Abstract
Polyethylenimine (PEI) is a polycation widely used for successful gene delivery both in vitro and in vivo experiments. However, different studies showed that PEI could be cytotoxic to transfected cells, and the mechanism of toxicity is poorly understood. Identification of PEI-interacting proteins may help in understanding the toxicity pathways. In this study, we investigated proteins that could interact with PEI in human colorectal adenocarcinoma cells (HT29). In order to identify the proteins interacting with PEI, PEI was immobilized to sepharose beads as solid matrix. The HT29 cell lysate were passed through the matrix. PEI-bound proteins were isolated, and further separation was performed by two-dimensional gel electrophoresis. After gel digestion, proteins were identified by matrix-assisted laser desorption/ionization–time-of-flight (TOF)/TOF mass spectrometry. Our data indicated that most of the identified PEI-interacting proteins such as shock proteins, glutathione-S-transferases, and protein disulfide isomerase are involved in apoptosis process in cells. Thus, although this is a preliminary experiment implicating the involvement of some proteins in PEI cytotoxicity, it could partly explain the mechanism of PEI cytotoxicity in cells.
Introduction
The concept of gene therapy for correction of genetic deficiencies was first introduced in 1972 by Friedmann and Roblin. 1 The successful gene therapy is largely dependent on the development of effective gene delivery vectors. Nowadays many different types of vectors have been developed for efficient delivery of therapeutic genes into living cells. 2 Generally, gene delivery vectors are classified into viral and nonviral vectors. 3 Viral vectors are very efficient in gene delivery; however, they suffer from many problems such as mutagenesis, toxicity, immunogenicity, capacity, and cost of large-scale production. These problems encouraged researchers to use nonviral vectors for gene delivery. Up to now, different types of nonviral vectors such as cationic polymers, 4 liposomes, 5 peptides, 6 cationic lipids, 7 and dendrimers 8 have been used for gene delivery. The cationic polymer polyethylenimine (PEI) has been widely used for gene delivery. 4,9 –11 Despite large studies on PEI-based vectors, the toxicity of this vector remains unknown. Although several mechanisms have been suggested that the toxicity of PEIs is related to direct membrane perturbation processes, 12 –14 it seems that PEI could induce mitochondrially mediated apoptosis. Studies in three clinically relevant human cell lines showed that PEI induced mitochondrial-mediated apoptotic cell death. The apoptotic process were linked to cytochrome c release, following activation of caspases 9 and 3. 14 Moreover, a recent study demonstrated that PEI mediated the mitochondrial membrane potential loss in a concentration- and time-dependent manner, impaired mitochondrial respiration, and also induced mitochondrial swelling. 15 During this process, PEI can interfere with different proteins within the cell. However, the detailed mechanisms of PEI-mediated mitochondrial damage and apoptotic cell death still remains to be discovered. As there has been no report on PEI–protein interaction within cells that could partly explain the suggested PEI cytotoxicity mechanisms, we employed proteomics technique to identify the proteins that may interact with PEI during the transfection process when PEI is released inside the cell. PEI 25 kDa was used in this study because it is the most widely used polymer for transfection studies and its cytotoxicity has been already proven. Therefore, we investigated proteins that could physically interact with PEI in human colorectal adenocarcinoma cells (HT29) in order to shed light into proteins potentially involved in the mechanism of PEI cytotoxicity. PEI-interacting proteins were isolated through affinity chromatography-based purification method. Isolated proteins were further separated using two-dimensional (2-D) gel electrophoresis. After gel digestion, proteins were identified using matrix-assisted laser desorption/ionization (MALDI)–time-of-flight (TOF)/TOF mass spectrometry (MS) technique.
Materials and methods
Materials
Branched PEI (bPEI 25 kDa) was obtained from Polysciences, Inc. (Warrington, Pennsylvania, USA). Cell culture reagents were purchased from Gibco (Gaithersburg, Maryland, USA). Deionized water was used for all experiments, and all other chemical reagents were obtained from Merck (Germany).
Cell culture
HT29 cells (ATCC, HTB-38) were cultured at 37°C in a humidified 5% carbon dioxide atmosphere in Roswell Park Memorial Institute medium, supplemented with 10% fetal bovine serum (FBS), penicillin at 100 U/ml and 100 µg/ml streptomycin.
Preparation of PEI-activated sepharose 4B complex
An affinity column was prepared by direct coupling of PEI on cyanogens-activated sepharose. For the coupling procedure, 0.5 g cyanogen-activated sepharose 4B (Sigma, St Louis, Missouri, USA) was suspended in 40 ml hydrochloric acid (HCl) to swell. The gel was washed five times with 50 ml coupling buffer (containing 50% dimethylformamide, 0.1 M sodium carbonate, 10 mM sodium hydroxide at 4°C), and 10 ml HCl, respectively. The swelled beads were directly coupled to PEI. Briefly, 100 nM PEI dissolved in coupling buffer and was added to beads at the ratio of 2:1 (v/v). The reaction mixture was incubated overnight at 4°C. The remaining unreacted epoxy groups were blocked by incubating and shaking beads with blocking buffer (1 M ethanolamine, pH 8) at 4°C overnight. 16
The control column beads were prepared by swelled epoxy-activated sepharose 4B without incubation with PEI. Finally, using 50 ml coupling buffer and acetate buffer (pH 4), the control and PEI-coupled beads were washed in four cycles and stored at 4°C until use. The coupling of PEI to sepharose beads was confirmed using PerkinElmer Paragin 1000 Fourier transform infrared (FTIR) spectrometer.
Protein extraction
Cell extracts were prepared by lysis of 5 × 106 cells in 500 μl radioimmunoprecipitation assay (RIPA) buffer containing 50 mM tris(hydroxymethyl)aminomethane (Tris)-HCI pH 7.4, 1% Triton X-100, 0.1% sodium dodecyl sulfate (SDS), 1 mM ethylenediaminetetraacetic acid, 1% sodium deoxycholate, 150 mM sodium chloride (NaCl), and 2 µl complete protease inhibitor cocktail (Sigma) for 15 min at 4°C. After sonicating (UP100H, Hielscher) for 40 s, homogenates were centrifuged (Hettich Universal 320R, Germany) at 12,000g for 15 min at 4°C, and protein content of supernatant was measured using a Bradford protein assay kit (Bio-Rad, Hercules, Californnia, USA).
Affinity chromatography
PEI-interacting proteins were isolated using affinity chromatography. Total proteins (2 mg/ml) were incubated with control beads for 10 min at 4°C, followed by a brief centrifugation. The supernatants were incubated with PEI-coupled beads for 10 min at 4°C. After centrifugation at 1000g for 1 min, pellet was collected and washed three times with 300 μl RIPA buffer. For elution of PEI-interacting proteins, 200 μl of 2 M NaCl was added to pellet obtained in previous step. The elution step was repeated three more times and fractions were pooled. After centrifugation, supernatant fractions–containing PEI-interacting proteins were collected and dialyzed against deionized water for 3 days at 4°C using a dialysis membrane with a 2000 Da cutoff (Spectra/Por, Houston, TX, USA) to remove electrolytes. Desalted target proteins were freeze-dried and stored at −80°C until use.
2-D Gel electrophoresis
Nonlinear immobilized pH gradient (IPG) strips (pH range: 3–10; Bio-Rad) were used for separation of different PEI-interacting proteins according to their isoelectric point. To a final concentration of 1 µg protein/1 µl, all the freeze-dried samples were dissolved in rehydration buffer-containing 2 M thiourea, 6 M urea, 50 mM dithiothreitol (DTT), 20% Bio-Lyte (Bio-Rad), and 2% 3-[(3 cholamidopropyl)dimethylammonio]-1-propanesulfonate (CHAPS). For isoelectric focusing (IEF) experiment, 300 µl of lysate was loaded to 17 cm IPG strips. The IPG strips were actively rehydrated for 12 h at 50 V. Active rehydration was followed by IEF using PROTEAN IEF cell (Bio-Rad) at 250 V for 15 min, linearly increased to 8000 V and kept at this voltage for 5 h.
After the IEF step, IPG strips were incubated for 15 min in 6 ml equilibration buffer-containing 6 M urea, 50 mM Tris-HCl, 2.5% SDS, 1.5% DTT, and 30% glycerol. Strips were re-equilibrated in 6 ml of the same buffer-containing 2.5% (w/v) iodoacetamide instead of DTT. For the second dimension, equilibrated IPG strips were subsequently placed on the top of 12% SDS-polyacrylamide gel to separate proteins based on molecular weight. Electrophoresis was carried out at 16 mA for 30 min and then 25 mA for 7 h in Tris-glycine-SDS running buffer. Gels were stained by MS-compatible silver staining method, and protein spots were excised from the gel and transferred to Center of Genomic Sciences at University of Hong Kong for in-gel digestion and identification PEI-interacting proteins. 17
In-gel digestion
Gel slices were destained, washed, and, subsequently, gel slices were reduced in 10 mM DTT solution and alkylated in dark using 55 mM iodoacetamide for 30 min at room temperature. After alkylation, the gel pieces were washed with 500 μl of 100 mM ammonium bicarbonate and digested using 12.5 ng/µl trypsin (Promega, Madison, Wisconsin, USA) for 30 min at 4°C. Then, trypsin was removed, and 20 µl of 1 mM ammonium bicarbonate was added and incubated at 37°C overnight. By adding two fractions of 5% formic acid and 50% acetonitrile (Sigma), digestion was stopped, followed by dehydration with 100% acetonitrile. The extracted peptide mixtures were dried down using a SpeedVac (ThermoScientific, Inc., Waltham, Massachusetts, USA) and then resuspended in 0.1% formic acid (Sigma), followed by desalting on ZipTip μC-18 (Millipore, Billerica, Massachusetts, USA).
Mass analysis and database searching for protein identification
The MS analysis was carried out using 4800 MALDI-TOF/TOF analyzer in positive ion reflector mode (ABSciex, Framingham, Massachusetts, USA). Typical spectra were obtained by minimum possible laser energy in order to maintain the best resolution in MS/MS mode and averaging 500 acquisitions in reflector mode. Precursor ions with a charge state of 1+ were fragmented via post-source decay (PSD). Mass data were analyzed using Mascot software version 2.1.0 (Matrix Science Ltd, London, UK). Data were BLASTed against both SwissProt and NCBInr databases. MASCOT search parameters were set as follows: fixed modification, carbamidomethyl (cysteine); variable modification, oxidation (methionine); taxonomy, human; MS/MS fragment tolerance, ±0.2 Da; precursor mass tolerance, 75 ppm; and peptide charge +1 monoisotopic. MASCOT cutoff scores >30 and confidence interval (CI) >95% were accepted.
Classifications and network interaction analysis for PEI- interacting proteins
PEI-interacting proteins were classified using Protein Analysis Through Evolutionary Relationships (PANTHER) classification system (http://www.pantherdb.org), according to biological function and process. Furthermore, the identified proteins were submitted to Search Tool for the Retrieval of Interacting Genes/Proteins (STRING) database (http://www.string.embl.de).
Results
In order to identify the target proteins of PEI in HT29 cells, PEI was covalently bound to agarose. Attachment of PEI to cyanogen-activated sepharose beads was confirmed by FTIR spectroscopy. IR spectra showed that absorbance band at 2040 cm−1 disappeared, following the reaction between PEI and cyanogens-activated sepharose, indicating the required reaction (Figure 1). To further reduce the possibility of nonspecific binding of non-PEI interacting proteins, cell extracts were incubated with control agarose beads. The unbound protein was incubated with PEI sepharose beads.

FTIR spectrum of cyanogen bromide-activated sepharose (a) and PEI–sepharose complex (b). FTIR: Fourier transform infrared; PEI: polyethylenimine.
PEI-interacting proteins were separated using 2-D gel electrophoresis and identified by MALDI-TOF/TOF MS. Several spots on silver-stained 2-D gels were detected as a PEI-interacting protein, but most of them were too low in abundance to let recognition by MS. Nineteen PEI-interacting proteins were identified using mass spectral data Mascot software (Figure 2). The information of identified proteins, including protein names, Swiss-Prot accession number, CI%, theoretical molecular weight (Da)/isoelectric point, protein score, sequence coverage (%), unique peptides detected, and sequence of identified peptide with the highest ion score are listed in Table 1. PANTHER protein ontology database was used for classification of proteins according to biological process (Figure 3). Identified proteins were classified into eight groups including, metabolic process, cellular process, immune system process, developmental process, response to stimuli, generation of precursor metabolites and energy, cell communication, and transport. Most of the proteins were involved in metabolic process. Moreover, to get a deeper insight into potential relationship among identified proteins, STRING database (http://www.string.embl.de) and a functional annotation space, Kyoto Encyclopedia of Genes and Genome (KEGG) pathway enrichment were used (Figure 4). STRING-illustrated protein–protein interaction network is shown in Figure 4(a). Moreover, based on the neighborhood, gene fusion, co-occurrence, co-expression, experiments, databases, text mining, and homology, 10 additional proteins were selected to find more predicted functional partners for PEI-interacting proteins (Figure 4(b)). Different line colors represent the types of evidence for the association between proteins. The relationships could be used for exploring the predicted interaction networks, suggesting new directions for future experiments (Figure 4(b)). The KEGG analysis showed that the identified proteins were enriched in protein processing, endoplasmic reticulum, glycolysis/gluconeogenesis, and antigen-processing and presentation pathways.

Two-dimensional gel electrophoresis of PEI-interacting proteins in HT29 cells extract. PEI: polyethylenimine; HT29: human colorectal adenocarcinoma cells.
Identified PEI-interacting proteins in HT29 by MALDI-TOF/TOF.
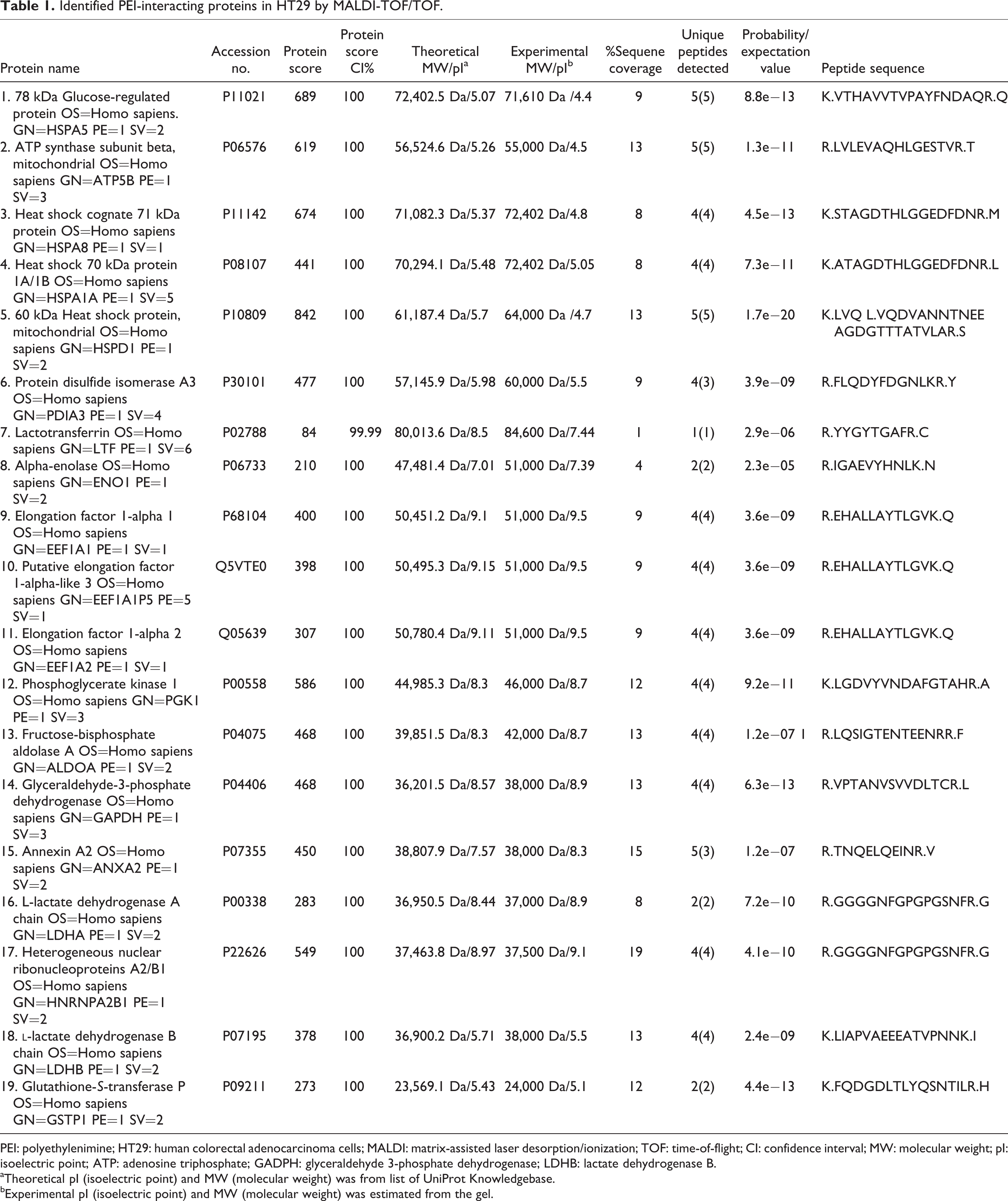
PEI: polyethylenimine; HT29: human colorectal adenocarcinoma cells; MALDI: matrix-assisted laser desorption/ionization; TOF: time-of-flight; CI: confidence interval; MW: molecular weight; pI: isoelectric point; ATP: adenosine triphosphate; GADPH: glyceraldehyde 3-phosphate dehydrogenase; LDHB: lactate dehydrogenase B.
aTheoretical pI (isoelectric point) and MW (molecular weight) was from list of UniProt Knowledgebase.
bExperimental pI (isoelectric point) and MW (molecular weight) was estimated from the gel.

Biological process classification of PEI-interacting proteins in HT29 cells through PANTHER classification. PEI: polyethylenimine; HT29: human colorectal adenocarcinoma cells; PANTHER: Protein Analysis Through Evolutionary Relationships.

Network analysis of PEI-interacting proteins in HT29 cells by STRING database. Nodes are proteins that are shown by gene names and colored lines show types of evidence for the association. (a) Protein–protein interaction modes between the identified proteins. (b) Interaction between identified proteins and more associated protein in cell. PEI: polyethylenimine; HT29: human colorectal adenocarcinoma cells; STRING: Search Tool for the Retrieval of Interacting Genes/Proteins.
Discussion
In recent years, PEI-derived nanoparticles have been widely studied both in vitro and in vivo and showed great promise as nonviral gene delivery systems. 4,9 Despite the fact that PEI was reported to be an efficient gene delivery vehicle, several studies indicated that PEI exhibited high cytotoxicity. 12,18,19 It was specifically suggested that PEI could induce apoptosis in transfected cells. 14,20 Although mechanisms of PEI cell death induction is poorly understood, recent studies by Hall et al. showed that the PEI 25 kDa, in a concentration- and time-dependent manner, facilitates leak of proton from mitochondria and inhibits the electron transport chain. Therefore, it seems that PEI could induce cell death through induction of apoptosis process. 21 The purpose of this study was to separate and recognize the proteins that physically interact with PEI in order to shed light into the mechanism of PEI cytotoxicity using proteomics analysis. Our data showed that 19 proteins could be considered as PEI target proteins in the HT29 cells. These proteins were classified into eight groups based on biological process. Four proteins (HSPA5, HSPA8, HSPA1A, and HSPD1), belonging to heat shock proteins (HSPs) family were identified as PEI-interacting proteins. HSPs are a group of proteins that are present in cells under normal conditions but accumulated in cells exposed to environmental stresses such as heat, high pressures, and toxic compounds. 22 HSPs, also known as chaperones, play an essential role in protein folding, assembly of multiprotein complexes, transport of proteins into organelles, control of signaling, and cell cycle. 23 On the other hand, HSPs have important protective function role against apoptosis. 24,25 Many studies showed that HSPs could block apoptosis by interfering with both the intrinsic and the extrinsic apoptotic pathways. 25 –27 Therefore, it seems possible that the direct interaction between PEI and HSPs could lead to apoptosis in target cells. Glutathione-S-transferases (GSTs) comprise a large group of detoxification enzymes that are present in cytosol and mitochondria. 28 Previous studies showed that overexpression of GSTs in cancer cells causes resistant to apoptosis induced by oxidative stress. 29,30 Incubation of cell extract with sepharose-bound PEI beads showed that PEI has affinity for GST. Interaction of GST with PEI may interrupt its function and induce apoptosis. Our study showed that heterogeneous ribonucleoprotein (hnRNP) was another target of PEI. hnRNPs are complexes of RNA and protein, which involve in a variety of cellular functions including transcription, RNA splicing, DNA repair, telomere biogenesis, and translation. Recent studies suggest that hnRNPs may also be involved in apoptosis inhibition, cell invasion, and angiogenesis. 31 –33 Thus, interaction of PEI with hnRNPs may interfere with anti-apoptosis activity of hnRNPs. Furthermore, our results showed that three proteins of PEI targets are involved in glycolysis process, including glyceraldehyde-3-phosphate dehydrogenase (GAPDH), phosphoglycerate kinase (PGK1), and fructose bisphosphate aldolase. Different investigations have reported that these proteins may play role in apoptosis process. 34 –36 PEI may be altering the cell viability by interacting with these enzymes. Also this study indicated that the PEI interacts with protein disulfide isomerase (PDI), which is a major protein of the endoplasmic reticulum that catalyzes disulfide bond formation in proteins and assists protein folding. However, several studies showed that overexpression of cytosolic PDI inhibited cell death after apoptotic stimulation. 37,38 In addition, PDI knockdown could activate apoptotic cascade, cause cytochrome c release from mitochondria, and activate caspases 6 and 9 pathway. 39 In summary, this study suggested that PEI can induce apoptosis by interacting with some proteins involved in apoptosis process.
This study has some limitations which need to be considered. To assess the effect of PEI on upregulation or downregulation of identified proteins inside the cell, proteomic approach can be useful for confirming the level of protein expression in PEI-treated cells in comparison with the control cells. Also evaluation function of some of the identified proteins in the apoptotic process should be carried out. This experiment will be performed in our next project.
Conclusion
The results of the present experiment suggested that the interaction of PEI with identified proteins that are involved in induction of apoptosis process could partly explain the mechanism of PEI toxicity in cells. Furthermore, PEI–protein interactions suggested by this study were linked to different cell processses, suggesting new directions for future experiment involving PEI cytotoxicity, thereby leading to the design of safer polycationic vectors for experimental and therapeutic interventions.
Footnotes
Conflict of interest
The authors declared no conflicts of interest.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was funded by Mashhad University of Medical Sciences, Mashhad, Iran. This study was part of PharmD thesis of MKh.
