Abstract
Biomedical research on mammals has traditionally neglected females, raising the concern that some scientific findings may generalize poorly to half the population. Although this lack of sex inclusion has been broadly documented, its extent within circadian genomics remains undescribed. To address this gap, we examined sex inclusion practices in a comprehensive collection of publicly available transcriptome studies on daily rhythms. Among 148 studies having samples from mammals in vivo, we found strong underrepresentation of females across organisms and tissues. Overall, only 23 of 123 studies in mice, 0 of 10 studies in rats, and 9 of 15 studies in humans included samples from females. In addition, studies having samples from both sexes tended to have more samples from males than from females. These trends appear to have changed little over time, including since 2016, when the US National Institutes of Health began requiring investigators to consider sex as a biological variable. Our findings highlight an opportunity to dramatically improve representation of females in circadian research and to explore sex differences in daily rhythms at the genome level.
Underrepresentation of females is a persistent problem in biomedical research and is particularly stark among preclinical studies using non-human mammals (Beery and Zucker, 2011; Woitowich et al., 2020). In addition, many preclinical studies either do not report the sex of biological samples or do not report results by sex (Mamlouk et al., 2020). Altogether, this lack of sex inclusion obscures the extent to which research findings generalize to roughly half the population (Clayton, 2016). Recognizing this issue, the US National Institutes of Health (NIH) created a policy in 2016 that requires investigators to factor biological sex into the design, analysis, and reporting of vertebrate animal and human studies (NOT-OD-15-102: Consideration of Sex as a Biological Variable in NIH-funded Research, n.d.).
Unfortunately, lack of sex inclusion is an issue in circadian research as well. For example, only 34% of studies on the non-visual effects of light in humans have included females (Spitschan et al., 2022). Even more striking, among studies on light and circadian phase-shifting in rodents published 1964-2017, only 7% included females (Lee et al., 2021). The severe underrepresentation of female rodents in circadian research may be a legacy of early observations of effects of the estrous cycle and estradiol on daily rhythms in hamsters (Morin et al., 1977; Takahashi and Menaker, 1980), although recent work indicates that female rats are not more variable than male rats in any neuroscience-related traits (Becker et al., 2016). Encouragingly, there is growing recognition of the importance of addressing sex bias and investigating sex differences in sleep and circadian rhythms (Spitschan et al., 2022; Joye and Evans, 2022). Although sleep and circadian research increasingly makes use of genomic techniques, sex inclusion trends in genomic studies of daily rhythms remain undescribed.
To address this gap, we first assembled a collection of publicly available transcriptome studies (bulk microarray or RNA-sequencing) from mice, rats, or humans that had samples from at least 3 times of day (Supplemental Tables S1 and S2). We downloaded the metadata for each study using the seeker R package (Schoenbachler and Hughey, 2022). From the metadata, we extracted the organism, tissue, time of day, and biological sex (where available) of each sample from each study. Where necessary, we obtained information on biological sex from the published article. For our analysis, we only included studies linked to a published article and considered one study as corresponding to one article. We did not filter studies or samples with respect to genotype, light-dark cycle, or any other experimental condition. Altogether, the collection comprised 123 studies in mice (7305 samples), 10 in rats (373 samples), and 15 in humans (191 subjects). We defined each study’s sex inclusion status as male only, female only, male and female (if each sample’s sex was identifiable), mixed (if each sample was based on pooled tissue from males and females, or if the article stated using both males and females, but each sample’s sex was not identifiable), or unspecified.
We examined sex inclusion by organism (Figure 1a and Supplemental Table S3). In mice, 91 (74%) of 123 studies included only males, whereas 23 studies (19%) included females, as female only, male and female, or mixed. In rats, all 9 studies having specified sex included only males. In humans, sex inclusion was more balanced, as 8 of 15 studies had samples from males and females. We observed similar trends when analyzing the collection based on numbers of samples (Supplemental Table S4). These results indicate that females are highly underrepresented in mammalian circadian transcriptome data.
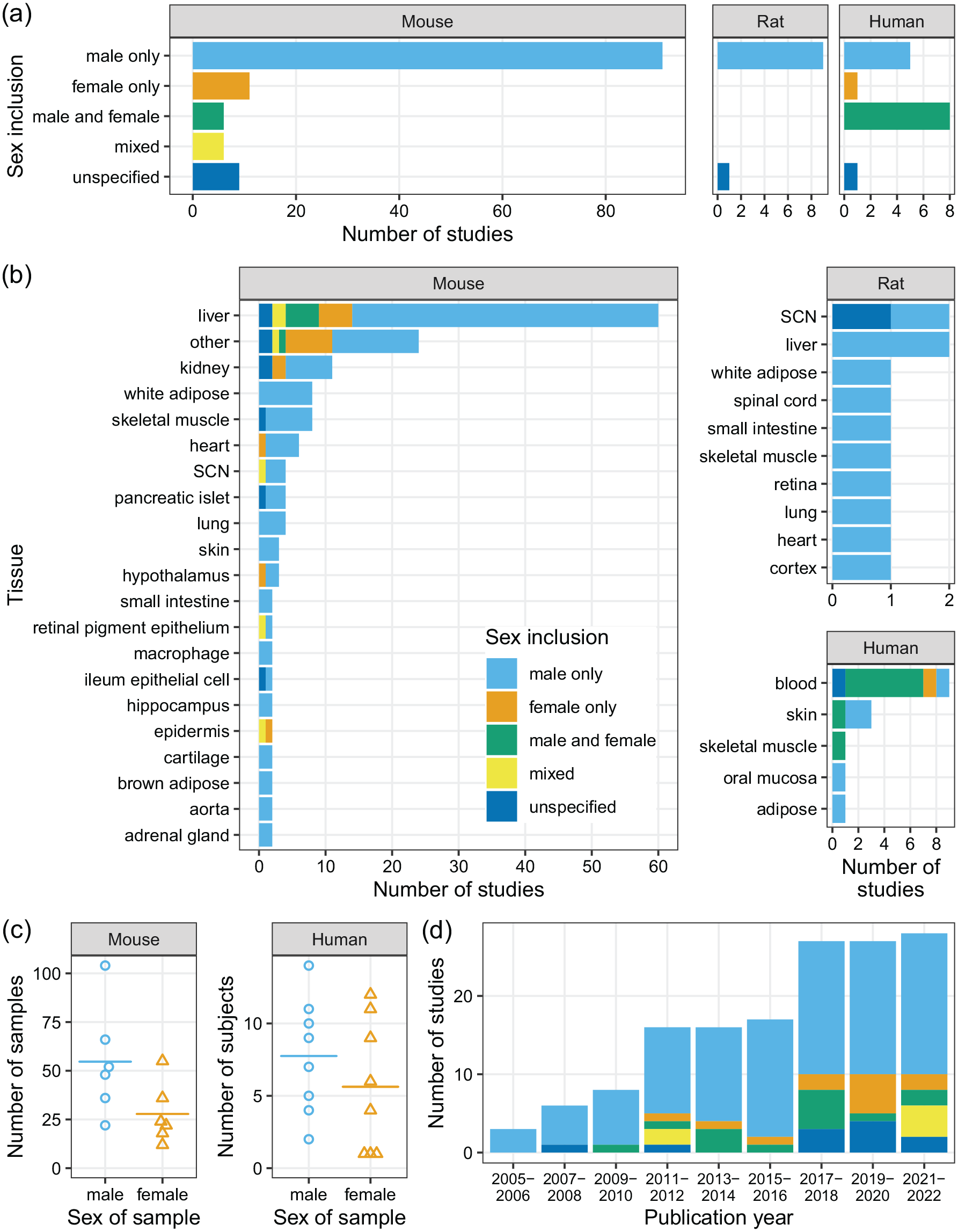
Sex inclusion in circadian transcriptome studies from mice, rats, and humans. Each study corresponds to a published article. (a) Barplots of number of studies by organism and sex inclusion status. The latter is also represented by color, which is consistent throughout the figure. (b) Barplots of number of studies by organism, tissue, and sex inclusion status. For mouse studies, the “other” tissue comprises tissues represented in only 1 study. Some studies included samples from multiple tissues, so the total counts in (b) could be more than the counts in (a). (c) Beeswarm plots of number of samples (per mouse study) and number of subjects (per human study) among studies whose sex inclusion status was male and female. Each point represents a study. Each horizontal line represents the mean for that group. (d) Barplot of number of studies (from all organisms) by sex inclusion status and publication year of the corresponding article. Abbreviation: SCN = suprachiasmatic nucleus.
We next examined sex inclusion by organism and tissue (Figure 1b and Supplemental Tables S5 and S6). In mice, 23 of 44 unique tissues had samples only from males, whereas 16 tissues had samples in some way from females (as female only, male and female, or mixed; Supplemental Table S7). Within liver, by far the most common tissue, 46 of 60 studies were male only, whereas 5 were female only and 5 were male and female (Vollmers et al., 2009; Yang et al., 2016; Sato et al., 2017; Weger et al., 2019; Mekbib et al., 2022) (Supplemental Table S8). In humans, the most common tissue was blood, for which 6 of 9 studies included males and females. However, both mouse and human studies including males and females tended to have more samples (or subjects) from males than from females (Figure 1c; paired differences of 26.8 ± 34.7 samples for mouse and 2.1 ± 5.0 subjects for humans).
We also examined sex inclusion trends over time, based on the publication year of each study’s corresponding article (Figure 1d and Supplemental Table S9). These results indicate that even as the overall number of studies has increased, the representation of females has remained low and roughly constant since 2011. Although the proportion of male-only studies appears slightly lower since 2021, whether this is noise or a sustained trend remains to be seen.
To explore these results further, we examined the articles for studies published in 2021-2022, which revealed 3 main findings (Supplemental Table S10). First, most studies whose sex inclusion status was male only or female only did not mention a justification. Second, the underrepresentation of females in transcriptome experiments was concordant with that of other experiments. Third, of the 9 studies that included males and females for any experiment, 2 reported results by sex (Jouffe et al., 2022; Mekbib et al., 2022).
Although the collection of studies we examined here is extensive, it is not exhaustive. First, it includes only transcriptome studies, not studies based on other genomic techniques, which are relatively less common and often performed on the same or similar samples. Second, the collection only includes studies in which time of day was an experimental variable, and thus excludes studies based on samples from post-mortem human donors, where information on time of day of death has revealed daily variation in gene expression and where donors may be more sex-balanced (Li et al., 2013; Chen et al., 2016; Ruben et al., 2018). Third, the collection does not include studies from less commonly used vertebrates or from invertebrates—despite the importance of fruit flies to circadian research. Nonetheless, our findings are consistent with those for biomedical research as a whole (Woitowich et al., 2020) and for circadian phase-shifting experiments (Lee et al., 2021).
Our study highlights the utility of compiling and standardizing the vast amount of publicly available circadian data (Pizarro et al., 2013; Ceglia et al., 2018), as also recently shown by a meta-analysis of circadian gene expression in mouse liver (Brooks et al., 2022). Indeed, a secondary finding of our analysis is the predominance of liver as a tissue source among mouse studies (amid the unsurprising predominance of mice overall). Although the liver may have somewhat more rhythmic genes compared with other tissues (Zhang et al., 2014), its current popularity is likely out of proportion to its importance in the mammalian circadian system, even among peripheral tissues. A similar argument applies to human blood, although here the primary driver is likely the relative non-invasiveness of drawing blood from live humans. The greater representation of females among human studies could be related to Institutional Review Board approval and participant recruitment. In any case, recent work on organoids and diurnal rodents may lead to more species and tissue diversity in circadian data sets (Rosselot et al., 2022; Bano-Otalora et al., 2021).
Given the few studies and tissues having sufficient data from males and females, we leave a meta-analysis of sex differences in daily rhythms for future work. Previous work indicates that some differences, such as those related to development and reproduction, may be large, whereas others may be more subtle (Kuljis et al., 2013; Weger et al., 2019; Joye and Evans, 2022). To this point, a sex-based analysis is not possible for studies whose sex inclusion is “mixed” (whether due to pooling or lack of labeling). Rigorous quantification of sex differences or sex-specific rhythms in circadian genomic data will entail not only proper experimental design but also use of statistically valid methods such as those for quantifying differential rhythmicity to avoid misinterpretation (Thaben and Westermark, 2016; Singer and Hughey, 2019; Weger et al., 2021; Pelikan et al., 2021).
Importantly, however, the ethical principle of fair representation—as well as the NIH’s policy on sex as a biological variable—does not require that all studies be powered to detect sex differences. Such a requirement could be challenging for circadian genomic experiments, which already involve a time-course, potentially in multiple conditions, on limited budgets. Instead, the principle and policy require that sex be factored into every step of research, from design to reporting.
In summary, females remain strongly underrepresented in circadian genomic studies. Given the duration of funding cycles and peer review, one might not expect the 2016 NIH policy to have an immediate effect. However, the apparent lack of justification for decisions on sex inclusion and infrequent reporting of results by sex, even in recent studies, points to an opportunity for us as a research community to do better. This opportunity seems especially relevant as the field moves increasingly from bulk to single-cell genomics. Improving sex inclusion could both improve our work’s generalizability and contribute to a growing understanding of the role of biological sex in daily rhythms.
At the reviewers’ request, we cite here all studies in the collection (Menger et al., 2005; Rudic et al., 2005; Oster et al., 2006; Yang et al., 2007; Miller et al., 2007; Bray et al., 2008; Almon et al., 2008a; Almon et al., 2008b; Hoogerwerf et al., 2008; Hughes et al., 2009; Zuber et al., 2009; Na et al., 2009; Keller et al., 2009; Vollmers et al., 2009; Zieker et al., 2010; Sukumaran et al., 2010; Balakrishnan et al., 2010; Bedolla and Torre, 2011; Sukumaran et al., 2011; Barclay et al., 2012; Hughes et al., 2012; Le Martelot et al., 2012; Li et al., 2012; Cho et al., 2012; Negoro et al., 2012; Nikolaeva et al., 2012; Spörl et al., 2012; Geyfman et al., 2012; Tsimakouridze et al., 2012; Koike et al., 2012; Menet et al., 2012; Paschos et al., 2012; Jouffe et al., 2013; Valekunja et al., 2013; Möller-Levet et al., 2013; Sutton et al., 2013; Gossan et al., 2013; Eckel-Mahan et al., 2013; Dyar et al., 2014; Archer et al., 2014; Du et al., 2014; Masri et al., 2014; Young et al., 2014; Haspel et al., 2014; Liu et al., 2014; Arnardottir et al., 2014; Zhang et al., 2014; Fang et al., 2014; Johnson et al., 2014; Hodge et al., 2015; Van Dycke et al., 2015; Pembroke et al., 2015; Renaud et al., 2015; Atger et al., 2015; Janich et al., 2015; Saleh et al., 2015; Zhang et al., 2015; Dudek et al., 2016; Yang et al., 2016; Wang et al., 2016; Erion et al., 2016; Masri et al., 2016; Rakshit et al., 2016; Gerstner et al., 2016; Kolbe et al., 2016; Kettner et al., 2016; Petrenko et al., 2017; Laing et al., 2017; Sobel et al., 2017; Mange et al., 2017; Husse et al., 2017; Fan et al., 2017;Fan et al., 2017; Castelo-Szekely et al., 2017; Wang et al., 2017a; Sato et al., 2017; Solanas et al., 2017; Wang et al., 2017b; Tognini et al., 2017; Quinault et al., 2018; Stubblefield et al., 2018; Yeung et al., 2018; Benegiamo et al., 2018; Mermet et al., 2018; Perrin et al., 2018; Kervezee et al., 2018; Hirako et al., 2018; Wittenbrink et al., 2018; Braun et al., 2018; Wu et al., 2018; Kinouchi et al., 2018; Brunet et al., 2019; Kervezee et al., 2019; Chaix et al., 2019; Weger et al., 2019; Christou et al., 2019; Resuehr et al., 2019; Fader et al., 2019; Cedernaes et al., 2019; Aras et al., 2019; Koronowski et al., 2019; Welz et al., 2019; Sato et al., 2019; Cederroth et al., 2019; Kuang et al., 2019; Nobis et al., 2019; Quagliarini et al., 2019; Pei et al., 2019; Seillet et al., 2020; Greenberg et al., 2020; Rijo-Ferreira et al., 2020; Dan et al., 2020; Levine et al., 2020; Wu et al., 2020; Brami-Cherrier et al., 2020; Guan et al., 2020; Tognini et al., 2020; Greco et al., 2020; Petrenko et al., 2020; Meng et al., 2020; Alzate-Correa et al., 2021; Lu et al., 2021; Weger et al., 2021; Zhang et al., 2021; Xin et al., 2021; Manella et al., 2021; Rubio-Ponce et al., 2021; Furlan-Magaril et al., 2021; Neumann et al., 2021; Milićević et al., 2021; Schwartz et al., 2021; Mishra et al., 2021; Præstholm et al., 2021; Aviram et al., 2021; Brown et al., 2021; Wei et al., 2022; Blacher et al., 2022; Maier et al., 2022; Trott et al., 2022; Jouffe et al., 2022; Greiner et al., 2022; Cresto et al., 2022; DeVera et al., 2022; Egstrand et al., 2022; Acosta-Rodríguez et al., 2022; Tsujihana et al., 2022; Mekbib et al., 2022; Petrus et al., 2022).
Supplemental Material
sj-csv-1-jbr-10.1177_07487304221134160 – Supplemental material for Sex Inclusion in Transcriptome Studies of Daily Rhythms
Supplemental material, sj-csv-1-jbr-10.1177_07487304221134160 for Sex Inclusion in Transcriptome Studies of Daily Rhythms by Dora Obodo, Elliot H. Outland and Jacob J. Hughey in Journal of Biological Rhythms
Supplemental Material
sj-csv-10-jbr-10.1177_07487304221134160 – Supplemental material for Sex Inclusion in Transcriptome Studies of Daily Rhythms
Supplemental material, sj-csv-10-jbr-10.1177_07487304221134160 for Sex Inclusion in Transcriptome Studies of Daily Rhythms by Dora Obodo, Elliot H. Outland and Jacob J. Hughey in Journal of Biological Rhythms
Supplemental Material
sj-csv-2-jbr-10.1177_07487304221134160 – Supplemental material for Sex Inclusion in Transcriptome Studies of Daily Rhythms
Supplemental material, sj-csv-2-jbr-10.1177_07487304221134160 for Sex Inclusion in Transcriptome Studies of Daily Rhythms by Dora Obodo, Elliot H. Outland and Jacob J. Hughey in Journal of Biological Rhythms
Supplemental Material
sj-csv-3-jbr-10.1177_07487304221134160 – Supplemental material for Sex Inclusion in Transcriptome Studies of Daily Rhythms
Supplemental material, sj-csv-3-jbr-10.1177_07487304221134160 for Sex Inclusion in Transcriptome Studies of Daily Rhythms by Dora Obodo, Elliot H. Outland and Jacob J. Hughey in Journal of Biological Rhythms
Supplemental Material
sj-csv-4-jbr-10.1177_07487304221134160 – Supplemental material for Sex Inclusion in Transcriptome Studies of Daily Rhythms
Supplemental material, sj-csv-4-jbr-10.1177_07487304221134160 for Sex Inclusion in Transcriptome Studies of Daily Rhythms by Dora Obodo, Elliot H. Outland and Jacob J. Hughey in Journal of Biological Rhythms
Supplemental Material
sj-csv-5-jbr-10.1177_07487304221134160 – Supplemental material for Sex Inclusion in Transcriptome Studies of Daily Rhythms
Supplemental material, sj-csv-5-jbr-10.1177_07487304221134160 for Sex Inclusion in Transcriptome Studies of Daily Rhythms by Dora Obodo, Elliot H. Outland and Jacob J. Hughey in Journal of Biological Rhythms
Supplemental Material
sj-csv-6-jbr-10.1177_07487304221134160 – Supplemental material for Sex Inclusion in Transcriptome Studies of Daily Rhythms
Supplemental material, sj-csv-6-jbr-10.1177_07487304221134160 for Sex Inclusion in Transcriptome Studies of Daily Rhythms by Dora Obodo, Elliot H. Outland and Jacob J. Hughey in Journal of Biological Rhythms
Supplemental Material
sj-csv-7-jbr-10.1177_07487304221134160 – Supplemental material for Sex Inclusion in Transcriptome Studies of Daily Rhythms
Supplemental material, sj-csv-7-jbr-10.1177_07487304221134160 for Sex Inclusion in Transcriptome Studies of Daily Rhythms by Dora Obodo, Elliot H. Outland and Jacob J. Hughey in Journal of Biological Rhythms
Supplemental Material
sj-csv-8-jbr-10.1177_07487304221134160 – Supplemental material for Sex Inclusion in Transcriptome Studies of Daily Rhythms
Supplemental material, sj-csv-8-jbr-10.1177_07487304221134160 for Sex Inclusion in Transcriptome Studies of Daily Rhythms by Dora Obodo, Elliot H. Outland and Jacob J. Hughey in Journal of Biological Rhythms
Supplemental Material
sj-csv-9-jbr-10.1177_07487304221134160 – Supplemental material for Sex Inclusion in Transcriptome Studies of Daily Rhythms
Supplemental material, sj-csv-9-jbr-10.1177_07487304221134160 for Sex Inclusion in Transcriptome Studies of Daily Rhythms by Dora Obodo, Elliot H. Outland and Jacob J. Hughey in Journal of Biological Rhythms
Footnotes
Acknowledgements
This work was supported by the US National Institutes of Health (R35GM124685 to JJH and F31GM143909 to DO).
Conflict of Interest Statement
The author(s) have no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Notes
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
