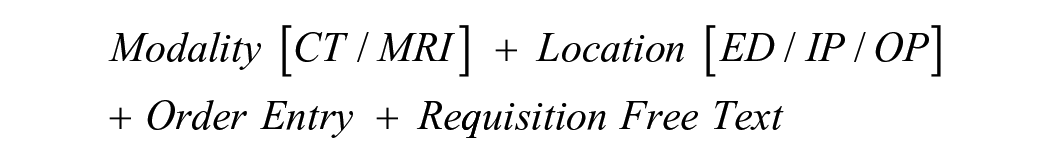Abstract
This is a visual representation of the abstract.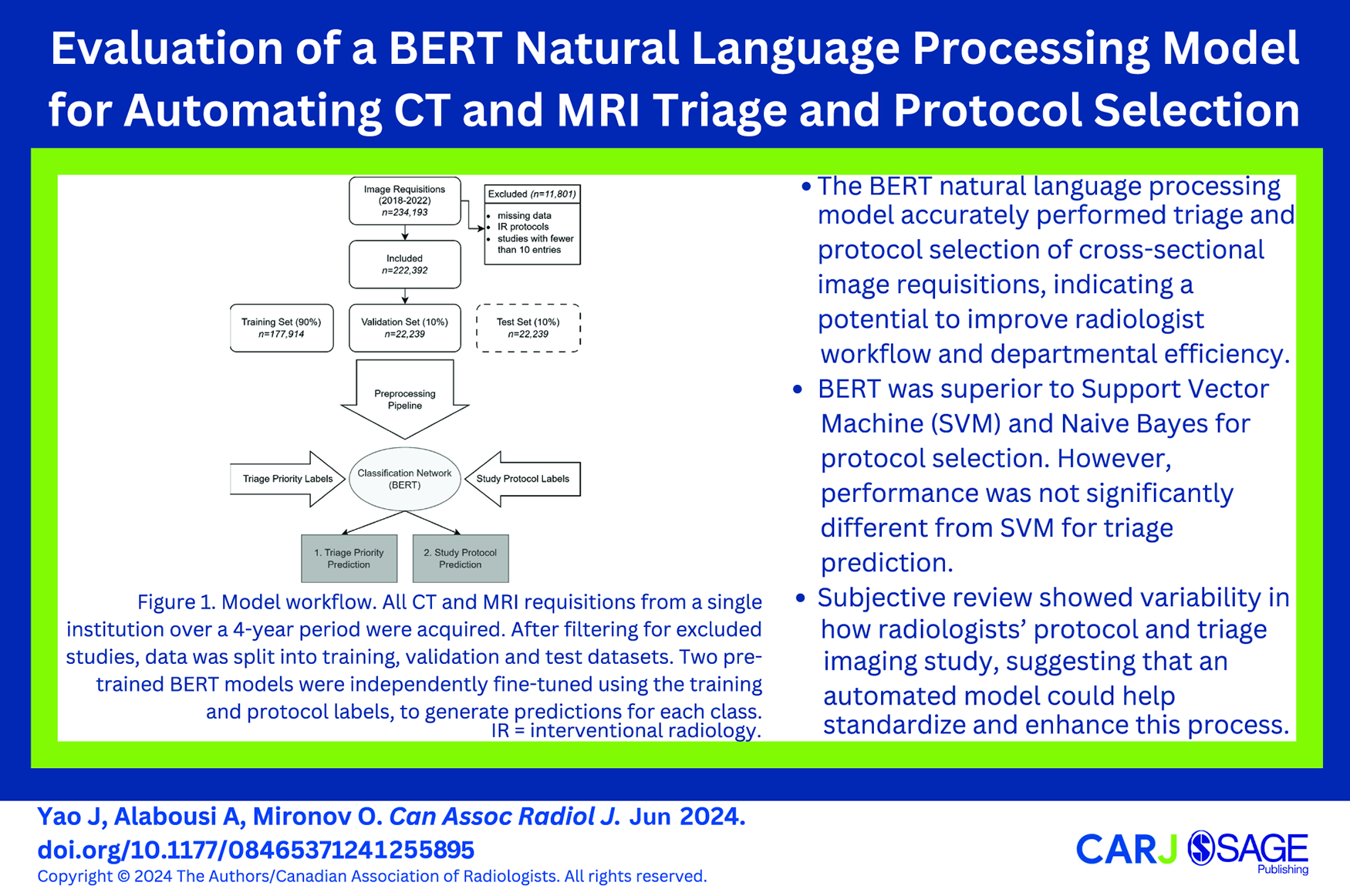
Introduction
In a busy clinical environment, imaging requisitions are often the sole avenue of communication between clinicians and radiologists. Due to the volume of imaging requests received daily, radiologists are limited in their ability to evaluate each request in depth and must rely on short vignettes to determine both the most appropriate imaging study and priority in which the study should be performed. However, the process of reviewing and approving these requests can be time-consuming and disruptive to the radiologist’s reading workflow. A survey of radiology trainees has acknowledged that protocolling is a source of exhaustion and burnout. 1 Additionally, errors such as study requests that are incongruent with the clinical indication, or incorrect study approval by the radiologist, may lead to non-diagnostic studies being performed, or longer patient wait times. 2 Automatic triaging/protocolling using artificial intelligence and machine learning has been suggested as a potential solution to this problem. Machine learning has been proposed for workflow optimization tasks in radiology, including order entry support, study protocolling, study triaging, and clinical decision support.3-5 These tasks are often accomplished using natural language processing (NLP) models, which can understand, interpret, and generate language. Advanced models, such as BERT (Bidirectional Encoder Representations for Transformers), are a recent development in the field, improving on traditional NLP using a masked language approach. 6 These models capture contextual information that may be lost in a traditional unidirectional (left-to-right or right-to-left) training, allowing for a better semantic understanding of text.
Previous studies have shown that natural language processing models such as BERT can be effectively used to automatically process text data including clinical requisitions and radiology reports.7,8 However, our study proposes the first comprehensive tool for both triaging and protocolling studies across a general cross-sectional imaging dataset, with the goal of approximating the diversity of protocols encountered during daily practice.
The objective of our study is to assess the accuracy of BERT for triaging/protocolling CT and MRI image requisitions. We compare our BERT model to 2 common traditional machine learning algorithms, Naive Bayes (NB) and Support Vector Machine (SVM). Finally, we assess model subgroup performance based on referral location and modality.
Methods
This was a retrospective study performed with approval from the Institutional Review Board of a large Canadian university. In keeping with the policies for a retrospective review, informed consent was not required. Studies were acquired at a single centre university-based database. The study was conducted according to the Checklist for Artificial Intelligence in Medical Imaging (CLAIM) guideline. 9 Consecutive deidentified CT and MRI requisitions performed between January 2018 and September 2022 were extracted from the electronic medical record (EMR). Requisitions originated from a combination of a large tertiary care hospital, emergency department, and outpatient/primary care requisitions. Protocols were clinically assigned by staff radiologists, radiology trainees, and medical radiation technologists under medical directives. A total of 234 192 studies were collected. Order entry details including requisition text, requested exam, and patient location (outpatient [OP], emergency [ED], inpatient [IP]) were acquired. Additionally, the radiologist-assigned Wait Time Information System (WTIS) triage priority (P1/emergent-P4/non-urgent) and protocol were compiled. The dataset was filtered for requisitions with missing fields and interventional-radiology specific protocols, which accounted for 11 654 studies. Duplicate protocols were consolidated, and protocols with fewer than 10 entries were excluded, reducing the number of unique protocols from 535 to 300. In total, 222 392 studies were included (Figure 1).
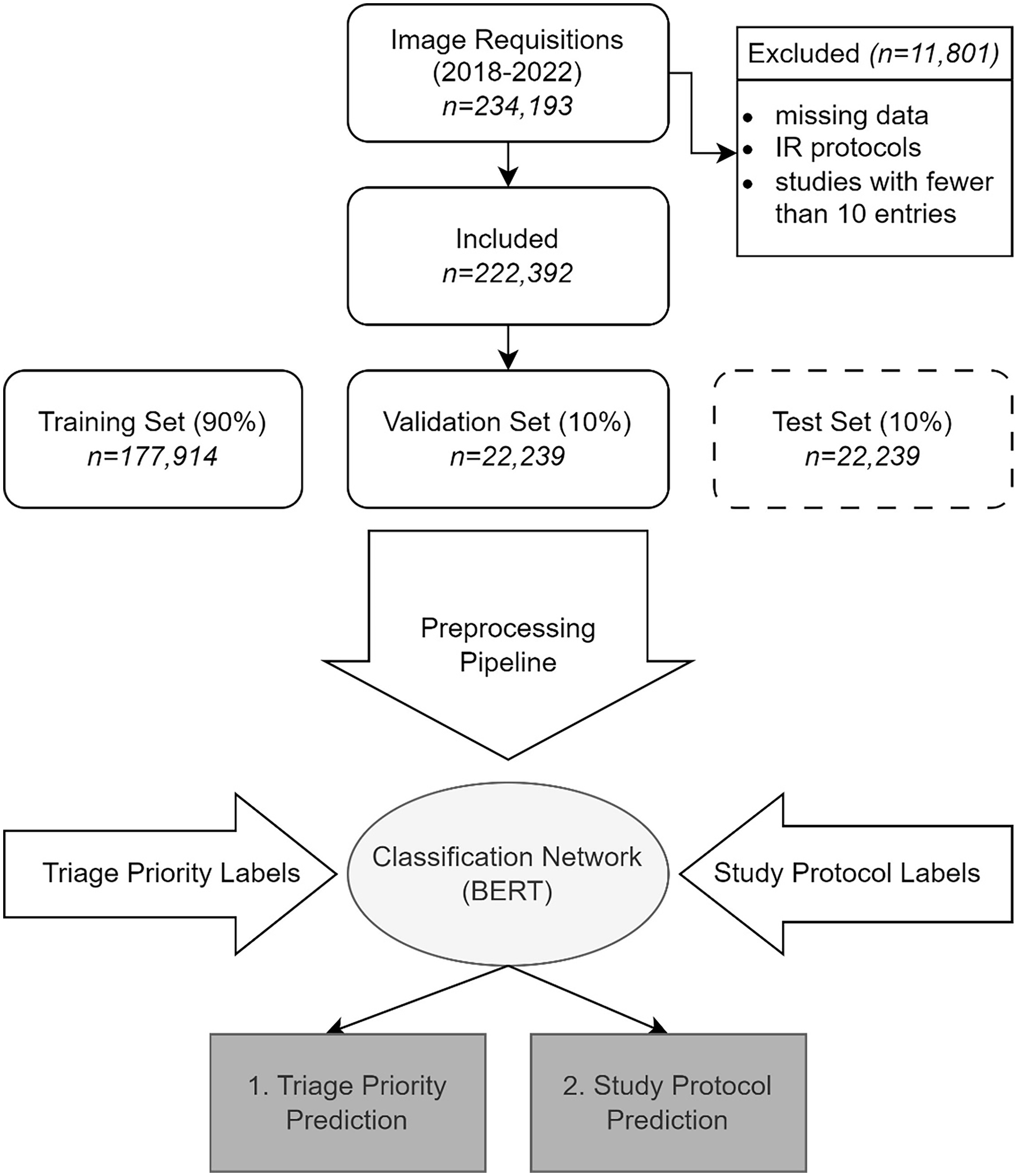
Model workflow. All CT and MRI requisitions from a single institution over a 4-year period were acquired. After filtering for excluded studies, data was split into training, validation, and test datasets. Two pre-trained BERT models were independently fine-tuned using the training and protocol labels, to generate predictions for each class.
Network Design
Studies were split into training, validation and test datasets based on stratified random sampling using an 80%-10%-10% split, respectively. Input text was structured as a string based on order entry details, as follows:
where order entry refers to the exam selected in the EMR, based on a pre-defined list of study categories (eg, MR head non standard, CT chest). The protocol and triage priority assigned at time of exam were used as ground-truth labels.
Data was pre-processed using standard techniques, consisting of lower casing, punctuation removal, stop word removal, and lemmatization. Dates and patient age were removed to improve model generalizability.
Two independent models for study protocol and triage prediction were developed. For both models, the BERTBase tokenizer of the Huggingface transformer library were used. 10 Models were fine-tuned on the training dataset, using the Adam optimizer with a learning rate of 5-e5 with warmup of 10% of training steps, and batch size of 12, for up to 10 epochs. Early stopping with a patience of 3 was used to prevent overfitting. The model was trained on a NVIDIA RTX 2070 using TensorFlow v2.10 and the transformers library v4.29.2.10,11
For comparative purposes with the BERT model, 2 traditional machine learning classifiers were trained: NB and SVM with a linear kernel. These classifiers utilized term frequency-inverse document frequency (TF-IDF) weighted bag-of-words features and were implemented using scikit-learn. 12 The same data splits were consistently employed across all models for training, validation, and testing.
Statistical Analysis
For all the models evaluated, key metrics of F1 score, precision, and recall were obtained. To accommodate imbalances in the dataset, both weighted and macro-averaged statistics were computed. The weighted statistics were derived from a weighted average of the outcomes for each ground-truth label, which is more sensitive to class imbalances. In contrast, the macro statistics were computed as a true average, unaffected by the size of each group. Pairwise comparison of the models was performed using a McNemar test with Bonferroni correction on the aggregated dataset. Micro-averaged, area under the receiver operating characteristic curves (AUROC), using a one-versus-rest approach, were also calculated for both models.
In addition to these measures, the models underwent assessment for multi-class protocol and triage suggestion, in the context of a workflow-support model to be used in conjunction with radiologist review. The top-1, top-3, and top-5 accuracy assessed for the protocol model, and the top-1 and top-2 accuracy were assessed for the triage model. A location and imaging modality-based sub-analysis was also conducted to better understand how the models perform in different clinical settings.
Results
Request Data
A total of 222 392 studies were used for model training and evaluation. Of these, 139 024 studies were CT scans (62.5%) and 83 368 were MRIs (37.5%). The distribution of the studies based on their originating departments was as follows: 48 447 (21.8%) from the emergency department, 24 306 (10.9%) from inpatient departments, and 149 639 (67.2%) from outpatient services. Based on imaging section, the distribution was as follows: 67 461 (30.3%) for abdomen, 3594 (1.6%) for breast, 34 679 (15.6%) for chest/cardiac, 30 647 (13.8%) for musculoskeletal (MSK), and 86 011 (38.7%) for neuro. The average requisition length was 10.38 words (standard deviation 7.87).
Of the 300 unique protocols, 116 (38.7%) were for CTs and 184 (61.3%) were for MRIs. By imaging section, 81 (27.0%) unique protocols were for abdomen, 3 (1.0%) for breast, 17 (5.7%) for chest/cardiac, 62 (20.7%) for MSK, and 137 (45.7%) for neuro. The frequency distribution of protocols was notably imbalanced. The top 5 most assigned protocols constituted 33.4% of the dataset, while the 25 most common protocols accounted for 67.7%. The most commonly used CT protocol was “CT Brain without Contrast,” and the most commonly used MR protocol was “MR Lumbar Spine” (Figure 2). The mean number of cases per protocol was 723 with a standard deviation of 2353.
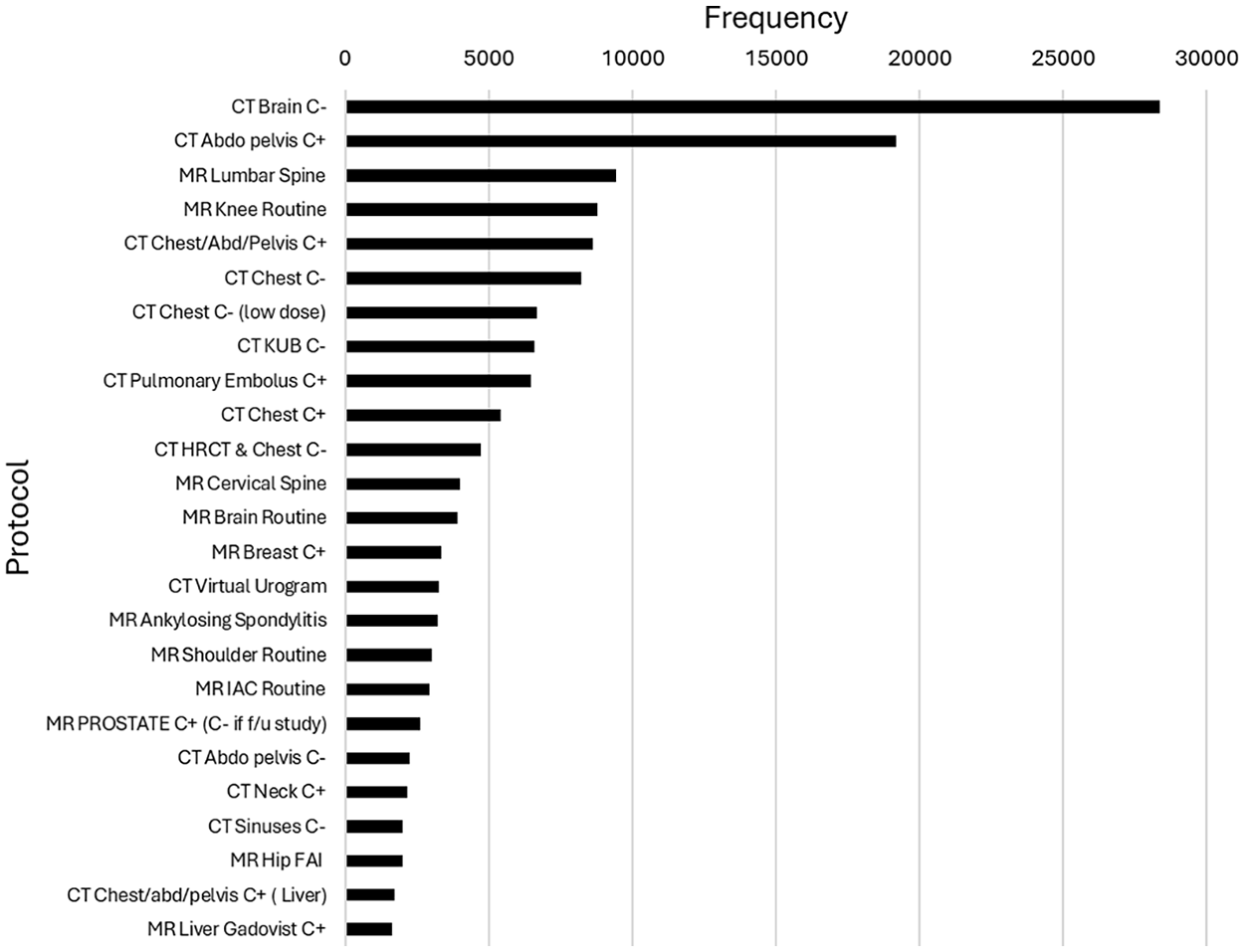
Top-25 most-commonly assigned study protocols.
The distribution of triage priority was also imbalanced. Most studies were triaged as P4 (50.1%), followed by P1 (27.5%), P3 (18.1%), and P2 (4.3%).
Model Performance
Weighted and macro-averaged statistics are presented in Table 1. For study protocol, the weighted F1 score on the test dataset was 0.901, while the macro F1 score was 0.658 (Table 1). For triage priority, the weighted and macro F1 scores were 0.844 and 0.754, respectively. Precision and recall are similarly high for both models. AUROC was 0.9995 for protocol and 0.9755 for triage selection (Figure 3).
Summary Statistics of F1, Precision, and Recall for the Protocol and Triage Networks, on the Test Dataset. Weighted- and Macro-Averaged Statistics Provided.
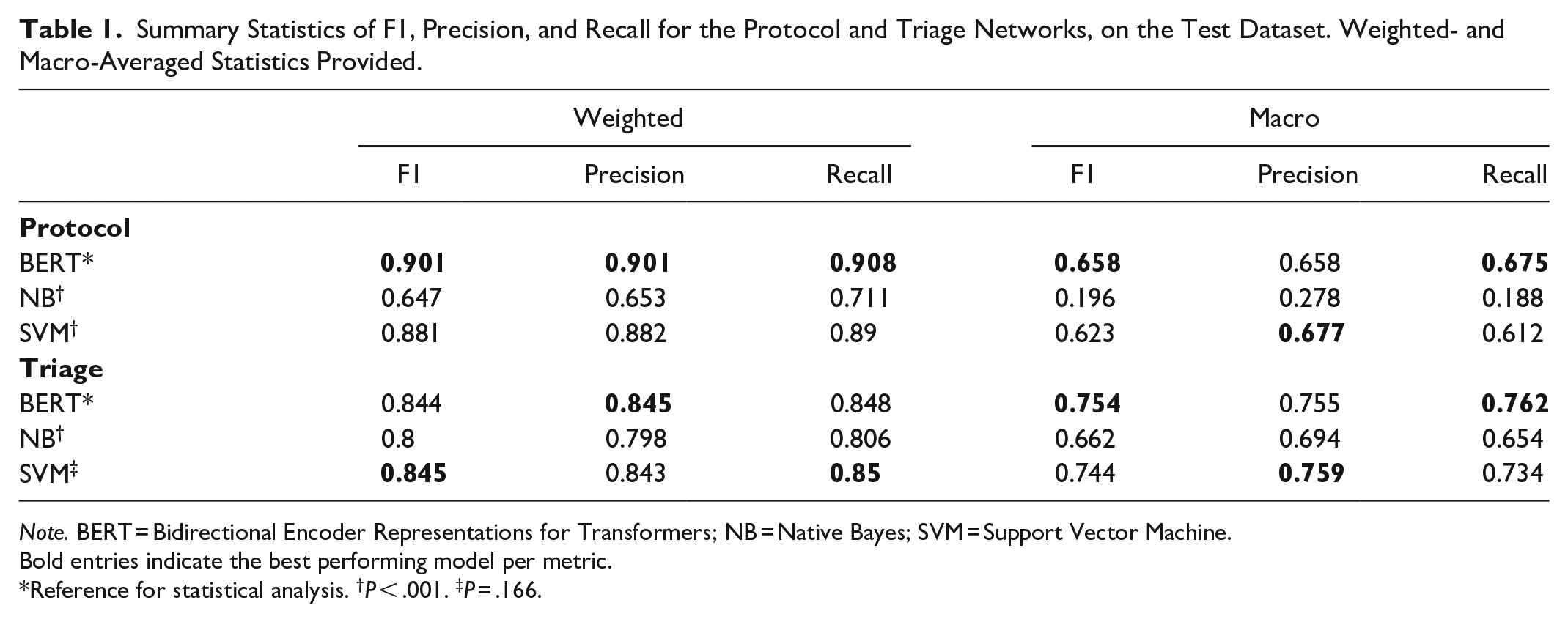
Note. BERT = Bidirectional Encoder Representations for Transformers; NB = Native Bayes; SVM = Support Vector Machine.
Bold entries indicate the best performing model per metric.
Reference for statistical analysis. †P < .001. ‡P = .166.
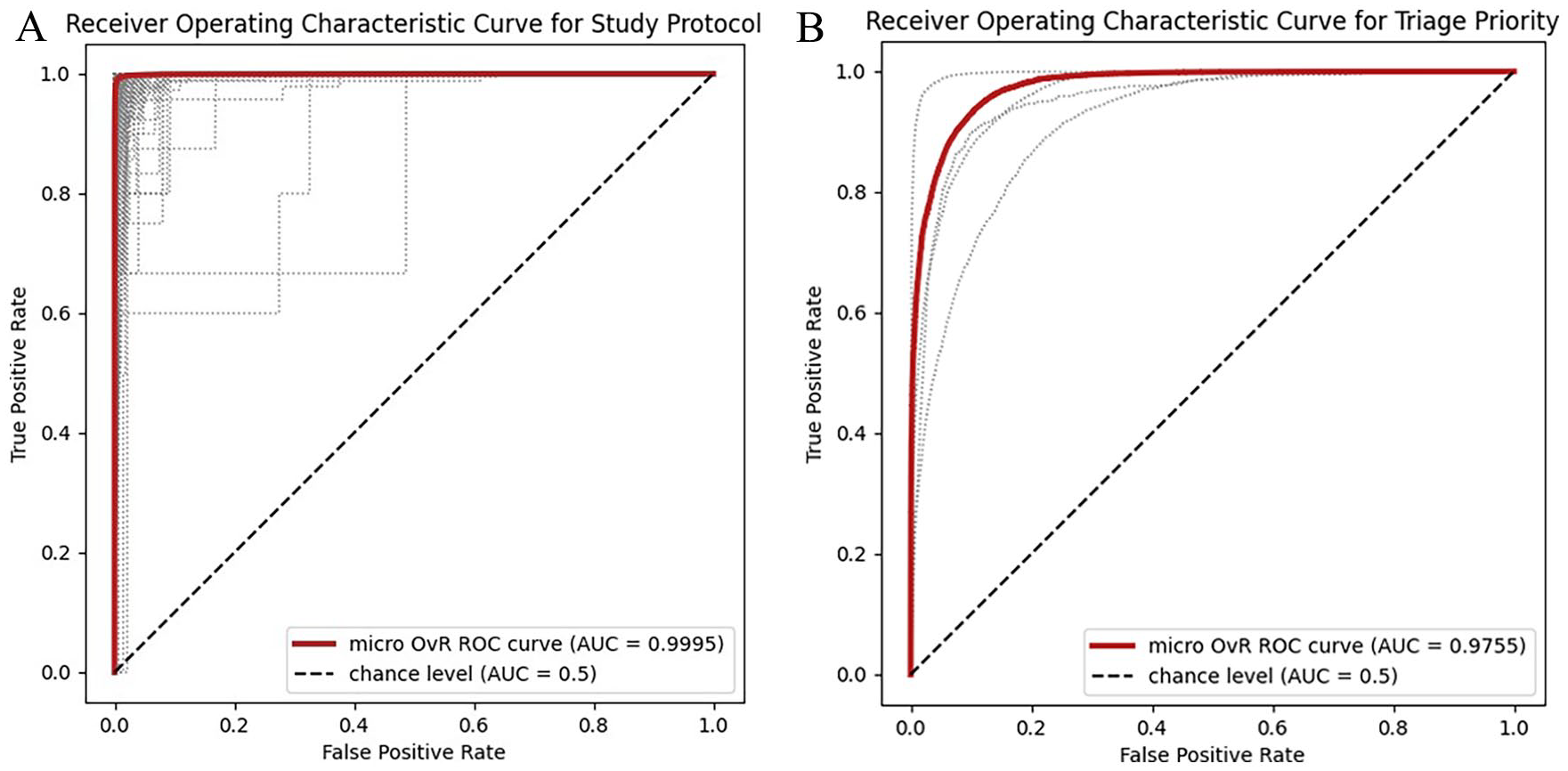
Receiver Operating Characteristic curves for (A) study protocol and (B) triage priority models. Area under the curve was calculated using a micro-averaged, one-versus-rest technique. Fine-dashed lines represent individual ROC curves for each label.
Compared to the traditional machine learning models, BERT had superior performance to SVM for the protocol selection task (P < .001), with higher weighted (0.901 for BERT vs 0.881 for SVM) and macro (0.658 for BERT vs 0.623 for SVM) F1 scores. For the triage task, the BERT model performance was not significantly different from SVM (P = .166) with weighted F1 of 0.844 for BERT and 0.845 for SVM. Both BERT and SVM models significantly outperformed the Naive Bayes models in both tasks (P < .001).
In the context of multiclass suggestion, the accuracies for the protocol model were 0.905 (top-1), 0.984 (top-3), and 0.991 (top-5). For the triage model, the accuracies were 0.848 (top-1) and 0.981 (top-2).
Table 2 displays outcomes stratified by subgroups. When stratified by location, patients coming from the emergency department had the highest F1 scores for both protocol selection and triage priority (0.957 and 0.986, respectively). By modality, CT achieved a higher F1 score for protocol (0.899 for CT vs 0.895 for MR), while MR achieved a higher F1 score for triage (0.825 for CT vs 0.871 for MR). By imaging section, breast and MSK achieved the highest F1 scores for protocol (0.983 for breast and 0.962 for MSK), while MSK and neuro achieved the highest F1 scores for triage (0.927 for MSK and 0.892 for neuro). Protocol F1 score was lowest for chest imaging (0.807), although F1 improved to 0.947 if the protocols “CT chest” and “CT chest (low dose)” were consolidated.
Subgroup Analysis of Weighted-F1, Precision, and Recall for the BERT Protocol and Triage Models, Stratified by Patient Location, Study Modality, and Imaging Section.
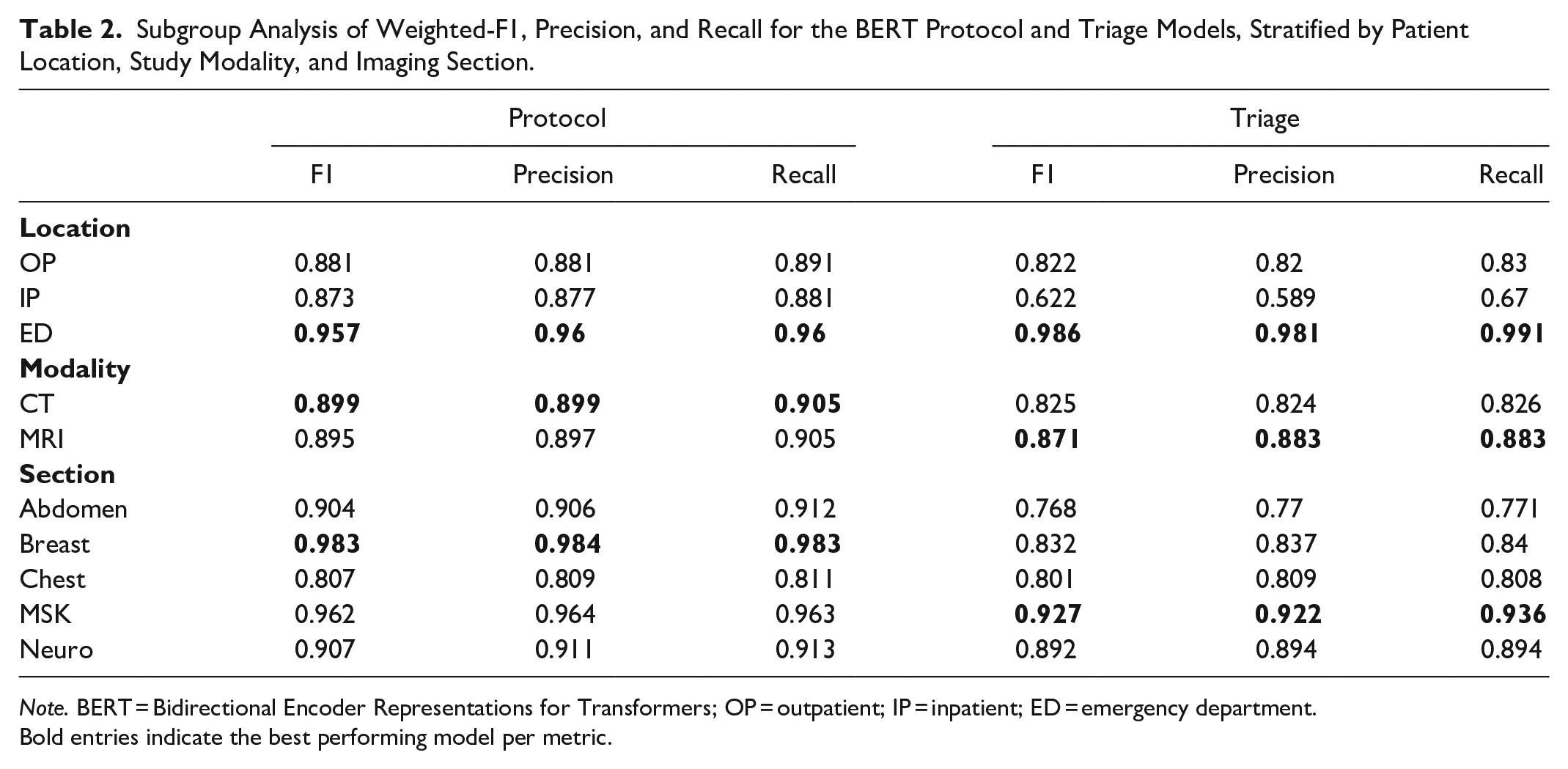
Note. BERT = Bidirectional Encoder Representations for Transformers; OP = outpatient; IP = inpatient; ED = emergency department.
Bold entries indicate the best performing model per metric.
Subjective Analysis
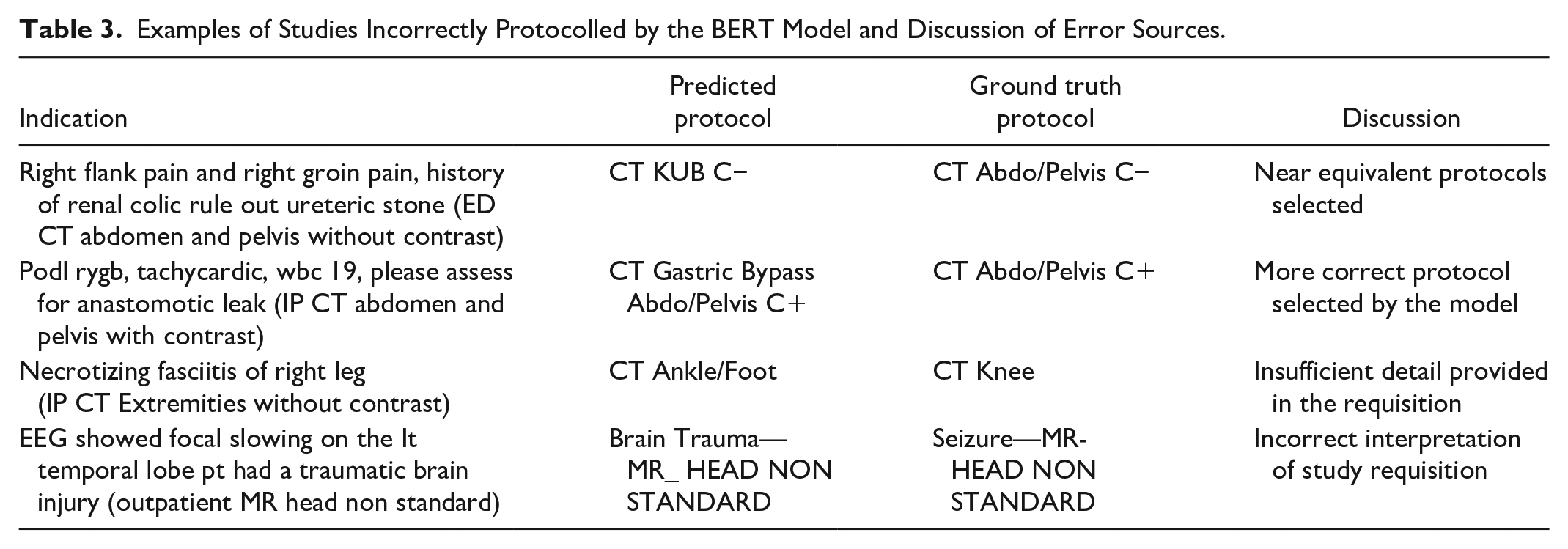
An analysis of discrepant classifications revealed several common factors contributing to inaccurate protocol and triage prediction (Table 3). For protocol, many causes were discrepant due to near-equivalent studies being selected. For instance, for a requisition to rule out ureteric stone, in one instance the ground-truth protocol was “CT Abdo/Pelvis C−,” while the model predicted “CT KUB C−.” These are near equivalent protocols, with no significant limitation caused by the substitution. In other instances, the +model suggested a more accurate protocol than the study performed, possibly related to human error in protocol selection. A frequent cause for discrepancy was vague requisitions. For example, in a requisition to rule out necrotizing fasciitis where no anatomic region was specified, the protocol was misclassified by the model due to the missing information. This highlights a limitation of the automated system, whereas a radiologist could access the patient’s electronic medical record or communicate with the clinical team to clarify ambiguities.
Examples of Studies Incorrectly Protocolled by the BERT Model and Discussion of Error Sources.
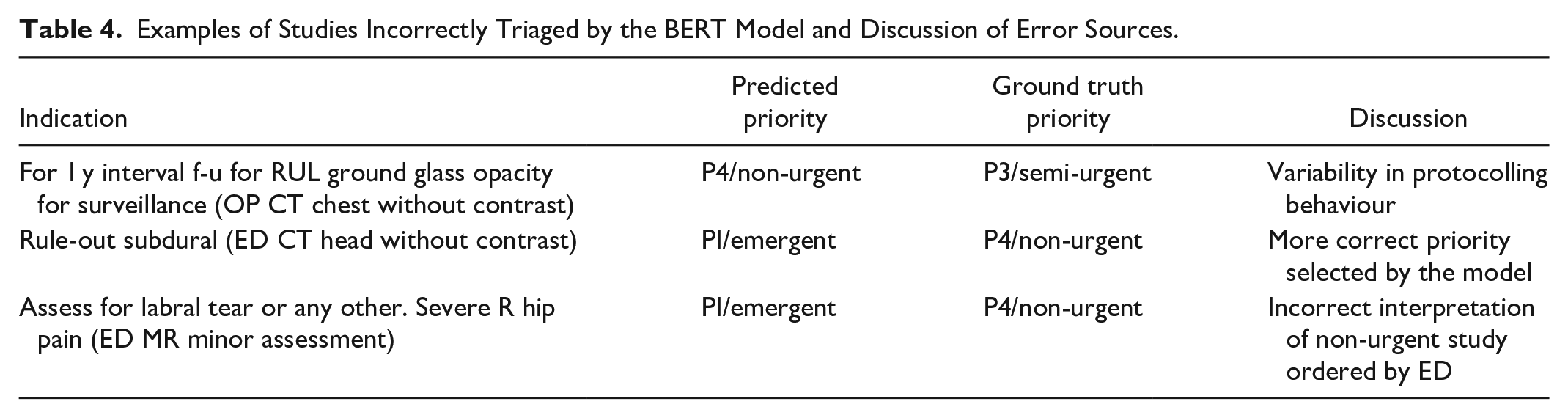
For triage, similar causes for error were identified (Table 4). One common source of error was the variability in how different radiologists prioritize cases. For example, in cases to follow up outpatient lung nodules, some were classified in the training dataset as P3/semi-urgent whereas others were classified as P4/non-urgent. This variability in the dataset likely mirrors the differing practices and judgment calls of individual radiologists, indicating that there is no uniform standard for classifying certain cases, which in turn affects the model’s prediction accuracy.
Examples of Studies Incorrectly Triaged by the BERT Model and Discussion of Error Sources.
Discussion
In this study, we demonstrated the accuracy of a BERT-based natural language processing model for triaging and protocolling CT and MRI studies. The model exhibited strong performance across various modalities and location-based subgroups, with particularly high accuracy in triaging emergency department (ED) studies. Our finding underscores the potential clinical utility of automated models for protocolling radiology examinations, thereby enhancing the efficiency of medical imaging workflows.
Previous models have assessed the performance of both NLP and traditional ML applications for automating radiology protocol selection with comparable results. Kalra et al developed a deep-neural network for protocolling CT and MRI requisitions belonging to 108 unique protocols, achieving similar weighted precision and recall to our model at 83.9% and 84.9%, respectively. 13 Lau et al evaluated the use of a BERT model for triaging CTs belonging to 25 classes, with similar results. 14 Other studies which have assessed performance of BERT models for neuroradiology and musculoskeletal specific exams demonstrated comparable performance to our sub-stratified results.15,16 For triage priority, few studies have specifically assessed the performance of machine learning models. One study has evaluated automatic radiology triage prioritization for MRI Brain using an SVM, achieving a precision 68.2% and recall of 73.7%. 17 The authors highlighted that triage priority is largely influenced by the referral demographics of the exam, which was consistent with our study results. We hypothesize that the relatively simpler triage classification task might contribute to the comparable performance we observed between the BERT and SVM models.
One notable finding was that BERT performance on minority classes was superior to both traditional models, as seen in the macro-averaged statistics. The ability of BERT to capture nuanced contextual information from word positioning may allow for superior performance on less common classes. 18 This may be advantageous in imaging, where accurate protocol adherence for specialized imaging can help to improve diagnostic certainty.
Clinically, there is strong desire from radiologists and clinicians for automated protocolling tools for diagnostic radiology to expedite imaging workflows. A recent Canadian multi-institutional survey found that 95% of clinicians would support an AI tool to prioritizes MRI requisitions. 19 Implementing such a tool within a decision support system could enable real-time feedback to clinicians regarding the suitability of their requisitions in line with ACR or regional guidelines. This approach could reduce the number of requisition rejections, expedite order entry, and enhance overall resource management. 20 Previous cost-analysis of automated protocol systems have suggested potential financial savings that could be offered by these models. 16
In addition to BERT models, large language model chatbots like ChatGPT are gaining traction, particularly with emerging applications in radiology workflows. 21 ChatGPT-4, like BERT, operates on a transformer architecture, enabling it to understand and generate text. A limited pilot study of CT protocolling with ChatGPT demonstrated promising results, with 84/100 protocols correctly identified. 22 However, the broader clinical application of these technologies is currently limited by challenges such as “hallucinations” (ie, the presentation of incorrect information as fact) and inherent unpredictability in responses. Future iterations of these models may mitigate these issues, potentially broadening their applicability in clinical settings.
Our study is subject to several limitations. The retrospective, single-centre design may not fully capture the heterogeneity and different practices at different medical centres, limiting the generalizability of our data. Furthermore, the model performance was evaluated against a ground-truth label of protocol and priority assigned clinically by individual radiologists and may be subject to inter-reader variability. While we used the BERTBase model for training and evaluation, specialized radiology-focused models like RadBERT 23 might achieve better results. Practical challenges in clinical integration, such as the need for continuous model maintenance and user-friendly interfaces, also pose significant hurdles. Finally, concerns about AI errors and a lack of transparency in model functioning are obstacles to the broader clinical adoption of these technologies.
In conclusion, our investigation reveals that NLP and BERT models are feasible tools for automatic protocol and triage of radiology studies, showing promise in enhancing radiologist workflow and increasing departmental efficiency. We hope that our preliminary findings will lay a foundation for further research and the broader application of these tools in clinical practice.
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.