Abstract
Long noncoding RNAs (lncRNAs) are the new class of transcripts and pervasively transcribed in the genome, which have been found to play important functional roles in many tissues and organs. LncRNAs can interact with target gene to exert their functions. However, the function and mechanism of lncRNA in cleft palate (CP) development remain elusive. Here, we investigated the role of lncRNA H19 and its target gene insulin-like growth factor 2 (IGF2) in CP of mice. All-trans retinoic acid (atRA) is a well-known teratogenic effecter of CP. After establishment of the CP mouse model using atRA in vivo, we found that the rate of CP in mice was 100%. The tail lengths of fetuses in atRA-treated mice were shorter than those of control mice from embryonic day (E)12 to E17. The expression of lncRNA H19 and IGF2 were embryo age-related differences between atRNA-treated and control mice. In addition, the the relationship between lncRNA H19 and IGF2 were negative correlation in the critical period of developmental palate. These findings suggest that lncRNA H19 mediate atRA-induced CP in mice.
Keywords
Introduction
Cleft palate (CP) is the most common congenital malformation in oral and craniofacial region. It may originate from disturbances at any stage of palate development, including defecting palatal shelf growth, delaying shelf elevation, failing shelf fusion, losing of medial edge epithelial cell death, postfusion rupture, or failure of mesenchymal consolidation. 1 The precise etiology and pathogenesis of CP are complex and poorly understood. All-trans retinoic acid (atRA) is the oxidative metabolite of vitamin A and essential for normal embryonic development. High levels of atRA can inhibit the growth of mouse embryonic palatal mesenchymal cells and fail palatal fusion, which leads to CP. 2 High levels of atRA can also abrogate medial edge epithelium cells apoptosis and degradation of the basal laminin, resulting in the failure of palatal fusion. 3 However, the mechanisms of atRA-induced CP have not been fully elucidated.
Long noncoding RNAs (lncRNAs) have been identified as the transcripts longer than 200 nucleotides of transcripts, 4 but they do not encode proteins 5 and are pervasively transcribed throughout the whole genome. 6 It has been shown that a lot of lncRNAs play critical roles in biochemical, cellular, and developmental activities. 7 The dysfunction of lncRNAs have been associated with various disorders including CP. 8,9 However, how the specific lncRNAs involved in atRA-induced CP have not been reported.
Long noncoding RNA H19 (lncRNA H19) is a 2.3-kb noncoding RNA (ncRNA), which is transcribed from the maternal allele and highly expressed prenatally and lowly expressed postnatally. LncRNA H19 is evolutionarily conserved at the nucleotide level of human and rodents and not translated into protein. 10 The lncRNA H19 gene is pervasively expressed in various tissues including palate in embryo. 11 However, whether the expression of lncRNA H19 in atRA-induced mouse CP varies with different developmental stages has not been examined yet. LncRNA H19 belongs to a conserved gene cluster on chromosome 7 in the mouse. The cluster contains the gene for insulin-like growth factor 2 (IGF2), and IGF2 locates 90 kb away from the lncRNA H19. Genetic evidence shows that the parent of origin-dependent expression patterns of the lncRNA H19 and IGF2 genes is coordinated in mouse. 12 IGF2 gene is also strongly expressed during embryogenesis and downregulated after birth. IGF2 gene codes for a growth factor that plays an important role in embryonic development and growth control. 13
In the present study, we speculated that lncRNA H19 may serve an important role in the process of atRA-induced mouse CP and thus investigated this effect in a mouse model of CP.
Materials and methods
Animals
A total of 24 two-month-old C57BL/6 female (F) and 10 male (M) mice were obtained from Henan Laboratory Animal Center. All experiments were carried out in accordance with the Experimental Animal Center Guide for the Care and Use of Laboratory Animals, and the Institutional Ethical Guidelines for Experiments with Animals. The mice were housed at constant temperature (22 ± 2°C) and relative humidity (50 ± 10%) with a 12-h light/dark cycle. The females were placed with the males (3F:1M) for approximately 18 h and checked for the presence of vaginal plugs, indicative of mating (embryonic day 0 (E0)).
Construction of CP model and sample collection
All-trans retinoic acid (atRA; Sigma, Saint Louis, Missouri, USA) dissolved in corn oil (Jinglongyu, Qinghuangdao, China), which was prepared to 10.0 mg/mL suspension and administered orally to each pregnant mouse (100-mg/kg body weight) in treated group on embryo day 10 (E10). The same volume of corn oil was administered to the control mice. On E12, E13, E14, E15, E16, and E17, the pregnant mice of both groups were sacrificed using the following procedures. The fetuses were first removed from the uterus of sacrificed mice. The length of the fetuses was measured by Vernier caliper (Mitutoyo, Tokyo, Japan). Then, the five palates of fetuses on E13.5, E14.5, and E15.5 were harvested from both groups under a stereomicroscope (Olympus, Tokyo, Japan), respectively, and immediately preserved in RNAlater solution (Ambion, Austin, Texas, USA) at 4°C and stored at −80°C for further RNA isolation. Also five palate samples from fetuses were collected at each time point (E13.5, E14.5, and E15.5) in corn oil (control) and atRA group, respectively, and fixed with 10% formalin for histological examination.
Quantitative real-time polymerase chain reaction (qRT-PCR)
The harvested palates of E13.5, E14.5, and E15.5 from the control and atRA-treated mice were lysed in Trizol lysis solution (Invitrogen, Carlsbad, California, USA). The total RNA of five palates were isolated according to the manufacturer’s instructions and finally dissolved in nuclease-free water. To detect the expression of lncRNA H19 and IGF2, first-strand cDNA was synthesized using the Reverse Transcription Kit (TakaRa Biotechnology, Kyoto, Japan), and amplified through RT-PCR with the SYBR Premix Ex Taq Kit (TakaRa) by ABI 7900 PRISM system (Applied Biosystems, Carlsbad, California, USA). 18 s rRNA was used as an internal control. The conditions of qRT-PCR for lncRNA H19 and IGF2 were as follows: polymerase activation for 15 min at 95°C, 40 cycles at 95°C for 15 s, 56°C for 20 s, and 72°C for 30 s. The threshold cycle (Ct) value of the PCR amplification curve of the target gene was analyzed. All primers were synthesized by Invitrogen Co. (Invitrogen, Carlsbad, California, USA). The primer sets used in qRT-PCR were as follows: 18 s rRNA-F: 5′-CGGACATCTAAGGGCATCA-3′, 18 s rRNA-R: 5′-AAGACGGACCAGAGCGAAA-3′; LncRNA H19-F: 5′-TACCCCGGGATGACTTCATC-3′, LncRNA H19-R: 5′-TATCTCCGGGACTCCAAACC-3′; IGF2-F: 5′-GTGTGTGTCAGCCAAGCATG-3′, IGF2-R: 5′-CAAATGTGGGGACACAGAGG-3′.
Statistical analysis
All data were compared using either double-sided Student t-test or one-way analysis of variance. Tukey’s post hoc test was used to determine the difference between groups. The choice of tests was made automatically using SPSS software, version 13.0 (SPSS, Chicago, Illinois, USA). All data were presented as mean ± standard deviation. The differences were considered statistically significant when *p < 0.05.
Results
The morphological alterations of control and atRA-treated mice
A dozen of C57BL/6 pregnant mice were exposed to atRA (100-mg/kg body weight) on embryonic day 10 (E10), and another dozen were exposed to corn oil (control). Multiple embryos were collected from each pregnant mouse. A total of 246 embryos were collected: 111 embryos from atRA-treated mice and 135 embryos from the control mice. The morphological alterations of all fetuses were observed under light microscope. Pregnant females of the control and atRA-treated mice were sacrificed from E17 (Figure 1(a)). In contrast, the tail lengths of fetuses in atRA-treated mice were shorter than those of control mice from E12 to E17 (Figure 1(b)).

Fetal morphology in the control and atRA-treated group. A single dose (100-mg/kg body weight) of at RA treated pregnant C57BL/6 mice and the same amount of corn oil was administered to the control groups. The fetuses of the control group and atRA-treated group were taken photograph on E17 (a). The short tails of fetuses were detected by Vernier caliper, and the tail lengths of atRA-treated group were shorter than that of the control group from E12 to E17 (b). Data were mean ± standard deviation of three replicate experiments. *p < 0.05, compared with corresponding control values. atRA: all-trans retinoic acid; E: embryonic day.
Histological alterations of palatal shelves in the control and atRA-treated mice
For the fetuses from atRA-treated mice, the detected rate of CP was 100%. Their morphological and histological alterations were examined. In detail, the budding of palatal shelves started to form horizontal growth on E13.5 in the fetuses from the control group (Figure 2(a)). In contrast, the formation of palatal shelves in the fetuses from atRA-treated mice grew vertically and could not lift to the level on E13.5 (Figure 2(b)). The palate shelves of normal fetuses started to contact on E14.5 (Figure 2(c)), while they were still separated and failed to lift on E14.5 in the fetuses from atRA-treated mice (Figure 2(d)). On E15.5, the palatal shelves of the fetuses from untreated mice were completely fused, but still separated with the fetuses from atRA-treated mice, indicative of CP formation (Figure 2(e) and (f)).

Histology analysis of palatal shelves in the control and atRA-treated group on the time points E13.5, E14.5, and E15.5. The palate shelves of fetuses elevated and reached a horizontal position above the dorsum of the tongue on E13.5 in control group (a). The palate shelves of fetuses delayed to elevate on E13.5 in atRA-treated group (b). The palate shelves of fetuses contacted at E14.5 in control group (c). The palate shelves of fetuses delayed to elevate at E14.5 in atRA-treated group (d). The palate shelves of fetuses contacted and fused each other in the end at E15.5 in control group (e). The palate shelves of fetuses failed to contact each other and resulted in CP in atRA-treated group (f). In the two group, hematoxylin and eosin staining was used (magnification, ×40). E: embryonic day; atRA: all-trans retinoic acid; PS: palate shelf.
The expression levels of lncRNA H19 at different developmental stages of CP
To study the expression patterns of lncRNA H19 parallel to the CP formation induced by atRA treatment, palate tissues on E13.5, E14.5, and E15.5 were used to test the expression levels of lncRNA H19. It was shown that the expression levels of lncRNA H19 varied with the development of palate (Figure 3). In detail, the expression levels of lncRNA H19 significantly were increased on E13.5 (1.20 ± 0.07 fold) and E15.5 (1.03 ± 0.04 fold), respectively, compared with the corresponding palate tissues of control fetuses. Interestingly, the expression level of lncRNA H19 was significantly reduced to 0.75 ± 0.04 on E14.5 compared with the corresponding control palate tissues.
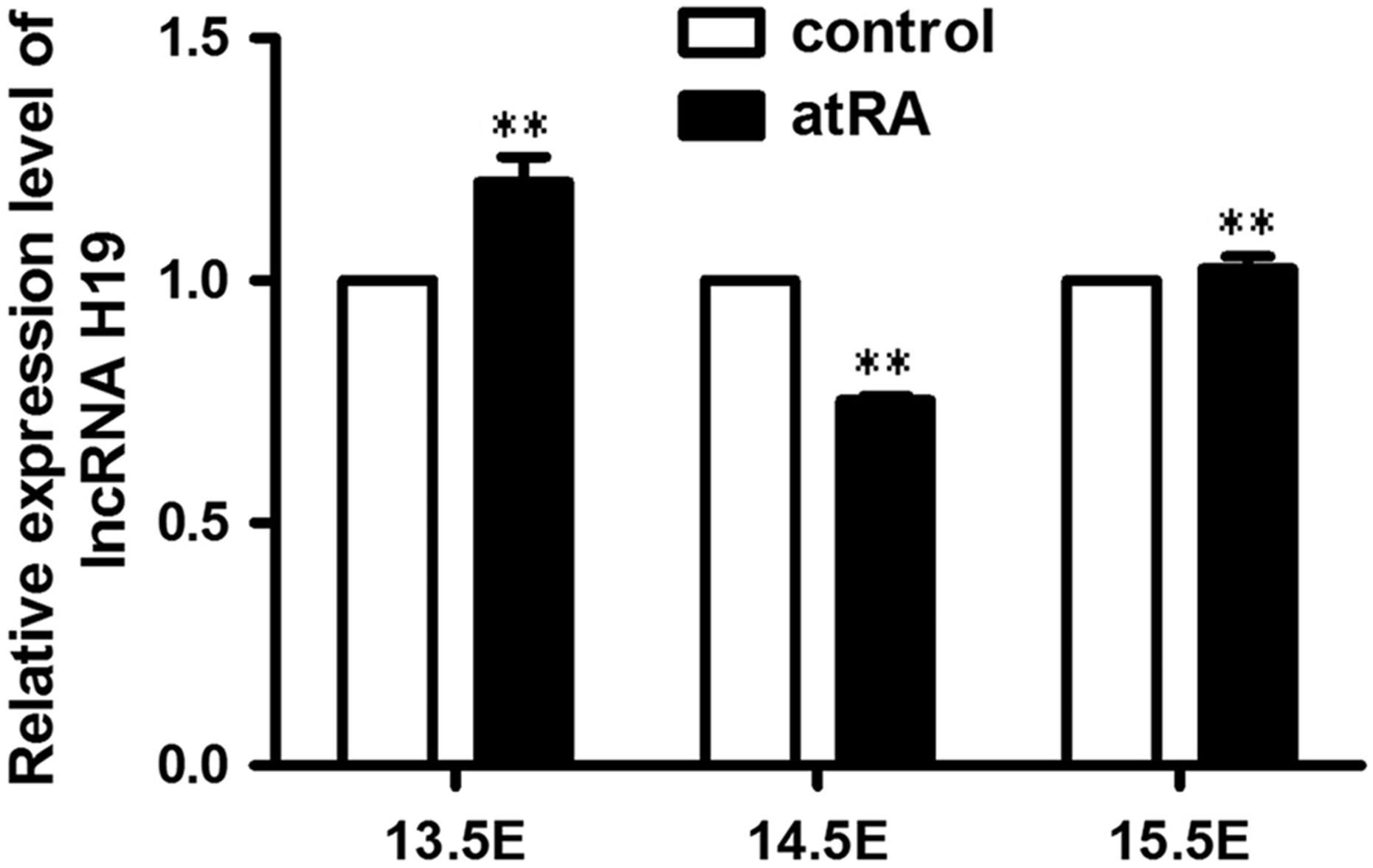
Relative expression of lncRNA H19 in the control (untreated) and atRA-treated groups on E13.5, E14.5, and E15.5. The expression levels of lncRNA H19 were determined by quantitative real-time polymerase chain reaction. Data were mean ± standard deviation of three replicate experiments. **p < 0.01 compared with the corresponding control values. atRA: all-trans retinoic acid; E: embryonic day.
The expression of IGF2 in the development of CP
To investigate the biological function of lncRNA H19, we examined one of the lncRNA H19 target genes, IGF2. IGF2 is a mitogen for a variety of cell types and required for normal embryonic growth. The expression of IGF2 was embryo age-specific during the development of palate (Figure 4). In detail, the expression levels of IGF2 gene significantly reduced to 0.66 ± 0.06 on E13.5, or 0.69 ± 0.08 on E15.5, respectively, compared with the corresponding palate tissues of control fetuses, but significantly increased on E14.5 (1.47 ± 0.42 fold) compared with the corresponding control palate tissues, respectively.

Relative expression of IGF2 in the control (untreated) and atRA-treated groups on E13.5, E14.5, and E15.5. The expression levels of IGF2 were determined by quantitative real-time polymerase chain reaction. The data are the mean ± standard deviation of three replicate experiments. *p < 0.05 or **p < 0.01 compared with the corresponding control values. atRA: all-trans retinoic acid; E: embryonic day.
Discussion
LncRNAs are the most abundant ncRNAs in the whole genome and have been demonstrated to have diverse biological functions. Documented studies have shown that lncRNAs play an important role in the development, 14 such as promoting pluripotency in embryonic stem cells 15 and regulating neuronal development. 16 In humans, CP accounts for one of the largest number of birth defects with a complex genetic and environmental etiology. However, few studies have been conducted to characterize the regulation of lncRNAs on the development of palate. LncRNA H19 gene was first identified as a CP gene in TGFβ3-knockout mice by RNA-Seq analysis. 9 While the lncRNA H19 involved in atRA-induced-CP has not been investigated yet. In this study, we established mouse CP model by atRA induced in our laboratory to define the role of lncRNA H19. First of all, the results demonstrated that some morphological abnormalities of fetuses in atRA-treated mice, such as short tails and CP, were similar with the findings of atRA-induced CP in rats. 1 Furthermore, the current study showed that the expression levels of lncRNA H19 varied with the stages of atRA-induced palatogenesis between the time points E13.5 and E15.5. As we know, the period from E13.5 to E15.5 is important for the development of palate. For example, both sides of the palates lift above the tongue on E13.5 and grow rapidly on E14.5, and begin to touch each other on E15.5. We found that the expression levels of lncRNA H19 in treated mice increased on E13.5 and E15.5, but decreased on E14.5. This was consistent with the results from previous study with CP of TGFβ3-knockout mice. 9 It has been revealed that the crucial stages of palatogenesis, including palatal convergence, adhesion, and fusion, usually occur from E13.5 to E15.5, 17 which implicated lncRNA H19 as the primary contributor to the development of CP induced by atRA.
As described previously, the lncRNA H19 may play an important role in the pathogenesis of CP induced by atRA. However, the mechanism of lncRNA H19-regulated CP in atRA-treated mice remains elusive. LncRNA H19 is crucial to the growth of embryonic and extraembryonic tissues throughout embryonic development, 18 while IGF2 is well recognized as an essential growth factor for proliferation and differentiation in various tissues during embryonic development. 19 To determine the mechanisms of lncRNA H19 and IGF2 interaction in atRA-induced CP, the expression levels of IGF2 gene at different stages of atRA-induced CP were also measured. It was found that the expression levels of IGF2 also showed embryo age-dependent pattern in the palatogenesis, which was unanimous that the expression levels of IGF2 mRNA showed embryo age-dependent pattern throughout rat gestation. 20 The expression levels of lncRNA H19 increased on E13.5 and E15.5, but reduced on E14.5. However, the expression levels of IGF2 decreased on E13.5 and E15.5, but increased on E14.5, and the phenomenon might be same that lncRNA H19 and IGF2 are two oppositely expressed imprinted genes located adjacent to each other at chromosome 7 in mouse. 21 Therefore, lncRNA H19 may play important roles in CP by interaction with IGF2. But the mechanism of lncRNA H19-mediated CP induced by atRA is still largely unclear. It was reported that the exposure of mouse preimplantation embryos to 2,3,7,8-tetrachlorodibenzo-pdioxin (TCDD) altered the methylation status of imprinted genes H19 and IGF2. 22 We speculated that the lncRNA H19 and IGF2 might be the similar patterns of methylation change in the stages of atRA-induced palatogenesis. Obviously, the precise mechanisms of lncRNA H19 and IGF2 in atRA-induced CP need to be further explored. In this study, we found that lncRNA H19 and the IGF2 genes were reverse expressed in atRA-induced cleft palatogenesis, which was consistent with the previous report on embryonic development. 23
Conclusion
The expression of lncRNA H19 and IGF2 were embryo age-related, and there was a negative correlation between atRNA-treated and control mice in the critical period of developmental palate. These findings suggested that lncRNA H19 mediates atR-induced CP in mice. However, further studies are necessary to discover the mechanism of lncRNA H19 in atRA-induced CP in mice.
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the grants from the National Natural Science Foundation of China (81502843).
