Abstract
The aim of the study was to examine whether antioxidant properties of 3,4,4′,5-tetramethoxystilbene (DMU-212) contribute to its anticarcinogenic activity and whether DMU-212 affects the expression of apoptosis-related genes. Two-stage model of hepatocarcinogenesis was used; male Wistar rats were challenged with N-nitrosodiethylamine (NDEA), 200 mg/kg body weight (b.w.), intraperitoneal, then phenobarbital (PB) in drinking water (0.05%) was administered. Simultaneously, DMU-212 was given per os at a dose 20 or 50 mg/kg b.w. two times a week for 16 weeks. DMU-212 caused a moderate decrease in hepatic thiobarbituric acid reactive substances and protein carbonyls concentration elevated in rats treated with NDEA/PB. The activity of antioxidant enzymes examined reduced by NDEA/PB treatment was not restored in rats coadministered with DMU-212. Effects of DMU-212 on messenger RNA (mRNA) expression of antioxidant enzymes in rats challenged with NDEA/PB were diversified; no changes in their protein expression were noted in any of the groups. The expression of 17,000 genes was analyzed by Affymetrix® Rat Gene 1.1 ST Array; 15 apoptosis-related genes were selected and validated by RT-q PCR. The combined treatment with NDEA/PB and DMU-212 increased the mRNA level of some genes driving mitochondria-mediated apoptosis, whereas the mRNA expression of some anti-apoptotic genes triggering receptor-mediated apoptosis was reduced. The expression of genes encoding caspases-4, -8, -9, and -12 was also increased in rats treated with DMU-212. Although antioxidant effect of DMU-212 in rats challenged with NDEA/PB was moderate, its potential anticarcinogenic properties were demonstrated as evidenced by modulation of apoptosis-related genes.
Keywords
Introduction
It has been shown that resveratrol (3,4′,5-trans-trihydroxystilbene) displays potent antioxidant and anti-inflammatory properties which play an important role in protecting against carcinogenesis. 1 However, its efficacy in living organism is limited because of a low bioavailability due to extensive biotransformation. Hence, trihydroxystilbene scaffold has become the subject of structural alterations to improve pharmacokinetic properties and anticancer activity. 2,3 One of the most promising resveratrol analogue investigated so far is 3,4,4′,5-tetramethoxystilbene (DMU-212), which shows potent anticancer activity superior to the parent compound. The mechanism of its activity includes a decrease in the expression of enzymes catalyzing activation of procarcinogens, induction of cell cycle arrest, and apoptosis. 2 –5 However, the data on anticancer activity of DMU-212 have been derived mainly from in vitro experiments and very few studies using animal models were performed thus far. 3,6
Oxidative stress may cause DNA, protein, and lipid damage leading to chromosome instability, mutation, and impairment of cell growth and thus it may result in carcinogenesis. 7
The response of cells to oxidative/electrophilic stress involves activation of defense mechanisms that lead to the induction of cytoprotective enzymes. Induction of genes encoding antioxidant enzymes is regulated by DNA element designated as antioxidant response element (ARE). Two proteins participate in the transcriptional activation of ARE-dependent genes: nuclear factor erythroid-2-related factor 2 (Nrf2) transcription factor and Kelch-like erythroid cell-derived protein with CNC homology (ECH)-associated protein 1 (Keap1). 8 The gene families regulated by Nrf2 pathway include phase II enzymes, antioxidant enzymes, NADPH-generating enzymes, and DNA repair enzymes. 9 Since increased oxidative stress has been implicated in the carcinogenic process, Nrf2-regulated antioxidant enzymes would be expected to play an essential role in reducing the level of reactive oxygen species (ROS)-derived DNA adducts and redox-sensitive signaling of tumor promotion. 10
In the attempt to determine whether anticancer activity of DMU-212 is related to its antioxidant activity, we investigated some parameters of antioxidant status in the liver of rats challenged with model carcinogen N-nitrosodiethylamine (NDEA). It is known that biotransformation of NDEA results in the generation of ROS. 11 As oxidative damage to RNA may cause errors in protein synthesis or dysregulation of gene expression, we examined the expression of both mRNA and protein of some antioxidant enzymes.
During tumorigenesis, cells develop mechanisms allowing them to escape their cancer suppression system. A characteristic feature of cancer cells is deregulation of the main apoptotic pathways. Some authors have suggested a cross-link between Nrf2 and some proteins involved in apoptosis stimulation. According to some data reported, Nrf2 inhibition is necessary for p53-mediated apoptosis. Another link between Nrf2 activation and apoptotic pathways is the Nrf2-mediated regulation of proteins from Bcl2 family. 12 Hence, we have examined the effect of DMU-212 alone and DMU-212 combined with NDEA/PB challenge on the expression of a panel of apoptosis-related genes in the liver of rats.
Materials and methods
Chemicals and antibodies
3,4,5,4′-Trans-tetramethoxystilbene (DMU-212) was synthesized in the Department of Chemical Technology of Drugs, Poznan University of Medical Sciences by partial methylation of trans-resveratrol as described by Murias et al. 13 Its structure and purity were confirmed by gas chromatography–mass spectrometry and nuclear magnetic resonance spectroscopy. 13
The reagent kit for protein carbonyls assay was purchased from BioCell Corp. Ltd. (New Zealand). Normal melting point agarose (Prona Agarose) was purchased from Hispanagar, Spain. All other chemicals were from Sigma-Aldrich (St. Louis, Missouri, USA) or from local chemical suppliers.
Primary and secondary antibodies against Nrf2, superoxide dismutase (SOD), glutathione peroxidase (GPx), catalase (CAT), glutathione reductase (GR), glutathione-S-transferase (GST) and lamin were supplied by Santa Cruz Biotechnology (Santa Cruz, California, USA).
Experimental design
Two-stage model of hepatocarcinogenesis, initiation by NDEA and promotion by PB, was applied. 14,15 Male Wistar rats, 240 ± 10% g, bred at the Department of Toxicology, Poznan University of Medical Sciences were used. The rats were housed in animal facility, at 22 ± 2°C with a 12-h light/12-h dark cycle, controlled humidity and circulation of air, and fed certified laboratory feed (Labofeed H, ISO 22000). Animals were divided randomly into five groups, eight animals in each group, and subjected to the following treatment. Rats in group II were administered DMU-212 by gavage at a dose 50 mg/kg b.w. 2 days a week for 16 weeks. To improve bioavailability, the compound was suspended in the 40% solution of 2-hydroxypropyl-β-cyclodextrin. 16 Groups III, IV, and V were given intraperitoneally a single dose of NDEA, 200 mg/kg b.w., followed by promotion with PB at a concentration of 0.05% in drinking water. Promotion was started 2 weeks after NDEA injection and continued till the end of the experiment, that is, for 14 weeks. Rats in groups IV and V were administered DMU-212 by gavage at doses 20 and 50 mg/kg b.w., respectively, 2 days a week for 16 consecutive weeks. Rats in group I (controls) were given vehicle by gavage (40% solution of 2-hydroxypropyl-β-cyclodextrin) 2 days a week for 16 weeks.
After 16 weeks following the NDEA injection, the animals were anesthetized by ketamine and xylazine, the blood was withdrawn from the heart, livers were excised, rinsed with ice-cold 1.5% potassium chloride and stored in −80°C until analysis. Cytosol and microsomal fractions of liver tissue were prepared by differential centrifugation according to the standard procedure as described previously. 17
The experiment was performed according to the Local Animal Ethics Committee (LAEC) guidelines for animal experimentation and permission from LAEC was obtained prior to the experiments.
Biochemical assays
The level of lipid peroxidation was determined in the liver microsomes and expressed as the thiobarbituric acid reactive substances (TBARS) concentration. 18 Reduced glutathione concentration was measured in liver homogenates with Ellman’s reagent. 19 Protein carbonyl concentration in the liver was determined by an enzyme-linked immunosorbent assay (ELISA) method according to the producer instruction.
Alkaline comet assay in the liver tissue was performed according to the method of Hartmann and coworkers. 20 The comets assessed under a fluorescence microscope were divided into five groups differing in the degree of DNA damage and the results were expressed as a total damage score for the sample. 21
Antioxidant enzymes were assayed in the liver cytosol. SOD activity was determined on the basis of the inhibition of spontaneous epinephrine oxidation. 22 CAT activity was determined by monitoring the rate of hydrogen peroxide decomposition. 21 GPx activity was assayed according to Mohandas and coworkers. 23 Hydrogen peroxide was used as a substrate. The disappearance of reduced form of nicotinamide adenine dinucleotide phosphate (NADPH) was a measure of the enzyme activity. GR activity was assayed by measuring NADPH oxidation in the presence of oxidized glutathione. Total GST activity was determined using 1-chloro-2,4-dinitrobenzene as a substrate. 23
Microarray expression analysis and validation by real-time quantitative PCR
RNA isolation was performed using TRI reagent (Sigma-Aldrich), and the RNA was then purified using columns (RNeasy Mini Kit; Qiagen, Germany). The RNA integrity and quality were checked on Bioanalyzer 2100 (Agilent Technologies, Inc., Santa Clara, Caifornia, USA). Total RNA from each sample was subjected to two round sense complementary DNA (cDNA) amplification (Ambion® WT Expression Kit; Thermo Fisher Scientific, Waltham, Massachusetts, USA). Obtained cDNA was used for biotin labeling and fragmentation by AffymetrixGeneChip® WT Terminal Labeling and Hybridization (Affymetrix, Santa Clara, California, USA). Biotin-labeled fragments of cDNA (5.5μg) were hybridized to Affymetrix Rat Gene 1.1 ST Array Strip (48°C/20 h). The array strips were scanned by Imaging Station of GeneAtlas System. Preliminary analysis of the scanned chips was performed using Affymetrix GeneAtlas TM Operating Software. The quality of gene expression data was checked according to quality control criteria provided by the software.
The results of microarray assay were validated by real-time quantitative PCR (RT-qPCR). RT-qPCR analyses were conducted in a LightCycler® Instrument 480 MultiwellPlate 96 (Roche, Mannheim, Germany) using a LightCycler 480 Probes Master kit. Target cDNA was quantified using the relative quantification method. In the case of negative control, cDNA was not added. For RT-qPCR analysis of selected transcript levels, 1 µL of total (20 µL) cDNA solution was added to the mixture of primers, probes and LightCycler 480 Probes Master kit (Roche, Mannheim, Germany). The quantity of transcripts in each sample was standardized by Actb and Gapd mRNA levels. Primer sites and probe sites were selected with the aid of the Universal Probe Library Assay Design Center of Roche Applied Science.
Nrf2 binding assay
The cytosol and nuclear extracts from rat liver were prepared using the Nuclear/Cytosol Fractionation Kit (BioVision Research, Milpitas, California, USA). Whole cell lysates were prepared from rat liver using radioimmunoprecipitation assay buffer. Nrf2 activation was assessed by an enzymatic immunoassay using the Nrf2 transcription factor ELISA assay kit (Active Motif, Rixensart, Belgium) according to the manufacturer’s instructions. Nuclear extracts were incubated in the oligonucleotide-coated wells where the oligonucleotide sequence contains the ARE consensus binding site (5′-GTCACAGTGACTCAGCAGAATCTG-3′). Then, wells were washed and incubated with the antibody against Nrf2. Subsequent addition of an HRP-conjugated secondary antibody provided a sensitive colorimetric read-out at 450 nm.
Western blot analysis
For the determination of the level of Nrf2, GST, SOD, CAT, GPx, and GR proteins the immunoblot assay was used. All the experiments were repeated three times. Whole cell lysates (GPx), nuclear extracts (Nrf2), and cytosolic proteins (Nrf2, GST, SOD, CAT, and GR) (100 μg) were separated on 12% and 10% sodium dodecyl sulfate polyacrylamide gel electrophoresis slab gels and proteins were transferred to nitrocellulose membranes. After blocking with 10% skimmed milk, the proteins were probed with rabbit polyclonal Nrf2, goat polyclonal antibody GST, goat polyclonal antibody SOD, rabbit polyclonal GPx, rabbit polyclonal GR, goat polyclonal CAT, and rabbit polyclonal lamin. The lamin protein was used as an internal control. As the secondary antibodies in the staining reaction, the alkaline phosphatase-labeled anti-rabbit IgGs were used. Bands were visualized with BioRad AP Conjugate Substrate Kit NBT/BCIP (BioRad Laboratories, Hercules, California, USA). The amount of the immunoreactive product in each lane was determined using the Quantity One software. Values were calculated as relative absorbance units (RQ) per milligram protein.
Immunohistochemistry and antibodies
Material for immunohistochemistry was fixed in Bouin solution for 24 h, dehydrated, and embedded in paraffin. For light microscopy, sections of 5 µm thickness were dewaxed and processed according to the indirect avidin–biotin/peroxidase (ABC) technique. In order to demonstrate the protein expression of carcinogenesis-related genes, the following antibodies were used: retinoblastoma protein (Rb1) (Santa Cruz Biotechnology, Heidelberg, Germany, sc-73598, diluted 1:100), monoclonal mouse anti-human anti-gamma-glutamyltransferase (GGT), Santa Cruz Biotechnology,sc-100746, diluted 1:100), monoclonal mouse anti-human anti-Bcl2 (Santa Cruz Biotechnology, sc-7382, diluted 1:100), monoclonal mouse anti-human anti-p53 (Novus Biologicals, Littleton, Colorado, USA; NBP1-4119, diluted 1:100), polyclonal rabbit anti-human anti-transforming growth factor (TGF) α (Novus Biologicals, NB-25561, diluted 1:50), monoclonal mouse anti-human anti-Ki67 (Dako, Copenhagen, Denmark, M7248, diluted 1:200). All antibodies were primarily tested and standardized on control rats tissues in order to check for the cross-species reactivity (human vs. rat).
The reactions were observed with a Nikon light microscope (Japan) and documented using a charge-coupled device (CCD) camera (arc6000c; Baumer Optronic, Radeberg, Germany). All the immunohistochemical reactions were performed blind on coded samples.
Statistical analysis of immunohistochemical findings
The statistical analysis of the level of protein expression was based on nonparametric Q Cochran test. It was used to verify the difference in the percentage of immunopositive cells noted in various groups of animals. Significance was defined at p < 0.05. On the basis of the results of H + E staining, which was analyzed through the use of MicroImage (Olympus, Japan) morphometric software, the content of reaction-positive cells was determined by comparing the area of immunopositive cells to the total area of organ surface present in the specimen. The statistical analysis was conducted using Statistica PL v 9.0 software.
Statistical analysis of biochemical and molecular assays
The GraphPad InStat statistical package (version 3) was used. The mean values and standard deviations were calculated. One way analysis of variance followed by the Student–Newman-Keuls test for multiple comparisons was used. A value of p < 0.05 was considered the limit of significance.
Results
Parameters of oxidative stress
In the liver of rats treated with NDEA/PB the increase in the level of oxidative damage markers was noted (Figure 1). Microsomal lipid peroxidation level expressed as TBARS increased by 50%, the content of protein carbonyl groups by 42% and DNA damage by 100%. Administration of DMU-212 to rats challenged with NDEA/PB caused a moderate attenuation of lipid peroxidation by 30% and 17%, and a decrease in the concentration of protein carbonyl groups by 16% and 30%, respectively, for doses 20 and 50 mg/kg b.w. However, the level of NDEA-induced DNA damage was not changed. DMU-212 alone administered at a dose 50 mg/kg did not modulate any of these markers.

Effects of DMU-212 on markers of oxidative damage in the liver of rats treated with NDEA/PB. Data from eight animals are presented as mean ± SD. * and # above the bar denote statistically significant difference from the control and NDEA/PB group, respectively; p < 0.05. NDEA: N-nitrosodiethylamine; PB: phenobarbital.
The content of GSH was decreased in the liver of NDEA/PB-treated rats by 33% and was restored in the rats administered simultaneously both doses of DMU-212. The compound tested alone caused slight but not statistically significant increase in GSH concentration (19%; Figure 1).
The activity of antioxidant enzymes examined was significantly decreased in the rats treated with NDEA/PB, by 32–77% (Table 1). A similar effect was observed for GPx and GR in the rats administered DMU-212 alone. Administration of the compound tested to the rats challenged with NDEA/PB did not affect markedly the activity of these enzymes as compared to the results in NDEA/PB group. The only changes observed included a further decrease in SOD activity (50%) in the rats treated with a lower dose of DMU-212 and a diversified response of GPx to the lower and the higher dose of the compound: 38% increase and 35% decrease, respectively. The other changes were of small magnitude and did not show statistical significance.
Effect of DMU-212 on the activity of antioxidant enzymes in liver of rats treated with NDEA/PB.a

SOD: superoxide dismutase, CAT: catalase; GPx: glutathione peroxidase; GR: glutathione reductase; GST: glutathione S-transferase; NADPH: reduced form of nicotinamide adenine dinucleotide phosphate; CDNB: 2,4-dinitrochlorobenzene; NDEA: N-nitrosodiethylamine; PB: phenobarbital.
aResults are mean + SD, n = 8. Control rats were administered vehicle. Values in brackets express % of change. Activity of enzymes is expressed as follows: SOD and CAT (U/mg protein); GPx and GR (nmol NADPH/min/mg protein); GST (nmol CDNB/mg protein).
b p < 0.05: the DMU-212 and NDEA/PB-treated groups are compared with controls.
c p < 0.05: the NDEA/PB-treated group is compared with the DMU-212 + NDEA/PB-treated groups.
Expression of mRNA and protein of antioxidant enzymes
A comparison of the results of mRNA and protein expression in the liver of rats revealed some discrepancy (Figure 2). In each experimental group, no significant changes in the expression of antioxidant enzymes protein as compared to control group were noted. On the contrary, the changes in relevant mRNA expression of some enzymes were observed. NDEA/PB treatment resulted in the increase in mRNA expression of SOD and GST by 44% and 49%, respectively. The expression of GPx mRNA was decreased by 40%, whereas that of CAT and GR remained unchanged. The effects of DMU-212 on mRNA expression in the NDEA challenged rats were diversified and depended on the dose of DMU-212. The lower dose of DMU-212 caused a decrease in mRNA expression by 70–92% as compared to that in the NDEA/PB group, except for GPx whose transcript level was increased by 38%. Administration of the higher dose of DMU-212 resulted in an increase in mRNA expression, by 24–69%, except for GPx whose transcript level was not changed.

Effects of DMU-212 on mRNA and protein of antioxidant enzymes expression in rat liver cells. (a) The RT-qPCR analysis of SOD-2, CAT, GPx-1, GR, and GST mRNA relative abundance. Data from six animals are presented as mean ± SD. * and # above the bar indicate statistically significant difference from the control and NDEA/PB group, respectively; p < 0.05. (b) Western blot analysis of SOD-2, CAT, GPX-1, GR, and GST protein levels; lane 1-controls, lane 2-DMU-212; lane 3-NDEA/PB; lane 4-NDEA/PB + DMU-212 (20 mg/kg); lane 5-NDEA/PB + DMU-212 (50 mg/kg). mRNA: messenger RNA: NDEA: N-nitrosodiethylamine; PB: phenobarbital; SOD: superoxide dismutase; GPx; glutathione peroxidase, CAT: catalase; GR: glutathione reductase, GST: glutathione-S-transferase; RT-qPCR: real-time quantitative PCR.
DMU-212 alone caused a marked decrease in the antioxidant enzymes mRNA expression by 24–95% (Figure 2).
Effect of DMU-212 on Nrf2 activation
To investigate the translocation of Nrf2 into the nucleus upon DMU-212 treatment, the levels of Nrf2 in cytoplasm and nuclear fractions were evaluated by Western blot analysis in the rat liver. As shown in Figure 3(b) and (c), the Nrf2 protein level increased slightly, by approximately 20% in the nucleus after the exposure to DMU-212 as compared to its level in NDEA/PB-treated rats, however, this difference was not statistically significant. Since the activation of Nrf2 is generally accompanied by the degradation of Keap1, the amount of Keap1 protein in cytosolic fraction was also measured. The amount of Keap1 protein was not affected by DMU-212 treatment (data not shown).
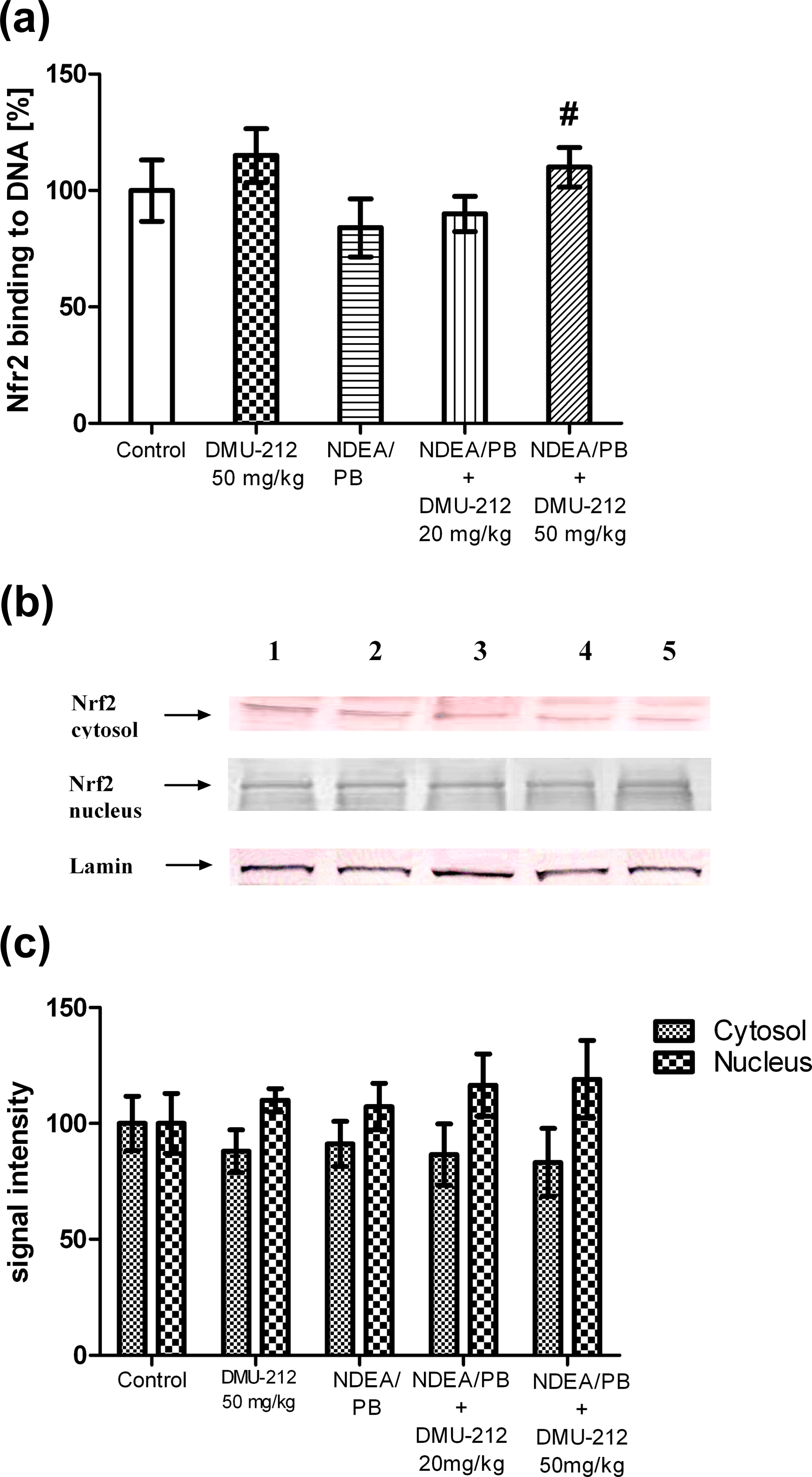
Effect of DMU-212 on Nrf2 activation in rat liver cells. (a) Binding of Nrf2 to ARE-containing oligonucleotides; (b) Representative immunoblots for the analysis of the cytosolic and nuclear level of the Nrf2 protein levels; lane 1-controls, lane 2-DMU-212; lane 3-NDEA/PB; lane 4-NDEA/PB + DMU-212 (20 mg/kg); lane 5-NDEA/PB + DMU-212 (50 mg/kg). (c) Densitometric analysis of the cytosolic and nuclear level of Nrf2 protein. Lamin was used as a normalization control. Data (mean ± SD) from six animals are presented (a, c), # above the bar denotes statistically significant difference from the NDEA/PB group, p < 0.05. NDEA: N-nitrosodiethylamine; PB: phenobarbital; Nrf2: nuclear factor erythroid-2-related factor 2; ARE: antioxidant response element.
The activation of Nrf2 was assessed by the measurement of its binding to a known ARE oligonucleotide sequence. The results show that the Nrf2 ARE binding was slightly increased in the DMU-212-alone treated rats (Figure 3(a)), whereas a statistically significant increase was observed in the rats given a combined treatment with DMU-212 (50 mg/kg) and carcinogen as compared to the level of binding in the NDEA/PB group.
The mRNA expression of apoptosis-related genes
In order to obtain a complete view on gene dynamics profile in the liver of rats treated with DMU-212 alone and DMU-212 combined with NDEA challenge, we performed the whole gene expression analysis by Affymetrix® Rat Gene 1.1 ST Array by which we examined expression of 17,000 different genes. The 15 genes were selected for results validation by RT-qPCR. The results from Rt-qPCR validation are presented in Figures 4 to 6.

Effect of DMU-212 on the mRNA expression of genes triggering mitochondria-mediated apoptosis in rat liver. mRNA relative abundance of Apaf-1, Diablo, Bak-1, Bad, Pten, and Stat1 was analyzed by RT-qPCR. Data (mean ± SD) from six animals are presented. * and # above the bar denote statistically significant difference from the control and NDEA/PB group, respectively; p < 0.05. mRNA: messenger RNA; RT-qPCR: real-time quantitative PCR; NDEA: N-nitrosodiethylamine; PB: phenobarbital.

Effect of DMU-212 on the mRNA expression of genes driving receptor-mediated apoptosis in rat liver cells. The relative abundance of Socs-2, Tnfsf10, Tnfrsf1a, Birc-3 and Tnfsf11 mRNA was assayed by RT-qPCR. Data from six animals are expressed as mean ± SD, * and # above the bar indicate statistically significant difference from the control and NDEA/PB group, respectively; p < 0.05. mRNA: messenger RNA; RT-qPCR: real-time quantitative PCR; NDEA: N-nitrosodiethylamine; PB: phenobarbital.

Effect of DMU-212 on mRNA levels of genes encoding Casp-4, -8, -9, and -12 in rat liver. mRNA relative abundance was analyzed by RT-qPCR. Data (means ± SD) from six animals are presented, * and # above the bar denote statistically significant difference from the control and NDEA/PB group, respectively; p < 0.05. mRNA: messenger RNA; RT-qPCR: real-time quantitative PCR; NDEA: N-nitrosodiethylamine; PB: phenobarbital.
The mRNA expression of genes driving mitochondria-mediated apoptosis is shown in Figure 4. The treatment with DMU-212 alone decreased Apaf-1, Stat-1, and Pten transcripts level by 52–75% as compared to those in the control group of animals. The differences in Diablo, Bak1, and Bad mRNA expression were not statistically significant. The NDEA/PB treatment regimen enhanced Stat1 and Pten mRNAs expression by 48% and 92%, respectively, as compared to control. Concomitantly, the expression of Bak-1 mRNA was decreased by 69%. The differences in Apaf-1, Diablo, and Bad transcripts level were not statistically significant. The combined treatment with NDEA/PB and the lower dose of DMU-212 suppressed the mRNA levels of Apaf-1, Diablo, Bad, Stat1, and Pten genes, except for Bak-1 whose transcript level was upregulated by 55%. Administration of the higher dose of DMU-212 led to a significant increase in the expression of all proapoptotic genes driving mitochondria-mediated apoptosis.
Figure 5 presents the mRNA expression of genes triggering receptor-mediated apoptosis. The application of NDEA/PB treatment protocol resulted in a slight increase in the mRNA expression of proapoptotic Tnfsf10 gene by 58% as compared to that in the control group. Concomitantly, proapoptotic Tnfrsf1a transcript level was reduced by 43%. No change in the mRNA expression of proapoptotic Socs-2 gene was observed. The differences in anti-apoptotic Birc-3 transcript level was not statistically significant. Concomitantly, the expression of Tnfsf11was decreased by 83%.
The compound tested alone caused a decrease in the Tnfsf10 and Tnfsf1a mRNAs expression by approximately 50% as compared to that in the untreated control. On the contrary, the Socs-2 and Tnfsf11 transcript levels were increased by 78% and 225%, respectively.
The combined treatment with NDEA/PB and DMU-212 in the dose of 20 mg/kg reduced Tnfsf10 and Socs-2 transcripts level by 78% and 67%, respectively, as compared to the group treated with NDEA/PB. Concomitantly, Tnfrsf1a mRNA expression was slightly increased by 36%. The differences in the mRNA expression of antiapoptotic Tnfsf11 and Birc-3 genes were not statistically significant. However, the higher dose of DMU-212 decreased Birc-3 and Tnfsf11 mRNAs expression by approximately 50%. The expression of proapoptotic Tnfsf10 and Socs-2 transcripts level was significantly increased by 158–200% as compared to the group treated with NDEA/PB. We did not observe any changes in the Tnfrsf1a mRNA expression.
Figure 6 shows mRNA levels of genes encoding Casp-4, -8, -9, and -12. Treatment with DMU-212 alone decreased the Casp-9 mRNA expression by 38%, whereas the transcript levels of Casp-4, -8, and -12 were not changed as compared to the control group. The NDEA/PB treatment increased the Casp-4 mRNA level by 96% in comparison with that in the untreated control, whereas the changes in Casp-8, -9, and -12 mRNAs expression were not statistically significant. Combining this protocol with the lower dose of DMU-212, resulted in a decrease in Casp-4, -8, -9, and -12 transcripts level by 70–85% as compared to NDEA/PB group. Concomitantly, the higher dose of DMU-212 significantly increased the mRNA expression of all investigated caspases.
Immunohistochemical analysis
Liver sections taken from NDEA/PB-exposed animals revealed the expression of GGT, p53, Rb1, and TGFα. GGT and p53 protein were expressed in hepatocytes in the closest proximity of the central vein of hepatic lobules (Figures 7(a) and (c)). Individual Rb1-positive cells were regularly spread within the tested liver (Figure 7(e)). TGFα-positive cells were present exclusively in the peripheral hepatocytes of hepatic lobules (Figure 7(g)). The expression of two proteins examined, namely Bcl2—a marker of apoptosis and Ki67—the product of suppressor genes was observed neither in the treated nor in the control animals (data not shown).

Representative immunohistochemistry images from rat liver. Photomicrographs depict (a) GGT, (c) p53, (e) Rb1, and (g) TGFα expression in rats treated with NDEA/PB. Concomitant treatment with DMU-212 resulted in a decreased immunostaining of (b) GGT, (d) p53 and (f) Rb1. TNFα expression was not attenuated by DMU-212 administration (h). NDEA: N-nitrosodiethylamine; PB: phenobarbital.
Coadministration of NDEA/PB and the lower dose of DMU-212 resulted in a decreased expression of GGT (43 ± 3.8% of positive cells in NDEA/PB-exposed animals vs. 12 ± 2.9% positive cells in cotreated animals; p = 0.002; Figure 7(b)) and p53 (22 ± 2.1% of positive cells in NDEA/PB-exposed animals vs. 9.0 ± 4.3% positive cells in cotreated animals; p = 0.04; Figure 7(d)). Interestingly, the higher dose of DMU-212 did not affect GGT expression, but resulted in the reduction of p53 expression (22 ± 2.1% of positive cells in NDEA/PB-exposed animals vs. 14 ± 2.7% positive cells in cotreated animals; p = 0.04; data not shown) and in the lack of Rb1 expression (Figure 7(f)).
TGFα expression in the NDEA/PB-initiated rats was not attenuated by DMU-212 (28 ± 5.2% of positive cells in NDEA/PB exposed animals vs. 27 ± 2.7% positive cells in cotreated animals; p = 0.988; Figure 7(h)).
In the livers of control animals as well as in those from the rats treated exclusively with DMU-212 no expression of GGT, p53, TGFα, and Rb1 protein was observed (data not shown).
Discussion
Parameters of oxidative stress
Pro-oxidant effects of a single NDEA dose were observed after 16 weeks following administration, which has been confirmed by other authors applying the same model of rat carcinogenesis. There are many reports on a marked increase in the level of hepatic lipid peroxidation, by 2–10 fold, 15,24 –27 however, in some studies the extent of the increase in TBARS level was similar to that in our experiment, 65–82%. 14,28,29 The magnitude of hepatic glutathione depletion due to NDEA treatment observed by other authors was also similar to that found in our study, 37–52%. 14,15,28,29 On the other hand, the extent of hepatic protein oxidative damage reported by others was much higher than that observed in our study, as evidenced by 2.6–3.5-fold increase in the content of carbonyl groups. 24,25
Long-term administration of DMU-212 to rats initiated with NDEA moderately decreased the oxidative damage to hepatic lipids and proteins, however, DNA damage was not attenuated. DMU-212 administration resulted in restoring the content of GSH depleted in NDEA/PB-treated animals up to the level observed in the control group.
In consistence with our findings, other authors have also reported a decrease in the activity of antioxidant enzymes in the liver of rats treated with NDEA/PB regimen, by 16–60%. 14,26,29,30
DMU-212 alone did not affect the markers of oxidative damage but decreased the activity of antioxidant enzymes (except for SOD), however, these alterations were statistically significant only for GPx and GR. Some authors have suggested that such an effect can be due to an enhancement of antioxidant status by exogenous antioxidant. 31 This interpretation can be confirmed by the decreased expression of mRNA of all antioxidant enzymes examined in the liver of rats treated with DMU-212 alone.
DMU-212 treatment did not restore the activity of antioxidant enzymes in the rats challenged with NDEA/PB except for GST whose activity slightly increased (statistically insignificant change) and GPx whose activity was enhanced in the rats receiving the lower dose of the compound tested.
It is generally accepted that the response of cells to oxidative/electrophilic stress involves the activation of a battery of defensive genes in a process mediated by Nrf2-Keap1 system. Under normal conditions, Nrf2 is sequestered in the cytosol by the actin binding protein Keap1. When oxidative stress becomes prevalent, the interaction between Nrf2 and Keap1 is disrupted leading to an increase in Nrf2 translocation to the nucleus. In the nucleus Nrf2 binds to ARE and initializes the transcription of a battery of cytoprotective genes. Antioxidant enzymes are one of the families of enzymes whose transcription is mediated by Nrf2 pathway. 9
In our experiment, no increase in Nrf2 translocation to nucleus due to pro-oxidant action of NDEA was observed. Similar results have been reported by Bishayee and coworkers 24,25 who did not find a difference between Nrf2 protein expression in control and NDEA-treated rats. Only slight, statistically insignificant, increase in Nrf2 translocation to nucleus was noted in rats given a combined treatment as compared to that in NDEA alone-treated rats. Similarly, DMU-212 very slightly enhanced Nrf2-ARE binding in rats challenged with NDEA/PB. This observation correlates with the lack of the induction of antioxidant enzymes proteins, which are controlled by Nrf2, however, an increase in three enzymes mRNA expression was revealed in the rats treated with carcinogen and the higher dose of DMU-212. Thus, it could be suggested that another mechanism independent of Nrf2 pathway was involved in enhanced expression of these enzymes mRNA.
It is interesting that all alterations in mRNA level of antioxidant enzymes were not paralleled with their protein expression which did not change significantly in any of the groups of animals. Moreover, there was almost no correlation between the alterations in mRNA expression and the activity of enzymes in the majority of experimental groups, with the exception of GPx for which the direction of changes was consistent in three groups of animals. Other authors have also reported that the activities of antioxidant enzymes do not always correlate with their mRNA and protein levels. It is suggested that a certain threshold of mRNA must be achieved before the protein can be translated. 32 Moreover, some posttranscriptional and posttranslational modifications including proteasomal degradation may be involved in these consequences. 33 Hence, the alterations in gene transcription not always affect protein levels and the enzyme activity to the extent suggested by the magnitude of transcriptional changes. 34
Collectively, the results discussed suggest the moderate antioxidant action of DMU-212 in the liver of rats challenged with NDEA.
Generally, the activation of Nrf2 by chemical compounds and natural products is considered to be beneficial, however, the dark side of Nrf2 has been also revealed. It is hypothesized that upregulation of Nrf2 signaling can increase the expression of Notch1, a molecule responsible for cell fate, leading to an increase in proliferation rate and a more malignant phenotype. 35 In the recently published review Ganan-Gomez and coworkers 12 have presented the data demonstrating that Nrf2 activation and apoptosis are antagonistic events since Nrf2 is able to directly activate the transcription of some antiapoptotic proteins. In this context, the lack of Nrf2 activation in the liver of rats challenged with NDEA/PB and treated with DMU-212 cannot be interpreted as unfavorable event since at the same time a number of proapoptotic genes have been induced.
Apoptosis-related genes
Carcinogenesis is a multistage process that leads to uncontrolled clonal proliferation, resulting in the evolution of normal cells into precancerous or cancerous cells. 36 The activation of apoptosis pathways can be considered as a principal mechanism of eliminating tumor cells by cytotoxic drugs. 37 There are two major signaling pathways that mediate apoptosis in response to cancer therapy: the extrinsic (receptor-mediated) and intrinsic (mitochondria-mediated) pathways. 38 Our previously published results of in vitro studies have demonstrated the ability of DMU-212 to evoke apoptosis in both mitochondria- and receptor-mediated manner. 5 Hence, to identify the transcripts unique to DMU-212-induced apoptosis pathway, in the present experiment we investigated the expression level of 15 apoptosis-related genes following transcriptome expression analysis. Similar to our previous study, 5 we noted that the expression pattern of these genes was specific to both extrinsic and intrinsic pathways of apoptosis. Apaf-1 is an important component of an oligomeric apoptosome, which cleaves procaspase-9, triggering mitochondrial apoptosis. 39 We found that although DMU-212 alone decreased Apaf-1 and Casp-9 mRNAs expression, it significantly upregulated their transcripts level after NDEA/PB treatment regimen. Moreover, the combined treatment with NDEA/PB and the higher dose of DMU-212 resulted in an increase in mRNA levels of Bcl-2 family genes: Bak-1 and Bad as well as other proapoptotic Diablo, Pten, and Stat1 genes. We observed a similar tendency in the expression of proapoptotic Socs-2, Tnfsf10, Tnfsf1a, and Casp-8 genes triggering receptor-mediated apoptosis. Conversely, the antiapoptotic Birc-3 and Tnfsf11 transcripts level was decreased after the combined treatment with NDEA/PB and DMU-212 in the dose of 50 mg/kg. Generally, we observed an increase in the expression of proapoptotic genes after the combined treatment with NDEA/PB and the higher dose of DMU-212. In contrast, the antiapoptotic transcripts level was decreased.
Immunohistochemistry
To determine whether DMU-212 affects early events in NDEA-induced hepatocarcinogenesis, some proteins, markers of neoplastic changes were examined in the liver of rats. It is known that induction of GGT in preneoplastic foci represents an early event in hepatocarcinogenesis. DMU-212 decreased the GGT immunoreactivity enhanced in NDEA/PB rats, however, only the lower dose was effective.
The p53 protein, a very efficient inhibitor of cell growth, is activated in response to stress signals leading to DNA damage. P53 inhibits cell growth by arresting proliferation or inducing apoptosis, thus preventing malignant transformation. 40 Consistently, the expression of p53 was enhanced in carcinogen-treated rats while the coadministration of DMU-212 decreased the p53 immunoreactivity. This might be interpreted as a result of an improved oxidative status in the rat liver, although the level of DNA damage, as evidenced by comet assay findings, was not reduced.
The Rb1 protein plays an important role in maintaining chromosomal stability, inhibits cell cycle in cells with DNA damage and acts as a tumor suppressor. 41 Thus, the increase in Rb1 immunopositivity in rats treated by NDEA/PB could be considered a defense mobilization of the cells. Recently antiapoptotic function of Rb1 has been revealed. 42 Disappearance of Rb1 expression might suggest that preneoplastic cell survival was suppressed by DMU-212 in rats challenged by carcinogen.
Several reports have demonstrated the relationship between TGFα expression and malignant transformation. 43 TGFα stimulates the proliferation of hepatocarcinoma cells by activating the signaling pathway of the epidermal growth factor receptor. As TGFα expression in NDEA-treated rats was not attenuated by DMU-212 administration, it could be suggested that DMU-212 in the doses tested does not suppress hepatic cell proliferation.
Results of our study show that some of DMU-212 affected end points displayed a biphasic dose–response. Similar duality in response to different doses/concentrations has been observed for resveratrol, a prototype compound. A number of experiments regarding this type of response, mostly for in vitro studies, have long been reported and presented in reviews by Calabrese et al. 44 and Mukherjee et al. 45 For example, it has been found that resveratrol biphasically affected proliferation of human endothelial cells with a low-concentration stimulation and a high-concentration inhibition; proliferation of the spleen cells was inhibited by high concentration of resveratrol, whereas at lower concentrations the proliferation was markedly increased; resveratrol at low concentrations enhanced the expression of heme oxygenase-1 in human aortic smooth muscle cells but at higher concentrations this effect was diminished.
Biphasic dose response has been also demonstrated in only few experiments in animal model. It has been found that high doses of resveratrol promoted atherosclerosis in hypercholesterolemic rabbits or caused renal toxicity in rats but simultaneously reduced cancer progression. 45 Resveratrol exerted a protective effect on indomethacin-induced gastric ulcer in mice at a low dose, however, the opposite effect has been noted at a higher dose. 46
Our experiment revealed that in rats challenged with NDEA/PB only the higher dose of DMU-212 enhanced the expression of apoptosis-related genes and antioxidant enzymes genes. Hence, it could be suggested that DMU-212 might be considered a potential anticancer agent only in higher doses; the lower doses can promote survival of cancer cells.
Summing up, our findings revealed that antioxidant effect of DMU-212 in the liver of rats challenged with NDEA/PB was moderate. Generally, the higher dose of DMU-212 caused an increase in the expression of proapoptotic genes and reduced the antiapoptotic transcript levels. Immunohistochemical analysis confirmed that DMU-212 can positively modulate some early events in NDEA-induced hepatocarcinogenesis.
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article. This study was supported by a research grant of the Polish State Committee for Scientific Research No. 1802/B/P01/2008/35.
