upper chamber of the transwell (Costar, Cambridge, USA), and another 600 µL complete medium was added to the lower chamber. After 24 h, cells retained in the inner side of the membrane were removed, and those migrated through the membrane were fixed with methanol for 20 min and stained with crystal violet solution for another 20 min. Finally, the visual fields of each chamber were randomly selected and photographed by an inverted microscope.
Enzyme-linked immunosorbent assay
Cell supernatant was collected to detect the level of IL-33 and IL-6 using human IL-33, IL ELISA kits (ABclonal, Wuhan, China), following the manufacturer’s instructions. Optical density values were measured at 450 nm. The concentration of each cytokines was calculated using standard curve of corresponding recombinant protein.
qPCR
Primer sequence used for qPCR.

Ribonucleic Acid (RNA)-sequence
Total RNA of cells in AR (n = 3) and Ctrl (n = 3) groups was collected by Trizol reagent (Invitrogen, Carlsbad, CA, USA), and isolated using RNeasy mini kit (Qiagen, Germany). Complimentary DNA (cDNA) libraries were synthesized following the protocols of TruSeq™ RNA Sample Preparation Kit (Illumina, USA), which included mRNA purification and fragmentation, cDNA synthesis and end-repair, as well as the PCR enrichment. Next, the libraries were quantified by Qubit®2.0 Fluorometer (Life Technologies, USA) and validated by Agilent 2100 bioanalyzer (Agilent Technologies, USA) to confirm the insert size and calculate the mole concentration. Cluster was generated by cBot with the library diluted to 10 p.m. and then sequenced on the Illumina NovaSeq 6000 (Illumina). Gene abundance was expressed as fragments per kilobase of exon per million reads mapped (FPKM). Stringtie software was used to count the fragment within each gene, and TMM algorithm was used for normalization. Differentially expressed genes (DEGs) were screened under the thresholds of |fold change (FC)| value >1.5 and p value < 0.05.
Bioinformatic analyses
Gene Ontology (GO) analysis (https://david.ncifcrf.gov/home.jsp) for biological processes, cellular components and molecular function and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis (http://www.genome.ad.jp/kegg) were performed, respectively, to annotated the biological role of the DEGs. Enrichment analysis was further performed and visualized using R package.
Cell transfection
AR-NFs seeded in 6-well with a confluence of 80% was prepared, then small interference RNA targeting BARX1 (si-BARX1) was transfected into the cells using lipofectamine 2000 (Invitrogen) following the instructions. After 48 h, cells were collected for qPCR verification. si-BARX1 (5’-GCU GGA GAA ACG CUU CGA GAA-3’) and the negative control (si-NC, 5’-UUC UCC GAA CGU GUC ACG UTT-3’) were synthesized by Geneseed Biotech (Guangzhou, China).
Statistical analysis
Results were presented as mean ± standard deviation. The statistical significance of differences between two groups was assessed unpaired t-test or one-way analysis of variance (ANOVA). Triplicate wells were prepared for each condition in each experiment. p < 0.05 was considered statistically significant.
Results
Phenotype detection of allergic rhinitis-nasal fibroblasts in proliferation, migration abilities, and production of cytokines
As shown in Figure 1(a), NFs isolated from two groups were both shuttle-shaped, and AR-NFs were slender spindle in morphology. MTT results showed that the proliferation rate of AR-NFs was significantly enhanced compared to the controls (Figure 1(b)). Transwell assay indicated that AR group-derived NFs showed a higher migration ability than Ctrl-NFs (Figure 1(c)). In addition, the supernatant IL-33 and IL-6 levels of AR-NFs were both significantly higher than those of the controls according to the ELISA analysis result (Figure 1(d)). Further, qPCR detection indicated that the mRNA expression levels of the two interleukins were also significantly enhanced in AR-NFs (Figure 1(e)), consistent to the ELISA result. Phenotype detection of AR-NFs in proliferation, migration abilities and production of cytokines. (A) Cellular morphology of NFs; (B) MTT assay was used to detect the proliferation rate of NFs; (C) Transwell assay was used to evaluate the migration ability; (D) ELISA assay was performed to detect supernatant IL-33 and IL-6 of NFs; (E) qPCR was used to detect mRNA expression levels of IL-33 and IL-6 in NFs. AR: allergic rhinitis; Ctrl: control; NFs: nasal fibroblasts. *p < 0.05.
RNA-seq analysis of differentially expressed genes in allergic rhinitis-derived nasal fibroblasts
A total of 729 DEGs, including 341 upregulated (red dots) and 388 down-regulated genes (blue dots), were identified in AR-NFs compared to Ctrl-NFs by RNA-seq in the thresholds of |FC| value >1.5 and p value < 0.05 (Figure 2(a)). |FC| value of most DEGs ranged from 1.5–4. A small amount of the DEGs was over eight and the dots could be found in the outer vertical border of Figure 2(a). The heap-map of DEG presented the expression profile of each NF sample, and the overall gene expression trend of three samples in each group was basically consistent (Figure 2(b)). The FC value of the top 20 DEGs were also listed in Table 2. As we could see, 70% of the DEGs were significantly upregulated, and BARX1 had the highest FC value (1705.1) (Table 2). In addition, the FC value of IL-33 reached 20.305, consistent with the increased production of IL-33 in ELISA result. RNA-seq analysis of DEGs in AR-NFs. (A) RNA-seq was used to screen DEGs in AR-NFs compared to Ctrl-NFs under the thresholds of |FC| value >1.5 and p value < 0.05, and volcano plot presented all the upregulated (red dots) and down-regulated genes (blue dots); (B) The fold change of all DEGs in each sample were presented in the heat-map. AR: allergic rhinitis; Ctrl: control; NFs: nasal fibroblasts. DEGs: differentially expressed genes. Top 20 differentially expressed genes in allergic rhinitis-nasal fibroblasts.

Transcription factor regulatory network of differentially expressed genes
For a better understanding the connections between those DEGs, TF regulatory network was constructed. As we could see in Figure 3, the network covered seven of the TFs and 41 of the targeted DEGs. And one of the TFs, BARXs, had 13 of target genes, including, CDK11A, LEPR, CTH, GBP2, Vascular cell adhesion molecule 1 (VCAM1), PTGS2, and PPP1R12B, et al. Another TF forkhead box L1 (FOXL1), with the log2 FC = −1.89, shared eight target genes with BARX1, and also regulated the transcription of another 19 genes. TF regulatory network of DEGs. TF: transcription factors; DEGs: differentially expressed genes.
Bioinformatic analyses of differentially expressed genes
Gene ontology enrichment analysis showed that many DEGs were involved in protein synthesis, including protein folding in endoplasmic reticulum, ATF6-mediated unfolded protein response, and PERK-mediated unfolded protein response (Figure 4). Some DEGs were enriched in positive regulation of vascular associated smooth muscle cell migration, and transforming growth factor beta receptor binding (Figure 4). Since little relation of the top 30 GO terms could be directly connected with the production of the cytokines, the keywords, “cytokine,” “chemokine,” and “immune,” were used to screen relevant biological processes and DEGs. As a result, six GO terms were screened out under the threshold of p value < 0.05 (Table 3). DEGs enriched in cytokine-related processes (GO:0034097, GO:0071345) reached ∼90–100, including some chemokine/cytokine and the receptors, IL33, CXCL6, IL7R, and CCR10, as well as some targets of BARX1, such as VCAM1, GBP2, CTH, PTGS2, and LEPR (Table 3). Additionally, 64 DEGs were involved in immune system development, including BARX1, and IL7R, and VCAM1 (Table 3). GO classification and enrichment analyses of all DEGs were presented in Supplementary Table S1. Top 30 GO terms of DEGs enriched in. circle, triangle, and square, represents biological process, cellular component, and molecular function. The size of the icon indicates gene number enriched in the GO term. The color of each icon indicates the q value. DEGs: differentially expressed genes. Differentially expressed genes enriched in relevant GO terms.

Kyoto encyclopedia of genes and genomes classification indicated that many DEGs were involved in immune and endocrine systems, and some DEGs were involved in cancer-related disease in the class of human disease, which might be related to the enhanced proliferation and migration abilities of AR-NFs (Figure 5). Moreover, 80 DEGs, ∼11% of all DEGs, were involved in signal transduction (Figure 5), indicating a comprehensive dysregulation of pathways in AR-derived NFs, compared to the controls. DEGs enriched in 20 sub-pathways of immune system, 23 sub-pathways of signal transduction, and another four pathways we interested in were listed in Table 4. PTGS2 and VCAM1 were both involved in NF-kappa B (NF-κB) and TNF signaling pathways, and protein phosphatase 1 regulatory subunit 12B (PPP1R12B) was associated with regulation of actin cytoskeleton, which belongs to cell motility pathway (Table 4). Enrichment analysis showed that, many DEGs were enriched in protein processing in endoplasmic reticulum and ECM-receptor interaction, and some of them were involved in hippo signaling pathway-multiple species and p53 signaling pathway (Supplementary Figure S1). KEGG classification and enrichment analyses of all DEGs were provided in Supplementary Table S2. KEGG classification of all DEGs. All the KEGG pathways were sorted into six groups marked in different colors. The length of each bar indicates gene number enriched in the pathway. DEGs: differentially expressed genes. Differentially expressed genes enriched in relevant KEGG pathways.

Silence of BARX1 reversed the phenotype of allergic rhinitis-nasal fibroblasts
To investigate the potential mechanism for the phenotype alteration of AR-NFs, qPCR validation was used to detect the FC of the top 5 DEGs. As we could see in Figure 6(a), the log2FC value of the 5 DEG was basically consistent to the RNA-seq result, and BARX1 had the highest validated FC value. Therefore, BARX1 was selected for further function investigation. Next, si-BARX1 was transfected to the AR-NFs, and qPCR validation showed that the expression level of BARX1 was decreased to ∼25%, indicating an effective interference of BARX1 in AR-NFs (Figure 6(b)). Phenotype analyses indicated that silence of BARX1 significantly inhibited the proliferation and migration ability of AR-NFs (Figure 6(c)). What is more, the supernatant IL-33 and IL-6 as well as the mRNA levels of the two interleukins were both reduced after silencing BARX1 in AR-NFs (Figures 6(d) and (e)). Silence of BARX1 reversed the phenotype of AR-NFs. (A) qPCR detection was used to verify the fold change of top 5 DEGs in AR-NFs compared to Ctrl-NFs; (B) qPCR analysis of the mRNA level of BARX1 in AR-NFs after silence of BARX1; (C) MTT assay detected the effect of silencing BARX1 on the proliferation ability of AR-NFs; (D) Transwell assay evaluated the effect of silencing BARX1 on the migration ability of AR-NFs; (E) ELISA assay detected the effect of silencing BARX1 on the production of IL-33 and IL-6 from AR-NFs; (F) qPCR was used to detect mRNA expression levels of IL-33 and IL-6 after silencing BRAX1 in AR-NFs. AR: allergic rhinitis; Ctrl: control; NFs: nasal fibroblasts. *p < 0.05.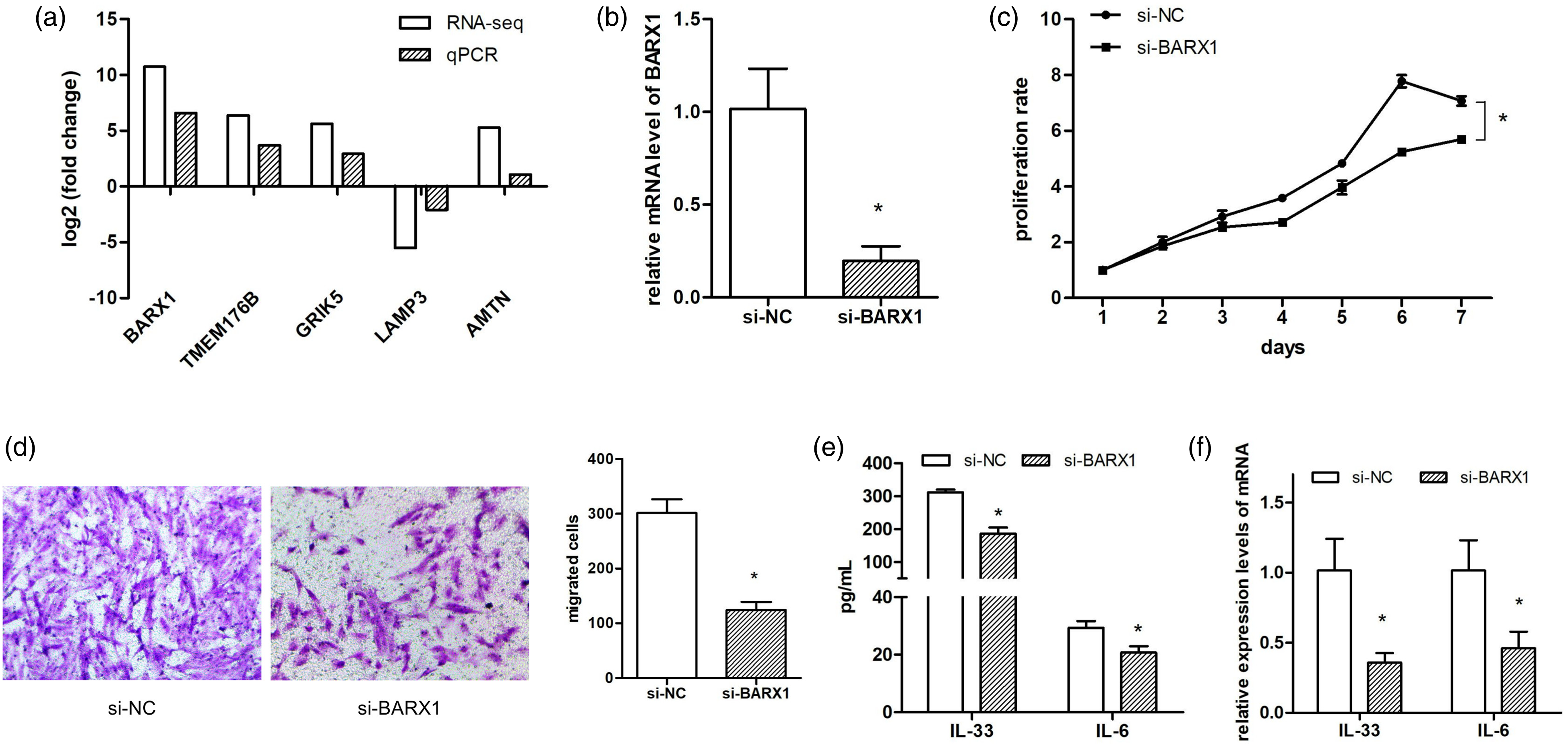
Discussion
Allergic rhinitis is a common chronic allergic respiratory disorder involved in various cells and mediators. The roles of the immune cells including DCs, T cells, and mast cells, have been well understood, 2 and the crosstalk between epithelial cells and immune cells is also well-clarified in the pathogenesis of AR. 5 However, the role of NFs in AR is less investigated. The present study uncovered the gene expression profile of AR-derived NFs for the first time, and also provided a potential treatment target to reverse the special phenotype AR-NFs.
Nasal fibroblasts are not only the major support cells of nasal mucosa but also could release various cytokines and chemokines to involve in the immune reaction. Several studies have proved that NFs are the important effector cells against the allergens and cytokines through some treatment to normal or AR-NFs. Different from the previous studies, our study focused on both normal and AR-derived NFs, and the direct identification of DEGs between the two NFs without any treatment helps to uncover the actual gene alterations in AR-NFs. Phenotype detection results indicated that AR-NFs showed a higher proliferation and migration abilities, compared to the controls, which is similar to the result of a previous study. 20 IL-33 is a member of IL-1 family and also a ligand for the receptor ST2, widely expressed on Th2 cells, mast cells, eosinophils and basophils, and epithelial cells. 21 Therefore, the increased release of IL-33 generally results in the production of proinflammation cytokines or the activation of the above cells. 22 IL-6 is recognized as a marker of allergic asthma, 23 and is also the major regulator in the differentiation and function of Th17 cells, 24 which play an important role in the Th2-mediated allergic response through the production of IL-17. 25 The elevated levels of supernatant IL-33 and IL-6 from AR-NFs indicates an extensive immune-regulatory and also important proinflammatory roles of NFs in the development of AR. Inhibition the expression of these cytokines released from NFs is helpful to alleviate the progression of AR.
RNA-seq identified 729 DEGs in AR-NFs compared to the controls. To date, there is no any reports about the gene expression profile of AR-derived NFs, and only the dysregulated genes in nasal mucosa tissue or airway epithelium cells from AR patients are identified in some researches.26,27 Under the thresholds of |FC| value >2 and p value < 0.05, a total of 127 mRNAs are screened out in three of AR nasal mucosa tissue compared to another three of healthy controls through microarray analysis. 26 Obviously, the DEGs identified in our RNA-seq result were far more than the 127 DEGs. Apart from the difference in sample sources (NFs vs nasal mucosa tissue), the major reasons were the differences in methods and thresholds. The original RNA-seq analysis result covers more than 40,000 genes, while only ∼26,000 probes are included in the mRNA microarray. Additionally, |FC| value used in our study is 1.5, lower than value of 2. The DEGs screened out in our NFs samples was also compared to the previous study. Interestingly, some genes such as IL33, ZFP90, MTHFD2, and BST2, were both dysregulated in AR nasal mucosa tissue 26 and our NFs samples, which indicate the coincidence of the two results to some extends.
The construction of TF regulatory network presented the regulatory relationships among the DEGs. One of the TF, BARX1 with highest FC value (1705.1) and various targets attracted our attention. Another TF, FOXL1, also play an important role in the complex transcription regulatory network, but its FC value was only 3.7, making it difficult to obtain a relative consistent validated result. Therefore, we focused on the top 5 DEGs, and qPCR validation showed that the resulted FC values were basically consistent the RNA-seq results. The selection of BARX1 for next confirmation is based on the considerations of both validated |FC| values and the direct or indirect biological roles. We checked the bioinformatic analyses of the top five candidates in the GO terms and KEGG pathways we interested in (Tables 3 and 4), and found few direct relationships of all the 5 DEGs. Different from other 4 DEGs without target genes, some target genes of BARX1 were enriched in cytokine-related biological processes and classical inflammatory pathways. Therefore, BARX1 with the highest validated FC value and the important biological roles were chosen out.
BARX1 is first studied in developmental biology and is proven to play vital roles in the development of molar teeth, stomach, 28 and muscle. 17 The role of BARX1 in immune or inflammation reactions is still a blank. Recent studies showed that BARX1 could also participate in the progression of various cancers including clear cell renal cell carcinoma, 18 endometrial carcinoma, 29 and gastrointestinal stromal tumor. 30 Overexpression or silence of BARX1 in cancer cells resulted in significant phenotype alterations in proliferation and migration abilities.18,29,30 Similarly, in the present study, silence of BARX1 significantly inhibited the proliferation and migration abilities of AR-NFs. Additionally, supernatant IL-33 and IL-6 levels were also reduced after silencing BARX1. Since little clues could be found between BARX1 and related bioinformatic annotation, we focused on its target genes.
For instance, VCAM1, GBP2, CTH, PTGS2, and LEPR were enriched in cytokine-related biological processes. The dysregulation of these genes might play some roles in the development of AR. KEGG pathway analysis showed that VCAM1 and PTGS2 were enriched in NF-κB and TNF signaling pathways. NF-κB pathway is an important mediator in the activation of immune and inflammation responses,31,32 and local NF-κB inhibition suppresses the allergic response in a mouse AR model along with reduced level of IL-6. 33 VCAM1 is a cell surface glycoprotein of the immunoglobulin gene superfamily, and is expressed in endothelial cells, epithelial cells, dendritic cells, 34 and fibroblasts. 35 The expression of VCAM1 could be induced by cytokine, and play an important role in recruiting immune cells to sites of active inflammation.34,36 Notably, it is reported that the activity of VCAM1 promoter could also be regulated by NF-κB (p65) in endothelial cells from atherosclerotic plaques, 37 and the upregulated VCAM1 in our RNA-seq result might be attributed to the activation of NF-κB pathway. Therefore, the increased IL-6 level in AR-NFs might be attributed to the activation of NF-κB pathway, and the reduced IL-6 in BARX1 silenced AR-NFs might be related to the inhibition of the pathway.
Another target gene PPP1R12B was enriched in cell motility-related pathway (regulation of actin cytoskeleton). PPP1R12B, one of the regulatory subunits of protein phosphatase 1 (PP1), is predominantly expressed in cardiac/skeletal muscle and brain. 38 Recent studies indicate that, PPP1R12B is closely associated with the cell migration behavior. For instance, loss of PPP1R12B causes breast cancer cells to exert higher traction forces, 39 and PPP1R12B knockdown significantly abrogated pseudopodium enriched atypical kinase 1-mediated tumor suppressive effects, including proliferation and migration abilities. 40 These evidences showed that PPP1R12B plays important roles in modulating cellular behaviors, and the role of BARX1 on the proliferation and migration behaviors of AR-NFs might be attributed to the transcriptional regulation of PPP1R12B.
In conclusion, our study provided a gene expression profile of AR-derived NFs, and proved that upregulation of BARX1 was responsible for phenotype alteration of AR-NFs. The potential mechanism of BARX1 might be related to the regulation of NF-κB pathway or the transcriptional regulation of PPP1R12B. Additionally, the DEGs identified in AR-NFs also provided a large number of candidate genes to further investigate the role of NFs in the pathogenesis of AR.
Supplemental Material
sj-pdf-1-het-10.1177_09603271211069038 – Supplemental Material for A comprehensive gene expression profile of allergic rhinitis-derived nasal fibroblasts and the potential mechanism for its phenotype
Supplemental Material, sj-pdf-1-het-10.1177_09603271211069038 for A comprehensive gene expression profile of allergic rhinitis-derived nasal fibroblasts and the potential mechanism for its phenotype by Zhengwen Li, Wentao Zou, Jingwen Sun, Shuang Zhou, Yue Zhou, Xiaojing Cai and Jiaxiong Zhang in Human & Experimental Toxicology
Supplemental Material
sj-pdf-2-het-10.1177_09603271211069038 – Supplemental Material for A comprehensive gene expression profile of allergic rhinitis-derived nasal fibroblasts and the potential mechanism for its phenotype
Supplemental Material, sj-pdf-2-het-10.1177_09603271211069038 for A comprehensive gene expression profile of allergic rhinitis-derived nasal fibroblasts and the potential mechanism for its phenotype by Zhengwen Li, Wentao Zou, Jingwen Sun, Shuang Zhou, Yue Zhou, Xiaojing Cai and Jiaxiong Zhang in Human & Experimental Toxicology
Supplemental Material
sj-pdf-3-het-10.1177_09603271211069038 – Supplemental Material for A comprehensive gene expression profile of allergic rhinitis-derived nasal fibroblasts and the potential mechanism for its phenotype
Supplemental Material, sj-pdf-3-het-10.1177_09603271211069038 for A comprehensive gene expression profile of allergic rhinitis-derived nasal fibroblasts and the potential mechanism for its phenotype by Zhengwen Li, Wentao Zou, Jingwen Sun, Shuang Zhou, Yue Zhou, Xiaojing Cai and Jiaxiong Zhang in Human & Experimental Toxicology
Footnotes
Author Contributions
JZ and XC designed the study.
ZL and WZ performed the experiments.
JS and SZ collected the data.
ZL and YZ analyzed the data.
All authors contributed to preparation of the article and approved the final version.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Ethics approval
This study was approved by the Ethics Committee of Shanghai Tenth Peoples’ Hospital, Tongji University, and written informed consent was obtained from all patients. All procedures conformed to the principles of the Declaration of Helsinki.
Data availability statement
The data supporting the conclusions of the study are available from the corresponding author upon request.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
