Abstract
Introduction
Pediatric low-grade gliomas (pLGGs) are a common type of brain tumour in children, accounting for about 40% of pediatric central nervous system malignancies. 1 Recent molecular evidence has linked pLGGs to changes in the mitogen-activated protein kinase pathway, which can be detected through invasive biopsy. However, there is a growing interest in utilizing noninvasive Machine Learning (ML)-based models that use medical images to identify molecular markers. 2 These models require accurate tumour segmentation in magnetic resonance imaging (MRI), which can be challenging if done manually due to various factors including the tedious and time-consuming nature of the task and inter- and intra-reader variability.
Existing research has focused on optimizing brain tumour segmentation in the adult population, but studies on ML-guided pediatric tumour segmentation are scarce. pLGGs exhibit significant differences in genetic alterations and characteristics compared to adult brain tumours, necessitating the development of varied treatment regimens and prognosis outlooks. 3 Furthermore, the accuracy of manual tumour annotation can be impacted by factors such as the quality of the MRI, neuroradiologist’s experience, and inter/intra-reader variability. 4 The BraTS 5 competition evaluates state-of-the-art methods for automated glioma segmentation in adults, with participants mainly focusing on deep learning approaches. Our segmentation model is developed based on ideas presented in notable approaches in the BraTS competition, including Myronenko’s 6 architecture that achieved a high Dice score of .88 for the whole tumour segmentation. We further enhanced the model by incorporating Jiang et al's two-stage segmentation model, 7 leveraging MRI scans in the initial stage to generate a coarse segmentation mask and refining it with another U-Net in the second stage. 3
In a recent study, Peng et al. proposed an automated workflows for tumour segmentation in pediatric high-grade gliomas using 3D U-Net architecture. 8 The model achieved a mean Dice score of .724 for both T2 hyperintensity and T1 contrast-enhancing (T1ce) segmentations, indicating strong similarity with manual segmentation masks. Additionally, the automated segmentation findings were used to assess tumour volume and bidimensional diameters, with high intraclass correlation coefficients for both T2 and T1ce segmentation masks. Radiomics has shown promising results in predicting molecular subtypes of pLGGs using Machine Learning (ML) models trained on radiomics features extracted from Fluid-attenuated inversion recovery (FLAIR) MRI scans. Wagner et al. 2 have shown that ML models trained on radiomics features extracted from FLAIR MRI scans can predict BRAF status in pLGG patients with a mean AUC of .75. This non-invasive approach eliminates the risks associated with biopsies and allows them to be performed more quickly, which is particularly beneficial for patients with unresectable tumours or those located in difficult-to-reach areas.
Measuring tumour volume using MRI is critical for assessing response in pediatric brain tumours, and current methods require manual evaluation by human experts, which is time-consuming and labor-intensive. Automatic volumetric segmentation has been preferable, but current approaches have received limited validation. 9
To address these challenges, this paper proposes a 2-step deep learning-based model for automating the pLGG segmentation process. Previous work has shown that the use of a two-step approach is an effective strategy, mitigating memory limitations while maintaining segmentation accuracy.10–12 The pLGG volume for both manually and automatically segmented tumours is determined, and their correlation is compared to assess the generated results by the proposed automated segmentation model. Additionally, an end-to-end radiomics-based molecular marker classification model is constructed to automate the classification of pLGG molecular markers and evaluate the performance of the suggested automated segmentation models.
Methods
Data
The study used preoperative brain FLAIR MRI scans from a well-characterized cohort of 288 children with pLGG between 2000 and 2021 from a pediatric hospital in Canada, making it one of the world’s largest cohorts of pediatric brain tumours. The most prevalent genetic alterations in this cohort were BRAF fusion (46%) and BRAF V600E mutation (19%) and additional uncommon molecular markers forming a third category for the purposes of this study. The raw MRI data was obtained from the respective Picture Archiving Communications System (PACS) and anonymized for further analysis, and manual tumour segmentation was performed by a fellowship-trained pediatric neuroradiologist using 3D Slicer (Version 4.10.2; https://www.slicer.org) and the manual placement of the volume of interest (VOI) was confirmed visually by a board-certified neuroradiologist. The manual segmentation of the VOI was only used in the training phase of the proposed model. Methods such as immunohistochemistry and droplet digital polymerase chain reaction were used to identify the BRAF fusion subtype and BRAF V600E mutation.
Ethics Approval and Consent to Participate
The Research Ethics Board (REB) of a pediatric hospital approved this retrospective study, and informed consent was waived by the “Research Ethics Board (REB) of The Hospital”. All methods were performed in accordance with the relevant guidelines and regulations.
Data Preprocessing
To have the same data dimensionality and coordinate system, 3D Slicer software13,14 was used to register all images, as well as the manually annotated ground truth segmentation masks, to the SRI24 atlas, which is an MRI-based atlas built upon normal adult human brain anatomy, 15 resulting in images and segmentation masks of size 240 × 240 × 155 voxels. Furthermore, skull stripping was performed using HD-BET 16 tool. To account for variable pixel intensity scales, N4 bias field correction prior to Z-score normalization was used to standardize the range of all image features. 17 Finally, pixel values were normalized to [0 - 1] range.
Data split
The study used multifold stratified cross-validation with 5-folds to ensure that the proportions of each molecular marker in each fold are the same as the proportions in the overall cohort. This method allowed us to have all accessible MRI scans in the test cohort once, enabling us to make robust predictions on all patients using our models, avoiding biased results based on (un)lucky training/testing splits.
pLGG Segmentation
The study introduces a two-step deep learning-based segmentation model as a solution to overcome GPU memory limitations encountered during the training phase. The first step downsamples the images, allowing for training of a U-Net model. This initial step generates a coarse segmentation mask at the patient level, serving as a foundation for subsequent fine-grained segmentation. Then, a VOI-based U-Net is trained using MRI 3D data patches of the tumour location to generate more precise segmentation masks. All MRI images were downsampled to 112 × 112 × 64 voxels for training the first model, and a patch of 112 × 112 × 112 voxels was extracted from each original image to prepare data for training the second model. The patch covers the whole tumour lesion and fits inside the GPU memory. The two models are referred to as model I and model II in the following sections. The proposed two-step segmentation approach is depicted in Figure 1. Proposed two-step network architecture for automated tumour segmentation.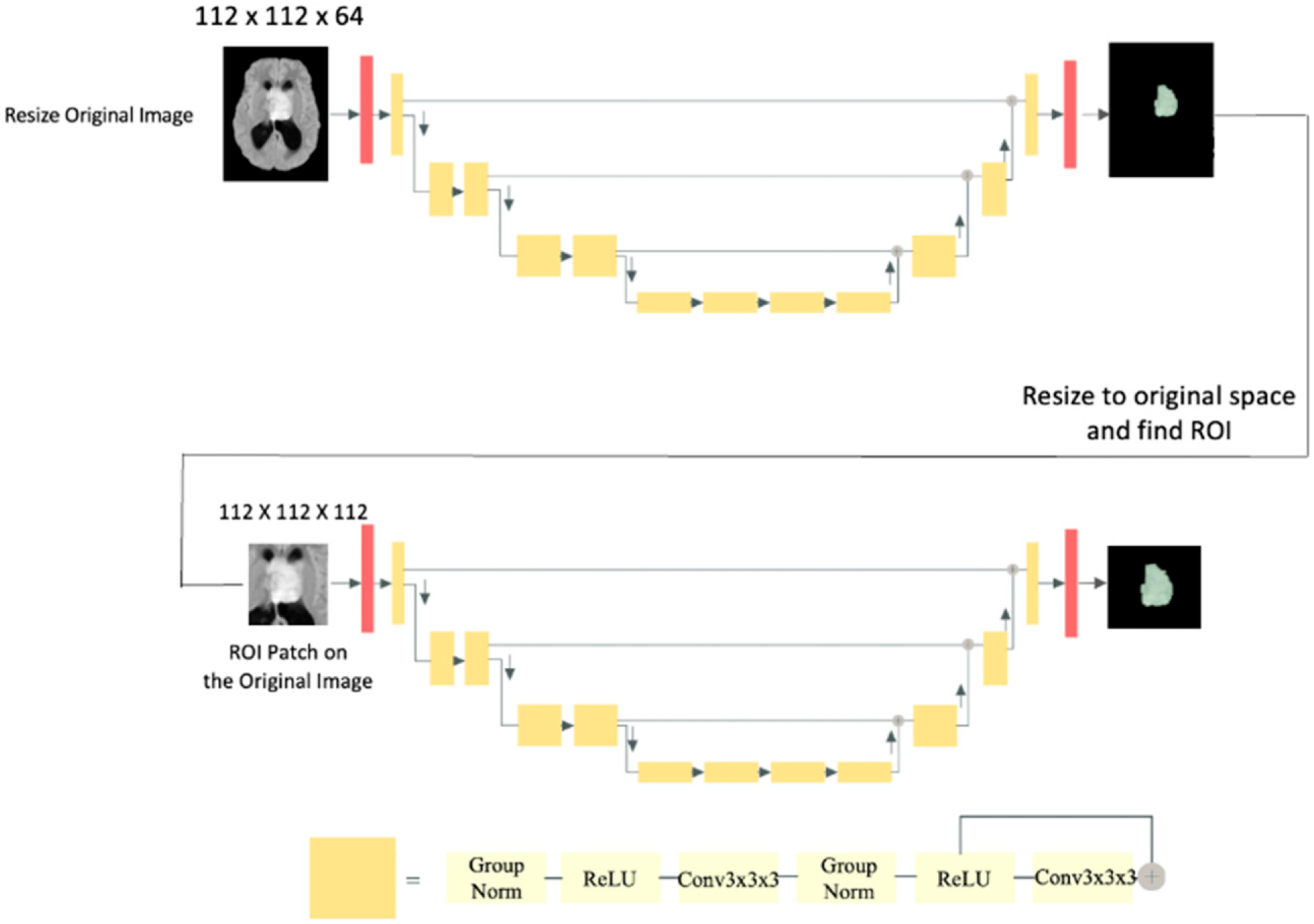
Model I
Initially, a single U-Net architecture was constructed to predict tumour segmentation masks. However, the GPU memory limit (on both NVIDIA V100 32 GB Tesla GPU with 128 GB memory and NVIDIA GeForce RTX 3090 with 56 GB memory) was reached by feeding the full-size 3D MRI scans as an input to this network. Through experimentation we learned that it is necessary to locate tumours in images first and then predict segmentations based on the identified region. However, object detection models such as Fast Region-based Convolutional Neural Network (Fast R-CNN) and Single Shot Detector (SSD)18,19 may not be time and computationally efficient in our studies since they must be trained on 3D images.
To roughly locate tumours, images were first downsampled and a U-Net segmentation model was trained using ResNet 20 blocks, which reduced image dimensions and increased feature size by a factor of two. The utilized architecture is similar to that presented by Myronenko. 6 The sigmoid function was used in the final layer to forecast the segmentation mask, and the final prediction was a segmentation mask of size 112 × 112 × 64 voxels that reflects the approximate tumour location in the MRI. Following that, the images were upsampled to their original size of 240 × 240 × 155 voxels and compared to manual segmentations conducted by neuroradiologists. The Adam optimizer and Dice loss were utilized to train the models based on best practices and literature.
Model I’s predictions are illustrated in Figure 2 (Right). Note that the automated segmentations produced by the model lack refined borders and require a more precise model for smoother tumour boundaries. Left: Single slice from sample MRI scans. Middle: Highlighted refined tumour segmentation masks generated by Model II. Right: Highlighted coarse tumour segmentation masks generated by Model I.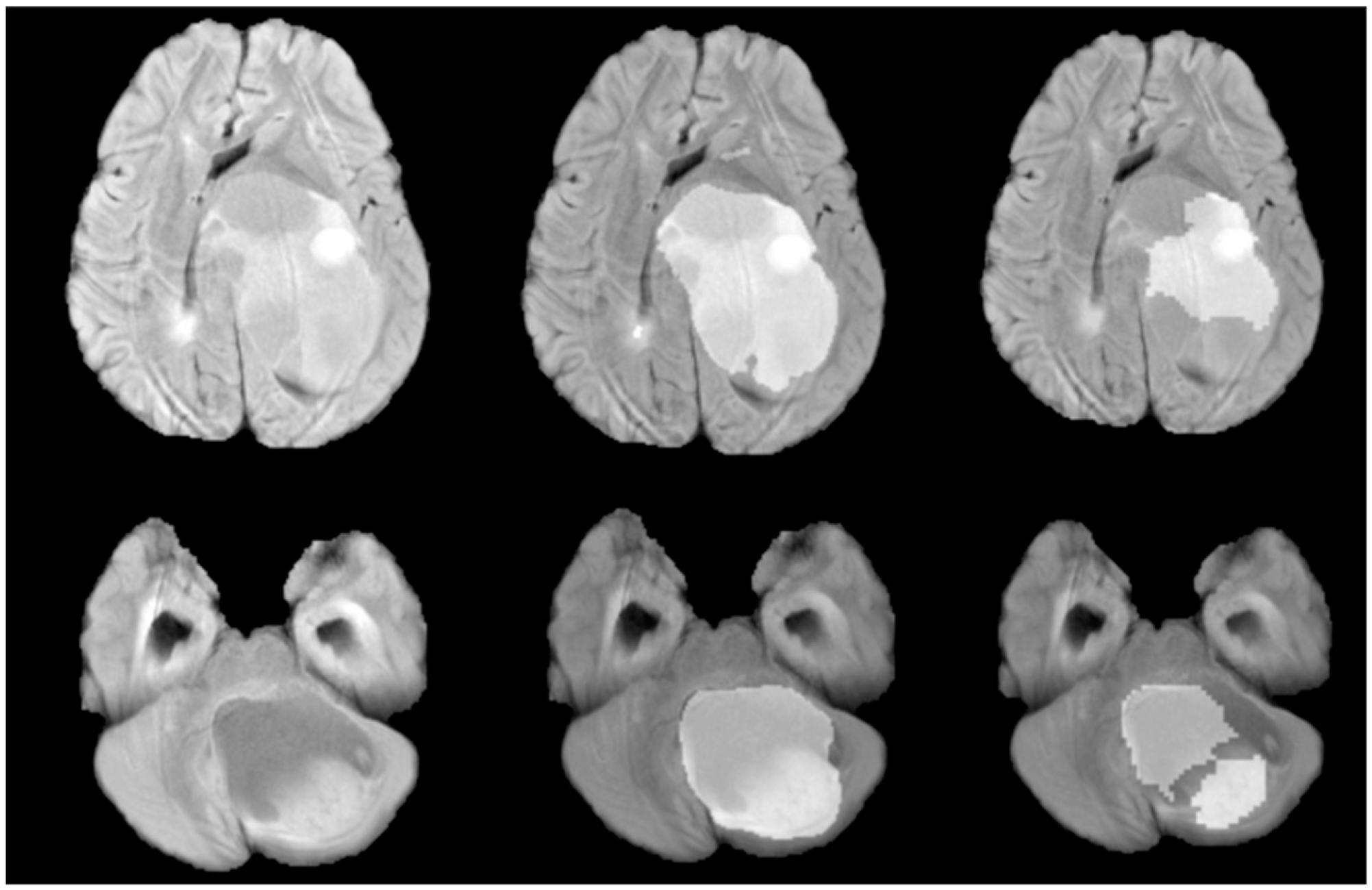
Model II
To refine the segmentation masks and address the shortcomings with the first model’s unrefined edges, a second U-Net-based model was trained using 112 × 112 × 112 patches. Using smaller patches helps train the model to focus more on the target (tumour mask), whereas using whole images adds irrelevant data to the model. The two-stage U-Net models were trained using 5-fold cross-validation to predict on the entire dataset and leverage the automated segmentations created on test sets in the downstream classification task. The model employs an encoder-decoder U-Net architecture, similar to Model I, but with patches (Model I ouput) centred around the middle of the tumour encompassing the whole tumour. The decoder generates an output of 112 × 112 × 112 voxels indicating tumour mask predictions. Improvements were seen in Model II when compared to Model I predictions as depicted in Figure 2 (Middle). Dice score was used for performance evaluation of both Model I and II.
pLGG Classification
The radiomics-based tumour classification approach selects the tumour region in an MRI scan using segmentation masks. However, manual tumour segmentation is costly and inefficient. Here we apply the output of the automated tumour segmentation model to an end-to-end radiomics pipeline for pLGG classification, eliminating the need for manual MRI contouring. This considerably increases the speed of the entire procedure and makes it more autonomous, which benefits both radiologists and patients. Three types of genetic alterations are classified in this work: BRAF V600E mutation, BRAF fusion, and other molecular markers that are rarer compared to the first two. The classification of pLGG tumours subtypes helps understand their unique characteristics, predict their behavior, and inform treatment decisions. It enables healthcare professionals to offer personalized therapies, monitor disease progression, and enhance outcomes for children with these brain tumours.
The proposed automated two-step segmentation model was used to generate tumour masks on the entire dataset. These segmentation masks that are independent of the radiologists’ manual tumour contouring, were used to create an end-to-end radiomics model for pLGG classification. Another radiomics model was created using the radiologists' manual segmentation masks. The classifications results were compared using an Area Under the Receiver Operating Characteristic (ROC) curve (AUC).
The PyRadiomics package was used to extract 3D radiomic features from each MRI scan. Random Forest was used as the classification model as previous research 2 has shown it performs best. The entire dataset from the proposed segmentation model was used in this step, allowing us to create classification models entirely based on the automated segmentation data. Classification performance was averaged across all instances in the dataset, making the final results more reliable.
End-To-End pLGG Volume Assessment
In the final experiment, we measured and compared tumour volume size by utilizing radiomic features on both manual and automated tumour segmentation masks. The manual measurement of treatment response and tumour growth is laborious and costly. This study aims to automate the assessment of tumour progression and treatment response by proposing an end-to-end pipeline that extracts tumour size automatically from MRI scans to allow neuroradiologists and oncologists to quickly assess the progression of patients' tumours at different time points.
Volume Features
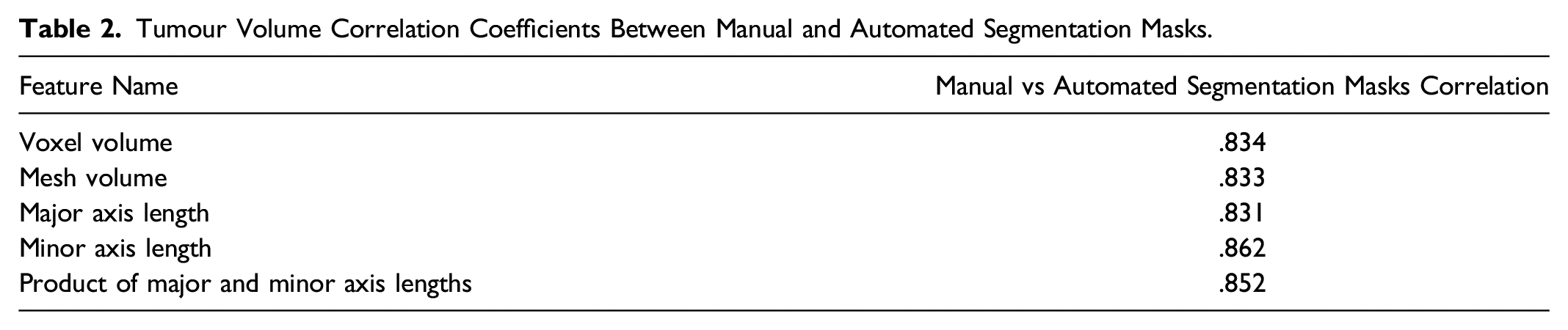
From the list of all radiomics features, 4 volume features were selected: Voxel volume, Mesh Volume, Major axis length, and Minor axis length. Voxel volume is approximated by multiplying the number of voxels in the VOI by the volume of a single voxel while mesh volume was derived from the VOI’s triangle mesh. Major and minor axis length yields the largest and second-largest axis length of the VOI-enclosing ellipsoid. The last two features were used to calculate the product of the 2D diameters for each tumour. Finally, to test our model’s segmentation findings, the correlation between selected features were calculated between the manually and automatically constructed segmentation masks.
Results
pLGG Segmentation
In 5-fold cross-validation, Dice score was computed on 288 MRI cases with pLGG and resulted in an average score of .752 using Model I. The second model enhanced segmentations and tumour borders and achieved an average Dice score of .795. The two-step segmentation method increased pLGG segmentation performance by 4.3% in the test set compared to the first model alone. The P-value from the t test between Model I and Model II equals .010, showing that Model II can significantly improve segmentation results compared to Model I.
pLGG Classification
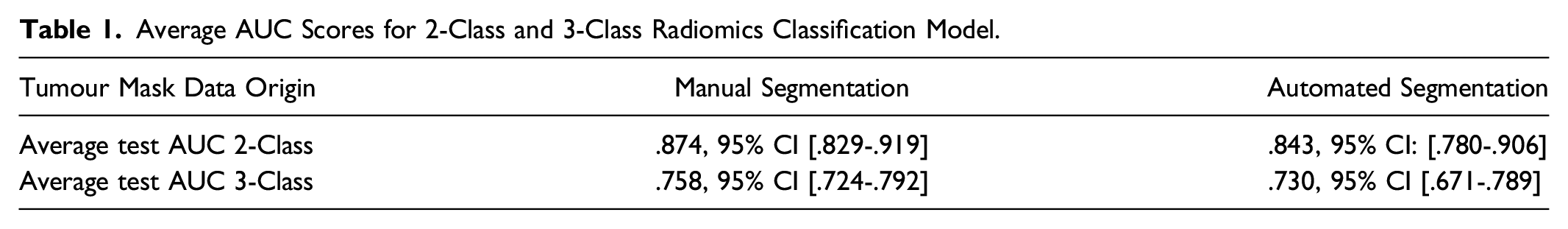
Average AUC Scores for 2-Class and 3-Class Radiomics Classification Model.
pLGG Volume Assessment
Tumour Volume Correlation Coefficients Between Manual and Automated Segmentation Masks.
Discussion
Manual tumour segmentation is costly and time-consuming, making it difficult to develop and deploy ML systems in clinical treatment. The development and application of ML systems for brain tumours require manual tumour segmentation and labelling, which is prohibitively expensive due to cost and time constraints. In addition, multiple radiologists annotating images in a dataset may result in discrepancies (i.e., inter- and intra-reader variability), causing the deep learning models to underperform. 22 Furthermore, labelling images places an additional burden on radiologists. It is anticipated that the automated tumour segmentation and classification algorithms will be capable of providing robust and consistent results while alleviating the workload of radiologists. This is primarily attributed to the absence of inter- and intra-reader variability commonly encountered in manual segmentation, which can contribute to enhanced accuracy and reliability. Automated brain tumour segmentation in adults has been heavily researched for the past decade,5,7,23 but few studies have focused on pediatric tumour segmentation due to insufficient data. 24
The differences in MRI signal characteristics between pediatric and adult brain tumours pose a challenge for utilizing the transfer learning method. To demonstrate this, a segmentation model was trained on the BraTS adult MRI dataset, achieving a validation score of .8. However, when the same model was fine-tuned using transfer learning on the pediatric dataset, it only yielded a Dice score of .37, emphasizing the need for specialized segmentation algorithms for pediatric brain tumours.
The main objective of this research paper is to develop an accurate segmentation model for pediatric low-grade glioma and compare its effectiveness with other ML-based models in tasks such as molecular subtype classification and volumetric analyses. This paper identifies challenges in producing patient-level segmentation masks using limited computing resources, where in our experience, the patch-based models produced false positives resulting in noisy masks. To solve this issue, this paper suggests a two-step segmentation model that first detects tumour location and provides a coarse segmentation mask, followed by a patch-based model that generates refined masks. The proposed 2-step method improved patient-level segmentation results by over 4% compared to traditional 1-step models and was used to produce automated segmentation masks for all MRI scans in test data.
Radiomics-based approaches can predict pLGG genetic markers (e.g., BRAF V600E mutation vs BRAF fusion) on MRI data. Tumour segmentation in MRI is necessary for radiomics model development, however, manual segmentation is tedious and time intensive. To make ML more practical in clinical settings, we need to build pipelines in an end-to-end fashion, performing automated segmentation prior to classification and volume assessment, which allows physicians to access ML outcomes without having to label or annotate data. To achieve this, we designed an end-to-end radiomics molecular subtype classifier and extracted tumour size features using our automatically generated segmentation masks, to estimate molecular subtypes and tumour volume size. These pipelines produced highly correlated results to models built using manual segmentation masks, demonstrating the effectiveness of our automated segmentation model beyond tumour mask generation.
During training, we alternate between using manual and automated segmentation masks to see how it affects the classification of MRI scans in test using automated segmentation masks. Results from this analysis showed that both training and testing on the automated segmentation masks performs better in comparison to training on the manual segmentation masks and testing on the automated segmentation masks (2-Class: AUC of .843 vs .765, 3-Class: AUC of .730 vs .687). Data consistency in training and test is crucial in achieving the best performance. For example, when data is labeled by two different readers, training a model with data from reader 1 and making predictions on data labeled by reader 2 can lower the performance compared to training and testing the model with data labeled by the same reader. 25
Despite the promising results, our work has some limitations that may lead to bias in our studies. This study is based on a relatively small dataset of MRI scans of pLGG patients from a single institution, which may introduce bias in the results. Future studies should expand the dataset by collecting data from more patients and institutions to minimize bias and enable external validation of the models. Additionally, this study currently focuses only on pLGG, hence, expanding the dataset by including diverse tumour types and fine-tuning the model would enable it to make predictions on a broader range of tumour types. Furthermore, having only one reader for manual segmentation can limit the generalizability of the models due to inter and intra-reader variability. Thus, having access to manual segmentation masks created by multiple readers would improve the models' validation and generalization.
Conclusion
In this paper, we introduced an automated pLGG segmentation method based on U-Net architecture. Using the resulting tumour masks of this study in ML-based tumour classification and volume calculation pipelines yields results that are comparable to manual segmentation. The proposed approach involves training a two-stage U-Net model for patient-level tumour segmentation and using the resulting tumour masks to train a radiomics model for tumour subtype classification. The tumour volume is then calculated from the radiomics features for both manual and automated segmentation masks, and the correlation between the two segmentation techniques is discovered to be strong, confirming the effectiveness of the proposed automated tumour segmentation method.
Footnotes
Declaration of Conflicting of Interests
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Funding
This research is funded by CCS/CIHR/BC Spark Grants in Novel Technology Applications in Cancer Prevention and Early Detection. This Project has been made possible with the financial support of Health Canada, through the Canada Brain Research Fund, an innovative partnership between the Government of Canada (through Health Canada) and Brain Canada, and of the Canadian Cancer Society.
