Abstract
Background and Objective
Multidrug and carbapenem resistant gram-negative bacilli bloodstream infections cause high mortality in intensive care units (ICUs). Predicting mortality can improve treatment and support end-of-life decisions. This study aimed to develop a machine learning model to predict mortality in ICU patients with these infections.
Methods
This retrospective cohort study was conducted at a tertiary care medical center between 2017 and 2023. Adult ICU patients with bloodstream infections caused by multidrug and carbapenem resistant Klebsiella pneumoniae, Pseudomonas aeruginosa, and Acinetobacter baumannii were included. Demographic, clinical, and laboratory data were collected. Mann-Whitney U and Chi-square tests were used to compare the groups. Multivariable analysis with binary logistic regression was used to identify mortality risk factors. Ten machine learning classifiers were evaluated using stratified 5-fold cross-validation, and model predictions were interpreted with SHapley Additive exPlanations (SHAP).
Results
197 patients were included, with a 15-day mortality rate of 48%. The Light Gradient Boosting Machine (LightGBM) classifier showed the best performance, with an AUROC of 0.94, AUPRC of 0.952, accuracy of 0.868, precision of 0.906, recall of 0.822, F1 score of 0.855, Matthews Correlation Coefficient (MCC) of 0.744, and Brier score of 0.131. SHAP analysis revealed coagulopathy, rapid access to antibiotics, septic shock, SOFA score, platelet count, C-reactive protein (CRP) level, and time-related parameters as the most important predictive features.
Conclusion
The LightGBM model showed promising results in predicting mortality in ICU patients. This model may support early intervention and assist in complex end-of-life decisions. This study was registered at ClinicalTrials.gov(https://clinicaltrials.gov/ct2/show/NCT06167083)
Keywords
Introduction
In intensive care units (ICU), infections caused by gram-negative pathogens that are resistant to multiple drugs (MDR) are common. MDR bacteria are defined as bacterial isolates resistant to at least one agent in three or more antimicrobial categories. 1 Pseudomonas aeruginosa, Klebsiella pneumoniae, and Acinetobacter spp. are the three main gram-negative MDR bacteria with carbapenem resistance and are responsible for up to 40% of all infections in the ICU.2,3 The World Health Organization (WHO) designated carbapenem-resistant Enterobacteriaceae, carbapenem-resistant Pseudomonas aeruginosa (CRPA), and carbapenem-resistant Acinetobacter baumannii (CRAB) as pathogens of critical importance in 2017. 4 In 2024, CRAB and carbapenem-resistant Enterobacterales persist as critical priorities on this list. Although CRPA is classified as a high priority disease, its mortality rate is high. These pathogens are developing resistance to new antibiotics and continue to become a significant global health concern. 5
The limited availability of carbapenems, which are considered last-line antibiotics for multidrug-resistant gram-negative bacterial infections, results in extended hospitalization, increased expenses, and higher death rates.3,6 Although mortality rates from antimicrobial resistance have decreased by 18% over the past decade with new antibacterial agents, according to the CDC, mortality rates remain high in middle- and low-income countries owing to difficulties in accessing new drugs. 7
Bloodstream infections (BSIs) caused by these strains can progress to sepsis and septic shock, resulting in high morbidity and mortality rates. Risk factors contributing to mortality have been described in the literature and prognostic scoring systems are available.
Machine learning (ML) is a subcategory of artificial intelligence (AI). This enables the development of algorithms that use mathematical methods to process the data. 8 Simple machine learning models, such as logistic regression, have been used for decades. 8 Digital storage of patient data and advances in AI have enabled the development of ML algorithms that provide better prognostic assessments than currently used scoring systems such as Sequential Organ Failure Assessment (SOFA) and Acute Physiology and Chronic Health Evaluation (APACHE) II, III, and IV. 9
The objective of this study was to develop a machine learning algorithm that can be used by clinicians to predict mortality in patients with bloodstream infections caused by multidrug and carbapenem resistant gram-negative bacteria (CR-GNB) in a single-center intensive care unit based on data from our patient population. Predicting a patient's prognosis can lead to the identification of more effective treatment modalities. 10 It may also allow time to discuss end-of-life decisions such as withholding, withdrawal, and do-not-resuscitate (DNR) in selected patients. 11
Patients and Methods
This retrospective cohort study was conducted in the intensive care unit of an 830-bed tertiary care medical center between June 1, 2017, and June 1, 2023, in accordance with the protocols of the Ethics Committee of the Kocaeli University Faculty of Medicine and the guidelines of the Declaration of Helsinki. The project was assigned the number 2023/226 and received approval under code GOKAEK-2023/12.31. This study was registered at ClinicalTrials.gov (https://clinicaltrials.gov/ct2/show/NCT06167083) and reported in accordance with EQUATOR (TRIPOD + AI) guidelines. 12 Informed consent was not obtained owing to anonymization of patient data.
The study population comprised adult patients of mixed socioeconomic levels and homogeneous ethnicity with bloodstream infections caused by multidrug and carbapenem-resistant Klebsiella pneumoniae (CRKP), CRPA, and CRAB. As bloodstream infections caused by carbapenem and multidrug resistant gram-negative bacteria are relatively rare, all eligible patients were included in the study. To ensure data diversity for machine learning, each patient was included only once, considering their unique clinical characteristics. Patients with multiple microorganism growth in the blood culture were excluded. The primary outcome variable was the 15-day bacteremia-related mortality rate in the ICU. The study included 96 patients who died and 101 patients who survived in the control group. The term “survival” was defined as either being alive 15 days after the collection of blood samples for culture or discharge from the intensive care unit with a recovery period at any point within 15 days.
Based on the literature, risk factors in patients with CR-GNB bloodstream infections were evaluated.6,13 Patient data were retrospectively assessed using the hospital's electronic medical record system. The records were carefully evaluated and digitized by an experienced clinician to prevent missing data. Records were reviewed to collect demographic data, including age, sex, and comorbid conditions such as obesity, diabetes mellitus, alcoholism, long-term steroid and other immunosuppressive drug use, chronic obstructive pulmonary disease (COPD), chronic liver and kidney disease, need for dialysis, heart failure, solid organ transplantation, solid organ tumor, hematogenous malignancy, COVID-19, history of cerebrovascular accident (CVA), and trauma, such as traffic accidents.
The evaluation focused on five time-based parameters. The metrics included the total length of hospital stay, total length of intensive care unit stay, total number of days on a ventilator, day on which blood culture was taken during the hospital stay, and the number of days between blood culture and discharge or death. The APACHE II score upon admission to the ICU, SOFA and Glasgow Coma Scale scores, presence of sepsis/septic shock and acute respiratory distress syndrome, and invasive procedures such as central line insertion, tracheostomy, and parenteral nutrition administration at the time of blood culture sampling were assessed. Major surgical interventions prior to blood culture sampling, source of bacteremia, access to appropriate effective antibiotics within 48 h, use of a polymyxin-based combination, and biochemical laboratory values were also recorded. The term “appropriate” antibiotic therapy is defined as a treatment regimen that includes at least one of the susceptible antimicrobial options within 48 h of blood culture growth, based on the results of in vitro susceptibility testing.
Bacteria were identified using matrix-assisted laser desorption/ionization coupled with time-of-flight mass spectrometry (MALDI-TOF MS; BioMérieux, France). The Vitec 2 system (BioMérieux, France) was used to assess carbapenem resistance. These tests were performed in accordance with the guidelines set by the European Committee for Antimicrobial Susceptibility Testing (EUCAST) in accordance with the recommendations of the Turkish Ministry of Health.
IBM SPSS for Windows version 29.0 (IBM Corp., Armonk, NY, USA) was used for all the statistical analyses. The Gaussian distribution was evaluated using the Kolmogorov-Smirnov test. Owing to non-normality, continuous variables were expressed as median and interquartile range (IQR). Categorical variables were presented as counts and percentages. The Mann-Whitney U test was used to compare the statistical significance between the groups. The chi-square test was used to examine associations among categorical variables. To identify the risk factors associated with mortality, univariable logistic regression analyses were first performed, and significant variables were included in the multivariable logistic regression analysis, which were then compared to those in the AI model. A P-value < .05 was considered to be statistically significant.
To ensure consistency throughout the dataset, the data collection protocol was conducted under the guidance of a clinician, following strict rules. A detailed analysis identified and addressed potential inaccuracies and data gaps. The Jupyter Lab environment served as a platform for data processing and analysis using the Python 3. Python 3 is popular owing to its efficiency, versatility, and extensive library ecosystems. Additional libraries, including Pandas, NumPy, Scikit-learn, Shapley Additive Explanations (SHAP), and Matplotlib, were used to enhance analytical accuracy.14–18 One-hot encoding represented subcategories of features, such as major surgery and source of bacteremia. To improve model performance and ensure comparability of classifiers, all features were normalized to the interval [−1, 1] using min-max scaling.
Model Training and Testing
To evaluate the prediction performance, ten different machine learning classifiers were used: Gaussian Process, Random Forest, Gradient Boosting, Extreme Gradient Boosting (XGBoost), Light Gradient Boosting Machine (LightGBM), CatBoost, Decision Tree, AdaBoost, Multi-Layer Perceptron (MLP), and Logistic Regression. Owing to the limited size of the study sample, a stratified 5-fold cross-validation approach was employed instead of the conventional training-test split. This technique separates the dataset into five equal portions while maintaining the distribution of the classes. Each subset was employed as the test data, and the remaining four subsets were used for the model training. Consequently, during the training and testing phases, the dataset was partitioned into 80% for training and 20% for testing.
LightGBM exhibited the best prediction performance among the classifiers. The LightGBM is a gradient-boosting framework that uses tree-based learning algorithms. It offers faster training speed, better efficiency and accuracy, and can process large-scale data. 19 To enhance the model performance, grid-based hyperparameter tuning was employed. Combining hyperparameter optimization with 5-fold cross-validation effectively reduces the risk of model overfitting. The hyperparameter tuning settings are listed in Supplementary Table 1.
The performance metrics included accuracy, precision, sensitivity (recall), F1 score, Brier score, and Matthews correlation coefficient (MCC). To evaluate model stability, these measurements were computed using 5-fold cross-validation techniques.
Additionally, the area under the receiver operating characteristic curve (AUROC) and area under the precision-recall curve (AUPRC) were used. This metric was emphasized because AUPRC was more descriptive than AUROC in unbalanced datasets.
The SHapley Additive exPlanations (SHAP) method was applied to explain the LightGBM predictions for clinical applications. 20 SHAP explains the features that the model emphasizes and their contribution to the predictions. SHAP visualizes how features contribute to the predictions. SHAP plots show the contribution of each feature to the predictive efficiency of the model. Each point represents a case in the dataset. Blue indicates low impact, whereas red indicates high efficiency. The configuration of the study is illustrated in Figure 1.
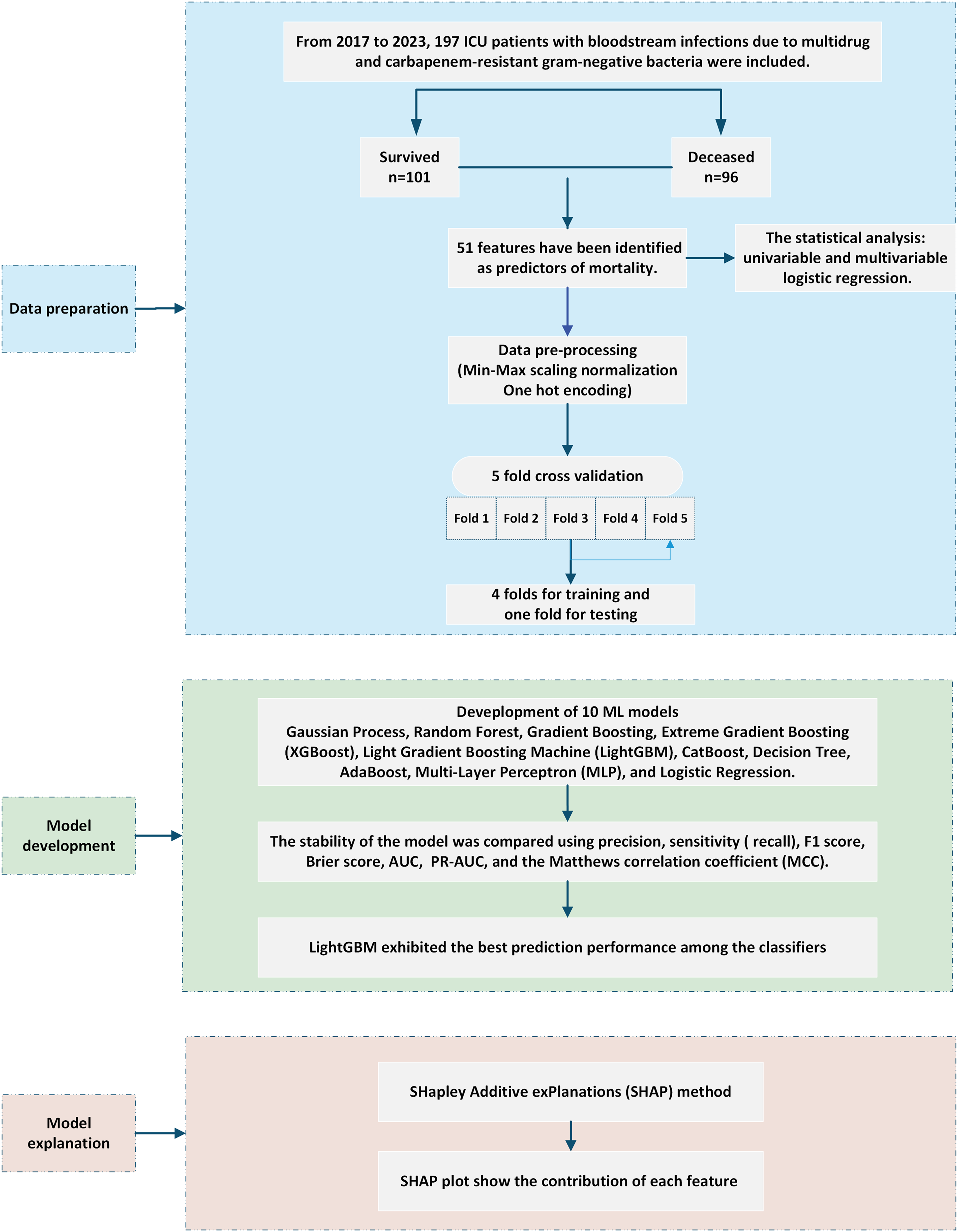
Flowchart of the Study Design.
Initially, subgroup analyses were planned according to microorganism type and patient age group (≤65 years and >65 years). Subsequently, the subgroup analyses were further refined based on key parameters identified as the most influential by the model and logistic regression analysis. Detailed subgroup analyses were performed using LightGBM.
Results
A total of 197 patients were included in this study. The most common microorganism was CRAB, and the most common cause of bacteremia was pneumonia. The basic demographic and predictive characteristics are outlined in Table 1 and a comprehensive version is provided in Supplementary Table 2.
Baseline Demographic Characteristics and Predictive Features.
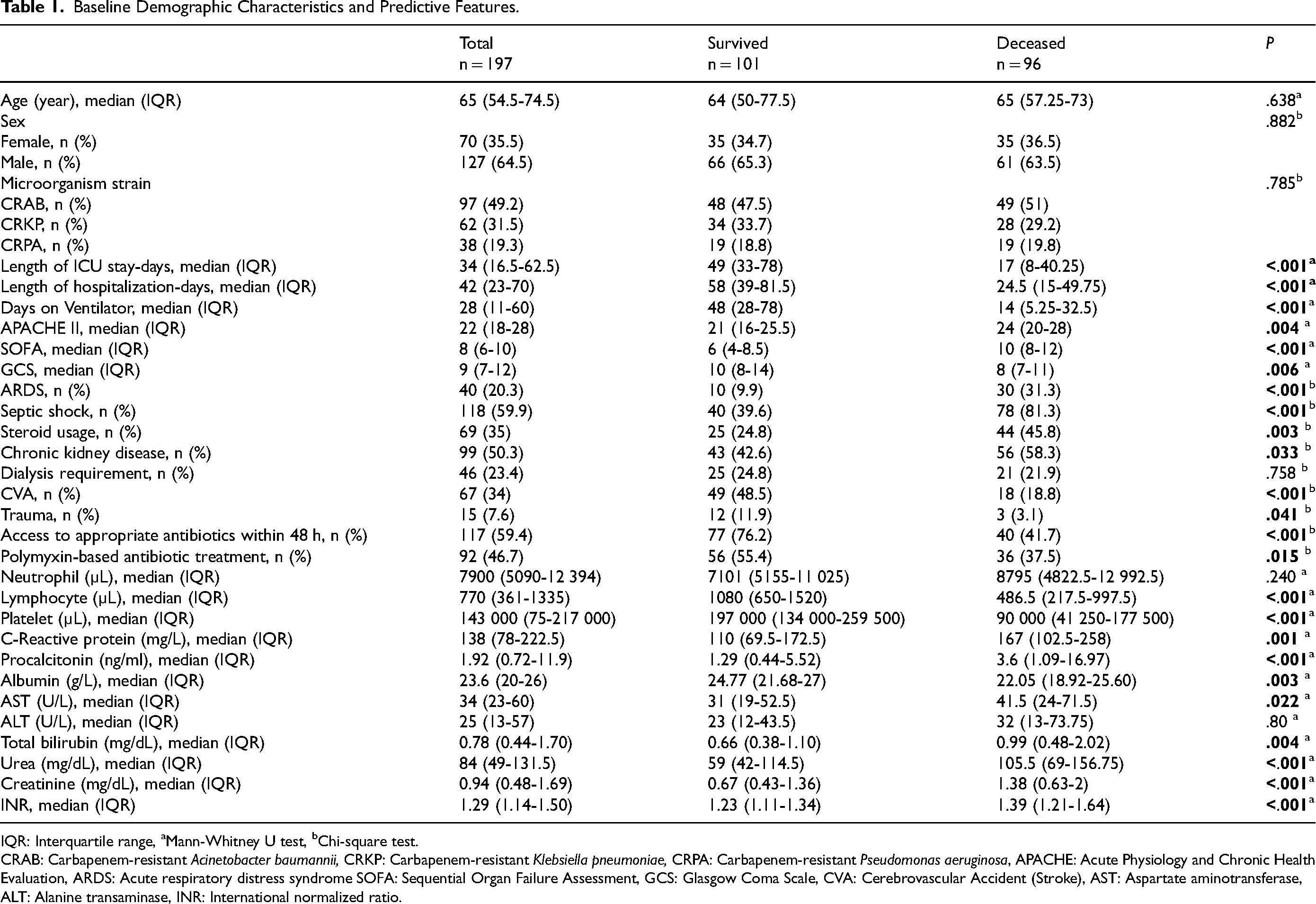
IQR: Interquartile range, aMann-Whitney U test, bChi-square test.
CRAB: Carbapenem-resistant Acinetobacter baumannii, CRKP: Carbapenem-resistant Klebsiella pneumoniae, CRPA: Carbapenem-resistant Pseudomonas aeruginosa, APACHE: Acute Physiology and Chronic Health Evaluation, ARDS: Acute respiratory distress syndrome SOFA: Sequential Organ Failure Assessment, GCS: Glasgow Coma Scale, CVA: Cerebrovascular Accident (Stroke), AST: Aspartate aminotransferase, ALT: Alanine transaminase, INR: International normalized ratio.
A logistic regression test was conducted to identify the significant risk factors associated with mortality. The results of both univariable and multivariable analyses are presented in Table 2.
Multivariable Analysis of Mortality Risk Factors Using Logistic Regression.
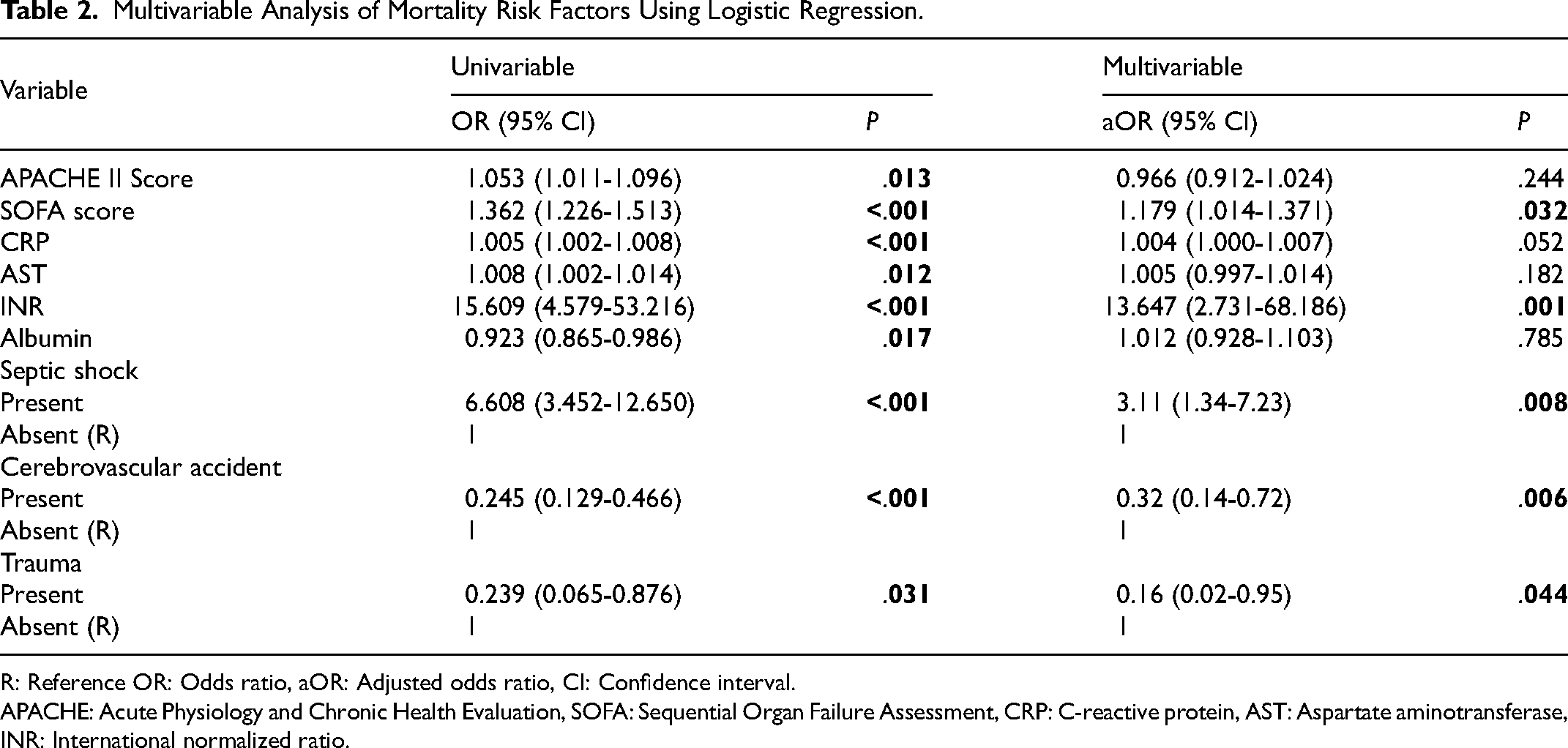
R: Reference OR: Odds ratio, aOR: Adjusted odds ratio, CI: Confidence interval.
APACHE: Acute Physiology and Chronic Health Evaluation, SOFA: Sequential Organ Failure Assessment, CRP: C-reactive protein, AST: Aspartate aminotransferase, INR: International normalized ratio.
Ten distinct classifiers were evaluated to validate the efficacy of the model. The performance metrics of all models are provided in Table 3. Figure Supplementary 1 illustrates the AUROC for all machine learning classifiers tested on the dataset.
Performance Metrics of the Evaluated Classification Models.
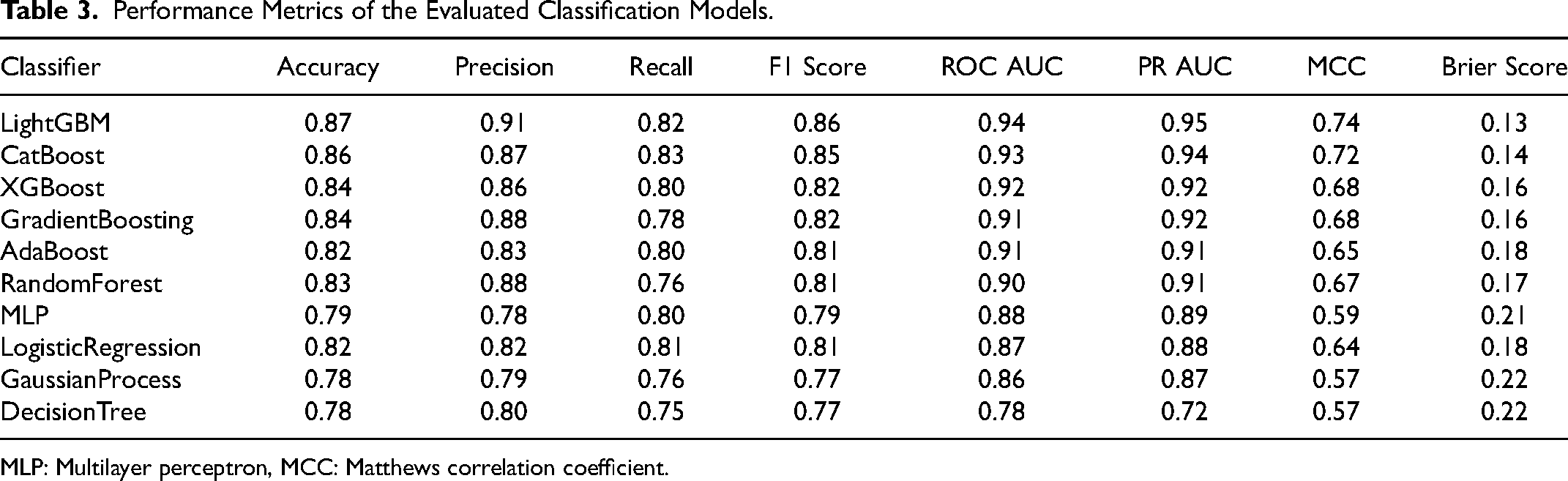
MLP: Multilayer perceptron, MCC: Matthews correlation coefficient.
The LightGBM classifier demonstrated the best performance and lowest Brier score among all the classifiers. The model employed a five-fold cross-validation approach, exhibiting an accuracy of 0.868, precision of 0.906, recall of 0.822, F1 score of 0.855, Matthews correlation coefficient (MCC) of 0.744, and Brier score of 0.131. Figure 2 depicts the individual and mean receiver operating characteristics for the LightGBM across all cross-validation folds. The mean AUC value was 0.94, with a standard deviation of 0.02.
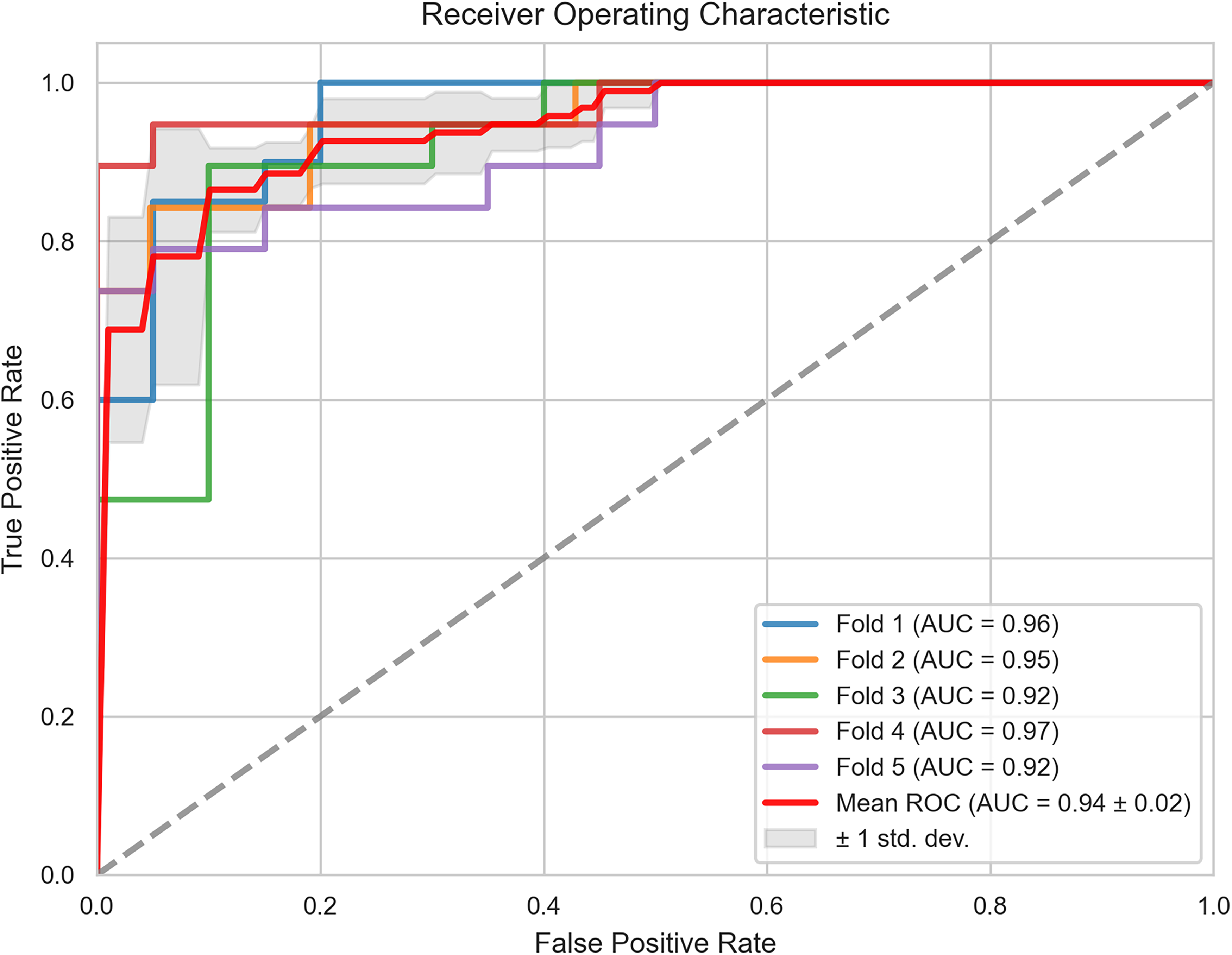
Receiver Operating Characteristic (ROC) Curves Were Generated Using Five-Fold Cross Validation. the Mean Receiver Operating Characteristic Curve was Derived from all Folds. the Shaded Region Represents the Standard Deviation.
Figure 3 illustrates the critical relationship between precision and recall (sensitivity), which is essential for understanding the balance between the true positive rate and predictive accuracy of the model. The AUC for the average precision-recall curve (PRC) was 0.95 ± 0.02, demonstrating high predictive accuracy across different cross-validation folds.
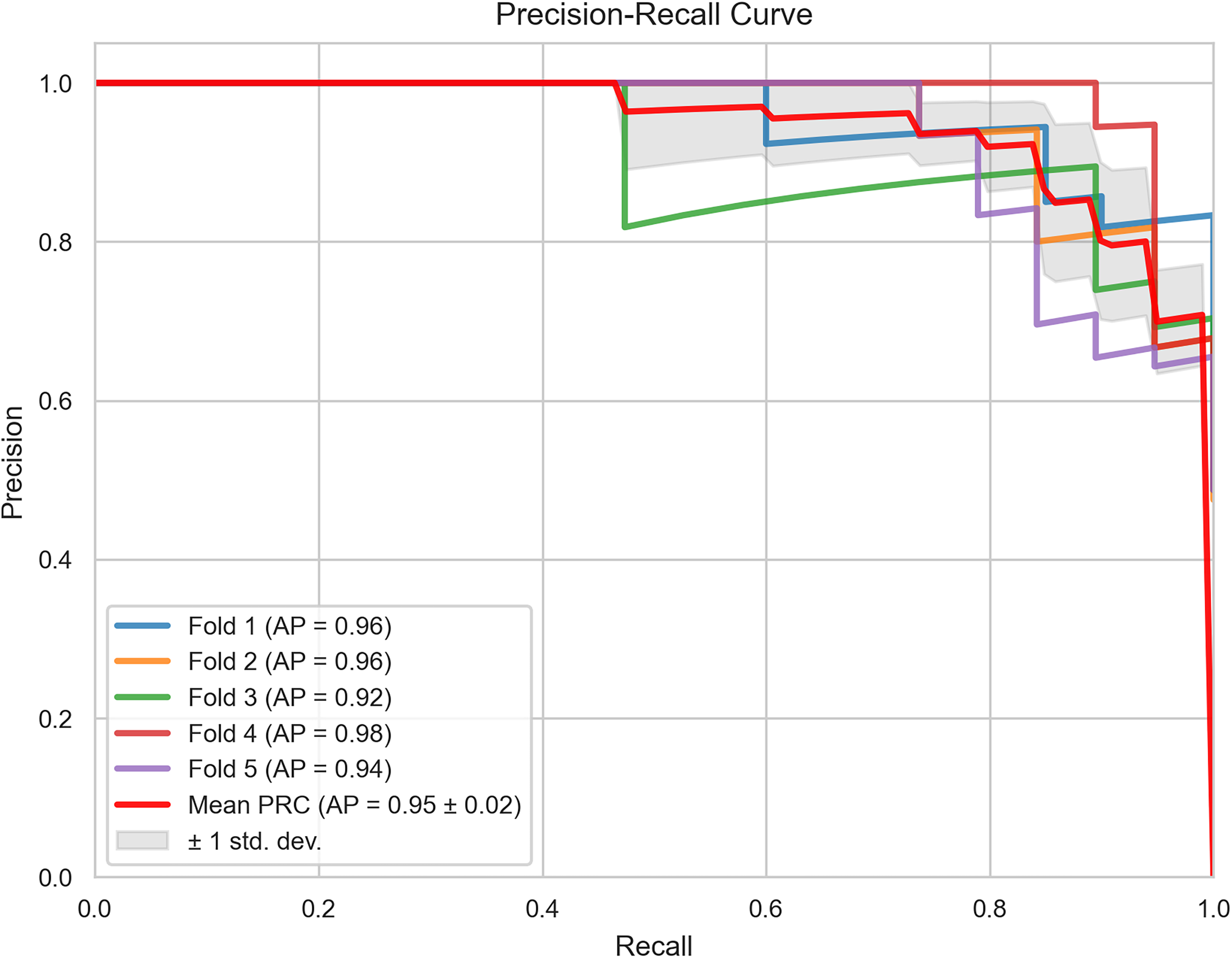
Precision-Recall Curves for all Folds are Presented Along with the Average Curve.
To clarify the prediction model of the LightGBM model, a SHAP plot is presented in Figure 4. The top-ranked variables, including the day of blood culture after hospitalization and length of hospitalization, contributed the most to the prediction of the model. Red dots indicate high efficacy. For example, for the SOFA score, the red dots on the right side of the x-axis indicate that higher values help predict mortality, whereas lower platelet values on the left side of the x-axis help predict mortality.
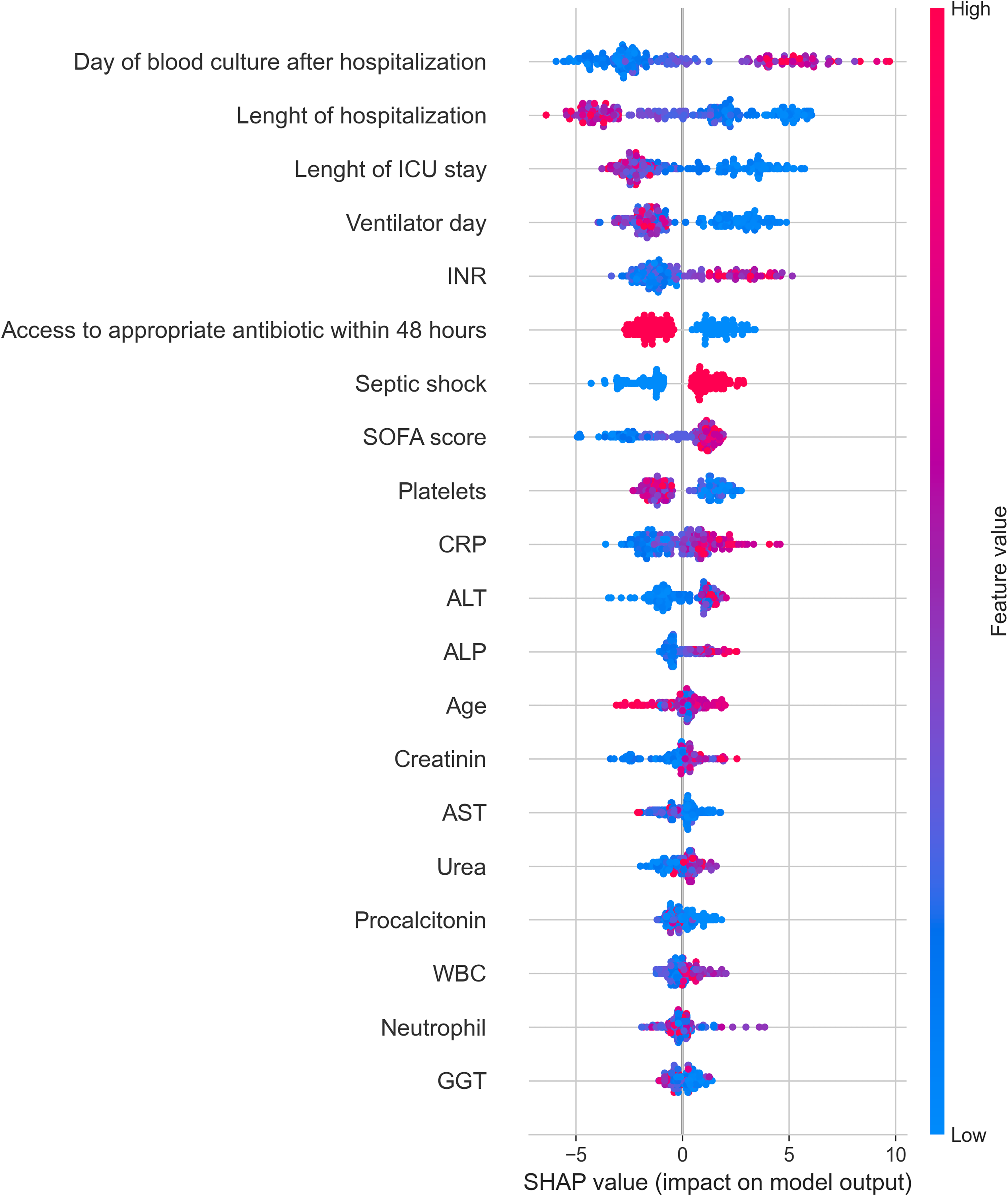
SHAP Summary Plot Showing Feature Importance and Impact on Model Predictions.
Multivariable logistic regression analysis identified the SOFA score, INR level, and the presence of septic shock as variables significantly associated with mortality. These parameters were also among the most influential features in the SHAP analysis. Accordingly, subgroup analyses were performed incorporating these clinical parameters together with age and microorganism type. For INR, a cut-off value of 1.5 or higher, previously reported as clinically meaningful for mortality prediction, and mortality prediction performance was evaluated using this threshold. 21 ROC curve analysis for the SOFA score is presented in Supplementary Figure 2. A SOFA score of 8 or higher demonstrated significant discriminatory power for predicting mortality, with a sensitivity of 79.2%, specificity of 62.4%, and an area under the curve of 0.77. Patients were therefore classified according to this threshold, and septic shock was evaluated as a binary variable. The results of the subgroup analyses are presented in Table 4.
Subgroup Analysis of the LightGBM Mortality Prediction Model According to Pathogen Type and Clinical Characteristics.
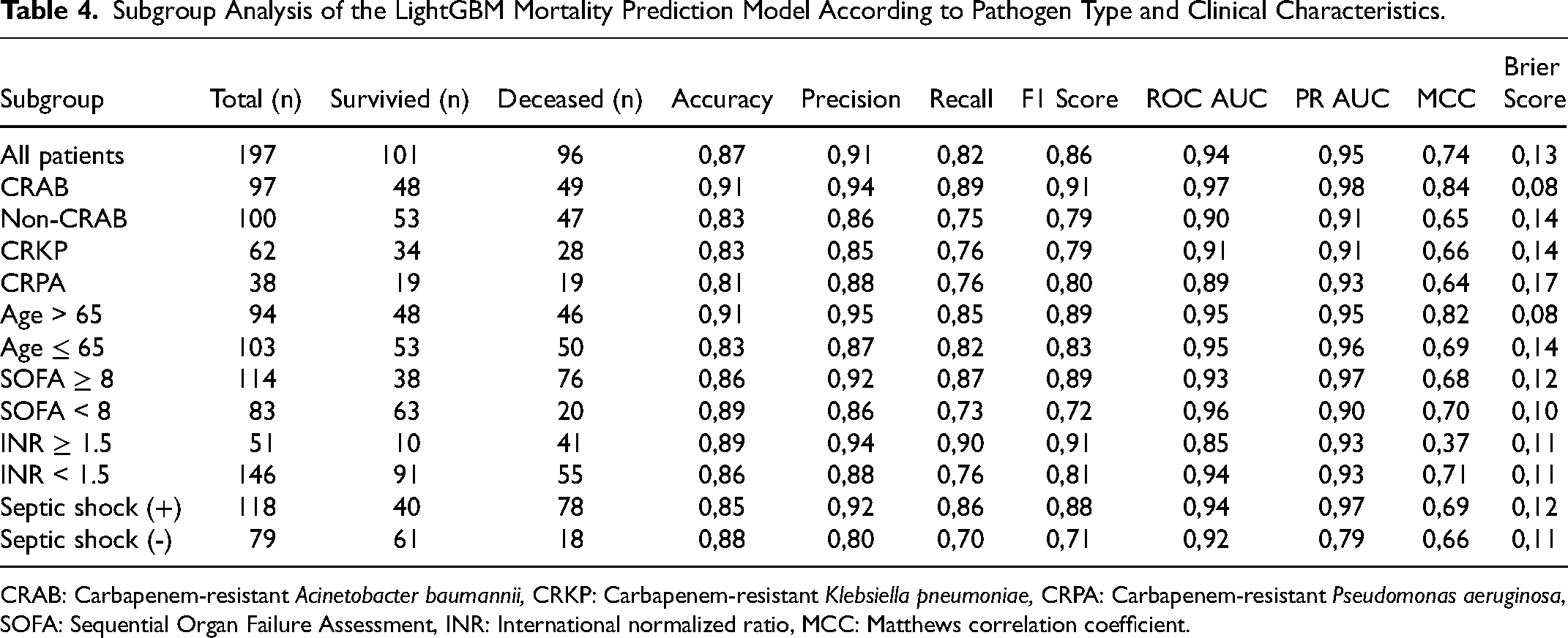
CRAB: Carbapenem-resistant Acinetobacter baumannii, CRKP: Carbapenem-resistant Klebsiella pneumoniae, CRPA: Carbapenem-resistant Pseudomonas aeruginosa, SOFA: Sequential Organ Failure Assessment, INR: International normalized ratio, MCC: Matthews correlation coefficient.
The Ceiba Tele ICU system (Ceiba Health, USA) is a continuous remote monitoring platform that is used in our hospital to track data and monitor intensive care patients, as mandated by national health authorities. The system continuously collects clinical and laboratory data, and all patients are assessed daily. Our model's features have been integrated into the Ceiba Tele ICU system's existing mortality prediction interface to support daily clinical risk assessment.
Discussion
There has been a global increase in infections caused by multidrug and carbapenem resistant gram-negative bacteria. 5 Predicting mortality enhances treatment efficacy and may aid in making critical decisions such as withdrawing life-sustaining therapies when appropriate. This study developed a supervised machine learning model using LightGBM to predict mortality. We explained the predictive features of the model using the SHAP and conventional statistical methods.
Machine learning models that predict mortality in sepsis are available in literature. Xie et al developed a model to predict 28-day mortality in total 3295, and 1131 of whom were deceased patients using the Medical Information Mart for Intensive Care-IV v2.0 (MIMIC-IV) dataset and used 16 predictive features. 22 MIMIC-IV is a public database derived from the electronic health records of the Beth Israel Deaconess Medical Center. 23 Among the various machine learning classifiers, the logistic regression model demonstrated the highest efficacy with an AUC of 0.806. 22 The AUC value of our model (0.94) yielded better results than the comparative study.
The other datasets also contained records of patients in the ICU. The eICU Collaborative Research Database (eICU-CRD) v2.0 contains 200 859 ICU admissions from 208 U.S. hospitals. 24 The Amsterdam UMCdb contains detailed information on 23 106 admissions of 20 109 patients to a single academic medical center in the Netherlands. 25
Several studies have been conducted using these datasets. Zhang et al developed a prediction model for in-hospital mortality in sepsis on 3535, of them 555 deceased patients from these three registries. The eXtreme Gradient Boosting (XGBoost) model with an AUC of 0.94, precision of 0.882, recall of 0.918, and F1 score of 0.937 was identified as a notable model. 26 According to our model, the AUC was equivalent, but the precision was lower (0.906). However, the recall (0.822) and F1 score (0.855) of the model in this study were superior.
While there are studies in the literature using large-scale databases, the detection rate of carbapenem-resistant microorganisms in databases containing sepsis data from intensive care units is low. For example, in a study using the MIMIC-IV database, carbapenem-resistant organisms were detected in only 183 of 4531 patients. 27 Furthermore, studies using these databases reported lower mortality rates than those observed in our study (48%).
Previous studies have suggested that treatment strategies may not substantially alter mortality prediction performance in CRAB infections when machine learning–based models are applied. In a 14-day mortality prediction study by Özdede et al, a machine learning model analyzed data from 128 patients with CRAB bacteremia, in which the Naïve Bayes classifier achieved the best performance with an area under the curve of 0.82. 28 In our subgroup analysis, the LightGBM model demonstrated excellent discriminatory performance for predicting CRAB-related mortality, with an AUROC of 0.97, exceeding the performance observed in the overall cohort. CRAB, classified as a critical priority pathogen by the World Health Organization, also represented a substantial proportion of our cohort, accounting for nearly half of the study population. 5 This higher representation may have enabled the model to better capture pathogen-specific mortality patterns, thereby improving its predictive performance.
Machine learning models operate as black boxes with predictions that are difficult to interpret. 29 We performed a SHAP analysis to understand which features had the greatest impact on the model output. The LightGBM highlighted 20 predictive features. The SHAP graph showed that coagulopathy, rapid access to appropriate antibiotics, septic shock, SOFA score, platelet count, C-reactive protein (CRP), alanine aminotransferase (ALT), alkaline phosphatase (ALP), age, creatinine, aspartate aminotransferase (AST), urea, procalcitonin, total leukocyte count, neutrophil count, gamma-glutamyl transferase (GGT) and time-related parameters were the main features used in our model. In the multivariable logistic regression analysis, the following factors were found to be significant: SOFA score (aOR: 1.179; 95% CI: 1.014-1.371; P = .032), coagulopathy (defined by the international normalized ratio; aOR: 13.647; 95% CI: 2.731-68.186; P = .001), and septic shock (aOR: 3.11; 95% CI: 1.34-7.23; P = .008). Our model can predict mortality in bacteremia more quickly and practically by using features that are not highlighted by logistic regression but noted in the literature.6,13
A Sankey curve diagram (Supplementary Figure 3) shows that death occurred within the first 10 days or after 50 days. According to our model, the first ten days are critical for ICU patients and their parameters should be closely monitored. For hospitalizations exceeding 50 days, decisions to withdraw life-sustaining therapy can be discussed considering medical, ethical, and legal issues.
The limitations of our study include its retrospective design, relatively small sample size, and inability to externally validate the model owing to the lack of suitable matching data. However, it should be noted that large datasets may not be representative of the patient populations at the centers. Therefore, future studies should aim to validate the proposed model in larger, prospective, multicenter cohorts and across different clinical settings to confirm its robustness, generalizability, and real-world applicability.
The absence of class imbalance in our dataset and the fact that the dataset was created by clinicians with at least five years of experience in the field, who followed ICU patients, increased the predictive potential of our model. The absence of missing values in our supervised dataset positively contributed to model performance and made the analysis process more reliable. To demonstrate the effectiveness of the proposed model, various performance metrics were evaluated.
The global prevalence of multidrug and carbapenem resistance has become critical, prompting the need for strategies to predict mortality. Our model aims to facilitate the timely administration of toxic drugs such as polymyxins, particularly in resource-constrained settings with limited access to novel antimicrobials. Therefore, it can help physicians to identify patients who require early aggressive interventions.
End-of-life decisions are frequently made in intensive care units and, unfortunately, can be challenging for clinicians. 30 Decisions made in situations in which a patient is at the brink of death are based on a combination of medical, local laws, and ethical considerations. For patients who have been hospitalized for over 50 days, have a low life expectancy, and have been warned by our model that mortality may occur, it can assist in making clinical decisions regarding the end-of-life.
Conclusion
Our machine learning model predicted mortality in ICU patients with multidrug and carbapenem resistant gram-negative bacilli bloodstream infections. The LightGBM classifier showed satisfactory predictive performance. SHAP analysis demonstrated model stability. This study showed the potential of machine learning to predict mortality risk. Our model was designed to support early interventions for critically ill patients and to assist in decisions regarding withdrawing life-sustaining therapies.
Supplemental Material
sj-docx-1-jic-10.1177_08850666261423499 - Supplemental material for Machine Learning in the ICU: Predicting Mortality in Patients with Carbapenem-Resistant Gram-Negative Bacilli Bloodstream Infections
Supplemental material, sj-docx-1-jic-10.1177_08850666261423499 for Machine Learning in the ICU: Predicting Mortality in Patients with Carbapenem-Resistant Gram-Negative Bacilli Bloodstream Infections by Özlem Güler, Volkan Alparslan, Burak İnner, Sibel Balcı, Ahmet Düzgün, Nur Baykara and Alparslan Kuş in Journal of Intensive Care Medicine
Footnotes
Acknowledgments
The authors gratefully acknowledge the Artificial Intelligence and Simulation Systems R&D Laboratory of the Kocaeli University for their support. The lab's advanced infrastructure is vital for implementing, optimizing, and deploying a machine learning model that is central to our research.
Author Contributions
Ö güler and V. Alparslan conceptualized the study methodology. Ö. Güler wrote the original draft of this manuscript. A. Düzgün curated the data from the patient files. İnner created software for the machine learning algorithm model and analyzed the data. S. Balcı analyzed data. N. Baykara and A. Kuş reviewed the manuscript as mentors. All authors have read and approved the final manuscript.
Consent to Participate
As our study was retrospective, permission to use patient data was obtained from the Kocaeli University Hospital, and no additional consent was obtained because the patient data were anonymized.
Data Availability Statement
The dataset was shared on e-picos.com. If requested, they could be forwarded via email.
Declaration of AI-Assisted Technologies Usage
During the preparation of this study, the author(s) used the DeepL program for language editing and improved readability. After using this tool/service, the author(s) reviewed and edited the content as needed and took full responsibility for the content of the publication.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical Considerations
This retrospective cohort study was conducted in the intensive care unit of Kocaeli University Hospital, Turkey, a tertiary care academic medical center, between June 1, 2017, and June 1, 2023. This study was conducted in accordance with the protocols of the Ethics Committee of the Kocaeli University Faculty of Medicine and the guidelines set forth in the Declaration of Helsinki. The project was assigned number 2023/226 and received approval under the code GOKAEK-2023/12.31. This study was registered at ClinicalTrials.gov (![]() ).
).
Funding
The authors declare that this research did not receive any financial support from any funding agency in public, commercial, or not-for-profit organizations. Only Kocaeli University has an open-access agreement with SAGE.
Supplemental Material
Supplemental material for this article is available online.
