Abstract
Circadian rhythms are daily oscillations in physiology and behavior that can be assessed by recording body temperature, locomotor activity, or bioluminescent reporters, among other measures. These different types of data can vary greatly in waveform, noise characteristics, typical sampling rate, and length of recording. We developed 2 Shiny apps for exploration of these data, enabling visualization and analysis of circadian parameters such as period and phase. Methods include the discrete wavelet transform, sine fitting, the Lomb-Scargle periodogram, autocorrelation, and maximum entropy spectral analysis, giving a sense of how well each method works on each type of data. The apps also provide educational overviews and guidance for these methods, supporting the training of those new to this type of analysis. CIRCADA-E (Circadian App for Data Analysis–Experimental Time Series) allows users to explore a large curated experimental data set with mouse body temperature, locomotor activity, and PER2::LUC rhythms recorded from multiple tissues. CIRCADA-S (Circadian App for Data Analysis–Synthetic Time Series) generates and analyzes time series with user-specified parameters, thereby demonstrating how the accuracy of period and phase estimation depends on the type and level of noise, sampling rate, length of recording, and method. We demonstrate the potential uses of the apps through 2 in silico case studies.
Keywords
Circadian rhythms are daily oscillations in physiology and behavior that continue in the absence of time cues and that have been widely observed in bacteria, archaea, fungi, plants, and animals (Hurley et al., 2016). In mice, for example, these rhythms can be observed in body temperature, locomotor activity, bioluminescent reporters of gene expression, and other measures (Leise et al., 2018). These different types of data vary greatly in waveform, noise characteristics, sampling rate, and length of recording. For instance, activity recordings often have high sampling rates, long durations, and square-like waveforms, while bioluminescence recordings tend to be sinusoid-like and only a few days in length. We developed 2 Shiny apps for exploration of these circadian data, enabling visualization and analysis of circadian parameters such as period and phase. Shiny is an R package supporting the creation of open-source interactive web apps (https://shiny.rstudio.com/). The motivation behind our apps is 2-fold: first, to create accessible tools for exploration and analysis of multiple forms of circadian data, and second, to develop an education resource for circadian data analysis. The original apps were developed by the 3 undergraduate authors L.C., L.K., and C.L. at Amherst College with extensive feedback and testing provided by undergraduates at Smith College, with the app development itself providing multiple valuable educational opportunities, including learning how to code in R and Shiny, how to communicate across the quantitative and life sciences, and how to create effective visualizations of data.
Five widely used estimation methods are included in the apps as a representative set of distinct analysis tools: the discrete wavelet transform (DWT), sine fitting, the Lomb-Scargle periodogram, autocorrelation, and maximum entropy spectral analysis (see Table 1 and the Methods section). The focus is on methods applicable to recordings of body temperature, locomotor activity, and bioluminescent reporters, such as PER2::LUC, with at least 3 days of data sampled at least once per hour. The analyses considered here are not aimed at, for instance, detection of circadian rhythms in transcriptome or proteome data, which tend to have short duration and sparse sampling and for which appropriate tools are freely available, including JTK_Cycle (Hughes et al., 2010), RAIN (Thaben and Westermark, 2014), MetaCycle (Wu et al., 2016), and LimoRhyde (Singer and Hughey, 2019), among others. Our educational apps complement existing circadian data analysis apps, such as Biodare (Zielinski et al., 2014), CATkit (Gierke and Cornelissen, 2016), Rethomics (Geissmann et al., 2019), and RhythmicAlly (Abhilash and Sheeba, 2019), by training users to be more knowledgeable in how best to use and interpret different types of analyses for a variety of data types.
Summary of methods available in CIRCADA apps.
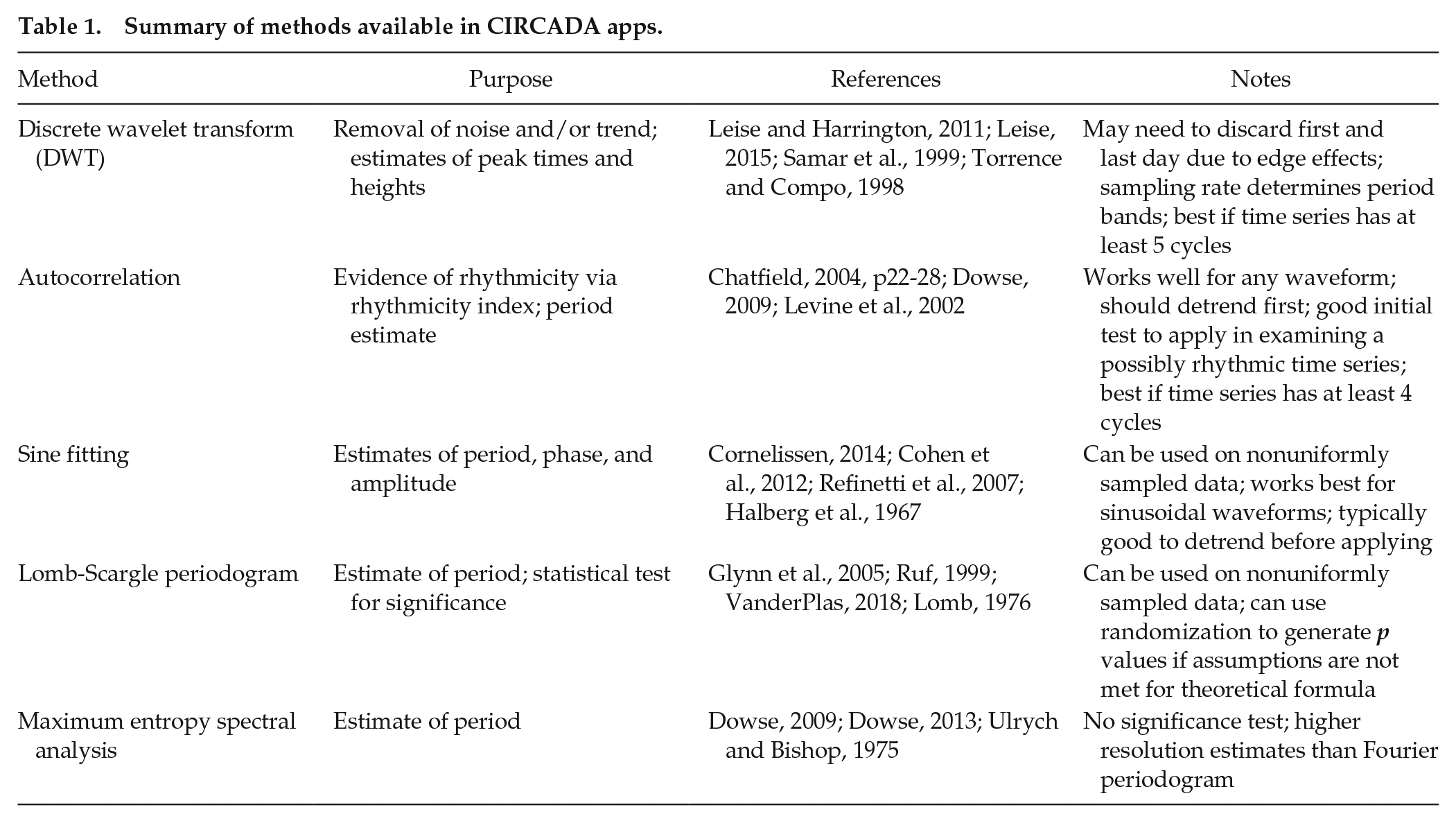
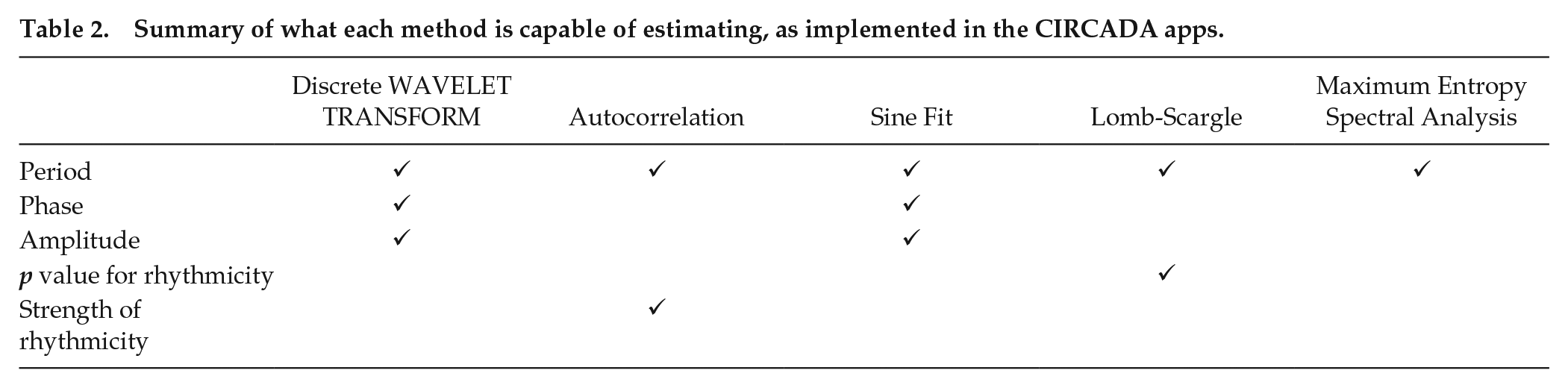
The 5 methods use quite different approaches to estimating rhythmic characteristics of time series and produce different subsets of the desired information about the time series, as shown in Table 2. All 5 methods generate estimates of the period (cycle length), but not all estimate the phase (reported as the peak time) and the amplitude (defined as half the peak-to-trough distance). The Lomb-Scargle method provides a
Summary of what each method is capable of estimating, as implemented in the CIRCADA apps.

The first app, CIRCADA-E (Circadian App for Data Analysis–Experimental Time Series; see Supplemental Fig. 1), allows users to explore a large set of data from Leise et al. (2018) with mouse body temperature, locomotor activity, and PER2::LUC recordings from multiple tissues (publicly accessible at https://osf.io/seyhp/). Samples from old and young mice under 24-h and 20-h T-cycles on normal and high-fat diets are available in these data. In our experience, finding large, interesting, public data sets in a usable form for undergraduate courses can be challenging. By directly interfacing a rich data set with multiple methods of analysis and visualization in a user-friendly app, CIRCADA-E supports a variety of training and educational activities. For instance, students learning to work in a circadian lab can practice generating and testing hypotheses about circadian rhythm measures using this curated set of data with built-in analysis. Possible student projects include comparing distributions of body temperature periods for animals under the 24-h versus 20-h T-cycles, comparing period and peak time estimates for body temperature versus locomotor activity rhythms, or practicing analysis of results from 2 × 2 or 2 × 2 × 2 factorial design experiments by testing for differences in the rhythmicity index of the SCN in vitro rhythms or in the amplitude of the body temperature rhythms. The app also facilitates direct comparison of the results of different methods to assess which might work best for a particular type of data and how accurate and reliable those results may be. Visualization is a major feature of the apps, with graphs of raw data, processed data, and outputs of each method, encouraging regular visual checking of whether results appear consistent with the data.
The second app, CIRCADA-S (Circadian App for Data Analysis–Synthetic Time Series; see Supplemental Fig. 2), generates and analyzes time series with user-specified parameters, listed in Table 3. CIRCADA-S can be used to explore how accuracy of period and phase estimation depends on sampling rate, length of recording, type and level of noise, and method used. This app also offers educational overviews and guidance for each method to provide further training support. As an example, Supplemental Figure 2 displays the CIRCADA-S Lomb-Scargle tab, which demonstrates how increasing the oversampling factor increases resolution of the period estimate and provides a link to a popup window with explanatory details about the method. Results from the 5 methods for a set of generated time series samples are summarized in the CIRCADA-S Summary tab, shown in Figure 1, including histograms of the period errors to facilitate comparison. For example, the DWT, autocorrelation, and Lomb-Scargle methods produce highly discretized period estimates, with the discrete values depending on the sampling interval length for DWT and autocorrelation and more heavily on the number of cycles for the Lomb-Scargle. In contrast, sine fitting and maximum entropy spectral analysis (MESA) generate more fine-grained estimates with resolution independent of sampling interval and number of cycles, with sine-fitting estimates more tightly clustered compared with MESA. Note that higher resolution does not necessarily correlate with higher accuracy of period estimation. As another example, Supplemental Figure 3 shows part of the CIRCADA-S DWT tab with an interactive demonstration of the DWT time series decomposition (which is also available in CIRCADA-E). There is also an Advice tab with suggested guidelines for using each method and brief glossary of the user-set parameter options.
User choices for generating synthetic data in CIRCADA-S.
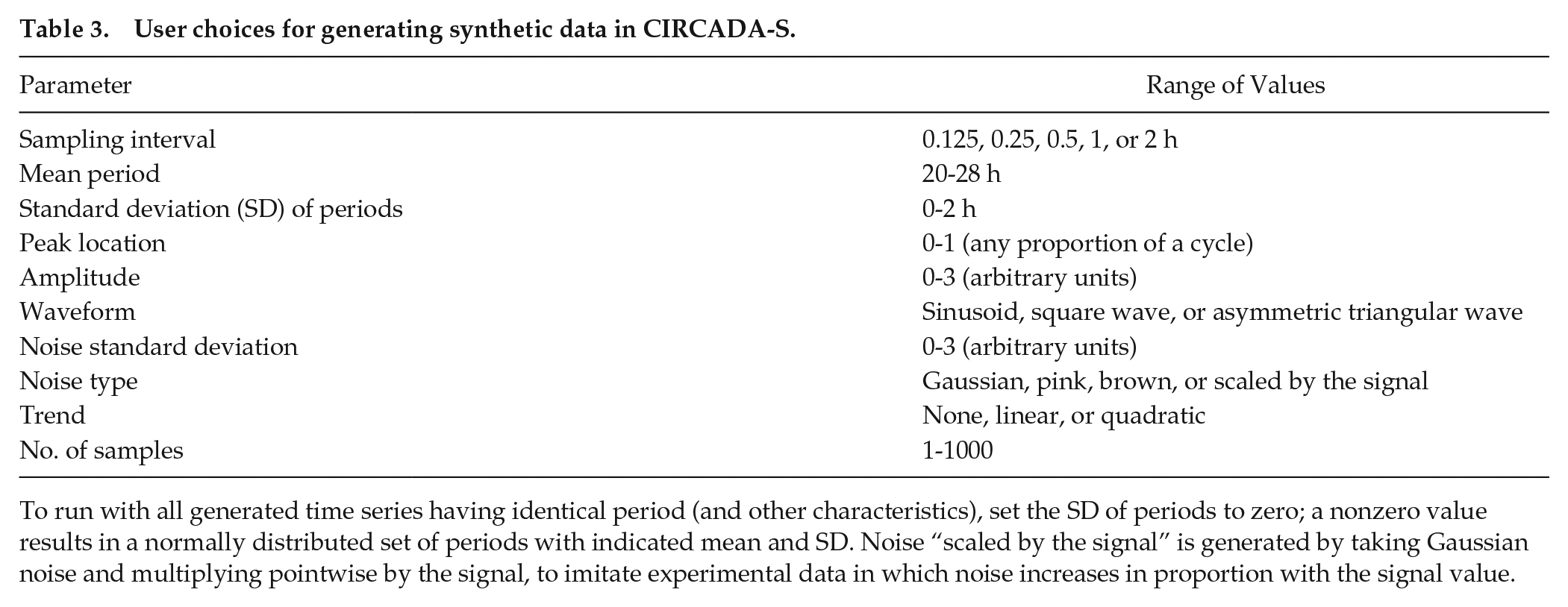
To run with all generated time series having identical period (and other characteristics), set the SD of periods to zero; a nonzero value results in a normally distributed set of periods with indicated mean and SD. Noise “scaled by the signal” is generated by taking Gaussian noise and multiplying pointwise by the signal, to imitate experimental data in which noise increases in proportion with the signal value.
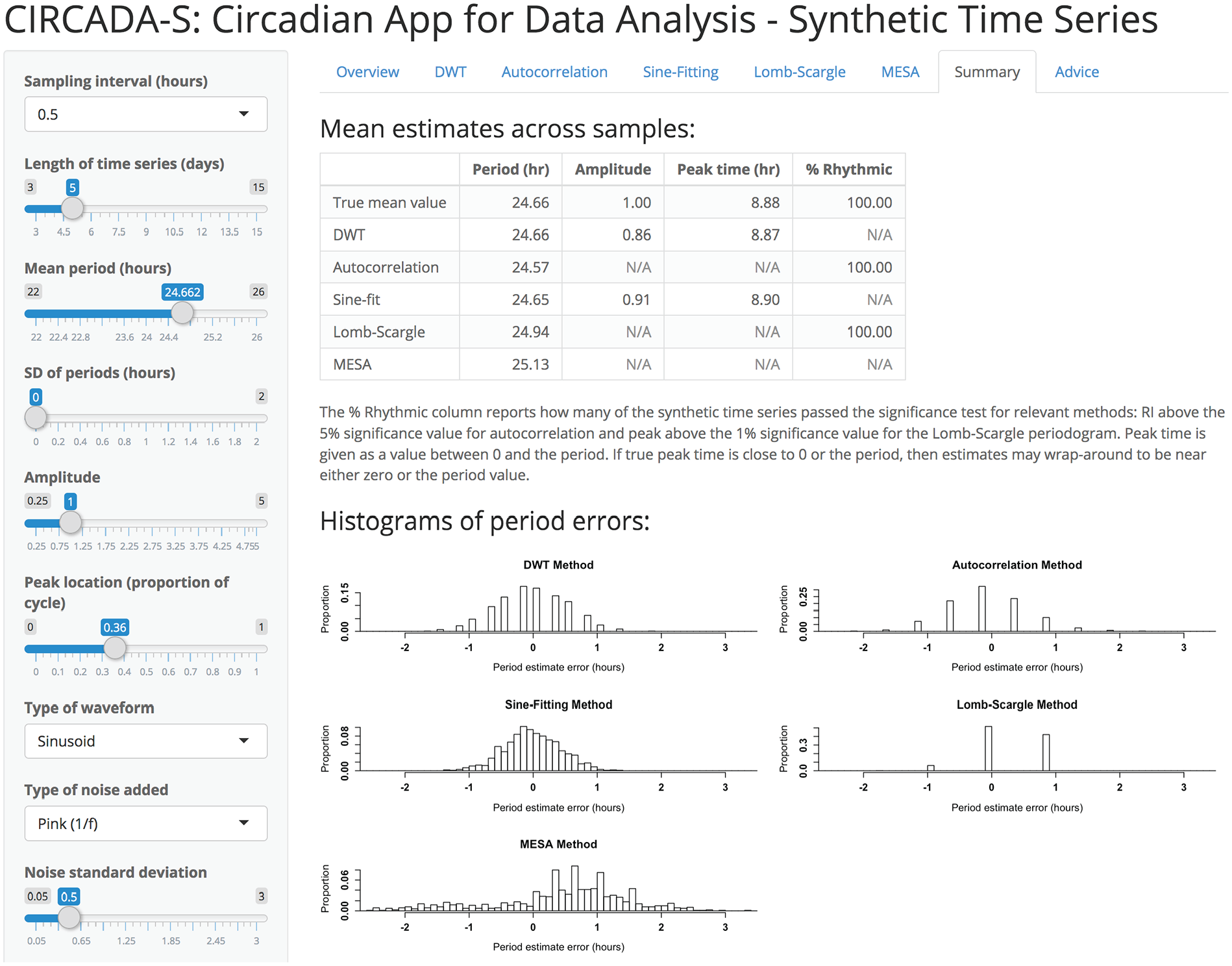

Partial screenshot of CIRCADA-S Summary tab, comparing results from the five methods for a set of 1,000 generated time series with the user-selected characteristics. The histograms of period errors demonstrate the differences between the methods. The DWT, autocorrelation, and Lomb-Scargle produce discrete estimates with possible values determined by the sampling interval and number of cycles, while sine-fitting and MESA produce finer-grained estimates.
Use in Varied Educational Settings
To better understand how these apps might be used, we applied them in a variety of educational settings. In a very introductory class, CIRCADA-S was used within a classroom demonstration to help students understand the Lomb-Scargle periodogram. In a more advanced course, CIRCADA-E was used by a student to address a research question about the interaction of light cycles on body temperature cycles. Within an undergraduate chronobiology research lab, all students completed an online exercise with brief quizzes to guide their exploration of CIRCADA-E. In a week-long chronobiology course for graduate students, CIRCADA-E was used to help students create simulated data with characteristics of their own data set, and results from the simulations were discussed in terms of how to select optimal data analysis tools. Students in all settings reported that this was a helpful tool and increased their understanding of methods to analyze rhythmic data. A sample online lesson, “Learning about the Analysis of Rhythmic Data,” which guides new users through the apps and methods, is posted on the Open Science Framework (OSF) site (https://osf.io/er6wj/), with checkpoint questions at regular intervals to support self-testing for comprehension.
In Silico Case Study: Sampling Rate Versus Duration
To demonstrate the effect of sampling rate versus duration of the time series on period estimate accuracy, we generated 1000 noisy sinusoids with a sampling interval of 0.125 h for 16 days. The time series were subsampled to generate larger sampling intervals, and days were dropped from the end to generate shorter durations. Results for the sine-fitting and Lomb-Scargle methods are shown in Figure 2 as examples. Each additional day in the time series tends to have a greater effect in increasing accuracy than does shortening the time step. For example, the period error in the sine-fit estimates is reduced by nearly 50% by going from 3 days to 5 days (requiring 36 versus 60 time points for 2-h sampling), while increasing the sampling from once every 2 h to 8 times an hour for 3 days (36 versus 576 time points) reduces the error by only about 16%.
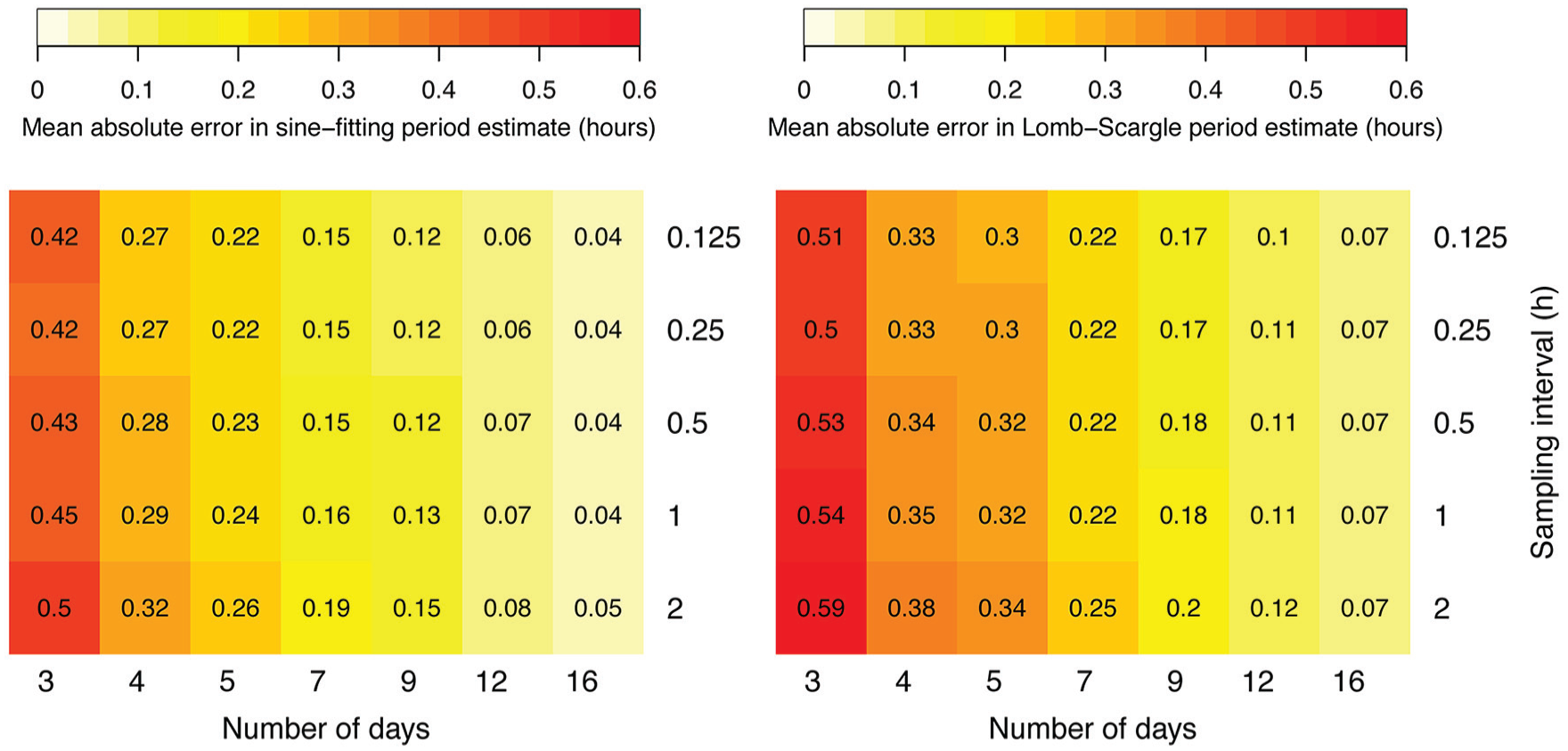
Errors in estimating period using sine-fitting (left) and Lomb-Scargle (right) methods for 1000 noisy sinusoids with normally distributed periods (mean 24 h, SD 0.5 h), amplitude 1, noise standard deviation 0.5 with pink noise, and quadratic trend, for indicated number of days and sampling interval.
In Silico Case Study: Statistical Power
CIRCADA-S can also be used to study how many samples might be necessary to achieve sufficient statistical power in an experiment, given a fixed duration and sampling interval. Sets of synthetic time series with characteristics selected to be similar to those of the experimental data can be generated to estimate the likely statistical power for different numbers of samples, for example, to detect a difference in period between 2 groups. See Figure 3 for an example showing that the addition of a cycle to the data can greatly strengthen the power and reduce the number of samples needed. Results of the app’s analysis are automatically downloaded to a file named CIRCADA-S-output.csv to facilitate this type of further exploration.
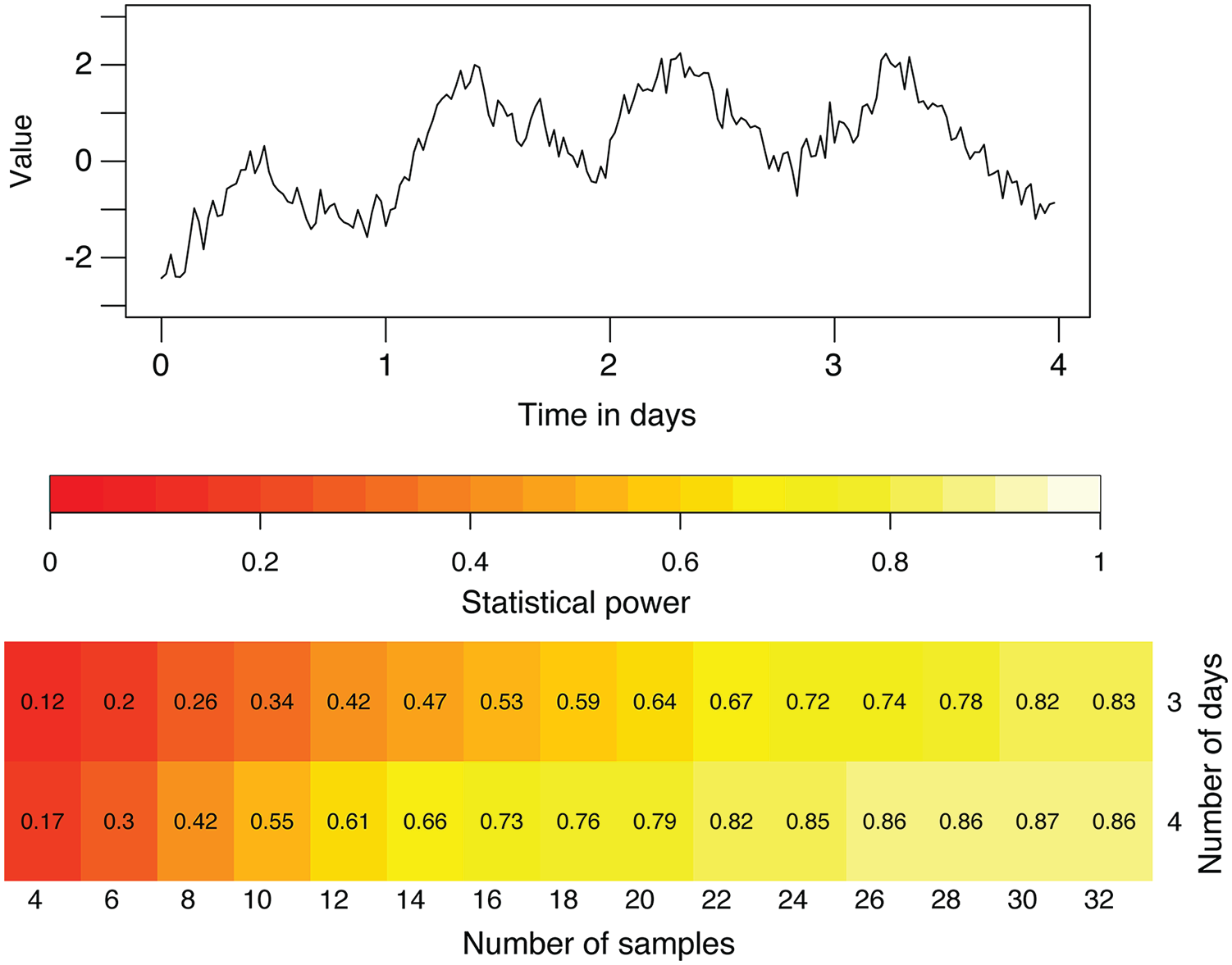
Power study using generated time series. Top: example of a 4-day noisy sinusoid. Bottom: Probability of detecting a significant different in sine-fit period between 2 groups for the indicated number of samples and length of time series (using t.test in R), where the true means are 23.35 and 23.85 h. Periods within each group are normally distributed, with a standard deviation of 0.25 h, amplitude of 1, noise is pink, with standard deviation is 0.5, trend is quadratic, and a sampling interval is 0.5 h. About 30 samples would be needed to obtain a statistical power of 0.80 using a 3-day duration, while increasing the duration to 4 days substantially decreases the required sample size to just greater than 20.
Running CIRCADA
To run the apps, download and save the compressed CIRCADA folder from https://osf.io/er6wj/ to your computer, then open CIRCADA-E.R or CIRCADA-S.R in RStudio and click “Run app.” All required packages will be installed as needed by running the app. A tutorial (GettingStarted.pdf) is available at https://osf.io/er6wj/ for learning the features of the apps and includes suggestions for training activities and projects. An online lesson is also provided on the OSF site.
Methods
The Shiny code for both apps is freely available at https://osf.io/er6wj/, with easily adaptable R code provided for each of the methods. Figures were generated using R 3.5.3 (https://www.r-project.org/) in RStudio 1.2.1335 (https://www.rstudio.com/). Experimental data used in CIRCADA-E are from Leise et al. (2018), using methods as described in that article where they overlap with those used in the app. The data are accessed from the OSF site https://osf.io/seyhp/ using the
Discrete Wavelet Transform
Time series are detrended using the DWT before applying other methods (with the exception that linear detrending is used if the time series covers 4 days or less). The apps use the function wavMRDSum from the
Autocorrelation
The R command acf from the
Sine Fitting
The R command nls (nonlinear least squares) from the
Lomb-Scargle Periodogram
The R command lsp is used to calculate the Lomb-Scargle periodogram, from the lomb R package (https://cran.r-project.org/web/packages/lomb). The highest peak corresponding in the period range of 16 to 34 h is chosen as the period estimate. Higher period resolution may be obtained by increasing the oversampling factor, as shown in the periodogram in Supplemental Figure 2; oversampling factor 8 is used for the period estimate. In the CIRCADA-E app, the significance of peaks is assessed using a randomization procedure via randlsp, which produces a more valid result when the theoretical assumptions may not hold, such as normally distributed uncorrelated noise. However, the randomization procedure is computationally intensive, so the CIRCADA-S app runs it for only a representative time series and uses the theoretical
MESA
The R command spec.ar from the
Generation of Random Time Series
Each sample time series in CIRCADA-S is the sum of a periodic function plus random noise plus trend. The periodic function has period
Packages Used by the CIRCADA Apps
Packages used include
Supplemental Material
SupplementalMaterial-JBR-19-0114.R1 – Supplemental material for CIRCADA: Shiny Apps for Exploration of Experimental and Synthetic Circadian Time Series with an Educational Emphasis
Supplemental material, SupplementalMaterial-JBR-19-0114.R1 for CIRCADA: Shiny Apps for Exploration of Experimental and Synthetic Circadian Time Series with an Educational Emphasis by Lisa Cenek, Liubou Klindziuk, Cindy Lopez, Eleanor McCartney, Blanca Martin Burgos, Selma Tir, Mary E. Harrington and Tanya L. Leise in Journal of Biological Rhythms
Footnotes
Acknowledgements
We thank Dominica Cao, Yilin Cui, Hanxiao Lu, and Hannah Wang, for feedback and beta testing of the apps. The undergraduate research of L.C., L.K., and C.L. was supported by the Clare Boothe Luce Foundation and the Amherst College Gregory S. Call Undergraduate Research Fund. This work was supported by National Institutes of Health grants GM126545 and P01AG009975.
Conflict of Interest Statement
The authors have no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Notes
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
