Abstract
The principal impediment to gene therapy is the development of efficient, nontoxic gene carriers that can handle and deliver foreign genetic materials into various cell types, including healthy and cancerous cells. Poly-
Introduction
Gene therapy has drawn more attention for its potential in the treatment of various genetic disorders.
1,2
The major challenge in this therapy regards the development of an efficient, nontoxic gene carrier with an ability of handling and delivering foreign genetic materials into many cell types, such as healthy and cancerous cells.
3
Among different delivery vectors that have been applied to this purpose, viral vectors appear a promising candidate, however immunogenicity and oncogenic effects restrict their applications in human gene therapy.
4
Hence, nonviral vectors, including cationic polymers, cationic liposomes, and their conjugates, with low immunogenicity, no size limit, low costs, and so forth can be used as an option of choice.
5,6
Exclusively, polymeric vectors can present suitable ligands for specific cell targeting.
7
Poly-
Therefore, using biocompatible PLL derivatives
14
or conjugation of PLL with other polymers, such as PEI opens up new venues in dealing with aforementioned issues.
8,9,15,24
Toxicity of cationic polymers attributed to many factors such as molecular weight, charge density, and structure of polymers along with other parameters, such as the time of exposure, nature of the targeted cell/tissue, and interaction with circulating agents in plasma during in vivo application.
28
In a study, a biocompatible PLL analog known as poly (a-[4-amino-butyl]-
Materials and methods
Materials
Branched-PLL (MWapproximately 30–70 kDa) was purchased from Sigma–Aldrich (Munich, Germany). Phosphate-buffered saline, Dulbecco’s modified Eagle’s medium (DMEM), fetal bovine serum (FBS), penicillin, streptomycin, and trypsin were products of GIBCO (Grand Island, New York, USA). pDNA encoding Renilla luciferase (pRL-CMV; Promega, Madison, Wisconsin, USA) was transformed into Escherichia coli bacterial strain DH5 and amplified in selective Luria-Bertani medium. Also, it was purified by a DNA-purification column purchased from QIAGEN (Hilden, Germany). N-[2-Hydroxyethyl]piperazine-N′-[2-ethanesulfonic acid] (HEPES) and 3-[4,5-dimethylthiazol-2-yl]-2,5-diphenyl tetrazoliumbromide (MTT) were obtained from Sigma-Aldrich (Munich, Germany). Low-melting-point agarose and regular agarose were purchased from Fermentas (Sankt Leon-Rot, Germany). Tris, sodium hydroxide (NaOH), hydrogen chloride, ethylenediaminetetraacetic acid (EDTA), dimethyl sulfoxide (DMSO), and Triton X-100 were purchased from Merck (Darmstadt, Germany).
Cell culture
Neuro2A mammalian cells (ATCC CCL-131) were grown in DMEM supplemented with 10% FBS, 100 IU/ml penicillin, and 100 µg/ml streptomycin at 37°C in a humidified atmosphere under 5% carbon dioxide. Cells in passage three were used for different assays and cells were seeded 24 h prior to the next use.
Preparation of PLL/DNA complex
PLL/DNA complexes (polyplexes) were prepared with two weight ratios of 2:1 and 6:1 (w/w). To control the size of polyplexes, HEPES buffered glucose (HBG; 20-mM HEPES, 5.2% glucose, pH 7.2) was used to prepare small nanosized polyplexes. Besides, HEPES buffered saline (HBS; 20-mM HEPES, 5.2% glucose, 0.5-M sodium chloride (NaCl), pH 7.2) was applied for larger polyplexes. 30
Polyplex size measurement
The hydrodynamic size of polyplexes at two C/P ratios was analyzed using a particle size analyzer Nano-ZS (Malvern Instruments, UK) in both salt-free (HBG) and salty (HBS) buffers. A dilution of vectors in 125-µl buffer was added to an equal volume of the same buffer containing DNA. After a 20-min incubation, polyplex sizes were measured in triplicates.
Viability assay
Evaluation of cytotoxicity was performed by MTT assay. The cells were seeded at a density of 1 × 104 cells/well in 96-well micro-assay plates and incubated for 24 h. 1 Then, different concentrations of polymer (0–500 µg/ml) and two weight ratios of 2:1 and 6:1 (w/w) polyplexes (in two buffer HBG and HBS) were added to the cells and underwent incubation for another 24 h. After that, 10 μl of MTT solution (5 mg/ml) was added to each well. The incubation was iterated for an additional 1 h at 37°C, and then, the medium was aspirated off, while 100 μl of DMSO was added to dissolve Formazan crystals. Absorbance was measured via a spectrophotometric microplate reader (Statfax–2100, Awareness Technology, Palm City, Florida, USA) at a wavelength of 590 nm (reference wavelength of 630 nm). The cell viability as percentages were calculated as follows
The OD590(sample) and OD590(control) represent the measurement from the wells treated with PLL/DNA polyplexes in two buffers, and the measurements related to the untreated wells. 1,14
Apoptosis assay
As well-documented, DNA fragmentation can be detected as an apoptotic signal in treated cells. Applying a quantitative DNA-binding dye, namely propidium iodide (PI), cells in the absence of DNA are expected to take up less stain and become visible on the left of the G1 peak. Apoptotic Neuro2A cells were stained by means of PI and then evaluated through flow cytometry to detect the sub-G1 peak. 8,24
Briefly, Neuro2A cells at an initial population of 100,000 cells/well were exposed to two weight ratios of PLL polyplexes 2:1 and 6:1 (w/w) polyplexes (in two buffer HBG and HBS). After 24 h, the cells were incubated overnight at 4°C in a hypotonic buffer (50-μg/ml PI in 0.1% sodium citrate and 0.1% Triton X-100) prior to flowcytometric (FACS) analysis using an FACS can flow cytometer (Becton Dickinson, San Jose, California, USA).
Comet assay for detection of DNA damage
A total of 106 cells/well were seeded in 12-well micro assay plate and incubated for 24 h before the procedure. Then, two concentration of free PLL) 4 and 12 µg/ml) and also two PLL/DNA complexes ratio of 2 and 6 (C/P 2 and C/P 6) were added to the cells, which further underwent incubation at 37°C for 24 h. Afterward, DNA damage studies were carried using comet assay method with some modifications. 31 For every concentration, two slides were prepared and immersed in cold lysis solution at pH 10 (2.5-M NaCl, 100-mM Na2EDTA, 10-mM Trizma base, 1% Triton X-100, 10% DMSO) and kept at 4°C for 60 min. Following lysis, the DNA unwinding occurred employing the electrophoresis buffer (300-mM NaOH: 1-mM Na2EDTA at pH 13.5) for 30 min. Thereafter, electrophoresis was conducted at a constant voltage of 23 V and 300 mA at 4°C. Slides were then neutralized in 0.4-M Tris (pH 7.5) for 5 min, stained with ethidium bromide (20 µg/ml), followed by a washing step with water. As many as 100 cell nuclei per slide were analyzed via a fluorescent microscope (Nikon, Kyoto, Japan), equipped with an excitation filter of 520–550 nm and a barrier filter of 580 nm at 400× magnification. Scoring was subsequently performed by an image analysis system (CASP9). Of note, the percent of DNA in the tail (i.e. the amount of DNA in the comet tail of damaged cells) was indicative of the extent of DNA damage caused by the study samples.
Statistical analysis
Data are presented as mean ± standard deviation (SD). The statistically significance difference was determined using student’s t-test. The values of p ≤ 0.05 were considered significant.
Results
Size of the complexes
Complex sizes were determined by dynamic light scattering 32 (Tables 1 and 2). As can be seen, the polyplexes formed in HBS buffer showed larger sizes in comparison with those in HBG buffer, suggesting that the addition of the physiological levels of NaCl (150 mM) to PLL/DNA complexes culminated in significant increases in the mean diameter of the complexes (e.g. C/P 2: 262.97 vs. 4178.0; 16-fold).
Size of polyplexes formed with plasmid DNA at indicated C/P ratios in HBG.

SD: standard deviation; PLL: poly-
Size of polyplexes formed with plasmid DNA at indicated C/P ratios in HBS.
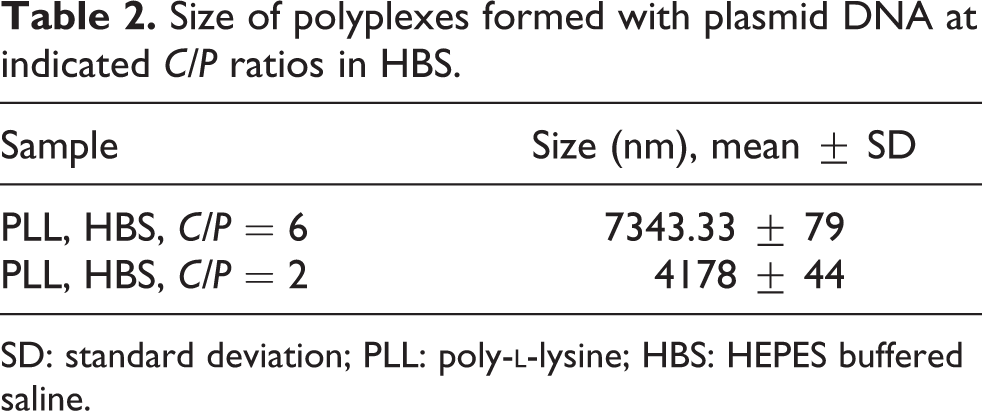
SD: standard deviation; PLL: poly-
Cytotoxicity
Figure 1 presents cell viability at different PLL concentrations. It was indicated that all PLL concentrations, except for 3.9 μg/ml, exerted a notably lowering effect on the number of Neuro2A cells (p < 0.001). When it comes to the influence of complex sizes on the cell viability, the results revealed that the largest complex (C/P 6 in HBS buffer) was associated with almost no negative impact on the cells. In other words, the complexes with 263- and 215-nm size caused 16.0 and 21.3% decline in the metabolic activity, respectively, in relation to the control (untreated cells) and larger complexes induced less toxicity (Figure 2).
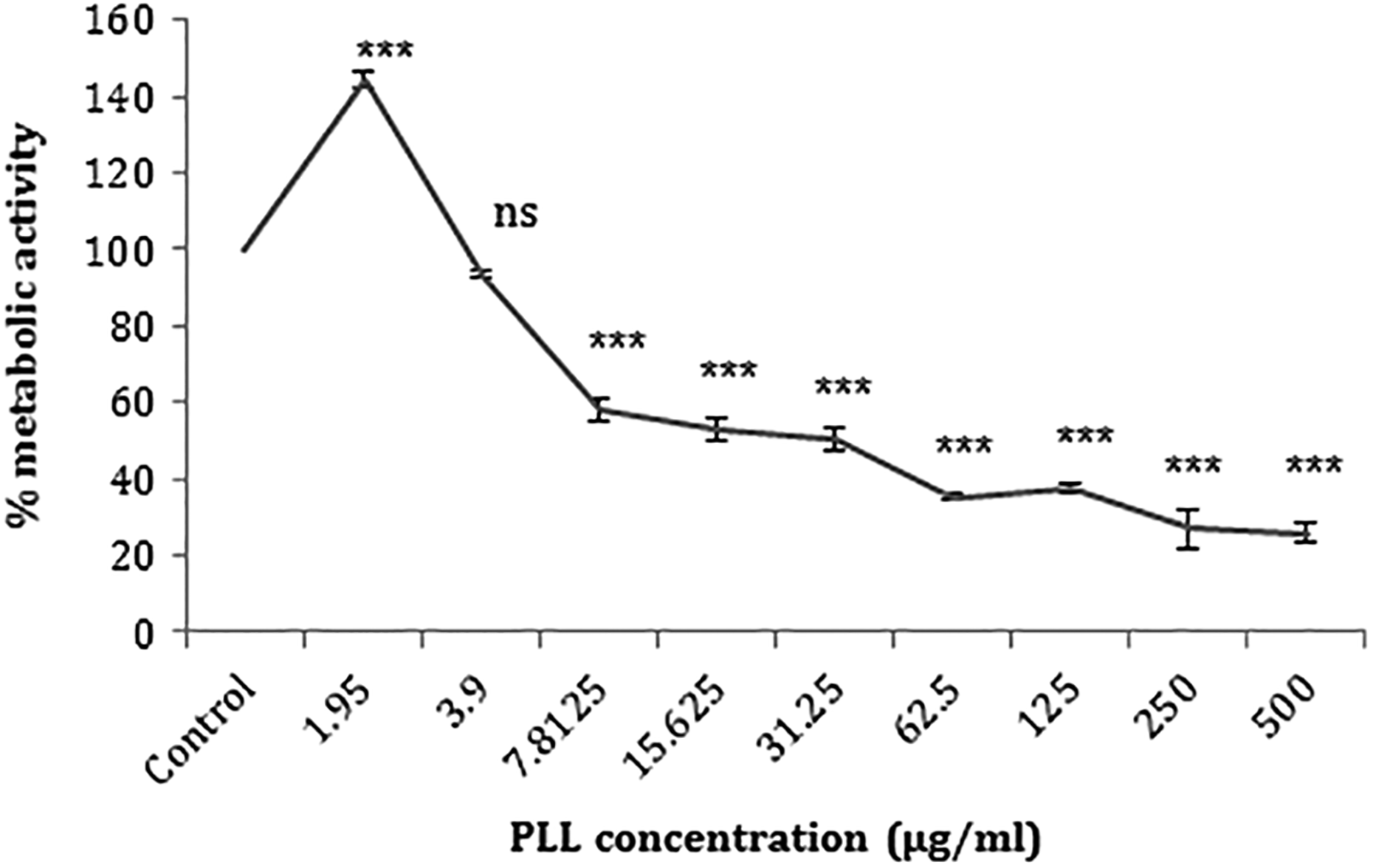
The effects of PLL concentration on the viability of Neuro2A cells are presented. Cells were treated with increasing concentrations of PLL for 24 h. The percentage cell viability (quantitated by MTT assay) was compared to the negative control (0 µM). Each point was expressed as a mean value ± SD in triplicate experiments. ***p < 0.001 as compared with control (ns: not significant). PLL: poly-l-lysine; SD: standard deviation.

Effect of PLL/DNA complexes size on the viability of Neuro2A cells after 4 h. Each point was expressed as a mean value ± SD in triplicate experiments. *** and ### indicate significant difference from untreated control. ***p ≤ 0.001. PLL: poly-
Apoptosis assay
PLL-induced apoptosis was examined using the PI flow cytometric analysis to determine the cell membrane integrity as well as DNA fragmentation. In Figure 3, the findings indicated the percentage of the sub-G1 peak after a 4-h treatment of the cells with different complexes. These data were representative of the peak area occurred prior to the G1 peak (Figure 4). Overall, the sub-G1 peak of the complexes was considerably lower than that of the control (p < 0.001). There was an inverse correlation between the polyplex sizes and the number of apoptotic cells; put it differently, with an increase in the size of PLL/DNA complexes from 215.1 nm to 7343.3 nm, the percentage of the sub-G1 peak diminished from 24.8% to 18.9%.

Flow cytometric analysis of cell membrane integrity and DNA fragmentation of Neuro2A cells treated with different sizes of PLL polyplexes (a: 215.1 nm, b: 262.97 nm, c: 4178 nm, and d: 7343 nm) after 4 h. ***p < 0.001 as compared with control. PLL: poly-

Flow cytometric analysis of cell membrane integrity and DNA fragmentation of Neuro2A cells treated with PLL complexes after 4 h. Histograms show the intensity of fluorescence in FL-2 channel, which corresponds to PI emission wavelength. Histogram representative of (a) PLL/DNA (C/P 2) in HBG buffer with 262.97 nm; (b) PLL/DNA (C/P 2) in HBS buffer 4178 nm; (c) PLL/DNA (C/P 6) in HBG buffer 215.1 nm, and (d) PLL/DNA (C/P 6) in HBS buffer 7343.33 nm. PLL: poly-
Comet assay
The single cell electrophoresis (i.e. COMET) assay was employed to determine the degree of DNA damage. Figure 5 displays the COMET assay images of Neuro2A mammalian cells exposed to two concentrations of PLL, 4 and 12 µg/ml, as well as two weight ratios of PLL/DNA complexes, C/P 2 and C/P 6. The untreated cells served as the control sample. It was demonstrated that all the four samples were observed with the significantly increased apoptotic tail as compared to the control (4.6%; p < 0.001; Figure 6).

The single cell gel electrophoresis (COMET assay) for detection of DNA damage induced by two concentration of free PLL; 4 and 12 µg/ml and also two PLL/DNA complexes ratio of 2 and 6 (C/P 2 and C/P 6). PLL: poly-
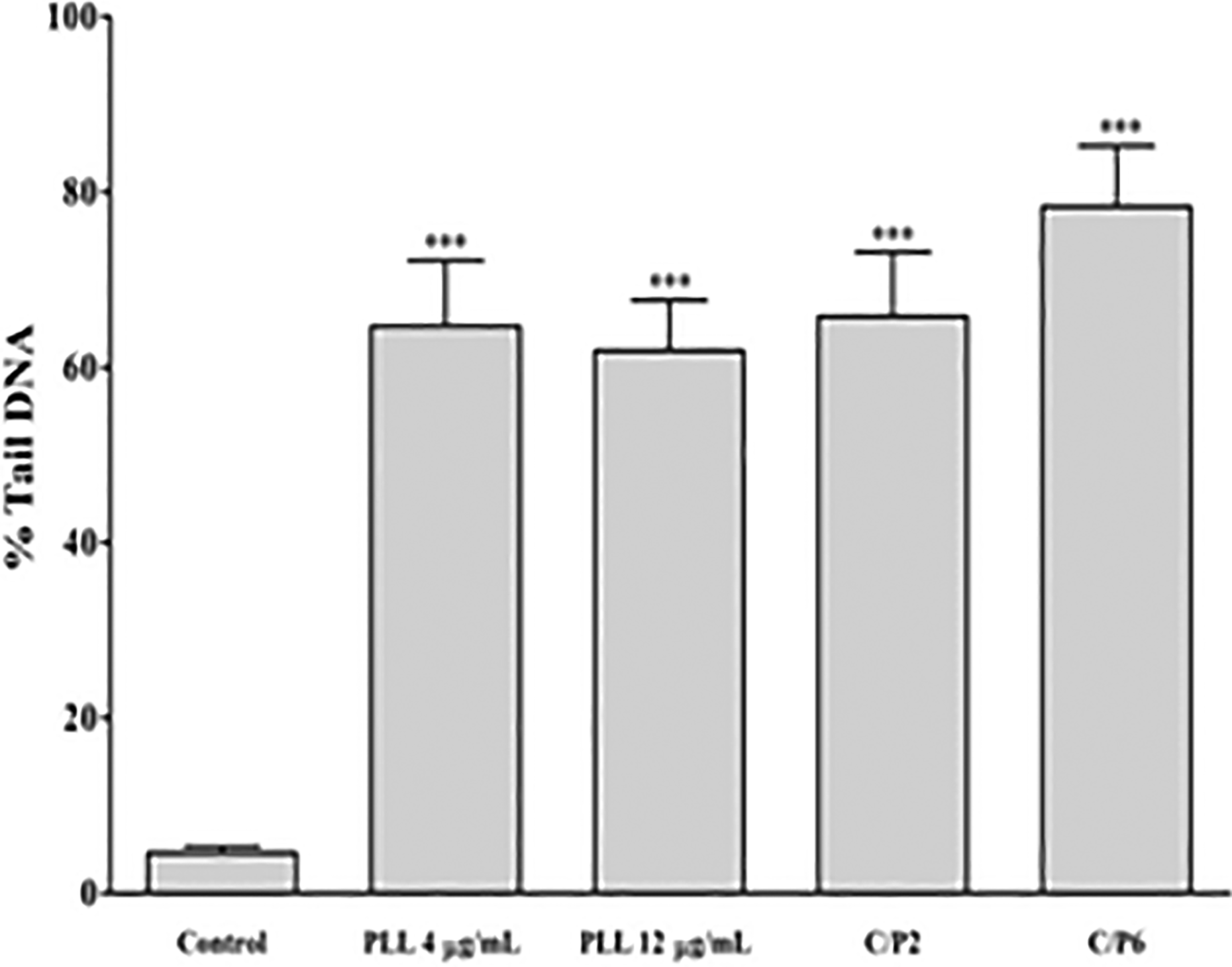
%Tail DNA induced by PLL in Neuro2A cells after 24 h. All data were represented as the means ± SEM of three independent experiments. ***p < 0.001: as compared with control. PLL: poly-
Indeed, free PLL carriers at the concentrations of 4 and 12 µg/ml induced approximately 64.7 and 61.9% tail DNA, respectively. Forming complexes with DNA also enhanced tail DNA content to 65.8% (C/P 2) and 78.4% (C/P 6).
Discussion
Among factors that should be considered for the use of polycations as gene delivery vectors, cell cytotoxicity has garnered more attention. 33,34 This study investigated PLL nanoparticle-induced toxicity at both cellular and molecular levels using three different testing methods, including MTT assay for metabolic activity, comet assay for genotoxicity, and flow cytometry for apoptosis detection. The findings of the MTT assay showed that cell death occurred at PLL concentrations higher than 7.8 μg/ml, which was corroborated by Strand et al.’s study that reported 10 μg/ml as an initial concentration associated with cell death. 35 The maximal reduction of cell viability (around 26.0%) was observed when PLL was utilized at the highest concentration (500 µg/ml). What is more, the effect of PLL on cell viability is dose dependent. As well-documented, cationic polymers, particularly those at higher molecular weights, tend to accumulate excessively onto the outer cell membrane, which, in turn, leads to toxicity and necrosis. 36 Indeed, the presence of positive charge makes cationic polymers, such as PLL binds to the negatively charged biological materials, including DNA, cell membrane phospholipids, cell membrane proteins, and blood proteins. 36 Therefore, molecular weight and charge density can determine the interaction between polycations with the cell membrane and the subsequent damage. 37
The efficacy and biosafety of nonviral carriers rely on their physicochemical and biological factors, namely the structure, ratio, charge, and size of DNA/vector complexes along with other parameters, such as time of exposure, type of the targeted cell/tissue, and interaction with circulating agents in plasma if applied as in vivo. 28 This is why nanoparticles in HBS buffer tended to form larger complexes than in HBG buffer and accordingly induced less cytotoxicity. Indeed, the hydrophobic aggregation of the complexes arises from increased solvent polarity that is more likely to cause increases in sizes. 25 Apoptosis is a programmed cell death that causes changes in cell morphology, including the translocation of phosphatidyl serine from the inner to the outer leaflet of a cell membrane, reduction of nuclear size, and elevation of DNA fragmentation. 38 The precise mechanism of cytotoxicity is not clear yet, but these effects are mostly mediated by electrostatic interaction of polycations with negatively charged cell membrane and followed by activation of intracellular signal transduction pathway and interfere with cell membrane integrity and function. 39,40 The cytotoxicity of PLL was not dedicated to lack of microtubules and microfilaments construction, so polymer uptake was not a compulsive condition for the observed cytotoxicity. 40 Fischer et al. reported that exposure of polymer with cells create cell membrane and succedent nuclear membrane disintegration, so it promoted DNA fragmentation as a remarkable feature of apoptotic pathway. 37,38 Due to Symonds et al. PLL polymer in both high and low molecular weight, capable of incepting apoptosis through mitochondrial pathway. Their result showed that PLL and PLL/DNA complexes cause late-phase mitochondrial apoptosis, which was exhibited in cytoplasm after 24-h exposure. 41 The results of the present study highlighted that PLL/DNA complexes induced apoptosis, which was consistent with previous findings reported by Symonds et al. that there are two kinds of cytotoxicity pertained to PLLs: early plasma membrane damage and late-phase apoptosis. 41 On the other hand, it was observed that high ratios of DNA/vector complexes lead to great levels of genetic instability as well as DNA damage in cells, characterized by DNA tail. 42,43 This was in agreement with other studies showing that such complexes at different C/P ratios increased genotoxicity. 44,45 Likewise, there have been some reports demonstrating that cationic polymers act as an apoptotic agent and augments the levels of caspase 3, caspase 9, and cytochrome c. 46,47 Akin to our findings, Choi et al. observed that cationic polymer-induced genotoxicity is dose dependent. Indeed, they concluded that with increasing the polymer concentration, the DNA damage mechanism has shifted from chromosomal damages to the necrotic pathway. 44 In another study in 2017, different cytotoxic effects of 20 types of polymer were investigated on blood and immunological cell. Due to their result, interaction of polymer molecules with cell membrane and its subsequent effects damaged cell membrane and induce apoptotic cell but not necrotic cells. 48 Chen et al. evaluated the effect of surface charge of hyper branched PLL on cell cytotoxicity in mice model. Their report confirm that positively charged surface was more susceptible for opsonin protein coverage; therefore, these surfaces more observable to phagocyte cells. 49
Conclusions
To the best of our knowledge, this is the first study dealing with the cytotoxicity and genotoxicity of PLL as gene-carrier polymers. These findings indicated that PLL/DNA complexes caused cytotoxic, apoptotic, and genotoxic effects in a dose-dependent and weight ratio-dependent manner, which also affected the size of polyplexes.
Footnotes
Authors’ note
This study was a part of the Pharm. D. thesis of E. Alinezhad-Mofrad.
Acknowledgements
The authors would like to thank the Vice Chancellor of Mashhad University of Medical Science for providing financial support to carry out this project.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Vice Chancellor of Mashhad University of Medical Science for carrying out this project under Grant No. 88596.

