Abstract
Deep learning techniques using convolutional neural networks (CNNs) have been successfully developed for various medical image analysis tasks. However, the skills to understand and develop deep learning models are not usually taught during radiology training, which constitutes a barrier for radiologists looking to integrate machine learning (ML) into their research or clinical practice. In this work, we developed and evaluated an educational graphical user interface (GUI) to construct CNNs for teaching deep learning concepts to radiology trainees. The GUI was developed in Python using the PyQt and PyTorch frameworks. The functionality of the GUI was demonstrated through a binary classification task on a dataset of MR images of the brain. The usability of the GUI was assessed through 45-min user testing sessions with 5 neuroradiologists and neuroradiology fellows, assessing mean task completion times, the System Usability Scale (SUS), and a qualitative questionnaire as metrics. Task completion times were compared against a ML expert who performed the same tasks. After a 20-min introduction to CNNs and a walkthrough of the GUI, users were able to perform all assigned tasks successfully. There was no significant difference in task completion time compared to a ML expert. The educational GUI achieved a score of 82.5 on the SUS, suggesting that the system is highly usable. Users indicated that the GUI seems useful as an educational tool to teach ML topics to radiology trainees. An educational GUI allows interactive teaching in ML that can be incorporated into radiology training.
Introduction
Artificial intelligence (AI) is a powerful image analysis tool that is becoming increasingly more prevalent in radiology. 1 Many radiologists have expressed interest in learning more about AI and applying it in their clinical practice,2,3 but the technical skills required to develop AI models are typically not taught in radiology residency, creating a large entry barrier. Consequently, few radiologists use AI models in their daily work.3-6 To make AI more accessible to radiologists, various graphical user interfaces (GUIs) have been developed to provide ease of access to AI and machine learning (ML) algorithms without requiring users to write code.4,7-9 However, these GUIs are typically designed for users that are already familiar with ML concepts and terminology, so their effectiveness may be limited for radiologists without a background in ML.3,5,9 Training radiologists to understand the capabilities and limitations of ML systems is an essential step to integrating ML systems into clinical radiology.10,11
In recent years, deep learning has become increasingly popular in the ML community, with various research groups demonstrating the effectiveness of convolutional neural networks (CNNs) in different radiology applications. 12 Although traditional ML algorithms require users to manually perform feature extraction prior to classification, deep learning methods can perform feature extraction and classification automatically, without requiring manually extracted features. 13 Convolutional neural networks, which are designed to learn spatial hierarchies of features through a backpropagation algorithm, have been demonstrated to be particularly effective in radiological applications. 12 Convolutional neural networks are composed of a series of convolution layers, pooling layers, and fully connected layers, which act together to identify key features in an image and output a probability for each class label. 12 Convolutional neural networks have been used for various medical image analysis tasks, with improved accuracies compared to other ML methods.12,14-16 Familiarity with state-of-the-art CNNs is considered beneficial for radiologists, since deep learning will likely impact their clinical practice in the future. 12 Interactive educational tools to teach ML and CNNs are, however, currently largely lacking in radiological education.
We therefore aimed to develop an intuitive and user-friendly GUI to construct CNNs without requiring any programming experience, allowing radiologists to build their own deep learning models, and serving as an experiential learning tool for radiology trainees to become more familiar with CNNs.
Methods
GUI Design
A GUI was developed in Python using PyQt and ML aspects were implemented using the PyTorch framework.17,18 The GUI consisted of a sequential workflow to build a CNN (Figure 1) without needing to write any code. The first step is data processing, where the user selects a dataset, resizes it, and splits the dataset into training and validation data. Next, the user specifies the CNN architecture by adding various convolution, pooling, and linear layers as well as specifying activation functions. Then, the user specifies the hyperparameter values, including the learning rate, batch size, and number of epochs to use during the training process. Alternatively, the user may perform a grid search to determine these values automatically. Lastly, the user trains the model and is presented with an evaluation page, containing various methods to assess the model’s performance, including the training curve, ROC curve, and confusion matrix. Throughout the GUI, default values were set to allow users without prior experience to choose hyperparameters to construct a basic functional CNN using the default values alone. These default values can be modified through a settings file. Tooltips are present throughout the GUI, ensuring that users can understand the system and the purpose of each hyperparameter. Suggested value ranges are also provided for certain hyperparameters. The results of trained models are displayed in one screen and users can save the models they build, as well as load pre-trained models. The workflow to construct a convolutional neural network is divided into 3 main groups. (A) Data processing group: users may select to upload a dataset, resize it, and choose the training/validation split. (B) Model construction group: users may specify the convolutional neural network architecture and model hyperparameters. (C) Training hyperparameters group: users may specify the hyperparameter values to use in the training process.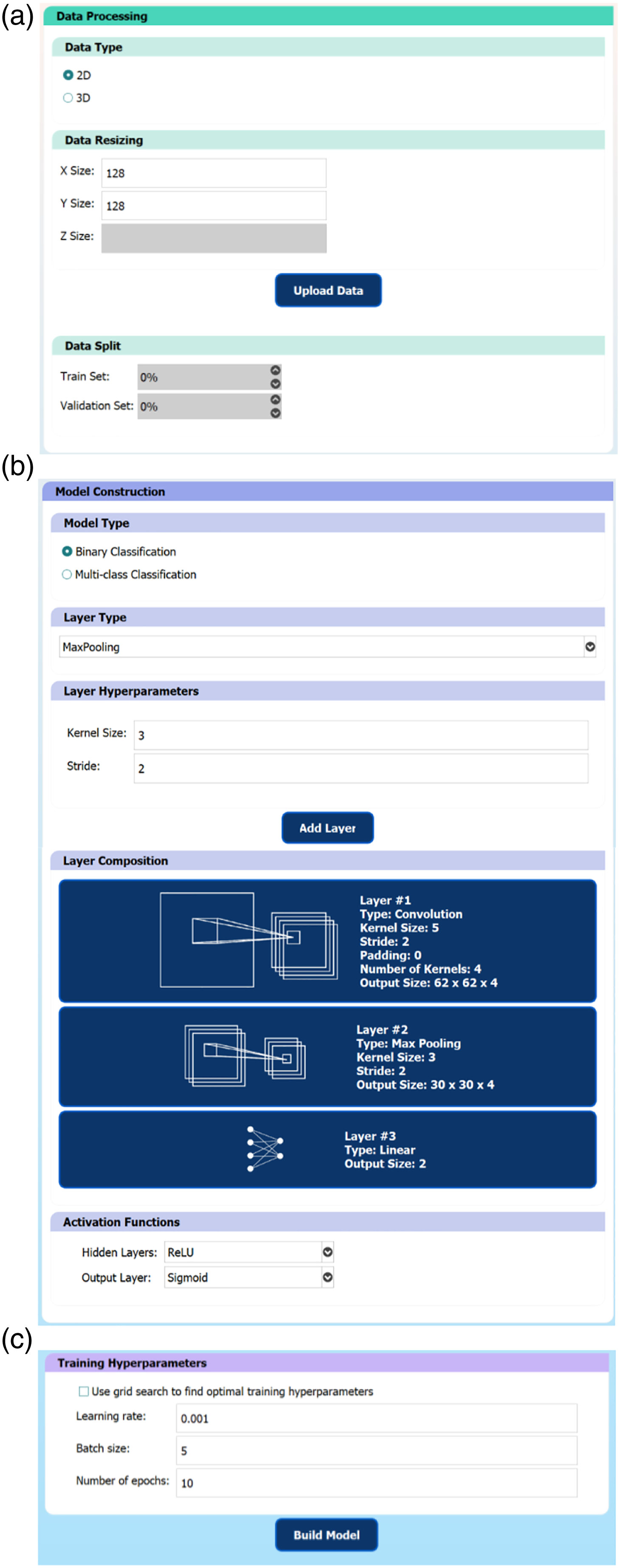
GUI Functionality
To verify the functionality of the GUI in constructing useful CNN models, a CNN was trained to perform binary classification on an open-source dataset containing 253 MRI (2D) slices of the brain,
19
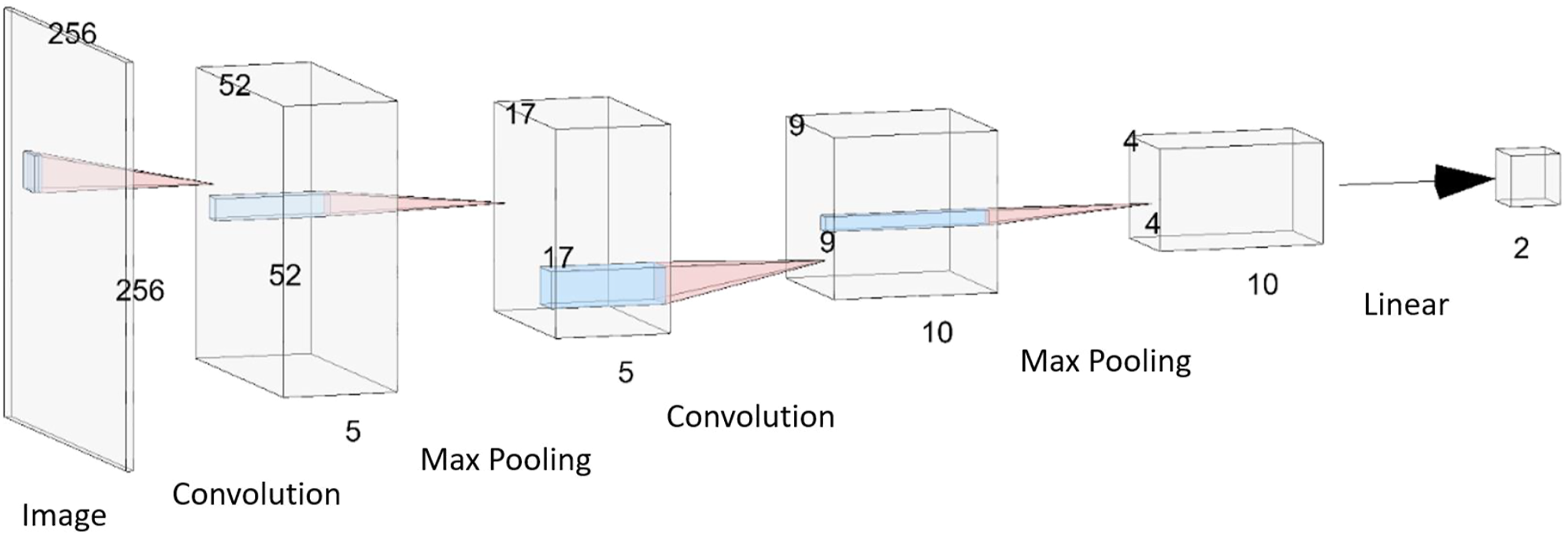
with the goal of classifying whether a tumor is present on an image or not. The dataset consisted of 155 images containing a brain tumor and 98 images without a tumor (but with various other mixed pathologies). Samples of the data are shown in Figure 2. Using the GUI, the images were resized to 256 × 256 pixels, a 70/30 training/validation data split was used, and a CNN model was constructed with the architecture shown in Figure 3. A grid search was used to determine the training hyperparameters and a binary classification threshold of 0.5 was used to generate the confusion matrix and calculate the model’s accuracy. Samples from the brain MRI dataset. (A) 2D slices of the brain with mixed pathologies but without any tumors. (B) 2D slices of the brain with the presence of a tumor. The 5-layer convolutional neural network model that was trained to perform binary classification on MR images of the brain to determine the existence of a tumor. The dimensions of each layer are indicated. The model consists of 2 sets of 2D convolution and max pooling layer pairs, followed by 1 linear layer to perform the classification. ReLU activation functions were applied after each convolution layer and a sigmoid activation function was applied at the output neuron of the final linear layer.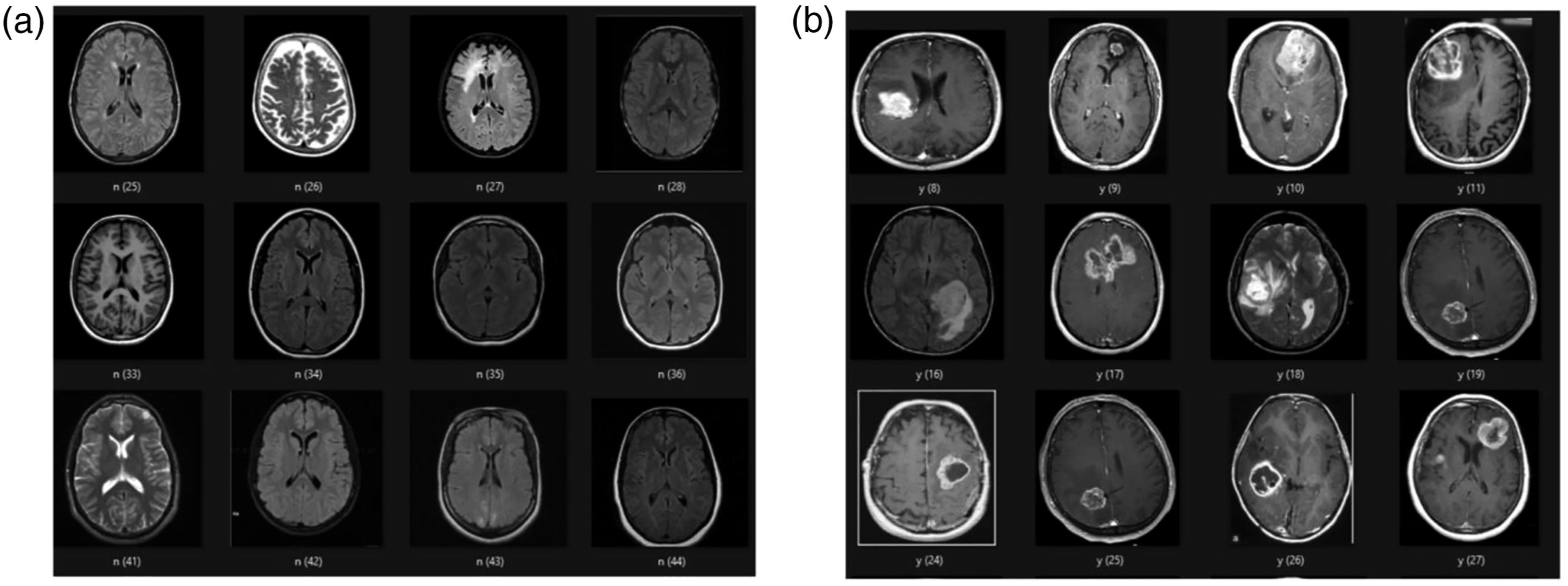
User Testing Procedure
To verify the usability of the GUI and assess its potential value as an educational tool in radiology, a user testing protocol was developed, and voluntary user-testing sessions were performed. A request for volunteers to test a ML-based software was sent out within the neuroradiology department of the Hospital for Sick Children. Five neuroradiologists and neuroradiology fellows (1 female, 4 males) were recruited, which is the typical sample size recommended for user testing experiments. 20 The participants had various levels of experience in radiology and ML. Two of the participants were neuroradiology fellows with no prior coding experience or exposure to ML concepts. Three participants were neuroradiologists with some prior knowledge of ML methods, but only one participant admitted to being comfortable implementing ML algorithms in code. Hence, the high-level learning objective was to introduce the fundamentals of CNNs and familiarize the participants with ML terminology, while gathering feedback on the educational value of the GUI. The user-testing protocol contained a brief introduction to the goal of the study, definitions for common ML terminology, some background information on CNNs, an overview of the testing session format, and specific steps for the user to follow using the GUI to: (a) build and train a specified CNN model on a prepared dataset and (b) load a pre-trained CNN model and comment on its performance. The 45-min test sessions took place on the Zoom video conferencing platform, 21 where Zoom’s remote-control feature was used to provide the participants with access to the GUI software.
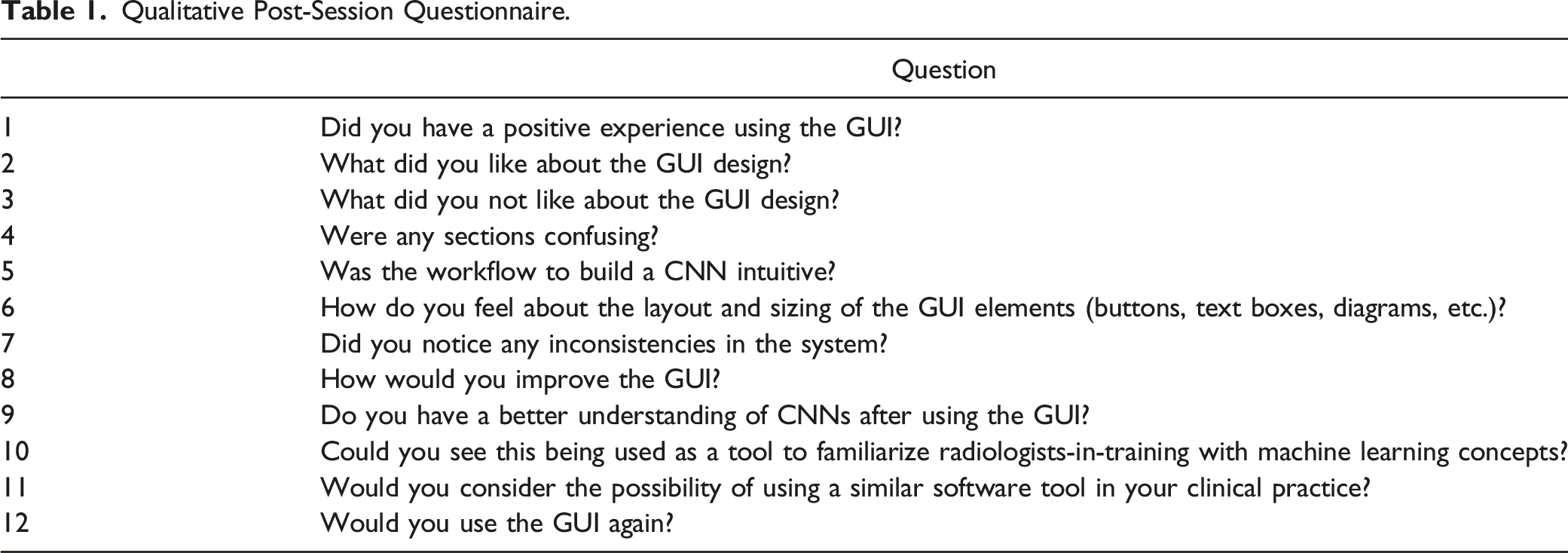
Qualitative Post-Session Questionnaire.

User Testing Assessment
In addition to the qualitative responses to questions in the test protocol (Table 1), a modified version of the System Usability Scale (SUS) was used to quantitatively assess the usability of the GUI. The SUS is a common method to assess the perceived usability of user-facing systems and has been used to evaluate the usability of many other ML tools.23-25 The SUS is a standardized questionnaire, where users are asked to rate their agreement with 10 statements, on a scale from 1-5, where higher scores indicate better agreement with the statement.26,27 The user’s scores on each item in the SUS can be summed (and normalized out of 100) to compute an overall score of the system’s usability, using the formula:
The user testing sessions were recorded with the participants’ consent. The recorded session videos were retrospectively reviewed to assess the system usability based on the analytical hierarchy process approach, where usability can be decomposed into a set of usability functions: task completion rate, number of errors, number of operations, time to perform tasks, and post session ratings. 29 Since all users were assigned the same tasks and they were completed with minimal errors (i.e., occasional misclicks and typos which would be expected under normal operating conditions), the time to complete the assigned tasks was selected as the primary metric to evaluate the GUI’s usability. Post session ratings were assessed through the SUS and qualitative questions.
A ML researcher in the field of medical imaging with 15 years of experience underwent the same process to serve as a benchmark for the task completion time.
Statistical Analysis
The mean and standard deviation values were calculated across the 5 user-testing participants for both the task completion time and SUS questionnaire responses. A 2-sample
Results
GUI Functionality
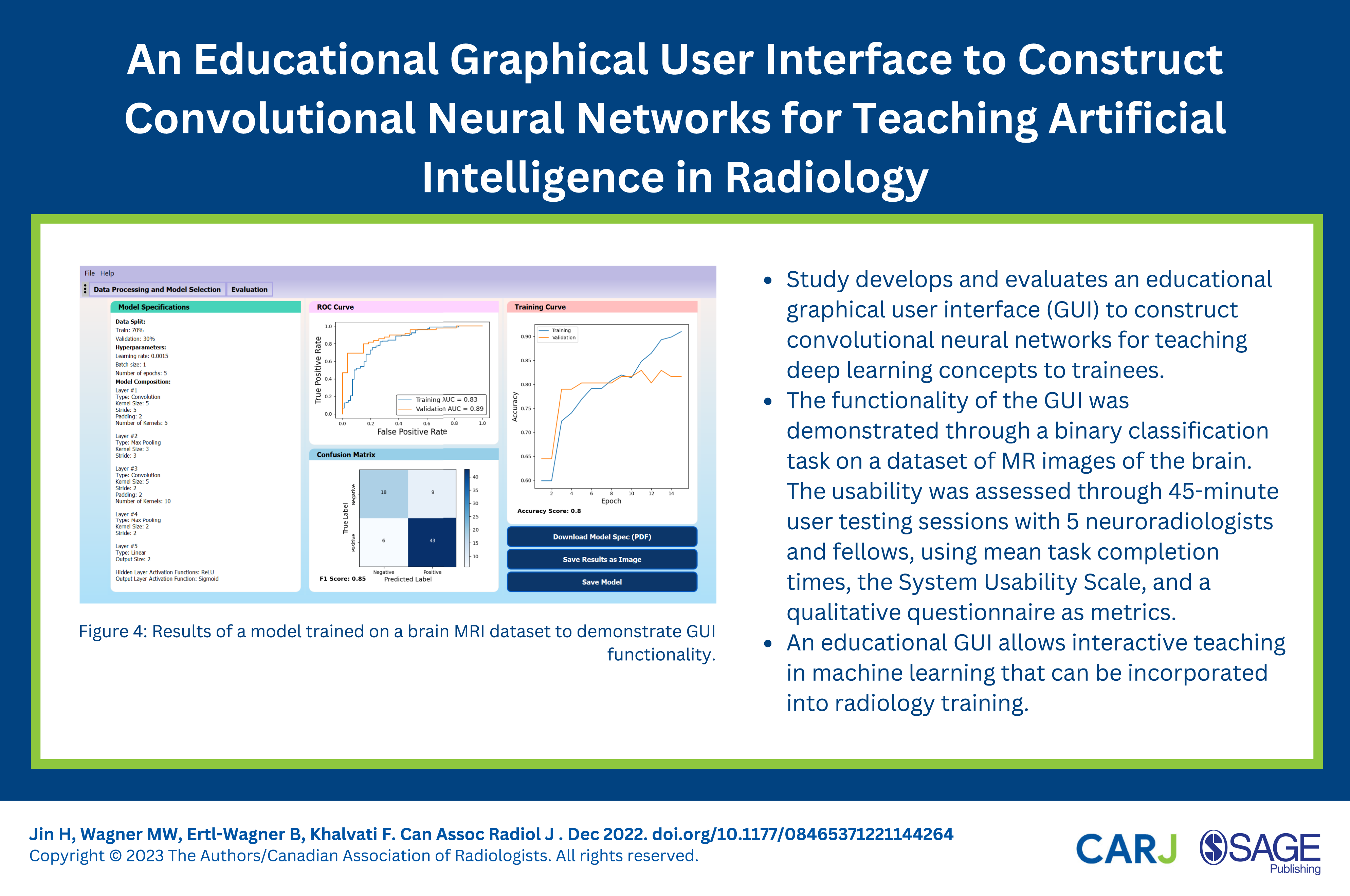
The CNN in Figure 3 was trained on the brain MRI dataset using a grid search to identify optimal training hyperparameter values (to maximize the validation AUC value). The model with the best performance used a learning rate of 0.0015, batch size of 1, and was trained for 5 epochs, achieving a validation AUC value of 0.89, F1-score of .85 and overall validation accuracy of 0.80. The evaluation page of the trained model is shown in Figure 4. These results suggest that the model is reasonably capable of solving the binary classification problem and can differentiate between images with and without a tumor. Results of a model trained on a brain MRI dataset to demonstrate graphical user interface functionality.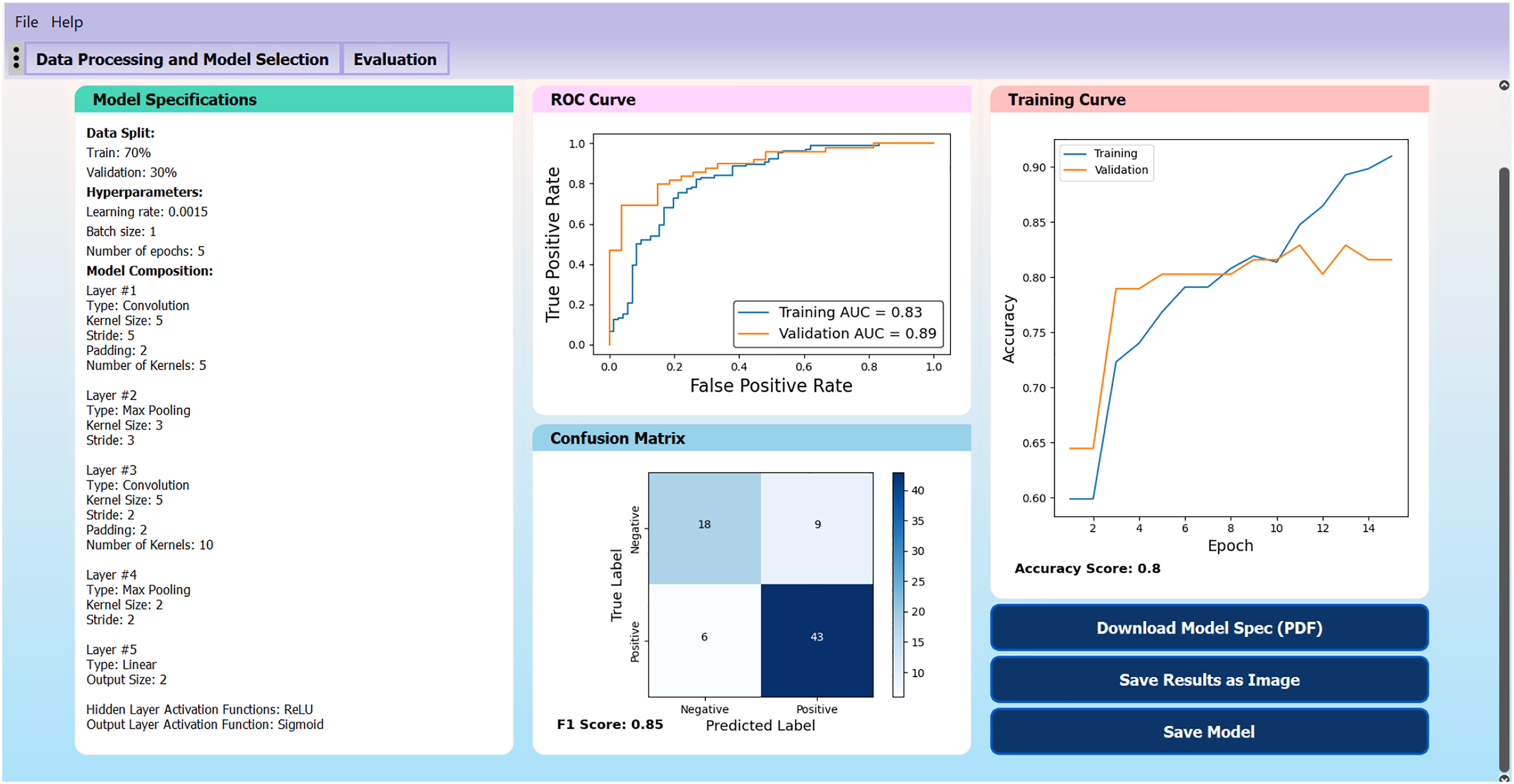
User Testing Results
Consolidated Task Completion Times Across 5 Users. The Benchmark Time Was the Time it Took a ML Researcher to Complete the Same Tasks.
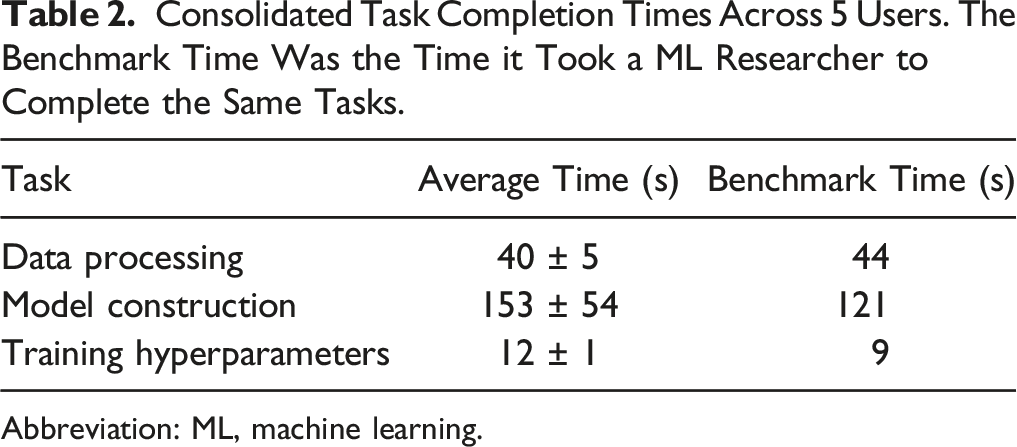
Abbreviation: ML, machine learning.
The model construction step took the longest amount of time and had the largest variance across different users, which was expected since the model construction step is the most involved part of the GUI and the testing protocol required users to construct a 5-layer CNN, with each layer being specified and added individually. Prior knowledge of CNNs and an understanding of how CNNs are structured did not lead to faster task completion times. In fact, the user who completed the model construction step the fastest had no prior ML background and the user who took the longest time had a strong ML background.
System Usability Scale Consolidated Responses. The Participants in Columns 1 and 3 Had No Prior ML Exposure. The Other Participants Had Some ML Background. The Participant in Column 4 Also Had Prior Experience Programming ML Algorithms.

Abbreviation: ML, machine learning.
In the responses to the qualitative questions, it became apparent that the training curve is likely an unfamiliar concept to most radiologists/fellows without a background in ML, and additional information on the model training process would have been helpful to improve their understanding. All participants indicated that they had a better understanding of CNNs after using the GUI and they could see the GUI being used as an educational tool to familiarize radiologists with ML concepts, perhaps accompanied by some classes on ML to provide the background knowledge to better understand the models. The participants without ML background mentioned that although the interface was easy to follow, how the selected hyperparameter values affected the model output was not as apparent (i.e., it was a black box) and they may encounter difficulty selecting hyperparameter values without the guidance of a technical expert. This challenge was also evident in the SUS responses to statements 4 and 10 (Table 3). To help address this limitation, after the user testing sessions, the option to perform a grid search through preset training hyperparameter values was added, to find optimal values for the learning rate, batch size, and number of epochs automatically, reducing the number of hyperparameters that the user would need to manually specify at the cost of increasing the training time. Generally, users agreed that the user interface is intuitive and beginner-friendly, and ML knowledge is not a pre-requisite to use the GUI.
Discussion
Our study demonstrates that an educational GUI allows interactive teaching in ML and CNNs for radiologist/radiologist-in-training users without prior experience in ML. User-testing feedback suggests that the GUI would be useful as a hands-on educational tool to teach ML to radiology trainees, and we expect that this hands-on-approach to teaching ML can be easily incorporated into radiology training. Our SUS score of 82.5 is comparable to or better than some other ML GUIs found in literature,23-25 suggesting that users would not need to spend much time learning the details of the GUI, and can instead focus on the ML aspects. To our knowledge, this study is the first to develop a GUI as a teaching tool for deep learning concepts where radiologists can construct their own CNN models. User testing feedback suggests that a major advantage of the current GUI design is its simple and intuitive workflow to build a CNN without assuming any programming background, making it accessible to a wide audience. Although many ML GUIs have been developed for radiology applications,4,7,8 a focus on evaluating the usability of the system as an educational tool for users without ML background appears to be lacking in the literature.
Future work may involve deploying the software in a classroom setting, which would help generate a more accurate assessment of the educational value of the GUI as a potential teaching tool for ML in radiology. Our GUI was designed to be a simple introduction to the fundamental concepts of CNNs. However, as users become more familiar with CNNs, they may look for more flexibility and options, so additional features should be added if it is to be used as a long-term educational tool. For example, different user modes depending on the user’s level of expertise could be added (i.e., beginner, intermediate, and advanced), with each mode providing additional flexibility and options to the user as they become more familiar with ML concepts. These options may include allowing users to specify which optimizer to use (Adam is the current default) and/or incorporate techniques such as early stopping, dropout regularization, and skip connections in their models. Options to control how the results are presented on the evaluation page could also be added, such as selecting what metrics to display, choosing how the classification threshold is determined, or incorporating techniques to improve the model interpretability. Data processing could also be improved to retain multi-channel input information (which is currently compressed to one channel for simplicity) and accept medical images in DICOM or NIfTI file format, instead of the current input data format of JPG, PNG, or numpy array images.
Incorporating ML in radiology requires a collaborative approach between radiologists and ML researchers and will be of crucial importance for the field of radiology in the future.11,30 There is a large body of research on the application of CNNs to solve image classification problems in radiology,12,14-16 but there is limited work translating these advances in ML into practical applications in radiology, leading to a pronounced translational gap. Radiologist involvement will likely be a crucial step to translate ML research into clinical applications, 30 so ensuring radiologists understand the fundamentals of ML techniques as well as the technical terms used by the ML community will be essential to the process. Currently, there is limited work on how to teach ML to radiologists, given their usually markedly different training background and skillset from the individuals who typically enter the field of ML. We propose to use our GUI as an educational tool, providing a platform where users can gain hands-on experience constructing CNNs, to support a ML curriculum in radiology training.
There are several limitations to our study. First, our sample size of users was relatively small for the user testing sessions and all participants were recruited from the same institution. Hence, the sample may not be fully representative of the entire radiologist/radiologist-in-training population and the results might not generalize well to other teaching institutions. Second, these user testing sessions were performed on a voluntary basis, so the sample may contain some inherent biases, as these users had volunteered their time to test the software, indicating that they were open to exploring new technologies and learning more about how ML could fit into the field of radiology, whereas radiologists who are not as open to the idea of using ML in radiology would likely not have volunteered to test the software system. Further user testing with a larger sample size and diverse audience would be helpful.
Conclusion
An educational GUI allows interactive teaching in ML and CNNs in radiology, even for radiologists without prior experience in ML. User-testing feedback suggests that the GUI would be useful as a hands-on educational tool to teach ML to radiology trainees, as important concepts are conveyed in this hands-on approach without the need for understanding or writing code.
Footnotes
Acknowledgments
We would like to thank Drs. Nikil Rajani, Dan Halevy, and Nicolin Hainc for participating in this study.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Chair in Medical Imaging and Artificial Intelligence, a joint Hospital-University Chair between the University of Toronto, The Hospital for Sick Children, and the SickKids Foundation.
