Abstract
This is a visual representation of the abstract.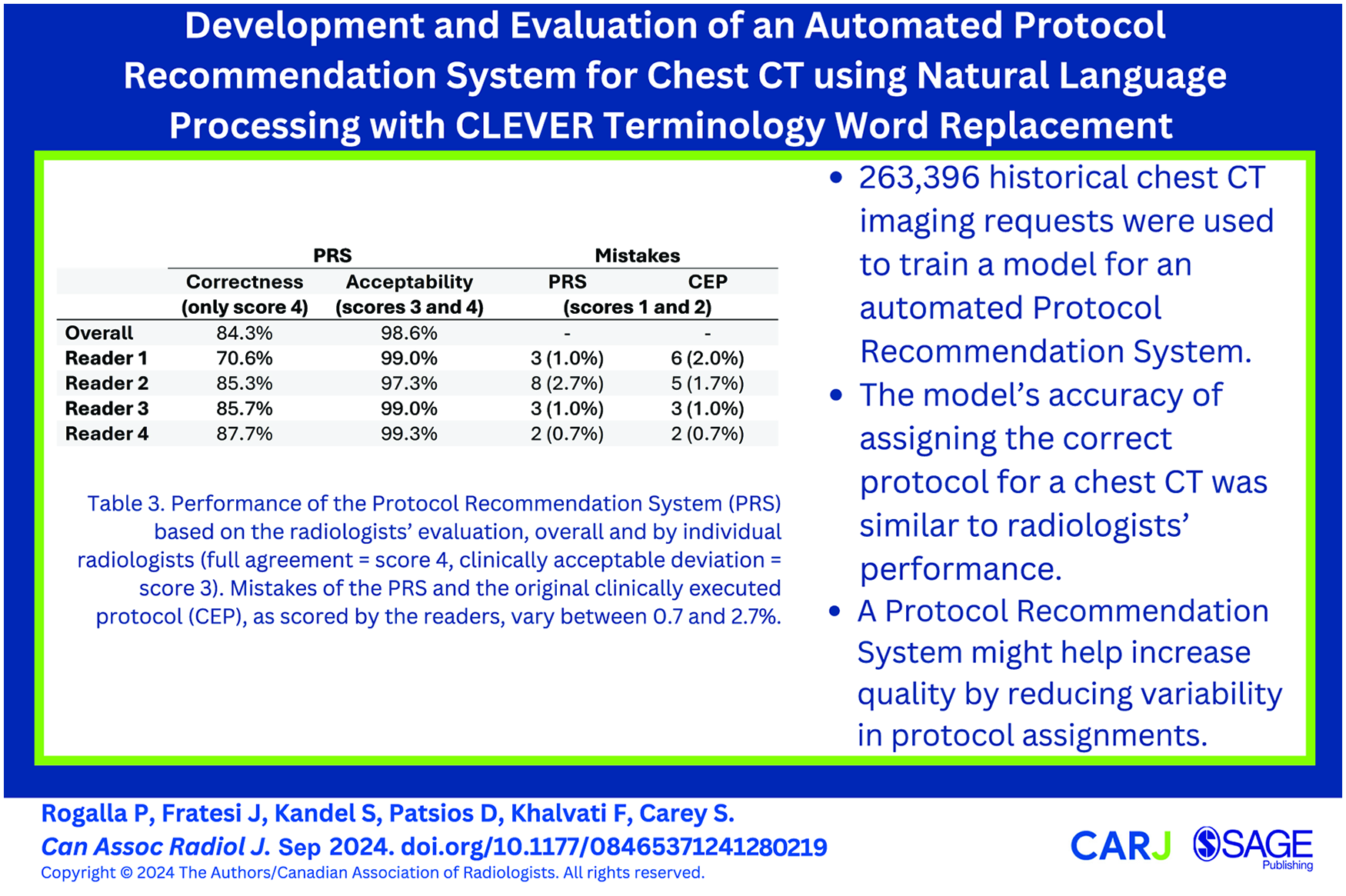
Introduction
Computed tomography (CT) imaging is a mainstay diagnostic tool for a wide variety of diseases. The ability of a radiologist to create an accurate interpretation is dependent on the CT images’ clear visualization of the associated pathology. For this reason, image quality remains an important research focus. The CT protocol used for an imaging procedure refers to the selection of various parameters of the scan which, for chest CT as well as other procedure types, are tailored to produce an image that is well-suited for the clinical indication. Some examples of these parameters include administration of contrast bolus, scan range, patient position, slice thickness, and CT tube settings. 1 The appropriate selection of these parameters is integral to producing a diagnostic-quality image, while inappropriately selected parameters can produce an unusable image and may even introduce health risks to the patient. CT imaging accounts for the majority of medical imaging-related radiation exposure, which underscores the importance of managing the radiation dose—image quality tradeoff. 2 Use of contrast medium may possibly pose risks as the kidney function can be impacted, in particular in patients with renal insufficiency, 3 although this is increasingly debated. Some patients may even have a history of allergic-like reactions to the contrast agent. In light of these factors, selecting appropriate CT imaging parameters is clinically important and requires expertise, necessitating that optimal protocolling be performed by trained radiologists in a clinical setting.
Protocolling is a monotonous and time-consuming task and can create disruptions for radiologists in a clinical setting. 4 While a single imaging request may only take around 1 minute to protocol on its own, the large volume of CT imaging procedures exacerbates the task. Machine learning techniques have become increasingly broadly applied to a large variety of medical fields and can achieve near-expert performance for a well-defined task. 5 The clinical workflow for protocolling, which involves free-text image request documents that are examined and then protocolled by a radiologist through the Radiology Information System (RIS), creates a fertile dataset for supervised natural language processing (NLP) techniques. 6 Automated protocolling systems have been proposed before, in MRI imaging,6,7 as well as abdominal CT, 8 and have shown potential to increase both the efficiency and quality of protocolling when used as a support tool for a radiologist. 9 We developed a protocol recommendation system (PRS) for chest CT and tested its performance against an optimized, reader-agreement reference standard. We hypothesize that such a system can help support the protocolling radiologist, making the task more efficient as well as reducing inter-reader variability.
Methods
Dataset Selection
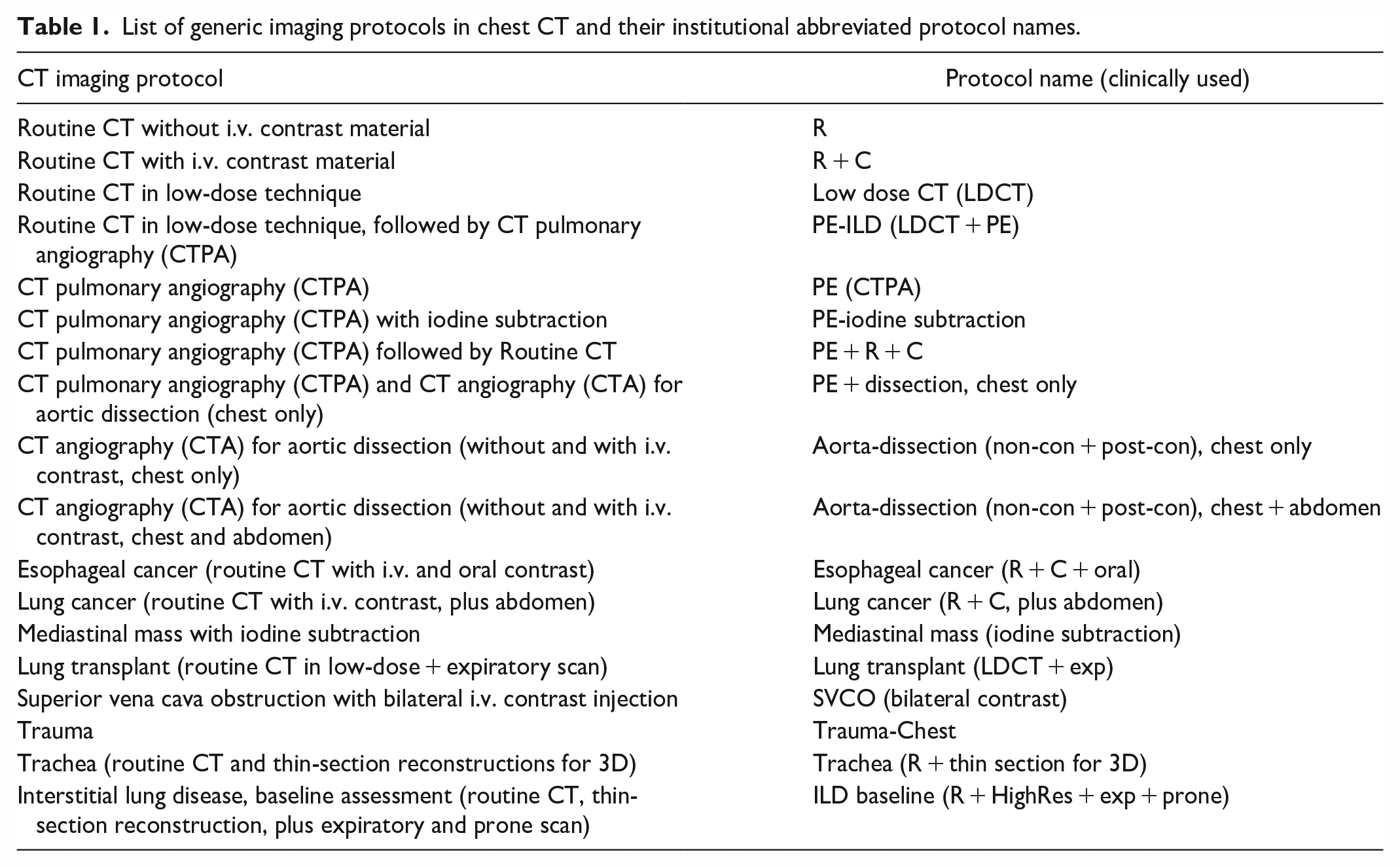
With research ethics board approval, a total of 322 387 chest CT records were extracted from the RIS from 2017 to 2022. Each record contained several different information fields pertaining to the imaging request; for this study, we aimed to predict the “Protocol” field exclusively from the free text associated with the request contained in the Hospital Information System (HIS) field. Returning patient records were not removed as patients may have shifting or changing clinical indications (eg, a cancer patient with acute chest pain may undergo a chest pulmonary angiography [CTPA] for suspected pulmonary embolism), and the comment field is manually entered by the referring physician in free-text style. Records that were missing either of these fields were removed, for instance, when no clinical information or comment was provided or the assigned protocol was not documented (n = 55 488), and records with infrequently occurring protocols (more than 150 protocols with less than 100 instances, n = 3503) were also removed from the dataset, leaving 18 unique protocols reflecting current clinical practice (Table 1). Three hundred randomly selected records were then held-out as an evaluation dataset, resulting in a total of 263 096 records available to train the model.
List of generic imaging protocols in chest CT and their institutional abbreviated protocol names.

Free-Text Pre-Processing and Classification Model
Firstly, a regular expression was used to remove the standardized text from the report template. CLEVER (CL-inical EV-ent R-ecognizer) word substitutions 10 were then applied to the remainder of the free text. The CLEVER text analysis tool has been clinically validated for information extraction from electronic medical records and has been used in similar works.11,12 In this case, the word substitutions help to reduce the size of the vocabulary by replacing different phrases with similar meanings with a single token across the dataset. Punctuation was subsequently removed. The text was then tokenized, and stopwords were removed using the nltk Python package. Very low-frequency words were then removed from the text, with the best threshold experimentally determined to be <50 occurrences in the entire dataset. Frequently occurring word pairs were concatenated with a hyphen to become a single token. The best-performing threshold was >1000 occurrences in the dataset, wich was also experimentally determined. A main motivator for this step was to preserve variations of the phrase “no contrast” as a single feature for the otherwise order-agnostic bag-of-words model. The bag-of-words model was then created using the term frequency-inverse document frequency weighting scheme. 13 The resultant sparse vector-space encoding was then used as the input to a multinomial logistic regression model with the “saga” solver and using the l2 penalty, trained using 10-fold cross-validation and implemented using the sklearn python package. A grid search was performed during the 10-fold cross-validation to determine the best-performing parameters for the pre-processing steps. The best-performing parameters were then used to train the model on the entire dataset. Figure 1 depicts the data processing pipeline.
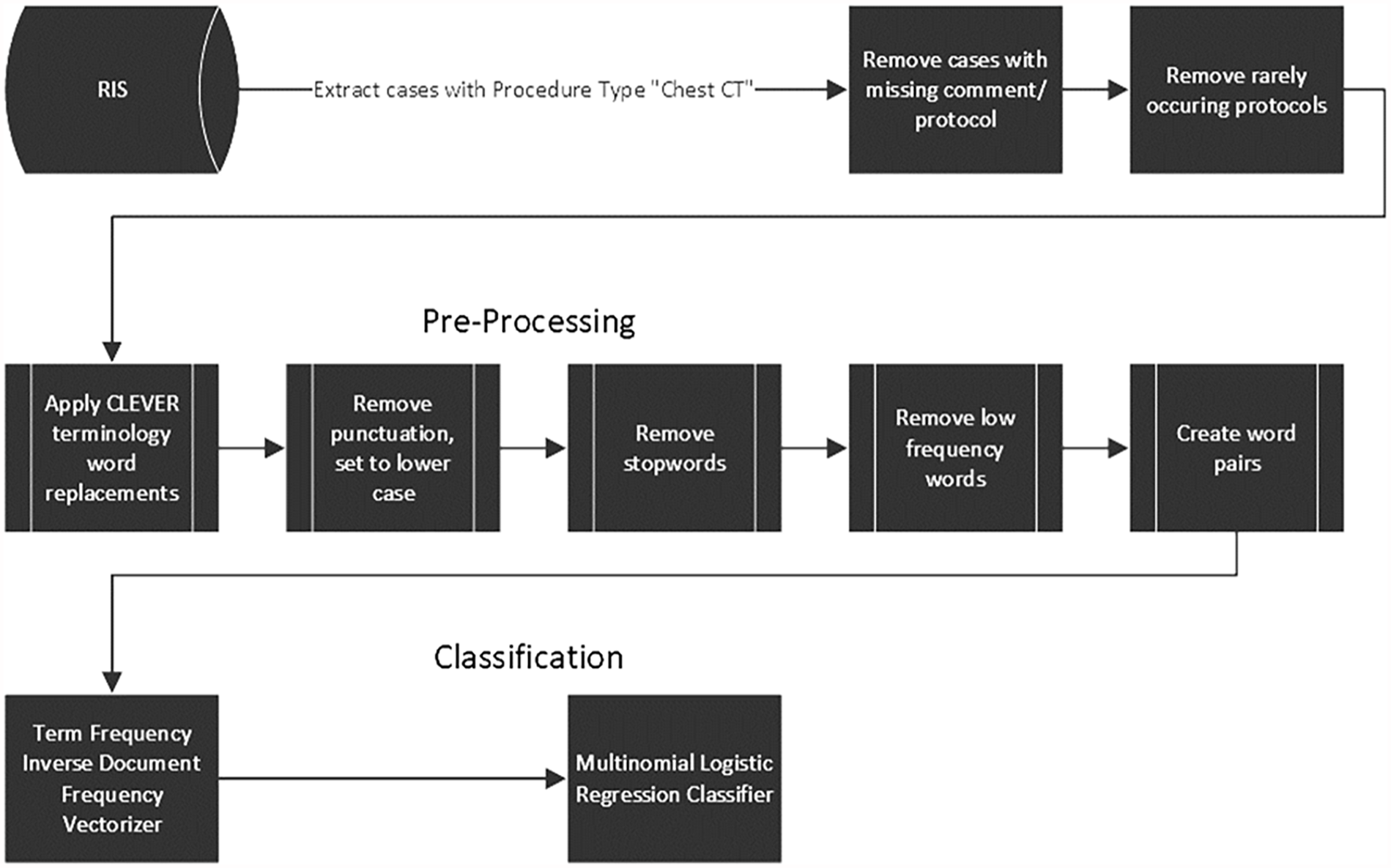
Flow chart depicting the data processing steps from the database to the classifier.
Clinical Evaluation
To evaluate the accuracy of the PRS and the clinically executed protocol (CEP) selected by a radiologist, the 300 cases held-out from the training dataset were used as an evaluation dataset. These cases were randomly sampled from the training dataset, but with the condition that each protocol occurs at a minimum of 10 times in the evaluation set. The distribution of protocols was different between the training and evaluation sets, as matching the distribution as closely as possible while also ensuring that each protocol is adequately represented would require the validation set to be much larger than 300 cases. Four fellowship-trained chest radiologists with 6, 15, 21, and 33 years of clinical experience, respectively, were asked to read through the clinical information for each of the evaluation cases and select one of the 18 protocols. After a reader made a selection, both the PRS’ and the clinically executed protocols were unblinded to the reader, who was then asked to grade both protocol selections against their own selection on a scale from 1 to 4 (1 = severe error, 2 = moderate error, 3 = disagreement but acceptable, 4 = agreement). The readers were allowed to change their minds after the PRS and the CEP were uncovered, and the change was recorded. The reference standard protocol for each case was eventually determined as the most frequently selected protocol by the 4 readers. In cases of a tie (eg, 2 readers select protocol A, 2 readers select protocol B), a judge reader (senior staff) would be used as a tiebreaker.
The protocol selections by the PRS and the CEP were defined as accurate when they matched the protocol of the reference standard. The accuracies of the PRS and the CEP were statistically compared using McNemar’s test. Two measures were determined from readers’ evaluation scores: the “correctness,” defined as the proportion of cases where the majority of readers assessed a grade of 4; and the “acceptability,” defined as the proportion of cases where the majority of readers assessed a grade of at least 3. The clinical performance of the PRS was assessed based on the readers’ evaluation scores, and the readers’ scores 1 and 2 were recorded to calculate the mistake rates of both the PRS and the CEP. The reliability of the readers’ evaluations was measured using Fleiss’ Kappa.
Results
Initially, the 300-case evaluation dataset contained a minimum of 10 cases for all 18 protocols. After the “reader-agreement” reference standard was created, all cases historically assigned to 2 of these 18 protocols (Lung Cancer, Mediastinal Mass) were re-labelled as another protocol by all readers, effectively removing those 2 protocols from the evaluation set. Similarly, 3 protocols (Trachea, PE-ILD, and ILD baseline) lost 1, 2, and 2 cases, respectively, in the final evaluation set because all readers re-labelled them with another protocol. The final distribution of protocols in both the training/test dataset, and the evaluation dataset (as determined by the “reader-agreement” reference standard) is shown in Table 2.
Overview of the Distribution of Protocols in Both the Train/Test Data and the Held-Out Evaluation Dataset, the Accuracy of the PRS and Clinically Executed Protocol Compared to the Ground Truth.
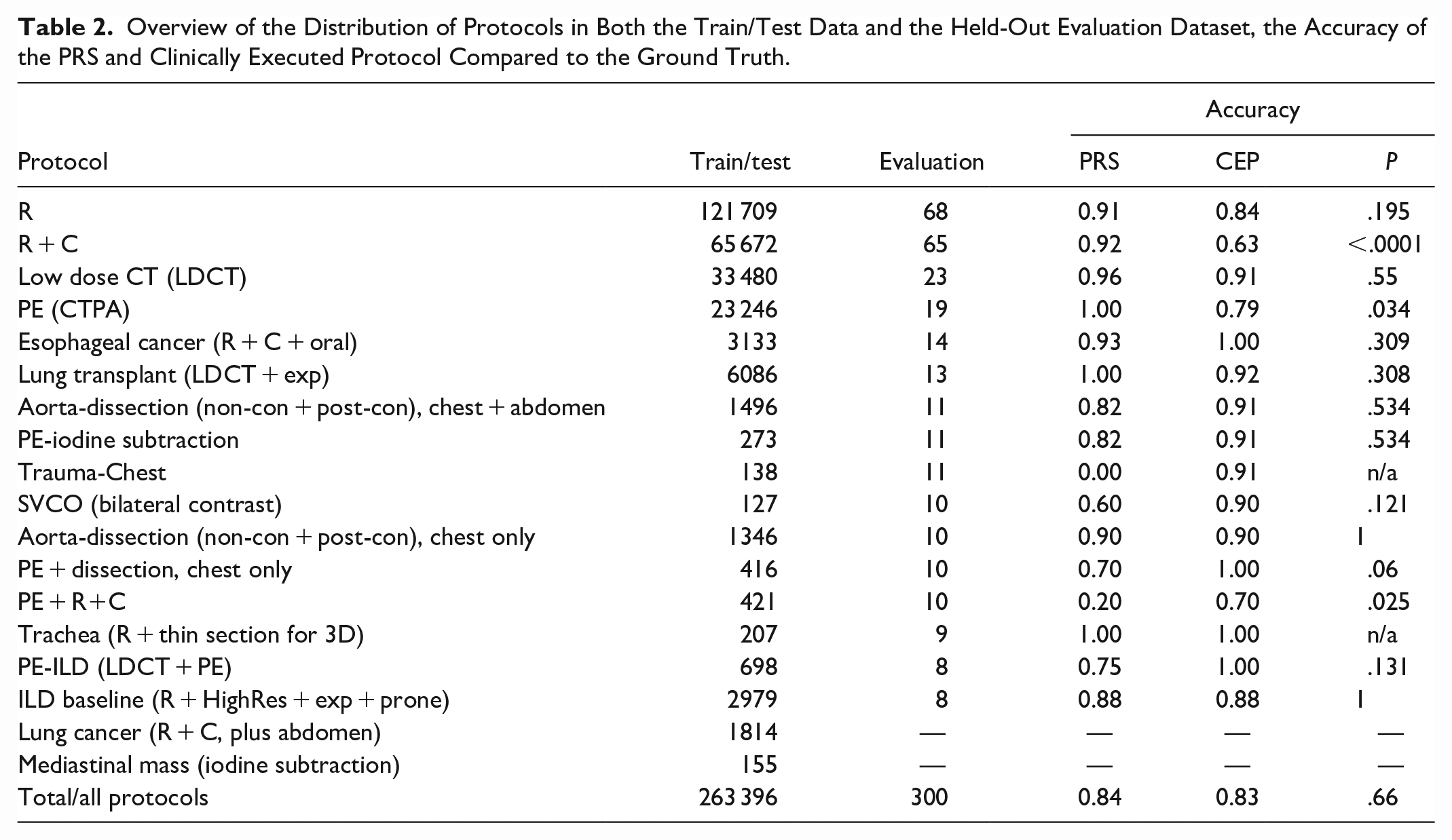
Only one reader changed their mind in 3 cases and selected different protocols after they had seen the PRS and CEP protocols. The accuracy of both the PRS and CEP for all cases was 84.3% and 83.0%, respectively. Table 2 shows the accuracies for each protocol. The PRS was found to perform significantly better in one protocol (“R + C”), with
Both the “correctness” and “acceptability” of the PRS and the CEP selection, calculated from the evaluation grades, are shown in Figure 2. Table 3 summarizes the performance of the PRS based on the readers’ scores, both overall and broken down by individual readers, as well as the readers’ classification of mistakes made by the PRS an CEP. For all protocols, the correctness of the PRS and the CEP was 84.3% and 83%, and the acceptability 98.6% and 99.3%, respectively. One reader classified 8 PRS selections (2.7%) as mistakes, and another reader classified 6 CEP selections (2%) as mistakes; a detailed breakdown of all 4 readers is also shown in Table 3.
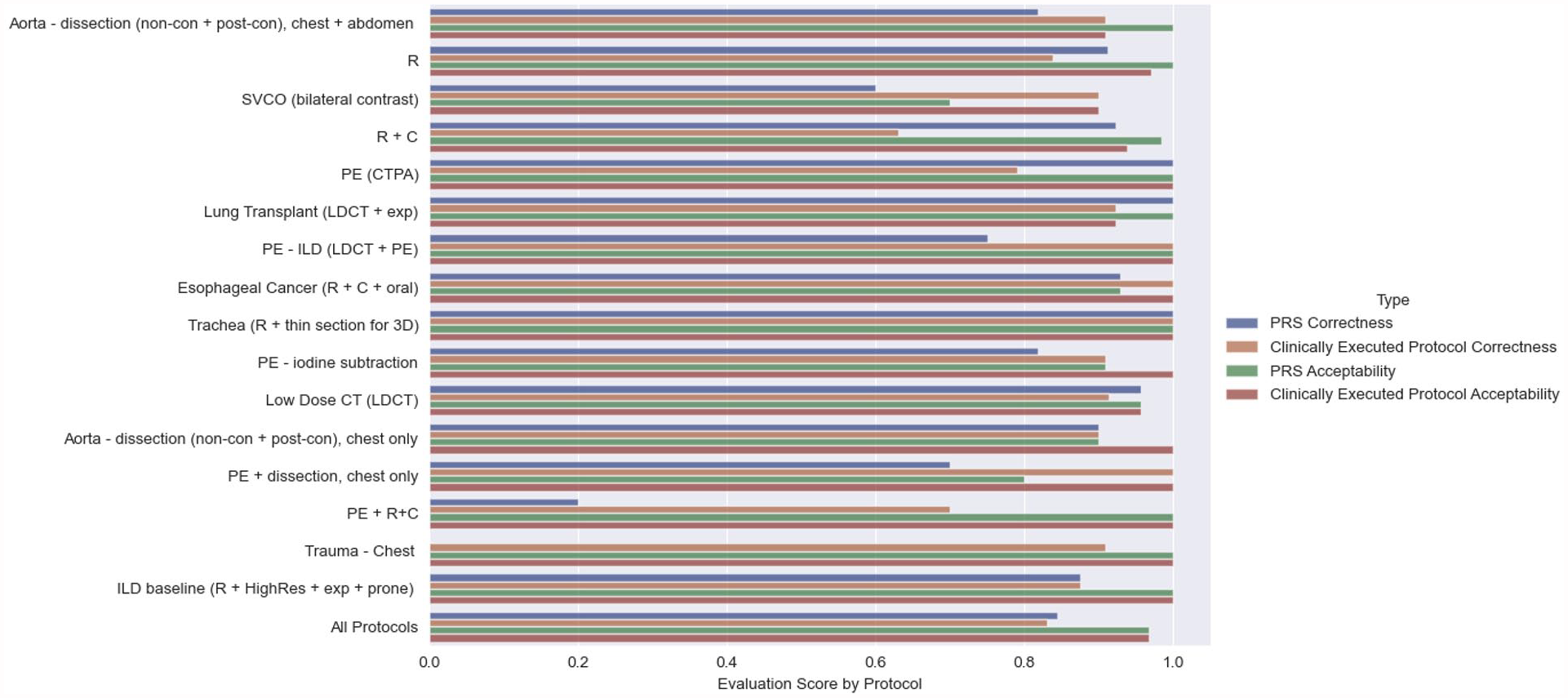
PRS-selected and clinically-executed protocols, as graded by readers. “Correctness” was defined as the proportion of cases where the majority of readers gave a score of 4. “Acceptability” was defined as the proportion of cases where the majority of readers gave a score of 3 or 4.
Performance of the PRS Based on the Evaluation Scores, Overall and by Individual Readers. Mistakes of the PRS and CEP, as Scored by the Readers, Vary Between 0.7% and 2.7%.
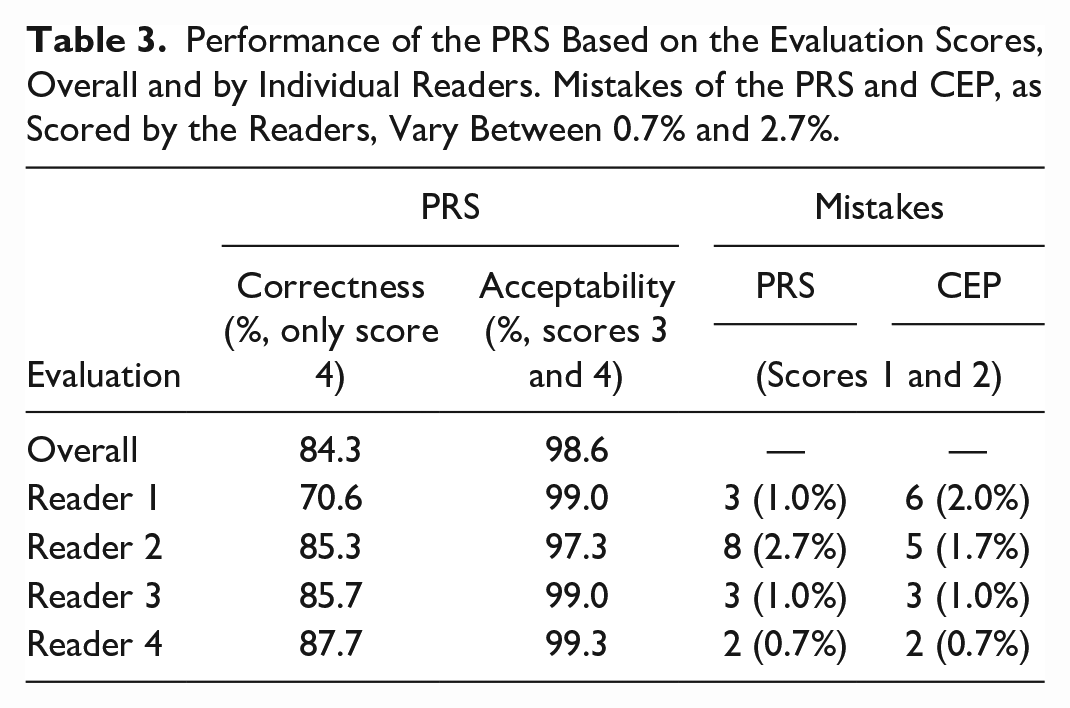
Figure 3 shows the level of agreement between readers for each case in the evaluation dataset, split by protocol. Overall, all 4 readers assigned the same protocol in only 203 out of 300 cases (67.6%), 3 readers agreed on the same protocol in 83 out of 300 protocols (28%), and in 15 cases, a judge reader needed to break the tie (5%). The Fleiss Kappa for all readers and protocols was 0.805. Supplement 1 shows example cases from the evaluation dataset.
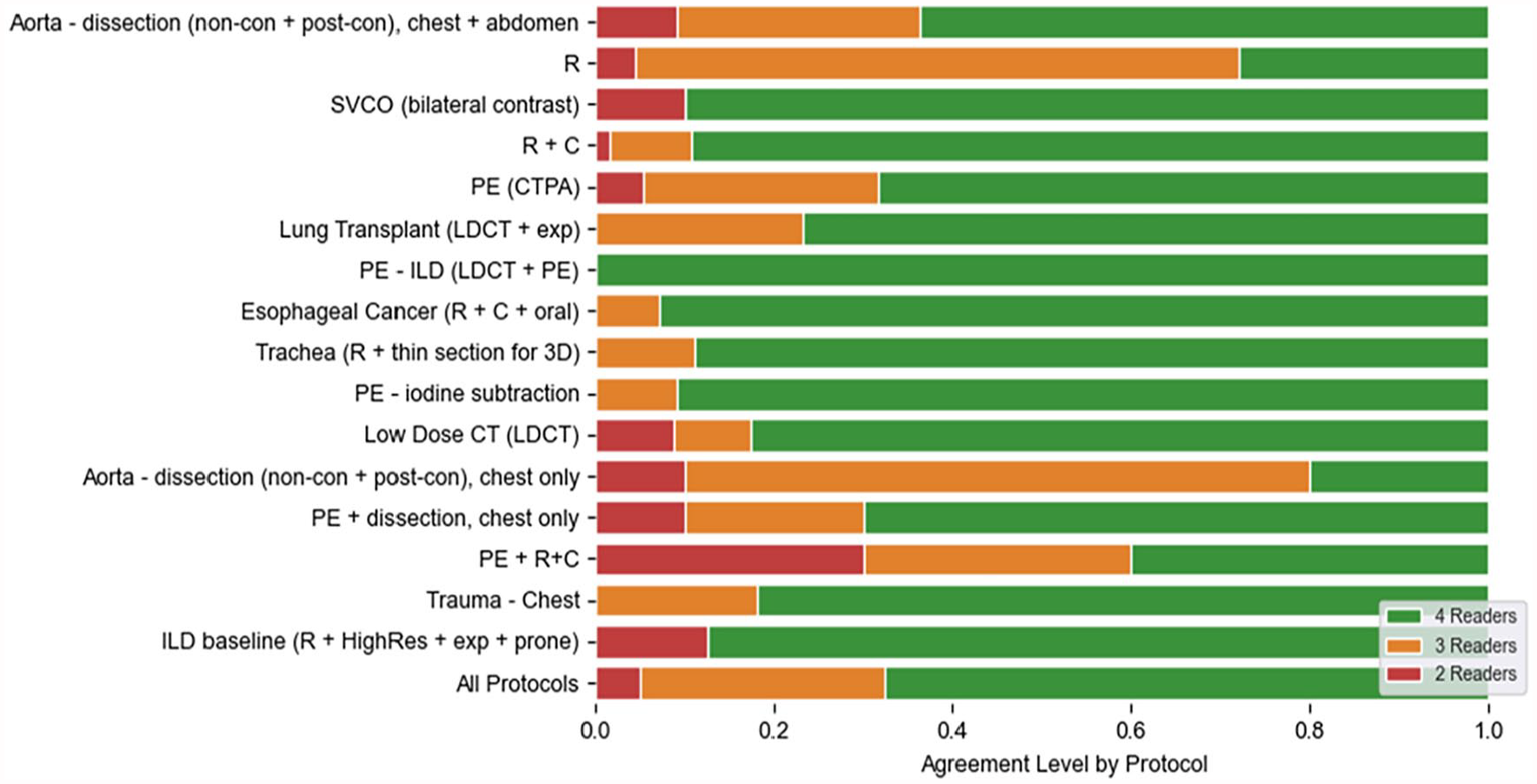
Level of agreement between readers, split by protocol.
Discussion
The overall accuracy of the automated PRS for chest CT matched the human performance. While the PRS’ overall accuracy of 84.3% against the reader-agreement reference standard leaves room for improvement, this performance was still slightly better than the clinically executed protocol (CEP) with an 83% accuracy, albeit not significantly. The CEP accuracy being as low as 83% is surprising, implying that a panel of radiologists will advise for a different protocol than was clinically executed in 17% of cases. The acceptability score of the CEP, however, was much higher at 99.3%, implying that these cases were not necessarily medical errors. The performance differed on a per-protocol basis, with the PRS performing significantly better on a routinely chosen chest protocol “R + C” (PRS: 92.3% vs CEP: 62.1%), but significantly worse on the protocol for chest trauma “Trauma- Chest” (PRS: 0% vs CEP: 90.9%) and for the biphasic protocol “PE + R + C” (PRS: 20% vs CEP: 70%).
The PRS’ superior performance for the routine chest with contrast protocol is surprising as labels in the training dataset were determined by the radiologist-chosen, clinically executed protocol. The differences between the “reader-agreement” reference standard and the clinically executed protocol may be caused by inter-reader variances or possibly by shifting clinical standards over time, as the evaluation dataset was sampled randomly from the training dataset across the entire 5-year window. The poorer performance for the 2 other protocols may be attributed to their lower frequency in the training dataset and the tendency of the PRS to default to the most commonly used protocol when the clinical history is sparse or of limited clinical relevance.
The training dataset (and evaluation dataset) was highly imbalanced, where the top protocol chest without contrast (“R”) accounted for approximately 40% of the dataset, whereas 11 of the 18 protocols each individually comprised less than 1% of the total. The PRS largely struggled with these lower-frequency protocols (such as “Trauma-Chest”—a rarely used protocol as our institution is not a trauma centre) but performed comparably or better than the clinically-executed protocol on the more frequent protocols.
Providing a meaningful clinical history and concise clinical query is fundamental to any patient transfer and imaging request.14-16 This is especially critical for imaging studies that can be conducted using various imaging protocols. 17 Additionally, certain protocols can significantly affect patient scheduling due to factors such as exam prioritization, equipment selection, scan time requirements, and patient preparation, underscoring the clinical importance of accurate protocolling. 18 While machine learning demonstrates superiority in uncertain scenarios, inadequate training data can render its output ineffective.19,20
We observed notable concurrent and temporal variability in radiologists’ protocol selection that should not be disregarded. First, the 4 readers who established the concurrent reference standard could only agree on the same protocol in 68% of the clinical requests. Second, 2 out of 18 protocols dropped out of the evaluation dataset because all cases were re-labelled to other protocols than the original, clinically executed protocols. Instead of using the historically executed protocols with cross-validation, which only one radiologist selected at that time, we opted to create a validation dataset based on a four-reader-agreement as a more robust reference standard for testing the PRS and human performance. The purpose was to mitigate against potential variability and errors in the historically executed protocols. Thus, the overall model performance would likely be enhanced with a sufficiently large dataset of cases annotated with this “reader-agreement” reference standard being available for training.
The process of protocolling incoming imaging requests is inherently repetitive and charitably monotonous, leading to a heightened risk of errors due to fatigue. Moreover, radiologists are frequently interrupted during image reading sessions to handle ad-hoc protocols or address emergency requests, which can divert their focus and potentially result in inaccurate image interpretation or protocol selection. 4 Despite the fact that protocolling a single CT examination may take less than a minute, the cumulative demand of timely protocolling all imaging requests prior to execution imposes a substantial burden on radiologists. Chung et al. conducted a multi-year evaluation of an automated protocolling system and demonstrated that it significantly reduced the radiologists’ protocolling workload and decreased the time from order entry to protocol assignment without increasing protocolling errors. 21
As medical imaging is integral to operations in high-reliability organizations (HROs), it is difficult to envision that relying solely on a machine learning model within a single-operator paradigm would address the key challenges of HROs, particularly the complexity, unpredictability, and constant change. 22 An automated PRS could help mitigate some of these risks when employed as a second opinion, akin to the use of a Computer-Aided Detection (CAD) system. 23 Alternatively, a “model-protocols-then-reader-validates” approach may alleviate the cognitive burden by simplifying the protocolling task to a validation task. More importantly, this approach has the potential to reduce variability in protocolling, ensuring that similar cases are managed consistently with minimal reliance on individual reader biases.
The PRS model necessitates continuous training to enhance its accuracy, alongside ongoing data curation efforts to adapt to the dynamic requirements of HRO environments. We observed a higher model performance for protocols extensively represented in the training dataset, indicating potential benefits from employing data augmentation techniques. 8 Protocols with fewer than 100 occurrences over a 5-year period were excluded from the analysis, yet these instances collectively constitute a substantial portion of the dataset, underscoring the importance of augmenting or curating available data resources.
In summary, the automated protocol recommendation system (PRS) for chest CT imaging requests achieved similar accuracy to human performance and may help radiologists master the ever-increasing workload by alleviating the burden of monotonous protocol selection, though challenges with less frequent protocols remain to be solved.
Supplemental Material
sj-docx-1-caj-10.1177_08465371241280219 – Supplemental material for Development and Evaluation of an Automated Protocol Recommendation System for Chest CT Using Natural Language Processing With CLEVER Terminology Word Replacement
Supplemental material, sj-docx-1-caj-10.1177_08465371241280219 for Development and Evaluation of an Automated Protocol Recommendation System for Chest CT Using Natural Language Processing With CLEVER Terminology Word Replacement by Patrik Rogalla, Jennifer Fratesi, Sonja Kandel, Demetris Patsios, Farzad Khlavati and Sean Carey in Canadian Association of Radiologists Journal
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
