Abstract
The circadian clock of Neurospora crassa is based on a negative transcriptional/translational feedback loops. The frequency (frq) gene controls the morning-specific rhythmic transcription of a sense RNA encoding FRQ, the negative element of the core circadian feedback loop. In addition, a long noncoding antisense RNA, qrf, is rhythmically transcribed in an evening-specific manner. It has been reported that the qrf rhythm relies on transcriptional interference with frq transcription and that complete suppression of qrf transcription impairs the circadian clock. We show here that qrf transcription is dispensable for circadian clock function. Rather, the evening-specific transcriptional rhythm of qrf is mediated by the morning-specific repressor CSP-1. Since CSP-1 expression is induced by light and glucose, this suggests a rhythmic coordination of qrf transcription with metabolism. However, a possible physiological significance for the circadian clock remains unclear, as suitable assays are not available.
Circadian clocks are widespread biological timing systems that orchestrate and coordinate biochemical pathways, physiology, and behavior of organisms in a time-of-day–specific manner. They are synchronized by daily recurring cues with the 24-h period of the earth’s rotation and thereby allow the anticipation of changes associated with the geophysical day-night cycle. The core of a circadian clock is based on cell-autonomous transcriptional-translational feedback loops (TTFLs) (Gallego and Virshup, 2007; Rosbash, 2009; Hogenesch and Ueda, 2011; Dunlap and Loros, 2017; Takahashi, 2017; Diernfellner and Brunner, 2020; Partch and Brunner, 2022).
The circadian clock of the filamentous fungus Neurospora crassa is driven by the transcription activator White Collar Complex (WCC), which supports rhythmic transcription of the core clock gene frequency (frq) and many clock-controlled genes (ccg’s) (Dunlap, 1999; Schafmeier et al., 2005). The intrinsically disordered FRQ protein (Pelham et al., 2020) dimerizes and assembles with FRQ-interacting RNA helicase (FRH) (Cheng et al., 2001, 2005; Hurley et al., 2013; Lauinger et al., 2014) and casein kinase 1a (CK1a) (Gorl et al., 2001; Marzoll et al., 2022) forming the FRQ-FRH-CK1a complex, FFC. The FFC inhibits and stabilizes the WCC by facilitating its phosphorylation by CK1a (Schafmeier et al., 2005, 2006). In the course of a circadian period, FRQ is progressively hyperphosphorylated (Querfurth et al., 2011), leading to its inactivation and degradation (He et al., 2003; Larrondo et al., 2015). The CK1a subunit of the FFC is the major kinase of FRQ (Querfurth et al., 2007, 2011; Marzoll et al., 2022) but other kinases also contribute to the phosphorylation (for review, see Diernfellner and Schafmeier, 2011; Diernfellner and Brunner, 2020). When the level/activity of FRQ declines, the previously phosphorylated WCC is reactivated by dephosphorylation by protein phosphatases including PP2a (Schafmeier et al., 2005) and PP5 (Cha et al., 2008). The reactivated WCC can now initiate a new circadian cycle of transcription of its target genes. At the frq locus, it binds about 1.2 kb upstream of the transcription start sites (TSS) to the so-called clock-box (c-box) (Froehlich et al., 2002) to initiate a new round of frq transcription.
The WCC is a heterodimeric transcription factor (TF) composed of a white collar-1 (WC-1) and -2 (WC-2) subunit (Cheng et al., 2002). WC-1 contains an LOV-domain photoreceptor that is activated by blue light (Froehlich et al., 2002). Light-activated WC-1 triggers dimerization of 2 WCC protomers (Malzahn et al., 2010), which then bind to a light response element (LRE) in the core frq promoter (Froehlich et al., 2002). The WCC dimer induces high levels of frq transcription and thereby synchronizes the circadian clock with light cues (Linden and Macino, 1997). Binding of the light-activated WCC to the LRE induces also transcription of a long noncoding RNA of unknown function that is transcribed toward the c-box (Kramer et al., 2003). In addition, a promoter located in the 3′ region of the frq gene directs transcription of an antisense RNA (Kramer et al., 2003) termed qrf (Xue et al., 2014). In constant darkness, qrf is rhythmically transcribed in antiphase to the overlapping frq sense RNA (Kramer et al., 2003; Xue et al., 2014). The qrf promoter contains an LRE, qLRE, and the light-activated WCC induces transcription of qrf RNA that interferes with and reduces expression of light-induced frq sense RNA about 2-fold (Xue et al., 2014).
The biological function of qrf transcription and its impact on the circadian clock are not understood. Kramer et al. (2003) replaced the qrf promoter at the EcoRV site located 219 bp downstream of the frq open reading frame (ORF) (see Figure 1a) by the 3′ UTR of the clock-controlled gene-2 (ccg-2) of Neurospora. This manipulation did not affect circadian clock function per se, but was associated with a small (2 h) but significant delay of the phase of the circadian conidiation rhythm. A later analysis by Xue et al. (2014) claimed, but did not show, that the EcoRV deletion did not completely abolish qrf transcription. They replaced the qrf promoter at the BssHII site 143 bp downstream of frq ORF with the quinic acid-inducible qa-2 promoter. When qa-2–dependent antisense transcription was either completely repressed or highly induced, the circadian conidiation rhythm of Neurospora was abolished. Xue et al. (2014) concluded that constitutive antisense transcription at a low level is mechanistically required for the generation of circadian transcriptional rhythms of frq sense RNA and therefore essential for the circadian clock. Finally, Li et al. (2015) detected a low-amplitude circadian rhythm of frq transcription when qrf RNA was expressed under control of the induced qa-2 promoter, suggesting that strongly induced qrf transcription did not fully inactivate the circadian clock.
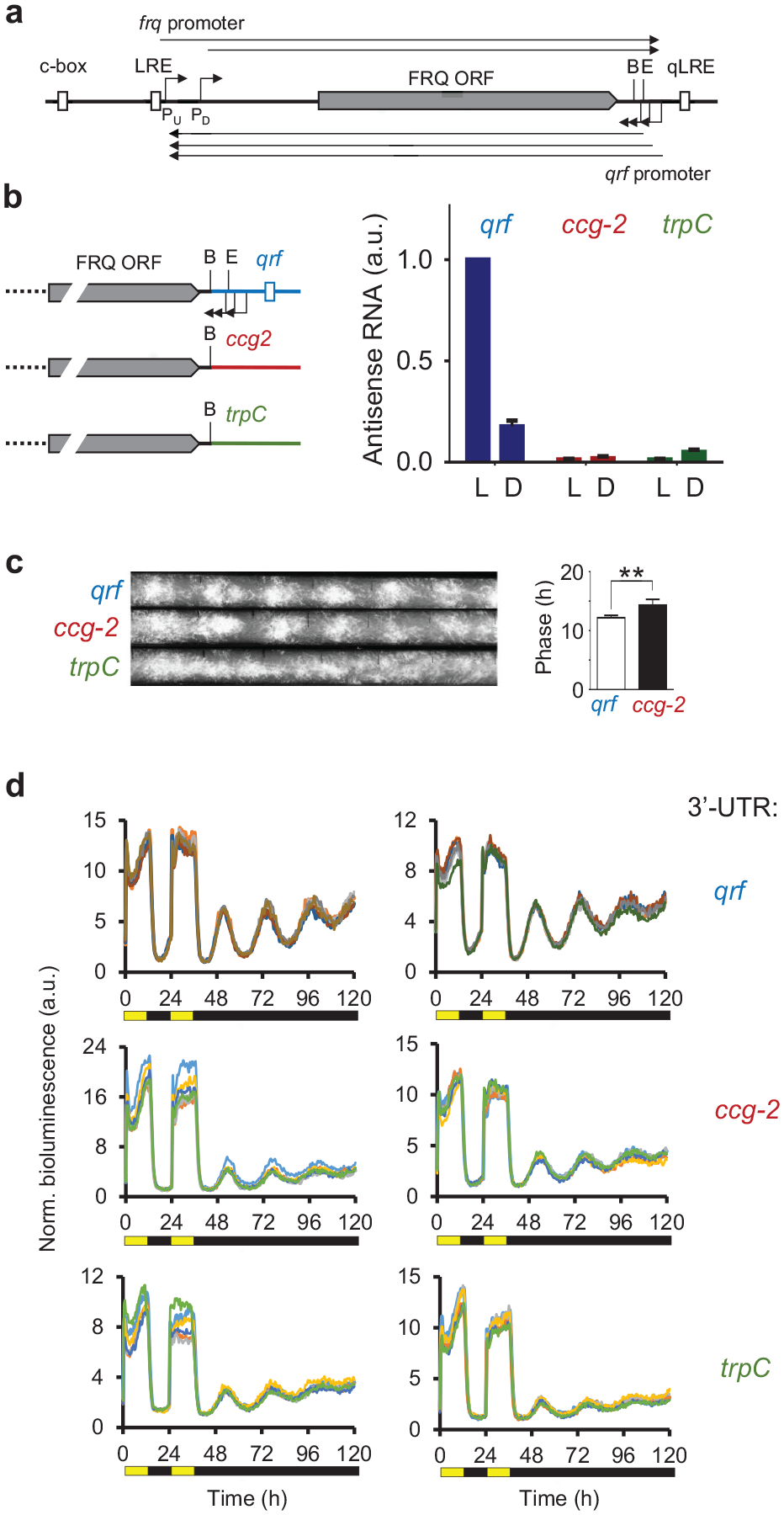
(a) Schematic of the frq locus. The arrows indicate transcription start sites and the filled box indicates the FRQ ORF. Start sites of the frq transcript were mapped between nt-1519 (PU) and nt-1170 (PD) relative to the first start codon (AUG) of the FRQ ORF (Colot et al., 2005; Diernfellner et al., 2005) and polyadenylation sites were mapped to between 54 and 265 bp downstream of the stop codon of the FRQ ORF (Kramer et al., 2003; Cemel et al., 2017) (http://neutra.bzh.uni-heidelberg.de). Start sites of qrf transcripts were mapped to positions 217, 241, and 278 bp downstream of the stop codon of the FRQ ORF (Kramer et al., 2003) and polyadenylation sites were mapped to between 1530 and 1500 bp upstream of the first start codon (AUG) of the FRQ ORF (Cemel et al., 2017) (http://neutra.bzh.uni-heidelberg.de). (b) (Left) Schematic representation of the qrf mutants. The WT frq locus and the frq genes with the ccg-2 and the trpC 3′ UTRs, respectively, are depicted. (Right) The frq genes with the different 3′ UTRs, qrf (WT control), ccg-2, and trpC were inserted into the his-3 locus of a Δfrq strains. Strains were grown in constant light (LL) or for 22 h in dark (DD22). RNA was prepared and frq expression was measured by RT-qPCR. Actin was used as an internal reference. Error bars represent ±SEM (n = 3). (c) Representative race tube assays of the indicated strains. frqwt: τ = 22.81 ± 0.19 h, Δfrq,frq-ccg-2: τ = 22.39 ± 0.70 h, Δfrq,frq-trpC: arrhythmic (n = 6). τ: period length. The bar plot shows the phases of the WT control and Δfrq,frq-ccg-2. Error bars represent ±SEM (n = 4-6). **Indicates p < 0.01. (d) Normalized bioluminescence recordings of frq-lucPEST reporter gene expression in Δfrq,frq-qrf and Δfrq,frq-ccg-2 and Δfrq,frq-trpC strains grown in 12-h light, 12-h dark, and 12-h light before release into constant darkness. For each strain, 5 technical replicates of 2 representative clones are shown. Abbreviations: ORF = open reading frame; LRE = light response element; qLRE = antisense LRE; WT = wildtype; B: BssHII site; E: EcoRV site; ccg-2 = clock-controlled gene-2; RT-qPCR = reverse transcription quantitative polymerase chain reaction.
We show here that the qrf promoter is dispensable for circadian rhythmicity. Rather, the circadian clock is highly sensitive to changes in the 3′-UTR of frq RNA that affect RNA turnover. We also show that the glucose- and light-induced transcriptional repressor conidial separation phenotype-1 (CSP-1) supports rhythmic expression of qrf in antiphase to frq.
Materials and Methods
Neurospora Strains and Culture Conditions
Neurospora wild-type strain was acquired from Fungal Genetics Stock Center (FGSC) with stock number #2489. For race tube assay, strains with ras-1bd mutation were used (Belden et al., 2007). Since the conventional frq deletion strain frq10 (Aronson et al., 1994) contains the enhancer site c-box and the qrf LRE, we created a full knockout of the frq locus in this study. The Δfrq bd, his-3 strain was created with the yeast in vivo recombination system as described previously (Colot et al., 2006) using the following knockout cassette primers:

The plasmids for the qrf knockout strains were created with the following primers:
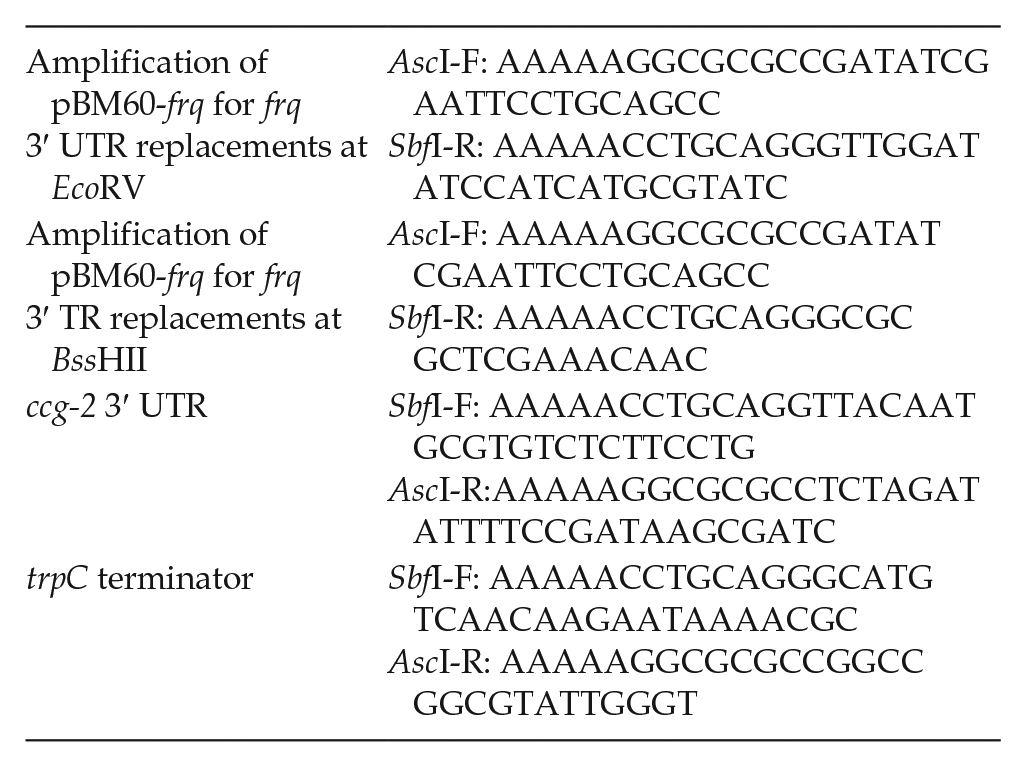
The Δfrq bd, his-3 (histidine auxotroph) strain was used for transformation. All constructs were targeted to the his-3 locus via homologous recombination. Briefly, 5- to 7-day-old conidia were harvested and washed and pelleted at 2600 g at 4 °C for 10 min with 50 mL of 1 M cold sorbitol. The conidial pellet was mixed with 1- to 2-μg linearized plasmid DNA and incubated on ice for 10 min. Electroporation was applied at 1.5 kV/cm, 25 μF, 600 Ω. The cells were immediately resuspended in 1 mL of 1 M cold sorbitol and plated onto Vogel’s solid media (1× Vogel’s, 1% [w/v] agar, 1× FGS [20% (w/v) sorbose, 0.5% (w/v) glucose, and 0.5% (w/v) fructose]). After incubation at 30 °C for 3-5 days, the single colonies were picked. N. crassa cultures were grown in standard growth medium (2% glucose, 0.5%
RNA Preparation and cDNA Synthesis
RNA was isolated with peqGOLD TriFAST (PeqLab). The reverse transcription was carried out with QuantiTect Reverse Transcription Kit (Qiagen) with indicated primers following manufacturer’s instructions. Relative transcript levels were quantified by quantitative real-time polymerase chain reaction (PCR) in 96-well plates with LightCycler 480 (Roche). The reaction was set by using qPCRBIO Probe Mix Hi-ROX (Nippongenetics) and TaqMan (5′: 6-FAM, 3′: TAMRA) or Universal Probe Library (UPL, Roche) probes. Primers and probes are listed below. Three replicates were used to calculate the mean threshold cycle (Ct) value. The relative enrichments were quantified relative to a housekeeping gene using the reverse transcription primers for cDNA synthesis:

And primers for qPCR are as follows:
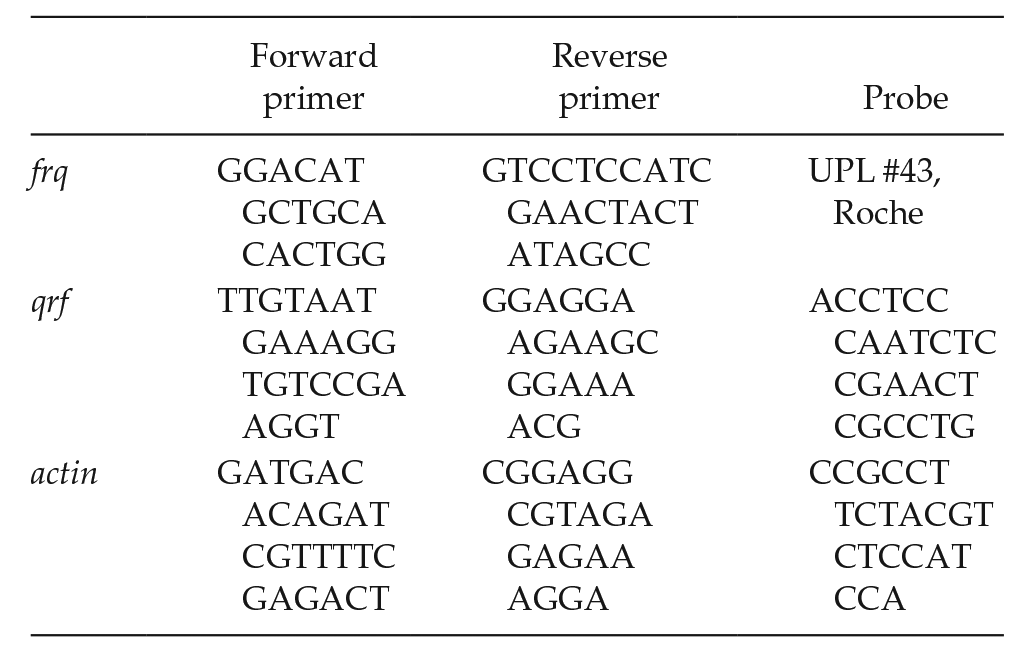
Race Tube Assay
The conidial suspension was inoculated from one end of the autoclaved glass tubes containing the media (1× Vogel’s, 0.1% glucose [w/v], 0.17% arginine [w/v], 50 ng/mL biotin, and 2% agar [w/v]) and was grown for 1 day prior to light to dark transfer. The conidial growth fronts were marked every 24 h.
Live-Cell Bioluminescence Monitoring
Sorbose medium (1× FGS [0.05% fructose (w/v), 0.05% glucose (w/v), 1% sorbose (w/v)], 1× Vogel’s, 1% agarose [w/v], 10-ng/mL biotin, and 75-μM firefly luciferin) was used for the live-cell bioluminescence assay; 96-well plates were inoculated with 3 × 104 conidia per well and incubated in dark at 25 °C for 2 days. The bioluminescence signal was recorded in constant darkness or in light-dark cycles at 25 °C for the indicated time windows with a multilabel plate reader. For dark recordings, the cells were synchronized by 1-h light pulse (LP) at 100-μmol photons m−2 sec−1 prior to the measurement in constant darkness; 0.3% glucose as carbon source was used when high glucose conditions are indicated. The light intensity titration (Figure 3e) was performed as described (Cesbron et al., 2015).
Results and Discussion
Antisense Transcription at frq Locus Is Not Required for Rhythmicity
Previous studies (Kramer et al., 2003; Colot et al., 2005; Xue et al., 2014) and RNA-Seq data (Cemel et al., 2017) indicate that the frq locus directs expression of multiple species of overlapping sense and antisense transcripts (Figure 1a). Light-induced expression of frq and its antisense transcript, qrf, are driven from their corresponding LREs, LRE and qLRE, respectively (Froehlich et al., 2002; He et al., 2002; Smith et al., 2010; Xue et al., 2014). The majority of sense frq RNA species terminate between the qLRE and the annotated TSSs of the qrf promoter (Belden et al., 2011), and qrf transcripts terminate within the region of annotated TSSs of the frq promoter (Figure 1a).
To analyze the impact of antisense transcription on the circadian clock, we replaced the qrf promoter with 2 different transcription termination sequences, the 3′ UTR of the ccg-2 of N. crassa (Kramer et al., 2003) and the trpC terminator of Aspergillus nidulans (Mullaney et al., 1985), respectively. The sequences were inserted at the BssHII site to ensure deletion of all mapped qrf TSSs (Figure 1b, left). The modified frq genes and an unmodified frq gene with its natural qrf sequence (frq-qrf control), respectively, were inserted into the Neurospora genome at the his-3 locus of a Δfrq strain (see the “MATERIALS AND METHODS” section). RNA directed from these genes was quantified by quantitative reverse transcription polymerase chain reaction (qRT-PCR) (Figure 1b, right). In the frq-qrf control strain, antisense RNA was expressed at a high level in light and at a ~5-fold lower level in the dark. In contrast, in the frq-ccg-2 and frq-trpC strains, the antisense transcript levels were significantly reduced, indicating that deletion of the qrf promoter at the BssHII site compromised the expression of antisense RNA in light and in dark.
We then assessed the conidiation rhythms of the frq-qrf control strain and the mutant strains lacking antisense transcription. The frq-qrf control strain and frq-ccg-2 strain exhibited rhythmic conidiation in constant darkness (DD), but the phase of conidiation of the frq-ccg-2 strain was delayed by 2 h (Figure 1c). The observed phase delay is consistent with a previous report (Kramer et al., 2003), in which the qrf promoter had been replaced at the EcoRV site by the 3′ UTR of ccg-2. In contrast, conidiation in DD was apparently arrhythmic in the frq-trpC strain, indicating that the circadian clock was compromised. Thus, despite frq-ccg-2 and frq-trpC strains being both deficient in antisense transcription, conidiation of frq-ccg-2 was rhythmic while the overt rhythm of frq-trpC was lost.
We then generated frq-qrf, frq-trpC, and frq-trpC strains that expressed in the tubulin locus a destabilized luciferase gene under control of the frq promoter (frq-lucPEST) and measured luciferase activity as described previously (Gooch et al., 2008; Cesbron et al., 2013). Both, frq-ccg-2 and frq-trpC strains supported rhythmic expression of frq-lucPEST, albeit with a lower amplitude than the frq-qrf control strain (Figure 1d). However, compared with the race tube analysis (Figure 1c), the frq-lucPEST reporter gene did not display differences in circadian phase between frq -qrf and frq-ccg-2 strains.
Since the frq mRNA terminates downstream of the mapped qrf TSSs (Belden et al., 2011), replacement of the qrf promoter by ccg-2 or trpC altered the 3′ UTR of the frq sense transcript. We therefore asked how the choice of the DNA sequence, which was used to replace the qrf promoter, affected the stability of frq mRNA. To assess RNA turnover under physiological conditions, we allowed accumulation of high levels of the frq RNA by growing mycelial cultures in constant light. The light-induced transcription of frq was then shut down by a transfer of the cultures to the dark and the decrease of frq RNA was then measured over a time course of 120 min (Figure 2a). The frq RNA levels in the frq-qrf control strain decreased with a half-time (t1/2) of 15 min, confirming that frq RNA is unstable (Diernfellner et al., 2005). The level of the frq-ccg-2 transcript decreased with t1/2 ~ 25 min, while the frq-trpC transcript was substantially more stable (t1/2 ~ 1 h). The data demonstrate, not surprisingly, that the 3′ UTRs affect the stability and hence the expression level of frq mRNA.
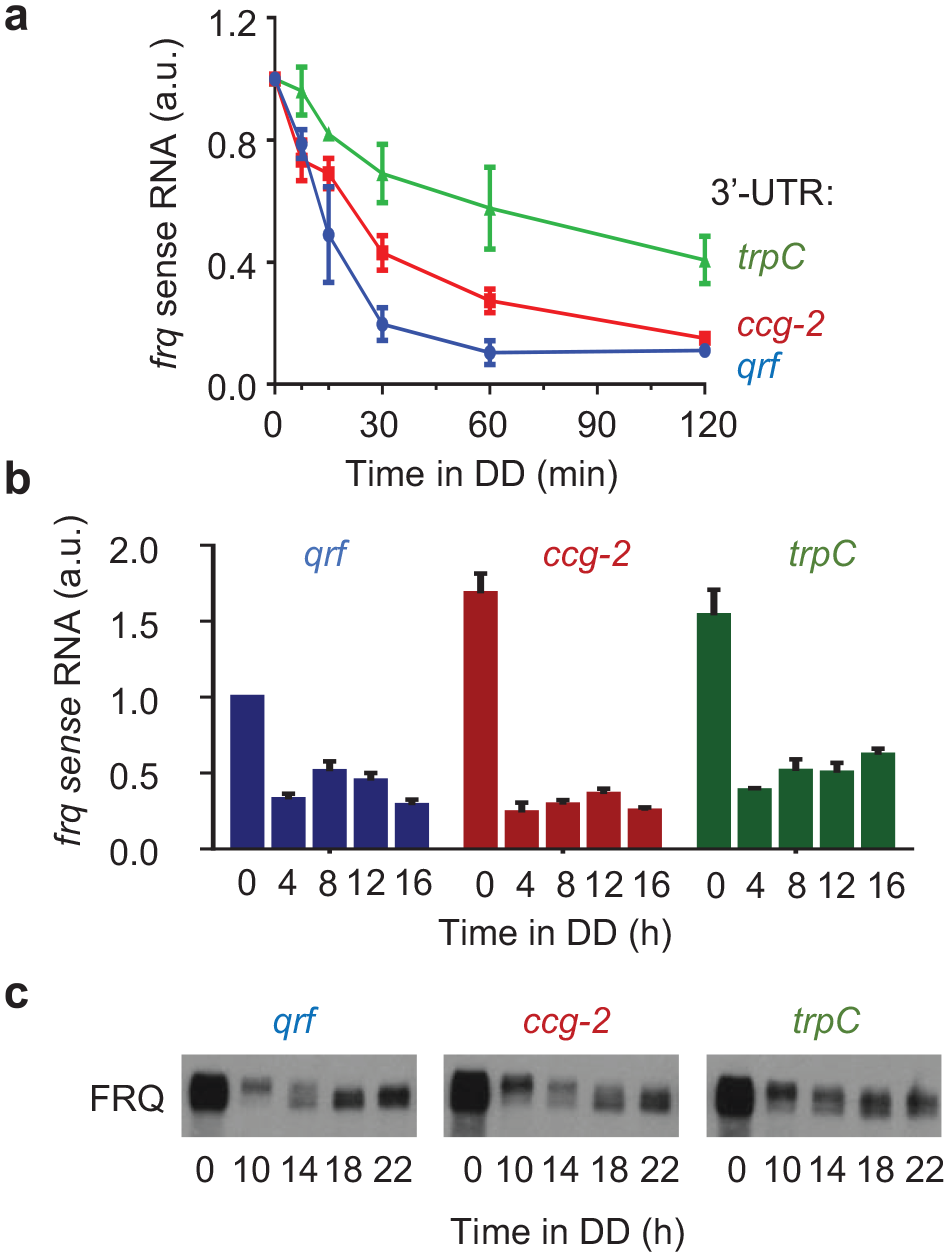
(a) Stability of frq mRNA in WT and qrf deficient strains. Strand-specific RT-qPCR results showing the frq mRNA levels in light (0 h) and in darkness at indicated time points (min). The transcript levels were normalized to the respective light value, and actin was used as an internal reference. Error bars indicate ±SEM (n = 3). (b) Rhythmic expression of frq in wt and qrf deficient strains. Strand-specific RT-qPCR results showing the frq mRNA levels in light (0 h) and in darkness at indicated time points (h). Relative frq levels were normalized to the wt levels in light (0 h). Error bars indicate ±SEM (n = 3). (c) Western blot analysis of FRQ expression profiles in the Δfrq,frq-ccg-2 and Δfrq,frq-trpC strains in DD at the indicated time points (n = 3). Abbreviation: RT-qPCR = reverse transcription quantitative polymerase chain reaction.
We then measured frq RNA and FRQ protein in light and after LD transfer of mycelial cultures (Figure 2b). In light (t = 0 h), frq transcript levels in the frq-ccg-2 and frq-trpC strains accumulated at a higher level than in the frq-qrf control strain, confirming previous reports that replacement of the antisense promoter results in accumulation of elevated levels of sense RNA (Xue et al., 2014). In constant darkness, the frq-qrf control strain expressed frq sense RNA in circadian fashion, with a peak about 12 h after transfer of the mycelial cultures into the dark (Figure 2b). FRQ protein displayed circadian abundance and phosphorylation rhythms (Figure 2c). In the frq-ccg-2 strain, frq RNA and FRQ protein also displayed circadian rhythms in DD (Figure 2b and 2c). In the frq-trpC strain, the levels of frq and FRQ in dark were elevated and did not display apparent rhythms on the time frame of the experiment (Figure 2b and 2c). Hence, the delayed conidiation rhythm of frq-ccg-2 and the arrhythmic conidiation of frq-trpC strains correlate with the expression profiles of frq and FRQ in these strains after LD transfer, but not with the phase and rhythm of the frq-lucPEST reporter.
Together, our results show that antisense (qrf) transcription at the frq locus is not required for rhythmicity of the core circadian clock. Rather, the circadian clock is sensitive to changes in the 3′ UTR that stabilize frq RNA, suggesting that rapid turnover of frq RNA appears to be critical for clock amplitude and therefore may influence overt rhythmicity.
Transcription Dynamics of qrf in Dark and in Response to Light
Next, we characterized the transcriptional properties of the qrf promoter using luciferase reporter constructs. Three TSSs of the qrf promoter were previously mapped, 2 between the BssHII and EcoRV sites and 1 immediately downstream of the EcoRV site (Figure 3a). To include all TSSs, we fused the qrf promoter immediately after the stop codon of the FRQ ORF (1117 bp) to luc-PEST (Figure 3a). The DNA sequence transcribed by the qrf promoter contains several ATG codons, and 3 of these are present in the 5′-UTR of the chimeric qrf-lucPEST transcription unit. We changed these 3 upstream ATGs in the reporter gene to CTGs to ensure optimal translation of lucPEST ORF. The modified qrf-lucPEST reporter supported efficient bioluminescence expression and exhibited a low-amplitude circadian rhythm (Figure 3b). We also generated lucPEST reporter constructs with qrf promoters truncated at the EcoRV and BssHII site, respectively (Figure 3a). In comparison with the transcriptional activity of the full-length qrf-lucPEST reporter, the bioluminescence levels supported by qrf promoters truncated at EcoRV and BssHII, respectively, were substantially reduced (Figure 3b). The residual activity was arrhythmic and similarly low for the EcoRV and BssHII truncations. The data suggest that truncations at either restriction site abolish qrf rhythms and transcription levels to the same extent.
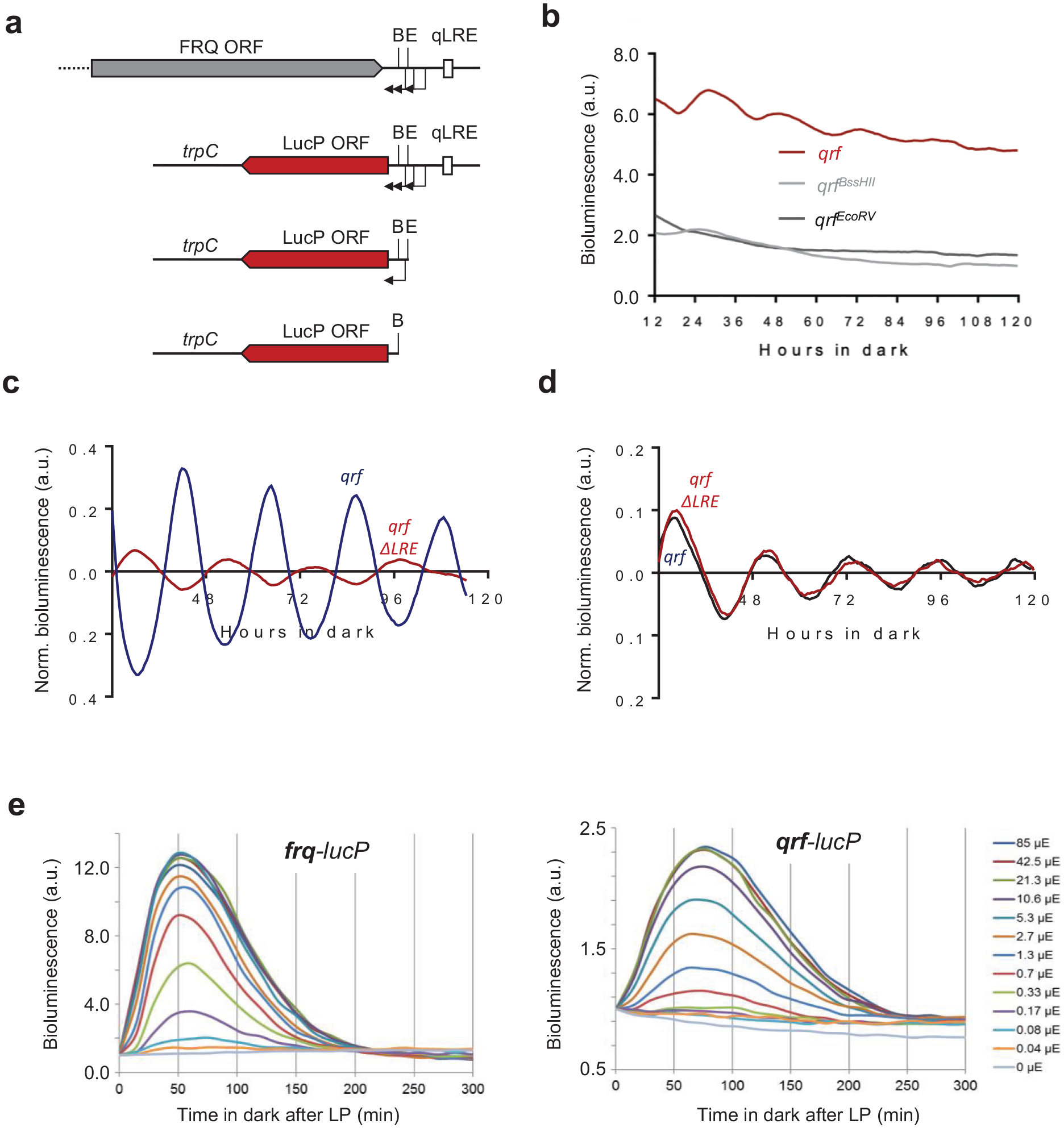
(a) Schematic representation of the luciferase reporter constructs together with the frq locus. The arrows indicate transcription start sites. The filled boxes represent FRQ ORF (gray) or LucP ORF (red). (b) Representative normalized bioluminescence recordings of luciferase reporters driven by the complete qrf promoter, truncated qrf promoters at E or B restriction sites. The reporters were expressed in wt bd. Two clones and their means are depicted. (c) Rhythmic qrf promoter activity. Normalized luciferase activity of the qrf promoter in comparison with the frq promoter activity. The bioluminescence activities were recorded in constant darkness after synchronization via light to dark transfer. (d) Normalized qrf- and qrfΔLRE-driven luciferase activities and in constant darkness. (e) Differential saturation of frq and qrf promoters. Strains expressing frq-lucPEST and qrf-lucPEST reporter genes were exposed to a 1-min LP of the indicated intensities. Luciferase activity at LP was used for normalization (n = 3). Abbreviations: ORF = open reading frame; B = BssHII site; E = EcoRV site; LP = light pulse; LucP = LucPEST (detabilized Luciferase); qLRE = antisense LRE.
We then compared the transcription rhythms of the isolated qrf (antisense) reporter with the rhythm supported by a frq (sense) reporter gene (Figure 3c). The expression levels of the qrf-lucPEST reporter oscillated in antiphase to the circadian rhythm of the frq-lucPEST reporter.
Light-dependent expression of qrf is controlled by the qLRE (Smith et al., 2010; Xue et al., 2014). To assess whether the qLRE is required for the antiphasic transcription rhythm in the dark, we mutated the sequence in the qrf-lucPEST reporter and measured luciferase expression in the corresponding reporter strain, qrfΔqLRE. The qLRE mutation did not affect the evening-specific circadian expression rhythm of the qrfΔqLRE promoter (Figure 3d).
Finally, to compare the responsiveness and sensitivity of the frq and qrf promoters to light, the frq-lucPEST and qrf-lucPEST reporter strains were exposed to 1-min LPs of different intensities and bioluminescence was then recorded for 300 min (Figure 3e). The frq-lucPEST reporter reached peak expression levels about 50 min after the LP, while the qrf-lucPEST reporter reached maximal expression levels after about 75 min. Furthermore, expression levels of the frq-lucPEST reporter saturated at LP intensities above 2.7-µmol photon m−2 sec−1, while the qrf promoter saturated at much higher LP intensities (>21.3-µmol photon m−2 sec−1). The data indicate that the qrf promoter is less responsive and less sensitive to light cues.
The qrf Promoter Is Controlled by CSP-1
In constant darkness, clock-controlled genes of Neurospora are rhythmically expressed in mainly 2 phases, subjective morning and evening, respectively (Bell-Pedersen et al., 1996; Sancar et al., 2015b). Morning-specific genes are expressed approximately in phase with the activity profile of the transcription activator WCC, while many evening-phased genes are regulated by the transcription repressor CSP-1 (Sancar et al., 2015b). Since the transcriptional activity of qrf is evening specific, that is, in antiphase to the rhythm of the WCC-driven frq promoter, we asked whether the qrf promoter is controlled by the morning-specific CSP-1 repressor. Analysis of CSP-1 ChIP-Seq (Sancar et al., 2011) and WC-2 ChIP-Seq (Sancar et al., 2015a) datasets revealed that CSP-1 binds overlapping with the LRE and qLRE of the frq and qrf promoters, respectively, and also close to the c-box of frq (Figure 4a). We have previously shown that frq expression is elevated and phase delayed in a Δcsp-1 strain (Sancar et al., 2011). To assess the impact of CSP-1 on the qrf promoter, we analyzed expression of qrf-lucPEST in a Δcsp-1 strain. Expression of qrf-lucPEST was elevated (p < 0.05), and its circadian rhythm was abolished. The data suggest that qrf is constitutively activated by at least one unknown TF. The activity of this TF is antagonized by CSP-1, which represses qrf transcription. Since CSP-1 is rhythmically expressed with a morning-specific phase, it generates an antiphasic evening-specific qrf rhythm (Figure 2d). Together, these results demonstrate that the isolated qrf promoter displays an evening-specific expression rhythm, which is dependent on the morning-specific activity of the repressor CSP-1.
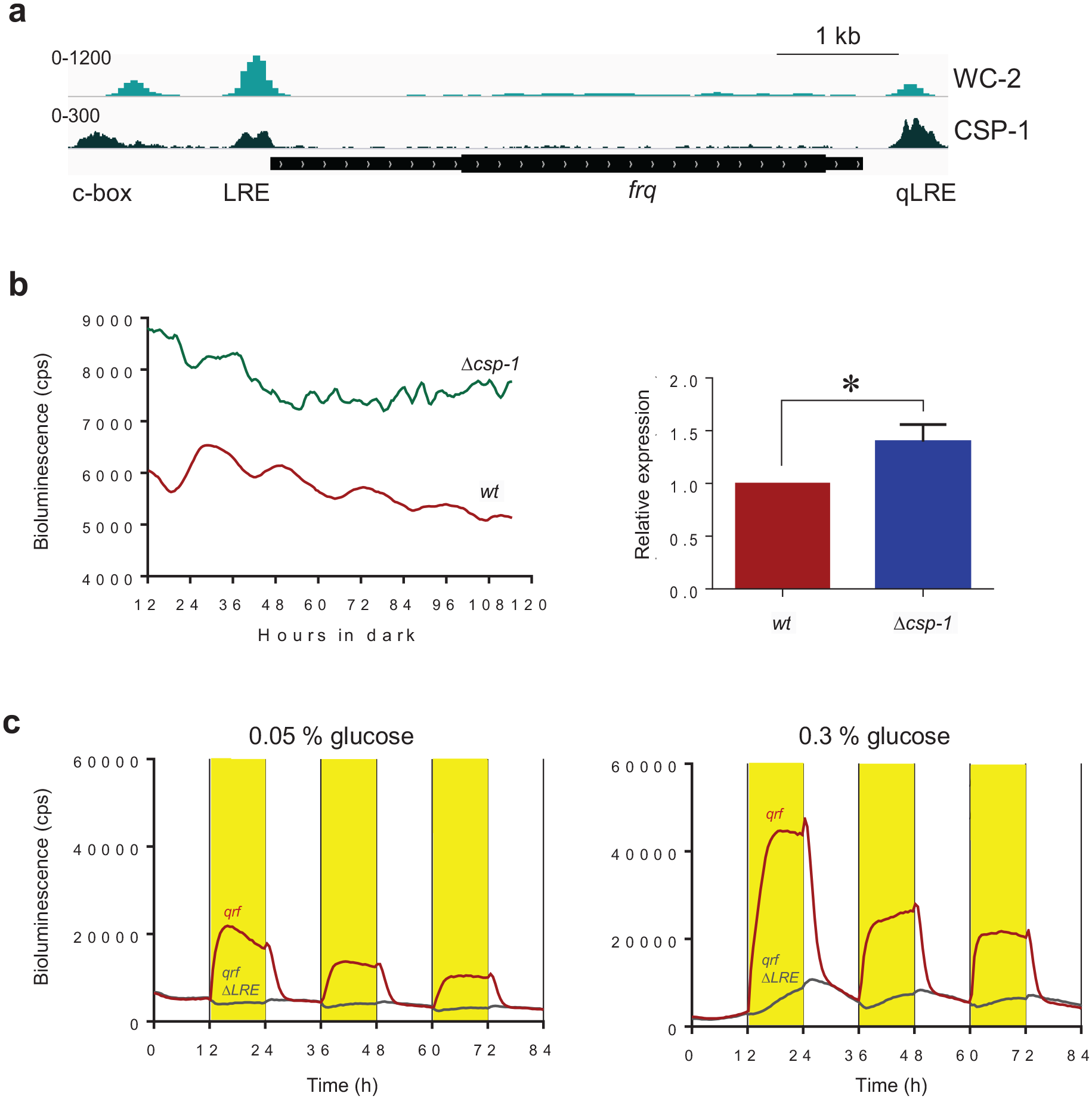
(a) WC-2 and CSP-1 binding at the frq locus. ChIP-Seq datasets were published in Sancar et al. (2011) and Sancar et al. (2015a), respectively. (b) Luciferase activity of qrf promoter in WT and Δcsp-1 strains. Following light to dark transfer, the bioluminescence activities were recorded in constant darkness. Intensity values of signals from all time points were added up and normalized to WT. The quantification of the 3 independent experiments is shown. Error bars represent ±SEM (n = 3). *Indicates p < .05. (c) Representative bioluminescence measurement of the qrf-lucPEST and qrfΔLRE-lucPEST reporter in low (0.05%) and high (3%) glucose concentrations in 12 h dark (white) to 12 h light (yellow) (LD) cycles. Abbreviations: LRE = light response element; ChIP = chromatin immunoprecipitation; WC-2 = white collar-2; CSP-1 = conidial separation phenotype-1; qLRE = antisense LRE.
CSP-1 is a short-lived repressor that is expressed in light-dependent fashion under transcriptional control of the WCC, and independently of WCC also in glucose-dependent fashion (Sancar et al., 2012). To characterize the impact of CSP-1 on qrf transcription, we examined qrf-lucPEST and qrfΔqLRE-lucPEST reporter strains. The strains were cultured in 96-well plates on agar medium with low and high glucose concentration. The cultures were exposed to repetitive 12 h:12 h LD cycles, and bioluminescence profiles were recorded for 84 h (Figure 4a and 4b). The qrf-luc-PEST strain displayed light-driven luciferase activity (bioluminescence) at low and high glucose concentration. On low glucose medium (Figure 3a, left), the bioluminescence levels decreased rapidly after the LD transition and approached dark levels within ~4 h. On high glucose medium (Figure 3b, right), the decrease in bioluminescence after the LD transition was biphasic: the initial rapid decrease of bioluminescence within the first 4 h in the dark was followed by a slower decrease during the 4- to 12-h time period after the LD transition. To our surprise, the bioluminescence supported by qrfΔqLRE-lucPEST was modulated by light in a glucose-dependent manner (Figure 4c). At low glucose concentration, qrfΔqLRE-lucPEST expression was essentially unaffected by light (Figure 4c, left). In contrast, at high glucose concentration in the medium, qrfΔqLRE-lucPEST expression displayed a specific response to light despite the absence of a functional qLRE (Figure 4c, right): after about 1 h in light, the bioluminescence supported by qrfΔqLRE-lucPEST increased steadily. After LD transition, the bioluminescence kept increasing for about 2 h and then declined steadily, resulting in a sawtooth-like bioluminescence rhythm that was delayed relative to the DL and LD transitions. Interestingly, the decrease in the dark of qrfΔqLRE-lucPEST expression superimposed with the slow decrease of qrf-lucPEST activity observed 4-8 h after LD transition (Figure 4c). The data suggest that light supports the expression of an unknown, glucose-dependent transcription activator of qrf, which accumulates steadily during the light phase and is degraded during the dark phase. Hence, the qrf promoter is regulated in a complex manner by at least two environmental cues, light and glucose. Light-dependent expression is directly controlled by the WCC via the qLRE, while glucose-dependent expression is controlled by an unknown activator and by the rhythmically expressed repressor CSP-1. Activity and presumably expression levels of both, activator and repressor, are dependent on light and glucose.
Taken together, our results demonstrate that antisense transcription is not required for the function of the circadian clock in constant darkness (Figures 1 and 2). Rather, the impact of qrf transcription on frq transcription in light via WCC (Kramer et al., 2003; Xue et al., 2014) (Figure 3), the direct impact of glucose via CSP1 on frq (Sancar et al., 2011) and qrf transcription (Figure 4a and 4b), and the indirect impact of glucose and light on qrf via an unknown transcription activator (Figure 4c) suggest that antisense transcription may help fine-tune and coordinate the light-dark phase of frq expression with the metabolic state of the fungus.
Footnotes
Acknowledgements
This work was supported by the Deutsche Forschung-sgemeinschaft (collaborative research center TRR186).
Conflict of Interest Statement
The author(s) have no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
