Abstract
Dysregulation of the long non-coding RNA prostate androgen regulated transcript 1 (lncRNA PART1) is involved in the tumorigenesis of various cancers. However, little is known about its function and molecular mechanism in the development of lung squamous cell carcinoma (LSCC). In this study, we examined the expression of PART1 in LSCC clinical tissue samples and cell lines, and gain- and loss-of-function experiments were performed to explore the function of PART1 in LSCC proliferation, invasion and migration. We found that PART1 was overexpressed in both LSCC tissues and cell lines. Functional studies revealed that PART1 knockdown significantly suppressed cell proliferation, invasion and migration but enhanced apoptosis in LSCC cells, whereas overexpression of PART1 showed the opposite results. Mechanistically, we identified that PART1 acted as a sponge of miR-185-5p, and sineoculis homeobox homolog 1 (Six1) was a direct downstream target of miR-185-5p. Moreover, restoration of miR-185-5p or silencing of Six1 partially abolished the oncogenic effect of PART1 in LSCC cells. Clinically, The areas under the receiver operating characteristic (ROC) curve of PART1, miR-185-5p, and Six1 were 0.7857, 0.7332, 0.8112, respectively. Notably, high PART1, low miR-185-5p, and high Six1 expressions were significantly associated with severe clinical parameters and were the independent risk factors for poor prognosis of LSCC patients. Thus, we concluded that the PART1/miR-185-5p/Six1 axis might serve as a novel biomarker for the diagnosis and treatment of LSCC.
Introduction
Lung cancer is considered as the most common malignancy and the leading cause of cancer-related mortality worldwide. 1 Non-small-cell lung cancer (NSCLC), including large-cell carcinoma, adenocarcinoma and squamous cell carcinoma, accounts for 80%–85% of all lung cancer, of which lung squamous cell carcinoma (LSCC), as the major subtype of pathological NSCLC, is mainly diagnosed in aged men and is closely associated with metastasis and relapse. 2,3 Although studies have revealed several genetic mutations of crucial oncogenes that lead to the pathogenesis of LSCC, new cases of LSCC are still increasing every year due to the lack of standard diagnostic strategies for its early stage. 4 Accordingly, a better understanding of the molecular mechanisms underlying the carcinogenesis of LSCC and the identification of novel biomarkers are required to develop effective therapeutic strategies. 5,6
High throughput sequencing analysis has shown that more than 80% of human genome is substantially transcribed as non-coding RNAs (ncRNAs) that lacks of protein coding ability, such as the well-known long non-coding RNAs (lncRNAs, >200 nucleotides) and microRNAs (miRNAs, 18–22 nucleotides). 7 Recently, intensive studies of ncRNAs have revealed their essential functions in regulation of diverse cell biological processes, including proliferation, metastasis, survival and genomic stability at the transcriptional and post-transcriptional levels. 8 lncRNAs that act as competing endogenous RNAs (ceRNAs) are able to sequester miRNAs (also called miRNA sponges) via competing with endogenous mRNAs to bind to the miRNA response elements (MREs) in a sequence-specific manner, which results in the reduction of combination between miRNAs and their target mRNAs, and thus weaken the suppressive function of miRNAs on the downstream target gene expressions. 9,10 To date, increasing evidence has identified the carcinogenic or tumor suppressive role of lncRNA-mediated ceRNA networks involved in NSCLC occurrence and dominating crucial cancer hallmark processes like proliferation, apoptosis, angiogenesis, immune escape, drug resistance, as well as epithelial-mesenchymal transition (EMT) and metastasis across NSCLC progression, 11 –13 accordingly, clarifying the molecular mechanism of the complex “ceRNA-lncRNA-miRNA-mRNA” networks underlying tumor biology will provide novel insight into the potential diagnostic and therapeutic biomarkers of NSCLC (Figure 1).

The schematic represented the ceRNA-mediated “lncRNA-miRNA-mRNA” network that regulated the initiation and progression of NSCLC. The depicted oncogenic or tumor suppressive role of “lncRNA-miRNA-mRNA” regulatory axis affected the major hallmark capabilities of NSCLC: proliferation, apoptosis, immortalization, angiogenesis, EMT and metastasis, drug resistance, genomic instability, immunological destruction.
lncRNA prostate androgen regulated transcript 1 (PART1) is located on chromosome 5q12 and is considered to be modulated by androgens and predominantly expressed in the prostate gland. 14 Recently, PART1 has been illustrated to be upregulated in several human cancers, including prostate cancer, esophageal squamous cell carcinoma and NSCLC. 15 –17 For instance, PART1 has been reported to serve as ceRNAs to sponge miR-635 and promote tumor progression in NSCLC. 18 However, the potential molecular mechanisms underlying PART1 on the regulation of LSCC cell proliferation and metastasis still remains largely unknown. Hence, the present study aimed to examine the function of PART1 in LSCC and elucidate the underlying mechanisms involved. In this study, the effects of PART1 on LSCC cell proliferation, invasion and migration were explored, and the potential regulatory relationships between PART1, miR-185-5p and Six1 were verified. Our findings might provide new therapeutic targets for patients receiving LSCC treatment.
Materials and methods
Clinical sample collection
Fifty-one paired LSCC tumor tissues and adjacent normal tissues were collected from patients diagnosed with LSCC who underwent surgical treatment at Shaanxi Provincial People’s Hospital (Xi’an, China). No patients had received any chemotherapy or radiotherapy before surgery. Written informed consent was obtained from all patients, and this study was approved by the Ethics Committee of the Shaanxi Provincial People’s Hospital and followed the declaration of Helsinki. The fresh tissues were immediately stored at −80°C for further analysis. Clinicopathological parameters were obtained from the patient’s records.
Cell culture
Human LSCC cell lines (H2170, H226, H520, SK-MES-1) and the normal lung epithelial cell line BEAS-2B were purchased form the cell bank of the Type Culture Collection of the Chinese Academy of Sciences (Shanghai, China). All the cell lines were maintained in RPMI-1640 media (Gibco, Waltham, MA, USA) containing 10% fetal bovine serum (FBS, Gibco), 100 U/ml penicillin, and 100 mg/ml streptomycin (Invitrogen, Carlsbad, CA, USA) in a humidified incubator with 5% CO2 at 37°.
Cell transfection
To knock down or upregulate PART1 expression, PART1 short hairpin RNA (5′-AAA CAG CCT CCT TCA AAA TGC AGC TTG ACT GTG ATT CAT GGC ACA AAG CTG GAA TAA CCC TGC CAG TTC T-3′) and negative control (5′-CCG GCA ACA AGA TGA AGA GCA CCA ACT C-GAG TTG GTG CTC TTC ATC TTG TTG TTT TT3′), pcDNA3.1-PART1 overexpressing plasmid and empty vector were synthesized commercially by GenePharma (Shanghai, China). Six1 siRNA (5′-UUU GCA AAG GCG AAA ACC GGA-3′) and siRNA negative control (5′-AGU CUA CGU AGA ACA AGG ACG-3′), miR-185-5p mimic (5′-UGG AGA GAA AGG CAG UUC CUG A-3′) and its negative control (5′-AUC AGU GAG CGA GCA GUA GUA G-3′) were obtained from RiboBio Co., Ltd. (Guangzhou, China). Cell transfection was performed in H2170 and H520 cell lines by using Lipofectamine 2000 (Invitrogen) following the manufacturer’s instructions. After transfection for 48 h, cells were collected for subsequent detection.
qRT-PCR
Total RNA from 51 paired clinical LSCC tissues or cell lines (H2170, H226, H520, SK-MES-1, BEAS-2B) was extracted with TRIzol reagent (Invitrogen) according to the manufacturer’s protocol. mRNA or miRNA was reverse transcribed into cDNA by using PrimeScriptII strand cDNA synthesis kit (Takara, Dalian, China). RT-qPCR analysis was performed using the SYBR Green Realtime PCR Master Mix (Toyobo, Nipro, Osaka, Japan) on the ABI 7500 qPCR system (Applied Biosystems, MA, USA). The specific primer sequences were designed as follows: PART1, forward 5′-CCT GT TTG GGG ATG CCT CAT-3′, reverse 5′-TGT GTC TCC TCT CCC GTA GG-3′; Six1, forward 5′-CCT AGG AAG CCG TCC TTT CTC C-3′, reverse 5′-AGA CCC TTT CCT CGT CTG GG-3′; β-actin, forward 5′-CTC TGC CGG AGA AAC AGG TG-3′, reverse 5′-GCG TCT CTT CAT CTC TCC CG-3′; miR-185-5p, forward 5′-TGG AGA GAA AGG CAG TTC CTG A-3′, reverse 5′-CAG GCG GTC TGG AGT-3′; U6, forward 5′-GAG ACG GGA ACG ACA AAC CT-3′, reverse 5′-TGG ACG AAG AGG ATT CGC TG-3′. β-actin and U6 were used for normalization. The relative expression levels of miRNA and mRNA were quantified by using the 2−△△Ct method.
Cell viability
Cell viability was determined by using the Cell Counting Kit 8 (CCK-8, Dojindo, Shanghai, China) according to the manufacturer’s instructions. Transfected H2170 and H520 cell lines were seeded into a 96-well plate at a density of 5 × 103 cells per well and incubated at 37° for 0, 24, 48, 72 and 96 h. Subsequently, 10 µl of CCK-8 solution was added into each well and incubated for 4 h at 37°. The optical density (OD) value was determined at 450 nm with a microplate reader.
Colony formation assay
To measure cell proliferation, 500 H2170 and H520 cells with different treatments were incubated in six-well plates for about 2 weeks until visible clones were formed. After washing with PBS, the cells were fixed with 4% paraformaldehyde for 15 min and then stained with crystal violet for 30 min at room temperature. The number of clones greater than 50 cells was calculated by an inverted microscope (Olympus, Tokyo, Japan).
Transwell assay
Cell invasion and migration assays were performed by using Transwell chambers (8 µm pore size, Corning, NY, USA). In brief, for migration detection, 1 × 105 transfected H2170 and H520 cells were suspended with 200 µl serum-free medium and seeded in the upper chambers. For invasion assays, the upper chambers were pre-coated with Matrigel (BD Bioscience, San Jose, CA, USA). Meanwhile, 600 µl medium containing 10% FBS was added to the lower chambers. After incubation for 48 h at 37°, H2170 and H520 cells remaining on the upper chambers were slightly removed with a swab, cells invaded or migrated into the lower chambers were fixed with 4% paraformaldehyde for 15 min and then stained with 0.1% crystal violet for 15 min. The cell numbers from six random fields of each chamber were photographed and counted under a light microscope (Olympus).
Western blot
Protein lysates were prepared from transfected H2170 and H520 cells, respectively, and extracted by using RIPA buffer (Pierce, Thermo Scientific, MA, USA), and the protein contents were measured with a BCA protein assay kit (Beyotime, Beijing, China). Proteins (10 μg) were separated by 12% SDS-PAGE and then transferred onto PVDF membranes (Bio-Rad, USA). After that, the membranes were blocked with 5% non-fat milk for 1 h at room temperature and incubated with primary antibodies against Six1, MMP-2, MMP-9, Vimentin, N-cadherin, E-cadherin (Abcam, Cambridge, MA, USA), β-actin at 4° overnight. After rinsing, the membranes were incubated with horse radish peroxidase (HRP)-conjugated secondary antibodies (Abcam) for 2 h at 37°. Protein bands were visualized with an enhanced chemiluminescence (ECL) system (Amersham Biosciences, NJ, USA) and quantified with ImageJ software (Rio-rad, CA, USA). β-actin was used as the internal control.
Cell apoptosis
Cell apoptosis was determined with the Annexin V-FITC/PI Apoptosis Detection Kit (Vazyme, Nanjing, China) following the manufacturer’s instructions. Briefly, the transfected H2170 and H520 cells were collected and centrifuged at 1000 rpm for 5 min at 4°, respectively. After rinsing twice with pre-cooling PBS, 2 × 105 cells were re-suspended with 100 µl 1× binding buffer. Subsequently, cells were incubated with 5 µl Annexin V-FITC and 5 µl PI staining solution for 10 min in the dark at room temperature. After mixing with 400 µl 1× binding buffer, cell apoptosis was examined with a FACS Calibur flow cytometer (Bection Dickinson, NJ, USA) within 1 h.
Luciferase reporter assay
The LncBase Predicted v.2 bioinformatics database was used to predict the potential target genes of lncRNA PART1 and miR-185-5-5p. PART1 luciferase plasmid containing wild-type (WT) or mutant (MUT) binding sites of miR-185-5p were constructed. Furthermore, the WT or MUT binding sites of miR-185-5p on the 3′-UTR of Six1 was cloned into the pmirGLO plasmid (Rromega, Madison, WI, USA) to construct the WT or MUT Six1 3′-UTR luciferase plasmid. The luciferase reporter plasmid and miR-185-5-5p mimics or negative control were co-transfected into H2170 and H520 cells by using Lipofectamine 2000 (Invitrogen), respectively. After transfection for 48 h, the relative luciferase activity was determined by using the Dual-Luciferase Reporter Assay System (Promega) according to the manufacturer’s protocol. Firefly luciferase activity was used for normalization.
RNA immunoprecipitation (RIP)
RIP experiments were performed by using the Magna RIP™ RNA-Binding Protein Immunoprecipitation Kit (Millipore, Billerica, MA, USA). Briefly, H2170 and H520 cells were pre-transfected with miR-185-5p or miR-NC for 48 h, respectively, and then the cells were incubated with RIP magnetic beads conjugated with anti-Ago2 antibody or IgG following the manufacturer’s instructions. The PART1 coprecipitated with Ago2 was determined by qRT-PCR.
RNA fluorescence in situ hybridization
The subcellular location of lncRNA PART1 in H2170 and H520 cell lines was detected by the RNA fluorescence in situ hybridization (RNA-FISH) kit (RiboBio, Guangzhou, China) method as previously described. 19,20 The nuclei were counterstained with DAPI (blue, RiboBio) and the fluorescence of PART1 (red) was observed under a laser-scanning confocal microscope (Leica, Germany).
Statistical analysis
The data were expressed as mean ± standard deviation (SD), and statistical analysis was performed by using GraphPad Prism 5.0. The differences between two groups were analyzed by Student’s t-test and differences among multi-groups were measured by ANOVA followed by Dunnett’s multiple comparison test. The relationship between PART1 and miR-185-5p or Six1 in LSCC tissues was assessed by Pearson’s correlation analysis. Receive operating characteristic (ROC) curves were constructed and the areas under the ROC curves (AUC) were evaluated to assess the diagnostic values of the PART1, miR-185-5p, and Six1. The 5-year overall survival rate of LSCC patients was determined by Kaplan-Meier survival analysis with the log-rank test, and the prognosis was evaluated by Cox’s proportional hazard regression model for multivariate analysis. Each experiment was performed at least three independent times. P < 0.05 was considered to indicate statistical significance.
Results
PART1 is upregulated in LSCC tissues and cell lines
qRT-PCR assay showed that PART1 expression was significantly overexpressed in LSCC tissues compared with the corresponding adjacent normal tissues (P < 0.01, Figure 2(a)). Furthermore, the expression level of PART1 was remarkably increased in LSCC cell lines H2170, H226, H520, SK-MES-1 compared with that in the normal lung epithelial cell line BEAS-2B (Figure 2(b)). Additionally, RNA fluorescence in situ hybridization (RNA-FISH) result revealed that PART1 was mainly located at the cytoplasmic region of H2170 and H520 cells (Figure 2(c)). Therefore, these data indicated that the elevated PART1 might be involved in the carcinogenic process of human LSCC.

PART1 was upregulated in LSCC tissues and cell lines. (a) Relative expression levels of PART1 in 51 paired LSCC tissues and adjacent normal tissues in LSCC samples were determined by qRT-PCR. (b) Relative PART1 levels in LSCC cell lines H2170, H226, H520, SK-MES-1 and normal lung epithelial cell line BEAS-2B. **P < 0.01. (c) Representative image of the subcellular location of PART1 in LSCC cell lines H2170 and H520 was detected by RNA-FISH. Red represents PART1; blue represents nuclei.
Knockdown of PART1 inhibits LSCC cell proliferation and apoptosis
To further elucidate the biological effects of PART1 in LSCC cells, H2170 and H520 cell lines were selected for the following gain- and loss-of-function investigations. qRT-PCR assay revealed that significant downregulation of PART1 was found in H2170 cells and marked upregulation of PART1 was observed in H520 cells with sh-PART1 and PART1 overexpression plasmid transfection, respectively (Figure 3(a)). Moreover, colony formation assay showed that the colony formation capacity was significantly repressed with PART1 knockdown in H2170 cells when compared with the sh-NC group, while it increased notably with PART1 overexpression in H520 cells (P < 0.01, Figure 3(b)). Similarly, CCK-8 assay also suggested that silencing of PART1 significantly suppressed cell viability in H2170 cells (P < 0.01), but ectopic expression of PART1 markedly promoted cell proliferative ability in H520 cells (P < 0.01, Figure 3(c)). Furthermore, the results of flow cytometry assay displayed that knockdown of PART1 induced the increased apoptosis rate in H2170 cells (P < 0.01), while H520 cells with PART1 overexpression showed a decreased apoptosis ratio (P < 0.05, Figure 3(d)).

Effect of PART1 on LSCC cell proliferation and apoptosis. (a) Relative expression of PART1 in H2170 cells with PART1 knockdown or in H520 cells with PART1 overexpression. The cell growth ability of H2170 cells with PART1 knockdown and H520 cells with PART1 overexpression was measured by colony formation assay (b) and CCK-8 assay (c). (d) The apoptotic rates of H2170 and H520 cells with PART1 knockdown or PART1 overexpression were determined by flow cytometry. *P < 0.05; **P < 0.01.
Knockdown of PART1 attenuated LSCC cell invasion, migration and EMT
To explore whether PART1 plays a role in the metastasis of LSCC cells, the invasion and migration of H2170 and H520 cells with PART1 overexpression or PART1 silencing were determined by Transwell assays. We found that downregulation of PART1 significantly suppressed the migration and invasion capacities in H2170 cells (Figure 4(a) and (b)). However, endogenous expression of PART1 remarkably accelerated the number of migrated and invaded H520 cells (Figure 4(a) and (b)). Furthermore, western blot assays revealed that the levels of EMT-related proteins, such as the epithelial marker E-cadherin, were apparently increased with PART1 silencing, but the mesenchymal markers Vimentin, N-cadherin, MMP-2 and MMP-9 were decreased in H2170 cells, whereas H520 cells with PART1 overexpression showed the opposite effects (P < 0.05, Figure 4(c)). Taken together, these findings suggested that the downregulation of PART1 could suppress migration, invasion and EMT in LSCC cells.

Knockdown of PART1 attenuated LSCC cell invasion, migration and EMT. Cell migration (a) and invasion (b) in H2170 and H520 cells with sh-PART1 or PART1 overexpressing plasmid interference were assessed by Transwell assay. (c) The expression levels of MMP-2, MMP-9, N-cadherin, Vimentin and E-cadherin were determined by western blot. *P < 0.05; **P < 0.01.
PART1 acted as a sponge to miR-185-5p
Increasing evidence has corroborated that lncRNA PART1 acts a sponge or a competing endogenous RNA (ceRNA) by regulating the expression and biological roles of miRNAs in cancers. LncBase Predicted v.2 bioinformatics database was carried out to predict the potential binding site for miR-185-5p on PART1 (Figure 5(a)). Accordingly, luciferase reporter assays suggested that the luciferase activity of WT-PART1 was notably decreased with miR-185-5p mimic transfection (P < 0.05), but no change in luciferase activity of the MUT-RART1 in both H2170 and H520 cells was observed (Figure 5(b)). Moreover, an anti-Ago2 RIP assay was performed to verify the association of PART1 and miR-185-5p in H2170 and H520 cells. The results showed that PART1 RIP was apparently enriched with miR-185-5p mimic transfection in both H2170 and H520 cells (P < 0.01, Figure 5(c)). Furthermore, qRT-PCR assay revealed that the expression levels of miR-185-5p were significantly reduced in both LSCC clinical samples and cell lines H2170, H226, H520, SK-MES-1 compared with those in normal groups (Figure 5(d) and (e)). Moreover, a negative relationship was found between PART1 and miR-185-5p expression in LSCC tissues (r2 = 0.182, P < 0.001, Figure 5(f)). Additionally, the level of miR-185-5p was significantly enhanced in H2170 cells with PART1 knockdown but distinctly reduced in H520 cells with PART1 overexpression (P < 0.01, Figure 5(g)). Subsequently, the expression level of miR-185-5p was overexpressed by miR-185-5p mimic transfection in H2170 and H520 cells (P < 0.01, Figure 5(h)), and we found that miR-185-5p upregulation significantly inhibited PART1 expression in H2170 and H520 cells (P < 0.05, Figure 5(i)), which further indicated that PART1 functionally interacted with miR-185-5p in LSCC cells.
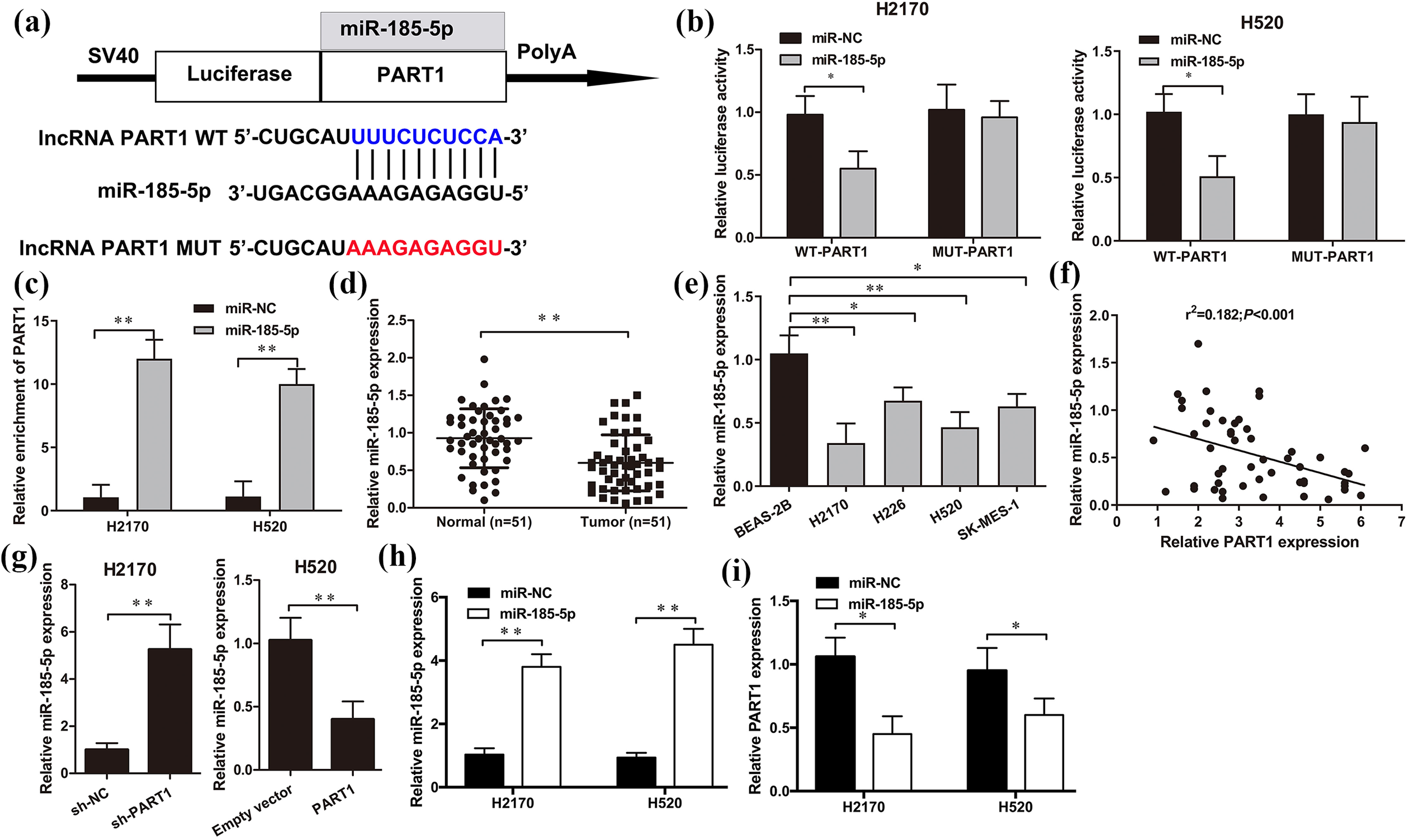
PART1 acted as a sponge to miR-185-5p. (a) Regions of putative interaction between PART1 and miR-185-5p were predicted by LncBase Predicted v.2 bioinformatics database. (b) Luciferase activity of H2170 and H520 cells with wild-type (WT)-PART1 or mutant (MUT)-RART1 or and miR-NC or miR-185-5p mimic cotransfection was measured by Luciferase reporter assay. (c) The enrichment of PART1 associated with Ago2 was detected by RIP assay in H2170 and H520 cells with miR-NC or miR-185-5p mimic transfection. miR-185-5p expression was downregulated in 51 paired LSCC clinic tissues (d) and cell lines H2170, H226, H520, SK-MES-1, BEAS-2B (e). (f) Pearson’s correlation analysis was performed to measure the relationship between PART1 and miR-185-5p in 51 paired LSCC clinic tissues. (g to i) The expression of miR-185-5p and PART1 were assessed by qRT-PCR in H2170 and H520 cells with sh-PART1, PART1, or miR-185-5p mimic transfection. *P < 0.05; **P < 0.01.
PART1 competitively binds to miR-185-5p by upregulating Six1 in LSCC cells
To further explore the underlying mechanism of the regulatory effects of PART1 on LSCC cells, the putative targets of miR-185-5p were predicted by TargetScan online software. Six1 was selected as a candidate target of miR-185-5p (Figure 6(a)) due to its crucial roles in carcinogenesis involved in multiple human tumors. The direct interaction between Six1 and miR-185-5p was confirmed by the luciferase reporter assay. As shown in Figure 6(b), the relative luciferase activity with miR-185-5p mimic and WT-Six1 3′-UTR co-transfection was notably suppressed in both H2170 and H520 cells when compared with the miR-NC group (P < 0.05). However, co-transfection of MUT-Six1 3′-UTR and miR-185-5p mimic failed to inhibit luciferase activity (Figure 6(b)). Moreover, qRT-PCR and western blot assays showed that the Six1 expression level was significantly reduced in H2170 and H520 cells with miR-185-5p overexpression (P < 0.05, Figure 6(c) and (d)). Additionally, to elucidate whether PART1 had an impact on Six1 expression, we detected the expression of Six1 in H2170 and H520 cells with either PART1 knockdown or PART1 overexpression by western blot method. As shown in Figure 6(e), Six1 was notably decreased by sh-PART1 interference compared with the sh-NC group in H2170 cells (P < 0.05), while Six1 was significantly upregulated by overexpression of PART1 in H520 cells (P < 0.05, Figure 6(e)). Moreover, we found that Six1 was remarkably overexpressed in LSCC cell lines H2170, H226, H520, SK-MES-1 (Figure 6(f)), and LSCC tumor tissues compared to adjacent normal samples (P < 0.01, Figure 6(g)) and a positive correlation was obtained between PART1 and Six1 in LSCC tissues (r2 = 0.215, P < 0.001, Figure 6(h)).

PART1 competitively binds to miR-185-5p by upregulating PART1 in LSCC cells. (a) The potential binding site of miR-185-5p in Six1 3′-UTR was predicted by the TargetScan database. (b) The luciferase activity was determined in H2170 and H520 cells with WT-Six1 3′-UTR or MUT-Six1 3′-UTR and miR-185-5p mimic or miR-NC co-transfection. The expression of Six1 was examined by western blot (c) and qRT-PCR (d) in H2170 and H520 cells with miR-185-5p mimic or miR-NC transfection. (e) The effect of PART1 on Six1 expression in H2170 and H520 cells was measured by western blot assay. (f) Six1 expression was upregulated in LSCC cell lines H2170, H226, H520, SK-MES-1 and (g) LSCC tissues and (h) was positively correlated with Six1 expression in LSCC tissues. *P < 0.05; **P < 0.01.
Diagnostic values of PART1, miR-185-5p and Six1 in LSCC
ROC curves were established to determine the diagnostic abilities of PART1, miR-185-5p and Six1 in LSCC specimens compared with adjacent normal tissues. As shown in Figure 7, the AUC for PART1 was 0.7857 (P < 0.01, 95%CI = 0.6949–0.8764), and the cut-off value was 2.850, with a sensitivity of 66.67% and a specificity of 86.27%; The AUC for miR-185-5p was 0.7332 (P < 0.01, 95%CI = 0.6336–0.8328), and the cut-off value was 0.7332, with a sensitivity of 72.55% and a specificity of 74.51%; The AUC for Six1 was 0.8112 (P < 0.01, 95%CI = 0.7230–0.8994), and the cut-off value was 2.750, with a sensitivity of 78.43% and a specificity of 78.43%. These results suggested that PART1, miR-185-5p, and Six1 had diagnostic capacities to distinguish LSCC tumor tissues from healthy controls.

ROC curve analysis of PART1, miR-185-5p and Six1 expressions for 51 cases of LSCC tissue samples from adjacent normal tissues.
The prognostic value of PART1, miR-185-5p and Six1 in LSCC patients
To explore the clinical significance of PART1, miR-185-5p and Six1 in LSCC, the 51 cases of LSCC patients were divided into high and low expression groups according to the median relative expression levels of PART1, miR-185-5p and Six1, respectively. As shown in Table 1, the overexpression of PART1 was positively associated with large tumor size (P < 0.01), advanced histological stage (P < 0.05), lymph node metastasis (P < 0.05) and poor differentiation (P < 0.05) in LSCC patients; partially consistent with PART1, increased Six1 expression was also highly correlated with large tumor size (P < 0.05), smoking history (P < 0.01), lymph node metastasis (P < 0.01) and differentiation (P < 0.01). Whereas, we found that miR-185-5p was down-regulated in advanced histological stage (P < 0.05), poor differentiation (P < 0.05), and metastasis tissues (P < 0.01). Additionally, the Kaplan-Meier analysis showed that the high expression levels of PART1 and Six1, and low expression levels of miR-185-5p have positive correlation with the worse overall survival of LSCC patients (P < 0.05, Figure 8). Furthermore, the multivariate Cox proportional hazard regression model analysis showed that high PART1, low miR-185-5p, high Six1 expression, differentiation, and lymph node metastasis were independent risk factors for the overall survival of LSCC patients (Figure 9).
Association between the clinicopathologic parameters and PART1, miR-185-5p and Six1 expression in LSCC (n = 51).

a P < 0.01.
b P < 0.05.

The Kaplan-Meier curves of the relative expression levels of PART1 (a), miR-185-5p (b), and Six1 (c) in patients with LSCC. *P < 0.05.

The forest plot of clinicopathological factors for the overall survival of LSCC patients based on multivariate Cox proportional hazard regression analyses.
PART1 exerted oncogenic effects on LSCC cells via the miR-185-5p/Six1 axis
To investigate whether the promoted effects of PART1 on LSCC cell aggressiveness was mediated by miR-185-5p/Six1 axis, the reduced miR-185-5p expression was rescued in PART1-overexpressing H520 cells with miR-185-5p mimic co-transfection (Figure 10(a)), meanwhile, the enhanced Six1 expression induced by PART1 was inhibited with si-Six1 or miR-185-5p mimic transfection in H520 cells (Figure 10(b)). Furthermore, we found that restoration of miR-185-5p partially reversed the promotional effects of PART1 overexpression on H520 cell proliferation, migration, invasion, and apoptosis (Figure 10(c) to (e)). Similarly, the stimulated effect of PART1 overexpression on cell proliferation, migration, invasion, and apoptosis was also notably abrogated with Six1 knockdown (Figure 10(c) to (e)) in PART1-overexpressing H520 cells. Therefore, these results suggested that miR-185-5p/Six1 axis participated in the promotional effect of PART1 in LSCC tumorigenesis (Figure 11).

PART1 exerted oncogenic effects on LSCC cells via the miR-185-5p/Six1 axis. (a) The expression of miR-185-5p in H520 cells with miR-185-5p mimic or pc-PART1 transfection was examined by qRT-PCR. (b) Six1 expression in H520 cells with si-Six1, miR-185-5p mimic or pc-PART1 transfection were detected by western blot. CCK-8 (c), Transwell assay (d), and flow cytometry (e) were performed in H520 cells. *P < 0.05; **P < 0.01.

The molecular mechanism diagram involved in PART1/miR-185-5p/Six1 axis regulating LSCC cell proliferation, invasion and migration.
Discussion
Although novel means for the diagnosis and treatment of LSCC have been developed in recent decades, unfortunately, biological markers appropriate for early detection and targeted therapy still remain ineffective in LSCC patients with poor prognosis. 21 LncRNAs such as MALAT1, ANRIL, SHNG1, and HOTAIR have recently been implicated to play essential roles in the pathogenesis of LSCC development due to their ability to manipulate progression and carcinogenesis in various malignancies in various malignancies. 22,23 . Recently, overexpression of PART1 has been reported to be associated with tumor recurrence and poor prognosis in patients with stage I–III NSCLC. 16 Similarly, in our cohort, we found that PART1 was remarkably upregulated in LSCC samples and markedly correlated with the large tumor size, advanced histological stage, lymph node metastasis, differentiation, and shorter overall survival, which were the crucial parameters associated with poor survival and prognosis in LSCC patients. Notably, the ROC diagnostic analysis is widely used to evaluate the diagnostic abilities of biomarkers, and the AUC area above 0.7 is commonly referred as favorable diagnostic value. 24 Consistent with the results from previous reports with the similar detection approach, 16,25,26 our study revealed that the AUC values of PART1, miR-185-5p, and Six1 were 0.7857, 0.7332, 0.8112, respectively, implicating that PART1, miR-185-5p, and Six1 might be effective diagnostic biomarkers for LSCC detection. The multivariate Cox model also demonstrated that high PART1, low miR-185-5p, and high Six1 expressions were closely associated with shorter overall survival of LSCC patients, which highlighted that high PART1, low miR-185-5p, and high Six1 expressions were the independent unfavorable prognostic factors for LSCC development.
To explore the biological effect of PART1 in LSCC, gain- and loss-of-function experiments were performed in LSCC cell lines. In the current study, we found that knockdown of PART1 in H2170 cells significantly repressed proliferation, invasion and migration and accelerated apoptosis in H2170 cells. In contrast, overexpression of PART1 showed the opposite effects in H520 cells. Notably, silencing of PART1 suppressed the expression of tumor invasion- and migration-related proteins such as MMP-2 and MMP-9. Also, knockdown of PART1 decreased epithelial-mesenchymal transition (EMT), the crucial step facilitating the metastasis of tumor cells, 27 by upregulation of the epithelial marker E-cadherin, and loss of mesenchymal markers in LSCC cells. Consistently, recent research has discovered that overexpression of lncRNA PART1 induces cell proliferation and prevents apoptosis in prostate cancer cells. 15 Furthermore, PART1 was also found to be upregulated in colorectal cancer cells, and promoted cell growth and metastasis in vivo and in vitro. 28 Accordingly, our findings demonstrated that lncRNA PART1 functioned as an oncogene in LSCC cell proliferation, invasion and migration.
Accumulating evidence has stated that dysregulation of lncRNAs serves as ceRNAs involved in the progression of tumorigenesis by sponging miRNAs. 29,30 The subcellular location of lncRNA in the cytoplasm has also been suggested to function as a post-transcriptional role. 31 In our study, PART1 was found mainly localized in the cell cytoplasm of LSCC cells, indicating its oncogenic role may through post-transcriptional regulation. Accordingly, bioinformatics analysis was performed and predict that multiple miRNAs were potential targets for PART1, particularly, miR-185-5p expression, which has been demonstrated to be involved in the progression of tumorigenesis via the lncRNA-miRNA interactions, 32 was identified as downregulated and inversely correlated with PART1 expression in clinical LSCC tissues. Evidence has suggested that in oral squamous cell carcinoma, lncRNA PDIA3P serves as a ceRNA for miR-185-5p to promote cancer progression. 33 In melanoma, downregulation of miR-185-5p abated the suppressive effect of lncRNA UCA1 knockdown on cell EMT and invasion. 34 Indeed, a recent study confirmed that miR-185-5p was also verified as a crucial target for lncRNA RNCR3 to promote cell proliferation and invasion in prostate cancer. 35 In this study, luciferase reporter assays and Ago2 RIP assays also demonstrated that PART1 could competitively bind to miR-185-5p in LSCC cells and were negatively regulated by PART1. Rescue experiments further determined that restoration of miR-185-5p exerted its suppressive functions on LSCC cell proliferation, migration and invasion via partial reversal of the biological effects of PART1. Therefore, these findings unveiled that PART1 served as a molecular sponge for miR-185-5p by directly targeting the complementary sequence of miR-185-5p in LSCC cells.
Sineoculis homeobox homolog 1 (Six1), as a homeodomain transcription factor, belongs to the Six family, which has been demonstrated to be significantly overexpressed in various human malignancies, and is closely related to metastasis, aggressiveness and poor prognosis. 36,37 With regard to the function of Six1 in the promotion of tumorigenesis via crosstalk with varying miRNAs in NSCLC, Liu et al. demonstrated that Six1 acts as the downstream target for miR-186-5p and was overexpressed in cisplatin-resistant NSCLC cells. 38 More importantly, it is worth mentioning that previous studies have demonstrated the reciprocal relationship of miR-185/Six1 in ovarian, breast and pediatric renal cancer cell lines. 39 We also identified Six1 as a direct target of miR-185-5p in LSCC cells. Additionally, Six1 was inversely correlated with miR-185-5p expression but positively correlated with PART1 expression in LSCC cells. Moreover, overexpression of miR-185-5p or knockdown of Six1 abrogated the promotional effects of PART1 on LSCC cell proliferation, invasion and migration. Taken together, these findings suggest that PART1 exerts oncogenic functions via the miR-185-5p/axis in LSCC and might be a valuable biomarker for the prognosis of LSCC patients.
In conclusion, the current study demonstrates that lncRNA PART1 is upregulated in LSCC tissues and cell lines, and PART1 exerts an oncogenic role in LSCC cell proliferation, invasion and migration through the miR-185-5p/Six1 axis. Therefore, these data will provide a better understanding of PART1-meidated LSCC carcinogenesis, and suggest the potential correlation of PART1/miR-185-5p/Six1 axis to the unfavorable prognosis of LSCC. However, our investigation on the lncRNA-mediated ceRNA regulatory network involved in LSCC progression is still in preliminary stages with limitations, further in-depth functional investigations with advanced genomic technologies are still needed to evaluate the accuracy of these diagnostic biomarkers with large clinical samples and to uncover the complex ceRNA molecular mechanisms involved in LSCC.
Footnotes
Acknowledgment
The authors show their gratitude to the “Shaanxi Natural Science Foundation Project.”
Data availability statement
The data that support the findings of this study are available from the corresponding author upon reasonable request.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by Shaanxi Natural Science Foundation Project [No. 2014JM2-8166].
Research ethics and patient consent
All clinical specimens were collected in accordance with the Ethics Committee of the Shaanxi Provincial People’s Hospital (No. 2015034, Date: 2015/03/10, Xi’an, China) and followed the declaration of Helsinki. Written informed consent was obtained from all patients involved.
