Abstract
The time-dependent degradation of core circadian clock proteins is essential for the proper functioning of circadian timekeeping mechanisms that drive daily rhythms in gene expression and, ultimately, an organism’s physiology. The ubiquitin proteasome system plays a critical role in regulating the stability of most proteins, including the core clock components. Our laboratory developed a cell-based functional screen to identify ubiquitin ligases that degrade any protein of interest and have started screening for those ligases that degrade circadian clock proteins. This screen identified Spsb4 as a putative novel E3 ligase for RevErbα. In this article, we further investigate the role of Spsb4 and its paralogs in RevErbα stability and circadian rhythmicity. Our results indicate that the paralogs Spsb1 and Spsb4, but not Spsb2 and Spsb3, can interact with and facilitate RevErbα ubiquitination and degradation and regulate circadian clock periodicity.
The circadian system controls physiological rhythms that let organisms anticipate the daily cyclic environmental changes associated with the time of day. The timekeeping mechanism that drives these rhythms, the mammalian circadian clock, consists of 2 interlocked transcription/translation feedback loops that function to produce robust 24-h rhythms of gene expression (Partch et al., 2014). The primary loop involves the genes Bmal1 and Clock (or its ortholog Npas2), three Period genes (Per1, Per2, Per3), and two Crypto-chrome genes (Cry1 and Cry2). CLOCK and BMAL1 are basic helix-loop-helix PAS-domain–containing transcription factors that form a complex that activates the transcription of the Per1-3, Cry1/2 genes as well as many circadian output genes predominantly during the day. PER and CRY in turn heterodimerize and translocate to the nucleus to inhibit their own transcription by interacting with and repressing the activity of the CLOCK-BMAL complex, completing a negative feedback loop (Reppert and Weaver, 2002; Partch et al., 2014). During the end of the night, the PER-CRY complex is degraded through ubiquitin-dependent pathways (Busino et al., 2007; Siepka et al., 2007), repression of CLOCK-BMAL is relieved, and the cycle begins again with 24-h periodicity (Partch et al., 2014). This timekeeping mechanism also drives rhythmic expression of RevErbα/β (Nr1d1/2), transcriptional repressors that regulate the circadian clock by driving the rhythmic expression of Bmal1, Clock/Npas2, and other genes expressed predominantly during the night (Preitner et al., 2002; Guillaumond et al., 2005; Crumbley et al., 2010; Crumbley and Burris, 2011). Both of these loops appear to be required for the proper functioning of the circadian clock (Reppert and Weaver, 2002; Bugge et al., 2012; Cho et al., 2012; Partch et al., 2014). Overall, proper timekeeping of this circadian clock system depends on the regulated expression and degradation of all these clock components (Stojkovic et al., 2014).
The ubiquitin-proteasome system (UPS) is the principal mechanism for the degradation of most proteins involved in various cellular processes (Castro et al., 2005; Vucic et al., 2011; Hammond-Martel et al., 2012). The direct role of ubiquitination in determining protein half-life is crucial for proteins with a daily rhythm (Stojkovic et al., 2014). Degradation of proteins via UPS involves two successive steps: tagging of the substrate protein by the covalent attachment of multiple ubiquitin molecules (conjugation) and the subsequent degradation of the tagged protein by the proteasome (Hershko and Ciechanover, 1998). The attachment of ubiquitin to the target protein requires a series of adenosine triphosphate–dependent enzymatic steps involving ubiquitin activating (E1), ubiquitin conjugating (E2), and ubiquitin ligating (E3) enzymes. E1 enzymes bind free ubiquitin and transfer it to E2 enzymes. E3 ligases, by interacting with the substrate protein to be degraded, facilitate the transfer of ubiquitin to the substrate using a variety of mechanisms (Hershko and Ciechanover, 1998). Importantly, the interaction between E3 ligases and the substrate protein is key in determining substrate specificity, directing which proteins are to be ubiquitinated at any given moment (Iconomou and Saunders, 2016). While many E3 ubiquitin ligases act alone, many others are found as components of much larger multiprotein complexes including the E2 and scaffolding and regulatory proteins (i.e., SCF complex; Kile et al., 2002). This highly regulated system interacts and regulates many cellular processes, including those of the circadian clock system.
Identifying E3 ligases that ubiquitinate specific substrates can be difficult and has mostly involved some form of protein interaction screen. Our laboratory developed a functional screening approach geared toward identifying E3 ligases capable of destabilizing any specific protein of interest (DeBruyne et al., 2015). Using this functional screen, we identified Seven in absentia2 (Siah2) and Sp1A/ryanodine receptor domain and SOCS box-containing4 (Spsb4) as candidate E3 ligases involved in the regulation of RevErbα stability (DeBruyne et al., 2015). While we have validated the role of Siah2, we had not rigorously explored the function of Spsb4 in regulating RevErbα stability. Here, we focus on exploring the role of Spsb4 and the entire SPSB family of E3 ligases in regulating RevErbα stability and overall circadian clock function.
Materials and Methods
Cell Culture and Transfection
Cells were maintained in Dulbecco’s modified Eagle’s medium supplemented with 10% fetal bovine serum (Invitrogen, Carlsbad, CA), 1x nonessential amino acid (Invitrogen), and penicillin/streptomycin/glutamine mix and cultured at 37 °C in 5% CO2 (DeBruyne et al., 2015). HA-tagged Spsb2 (Cat. No. HG14695-CY) and Spsb3 (Cat. No. HG16867-CY) plasmids were obtained from Sino Biological (Beijing, China). Plasmid transfection of AD293 cells was performed with FugeneHD (Promega, Madison, WI) according to the manufacturer’s protocol (Bugge et al., 2012). Lipofectamine 2000 (Thermo Fisher, Waltham, MA) was used for small interfering (siRNA) transfections as previously described (Baggs et al., 2009). A negative control siRNA (All-Stars Negative Control siRNA; Qiagen, Hilden, Germany) was used to ensure molar equivalence of siRNAs across all conditions. An equal mixture of 2 Qiagen Spsb1 siRNAs (Hs_Spsb1_1 and Hs_Spsb1_2), 2 Spsb2 siRNAs (Hs_Spsb2_1 and Hs_Spsb2_2), 2 Spsb3 siRNAs (Hs_Spsb3_1 and Hs_Spsb3_2), as well as 2 Spsb4 siRNAs (Hs_Spsb4_1 and Hs_Spsb4_2) was used, constituting a total of 10 pmol per 35-mm dish unless otherwise indicated. Cells were incubated for ~48 h after transfection before the start of the experiments.
Western Blot Analysis
Cells were lysed and processed for western blotting as described previously (DeBruyne et al., 2015). Membranes were incubated with the following antibodies: anti-Flag (Cell Signaling, Danvers, MA; Cat. No. 14793S), anti-RevErbα (Cell Signaling, Cat. No. 13418S), anti-HA (Cell Signaling, Cat. No. 3724S), anti-ubiquitin (Cell Signaling, Cat. No. 3933S), anti-GAPDH (Santa Cruz Biotechnology, Santa Cruz, CA; Cat. No. sc-25778), anti-β-tubulin (Cell Signaling, Cat. No. 5346S), and anti-rabbit horseradish peroxidase linked secondary antibody (Cell Signaling, Cat. No. 7074S). Band intensities were quantified from ImageQuant LAS 4000 (GE Life Sciences, Chicago, IL) images using ImageJ (National Institutes of Health, Bethesda, MD; Figs. 1A and 2A) or from images captured using an Odyssey Fc and quantified Image Studio (LI-COR instruments; all other blots).
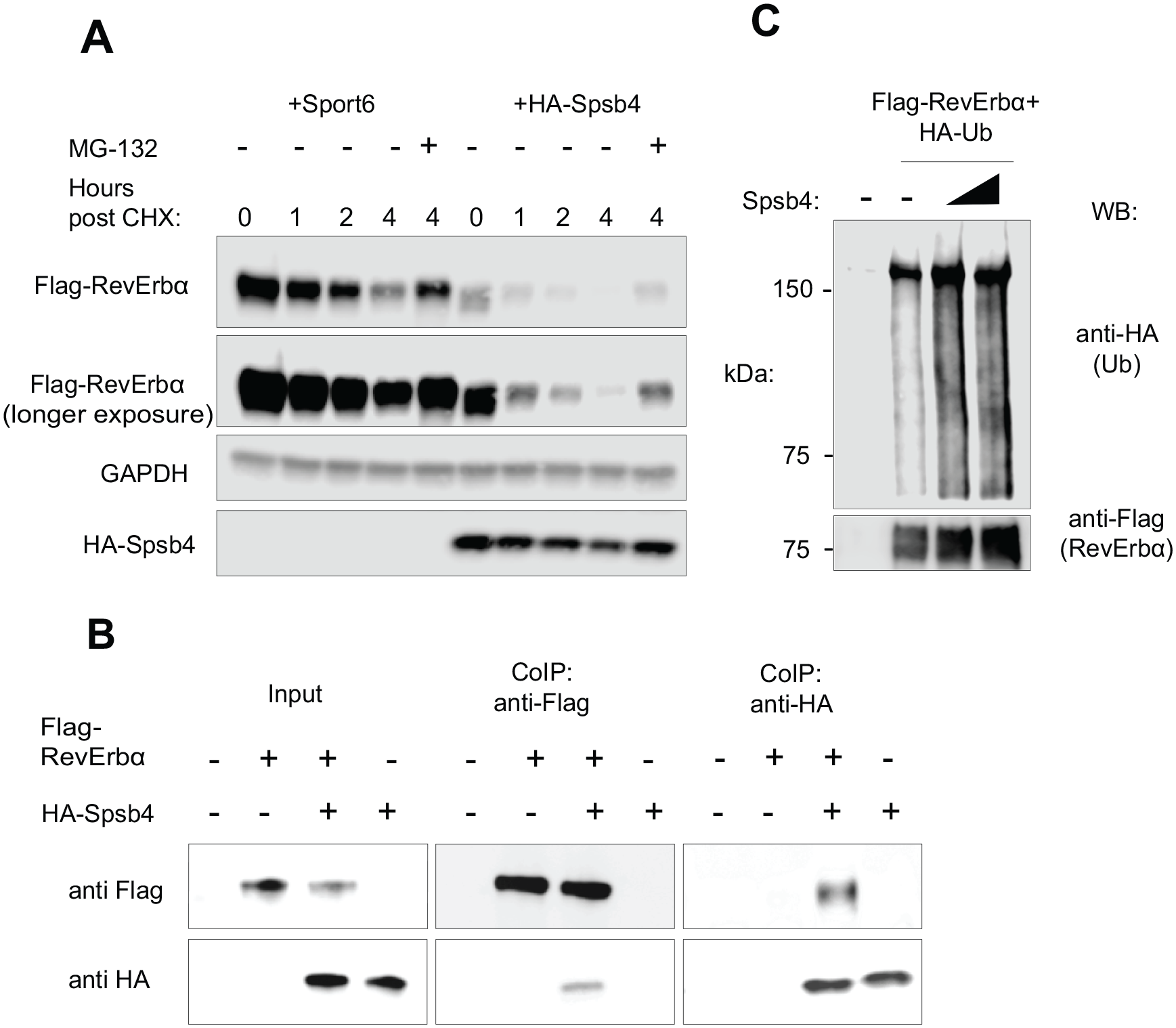
Spsb4 interacts with and ubiquitinates RevErbα. (A) RevErbα was coexpressed with either an empty vector (Sport6) or Spsb4 at equal concentrations in AD293 cells. Cells were then treated with cycloheximide and indicated cells with MG-132 for 4 h before lysis. Total cell lysates were subjected to western blotting analysis with anti-Flag, anti-GAPDH, and anti-HA antibodies. (B) Interactions of RevErbα and Spsb4 by anti-Flag and anti-HA immunoprecipitation of AD293 cells. Cells were transfected with Flag-RevErbα and HA-Spsb4. Forty-eight hours after transfection, MG-132, a proteasome blocker, was added to allow for accumulation of substrate-E3 ligase complexes. (C) Immunoblot analysis of lysates from AD293 cells transfected with plasmids for Flag-RevErbα, HA-ubiquitin, and increasing concentrations of Spsb4 (1 and 2 µg), followed by immunoprecipitation with anti-Flag and analysis via immunoblot with anti-Ub ab.
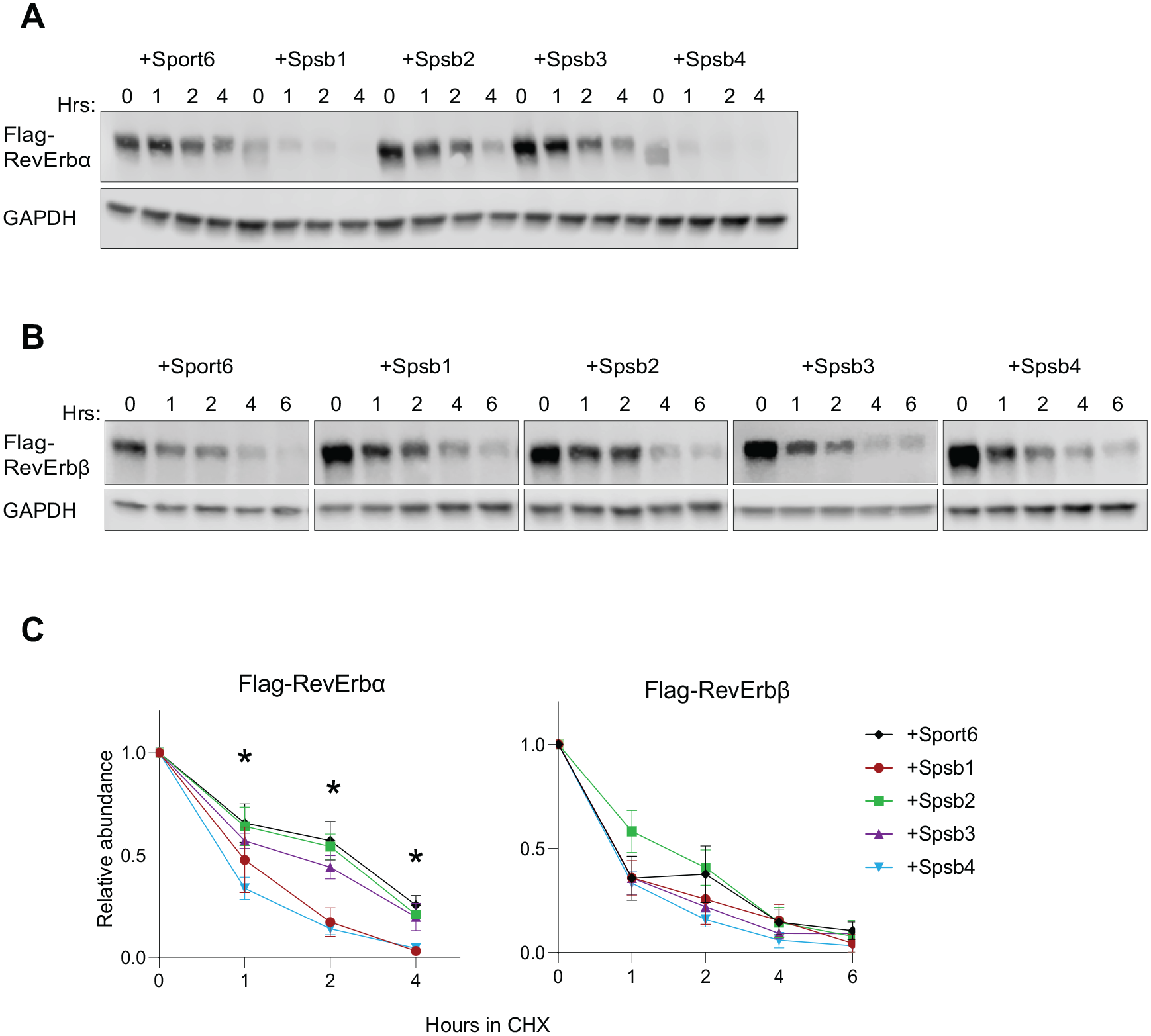
Spsb1 and Spsb4 destabilize RevErbα but not RevErbβ. Representative western blot of AD293 cell lysates illustrating the stability of (A) Flag-RevErbα or (B) Flag-RevErbβ when coexpressed with Sport6 or Spsb1-4 in a cycloheximide (CHX)–chase experiment (hrs = time with CHX). (C) Quantitation of western blot data from experiments of Flag-RevErbα or Flag-RevErbβ normalized to GAPDH and plotted relative to the time 0 point. Data are mean ± standard error of the mean of n = 3 to 7 independent experiments for Flag-RevErbα and n = 3 independent trials for Flag-RevErbβ, except for Spsb3 (n = 2). Two-way analysis of variance (ANOVA) reveals a significant time × E3 ligase interaction on RevErbα stability for Spsb1 and Spsb4 (p < 0.001) but not Spsb2 or Spsb3 (p > 0.5). *Differences at individual time points between Spsb1/Spsb4 and the Sport6 control (p < 0.05) using Sidak’s multiple comparison test. There was no significant effect of Spsb E3 ligases on Flag-RevErbβ stability (p > 0.05 for all 2-way ANOVA outputs).
Immunoprecipitation/Ubiquitin Assay
For immunoprecipitation, AD293 cells in 6-well plates at 60% to 70% confluence were transfected with the indicated plasmids. Approximately 48 h after transfection, MG132 (Cell Signaling), a proteasome blocker, was added to the cells at a final concentration of 5 µM for 4 h. Cells were then washed with phosphate-buffered saline and protein extracts prepared by incubation with a commercially available RIPA buffer (Boston BioProducts, Ashland, MA) at 4 °C. Lysates were incubated either with Flag M2 agarose beads (Sigma, St. Louis, MO) or HA-Tag sepharose beads (Cell Signaling). The beads were washed 4 times and immunoprecipitates eluted from the beads by boiling in protein loading dye at 90 °C. For detecting ubiquitination, 1.5 µg and 3 µg of constructs expressing RevErbα and HA-UB, respectively, were transfected. Two 10-cm dishes were used to obtain 1 lysate. In addition, 2 mM of N-ethylmaleimide was added to the RIPA lysis buffer and wash buffer to block the activity of isopeptidases and deubiquitinating enzymes.
RNA Isolation and Quantitative Polymerase Chain Reaction
U2OS cells were lysed using Trizol (Invitrogen) and stored at −80 °C. RNA was extracted according to the manufacturer’s instructions and resuspended in 30 µL of RNA-free water. Reverse transcription and quantitative polymerase chain reaction using SYBR green reagent, SSO Advance (Bio-Rad, Hercules, CA) were performed as previously described (Baggs et al., 2009; DeBruyne et al., 2015). Primers for Spsb family mRNAs were obtained from Qiagen (Hs_SPSB1_1_SG, Hs_SPSB2_1_SG, Hs_SPSB3_1_SG, and Hs_SPSB4_1_SG) and validated for amplification efficiency using a cDNA dilution series and for specificity in siRNA knockdown experiments (Suppl. Fig. S1). Other primers used have been validated previously (DeBruyne et al., 2015). GAPDH was used as an internal control for normalization. Data were analyzed using the 2−ΔΔCt method, normalizing against the average of all relevant experimental control samples.
Cell Synchronization
Bmal1-Luc U2OS cells were synchronized with dexamethasone prior to bioluminescence recordings. U2OS cells used for the detection of endogenous RevErbα and mRNA expression levels were synchronized with 50% horse serum (Atlanta Biologicals, Flowery Branch, GA; Cat. No. S12150). Both methods were validated to have similar efficiency in synchronizing cells (Suppl. Fig. S2).
Bioluminescence Recordings
Bioluminescence was measured continuously for at least 7 days using a LumiCycle (Actimetrics, Wilmette, IL) from a Bmal1-Luc U2OS cell line (Vollmers et al., 2008). Cell cultures were synchronized with dexamethasone and maintained in LumiCycle media (Yamazaki and Takahashi, 2005; DeBruyne et al., 2015). Data were analyzed using the LumiCycle Analysis software package (Actimetrics) as previously reported (DeBruyne et al., 2015).
Results
Spsb4 Behaves as an E3 Ligase for RevErbα
Our previous studies largely focused on simply validating Spsb4 as a “hit” in our E3 ligase-substrate screen (DeBruyne et al., 2015). In our first experiments, we focused on assessing whether Spsb4 displayed characteristics expected if it was truly an E3 for RevErbα. First, we determined if Spsb4-mediated RevErbα degradation could be blocked by inhibiting the proteasome with MG132 in a robust cell-based degradation assay. Cells were co-transfected with constructs expressing RevErbα and SPSB4 or an empty Sport6 vector. Forty-eight hours after transfection, cells were treated with cycloheximide (CHX) for the hours shown to block new protein synthesis for up to 4 h (Fig. 1A). Simultaneously, the indicated cells were also treated with MG-132 for 4 h. Lysates were then prepared and processed for western blotting. If the degradation of RevErbα was proteasome mediated, we expected to see an increase in RevErbα levels in the cells treated with MG-132 as proteins destined for the proteasome will not be degraded. Indeed, the robust destabilization of RevErbα by Spsb4 in this assay was substantially blocked in MG132-treated cells (Fig. 1A). The rapid degradation of RevErbα mediated by Spsb4 compared with controls, and its block by proteasome inhibitors, confirms our previous results (DeBruyne et al., 2015) and suggests that Spsb4 is directing RevErbα for degradation by the proteasome, one of the hallmark characteristics of an E3 ligase-substrate interaction.
Another essential feature of an E3 ligase is that it can interact with and ubiquitinate its substrates, which subsequently targets them for proteasomal degradation (Hammond-Martel et al., 2012). We therefore asked whether Spsb4 could also detectably interact with and ubiquitinate RevErbα in the same context in which Spsb4 robustly degrades RevErbα. Indeed, we were able to readily and specifically detect Spsb4 within RevErbα immunoprecipitates, as well as RevErbα in Spsb4 immunoprecipitates (Fig. 1B), indicating that these proteins can interact in the same complexes. Furthermore, the presence of Spsb4 greatly enhanced RevErbα ubiquitination in cell-based ubiquitination assays (Fig. 1C). Combined, our data indicate that Spsb4 can interact with RevErbα and cause its ubiquitination and degradation by the proteasome, suggesting that Spsb4 can act as a RevErbα E3 ligase, at least in a cell-based setting.
A key feature of UPS is that there is a high degree of multiplicity, in which a single E3 ligase might have more than 1 protein substrate (Nalepa et al., 2006; Iconomou and Saunders, 2016). Indeed, Spsb4 and other paralogs, Spsb1 and Spsb2, have also been found to regulate inducible nitric oxide synthase (Kuang et al., 2010; Nishiya et al., 2011) and transforming growth factor–β receptor II (Liu et al., 2015). Therefore, we tested to see if Spsb4 showed any specificity to RevErbα among mammalian clock proteins. We used our cell-based degradation assay to test this by transfecting AD293 cells with constructs expressing Flag-tagged Bmal1, Per1, and Cry1 as well as an empty Sport6 (negative control) or Sport6-Spsb4 vectors. We found that Spsb4 destabilized only Flag-RevErbα, whereas it had no effect on the stability of other Flag-tagged core clock proteins (Suppl. Fig. S3). Thus, among core clock proteins, Spsb4 appears to be a selective regulator of RevErbα stability, likely as an E3 ubiquitin ligase.
Spsb1 and Spsb4 Induce Proteasomal Degradation of RevErbα but Not RevErbβ
Mammalian genomes contain 4 paralogous genes expressing four SPRY domain- and SOCS box-containing proteins, SPSB1-4 (also known as SSB1-4). These proteins are characterized by a central SPRY domain and a C-terminal SOCS box, suggesting that SPSB proteins may function as a substrate-binding component of an ElonginC–Cul2–SOCS box E3 ubiquitin ligase complex (Kleiber and Singh, 2009). SPRY domains function as protein-protein interaction modules, and in SPSB proteins, they act as adaptors that bring the SOCS box-associated E3 ubiquitin ligase complex into close proximity with its substrate (Kile et al., 2002; Iconomou and Saunders, 2016).
The evolution of the Spsb gene family in vertebrates likely involved 3 duplication and divergence events resulting in 4 Spsb genes (Kleiber and Singh, 2009). Although the specific family members seem to be highly conserved across species, paralogs within species are relatively dissimilar (Kleiber and Singh, 2009). For instance, Spsb3 shares only 18% amino acid similarity with Spsb1 in vertebrates, and Spsb2 shares 44% sequence similarity with Spsb1 in mice (Kleiber and Singh, 2009). However, mouse Spsb1 and Spsb4 share 75% amino acid similarity (Wang et al., 2005). Across vertebrate species, there is a 92% and 89% similarity among Spsb1 and Spsb4 genes, respectively, possibly highlighting the functional importance for the conservation of their sequences (Kleiber and Singh, 2009). In addition, each of the 4 proteins has maintained their domain structure and sequence (Hilton et al., 1998), suggesting that there might be a functional redundancy between the more similar Spsb genes (Wang et al., 2005).
We therefore asked if other SPSB family members can also target RevErbα for proteasomal degradation. We first examined whether the overexpression of Spsb1-3 accelerated RevErbα degradation in CHX chase assays as previously described (see Fig. 1A; DeBruyne et al., 2015). In the control cells, transfected with an empty Sport6 vector, the overall RevErbα protein abundance was relatively stable over the 4 h of CHX treatment (Fig. 2A, C). In contrast, we observed that RevErbα was readily degraded in the presence of SPSB1 and SPSB4 but not SPSB2 and SPSB3 (Fig. 2A, C). Comparing RevErbα abundance at the initial time point 0 for each condition shows that Spsb1 and Spsb4 appear to degrade RevErbα prior to CHX addition, suggesting that the rate of degradation of RevErbα observed in cells expressing SPSB1 and SPSB4 may be an underestimation. Moreover, we further validated that SPSB2 and SPSB3 could not destabilize RevErbα using independently derived constructs expressing HA-tagged proteins expressed at levels comparable with SPSB4 (Suppl. Fig. S4). We also confirmed that this is not a cell-specific effect; SPSB1 and SPSB4 can also degrade RevErbα in U2OS cells (Suppl. Fig. S5). Overall, these results suggest that SPSB1, but not SPSB2 or SPSB3, is similar to SPSB4 in its ability to facilitate RevErbα degradation, consistent with the sequence similarities and evolutionary relatedness across the Spsb gene family.
We also asked if SPSB1 and SPSB4 (or SPSB2-3) could also target the RevErbα paralog RevErbβ for degradation, using the cell-based assay. Much to our surprise, REV-ERBβ stability was not altered by co-expression of any SPSB1-4 proteins, including SPSB1/SPSB4 (Fig. 2B, C). Although both RevErbα and RevErbβ exhibit rhythmic gene expression and are regulated posttranscriptionally by binding of heme (Raghuram et al., 2007; Yin et al., 2007), these data add to the notion that these paralogous proteins might be regulated differently. For instance, an N-terminal GSK3β site that is present in RevErbα and controls its interaction with E3 ligases and proteasomal degradation (Yin et al., 2006; Yin et al., 2010) is absent in RevErbβ (Bugge et al., 2012). This differential regulation could highlight a mechanism by which the clock is protected from perturbations associated with dysregulation of either RevErbα or β.
Spsb1 and Spsb4 Regulate Endogenous RevErbα Stability and Clock Function
Posttranslational events, such as time-dependent degradation, contribute to the generation of daily oscillations in clock gene products (Lee et al., 2001). Likewise, RevErbα/β protein abundance levels follow robust rhythmicity in most tissues and synchronized cell cultures (Preitner et al., 2002). This cyclic accumulation of RevErbα imposes circadian regulation of Bmal1 transcription and, in turn, governs overall clock function. For instance, continuous overexpression of RevErbα inhibits transcription of the Bmal1 gene, thereby disrupting the clock (Kornmann et al., 2007). Similarly, depletion of RevErbα resulted in significantly shorter period length in animals (Preitner et al., 2002), and genetically removing both RevErbα and RevErbβ eliminates clock function (Bugge et al., 2012; Cho et al., 2012). Finally, a delay imposed by RevErbα’s repression of Cry1 expression not only appears to be required for overall clock function but also plays a direct role in regulating the period of the clock (the longer the repression of Cry1 by RevErbα, the slower the clock function; Ukai-Tadenuma et al., 2011). These studies strongly imply that disrupting the rhythmicity of RevErbα abundance, either by altering its expression or degradation, likely alters the function and periodicity of the circadian clock. Thus, we next sought to determine if the SPSB proteins were essential for normal cycling of RevErbα protein levels and overall function of an endogenous circadian oscillator.
To examine the roles of SPSB proteins in overall clock function, U2OS cells containing the Bmal1-luc circadian reporter (Vollmers et al., 2008; DeBruyne et al., 2015) were transfected with siRNAs corresponding to each Spsb1-4 mRNA and subjected to kinetic luminescence imaging for 7 days following synchronization with dexamethasone (Vollmers et al., 2008; DeBruyne et al., 2015). Overall, these results were very consistent with their effects on RevErbα stability (Fig. 2). We found that knockdown of either Spsb2 or Spsb3 had little effect on period, lengthening it by ~0.7 h in either case (Fig. 3). In contrast, knocking down Spsb1 and Spsb4 each alone significantly lengthened circadian period by 2.2 ± 0.3 and 1.4 ± 0.2 h, respectively (Fig. 3). Thus, the two Spsb family members that robustly degrade RevErbα are also involved in regulating circadian period.
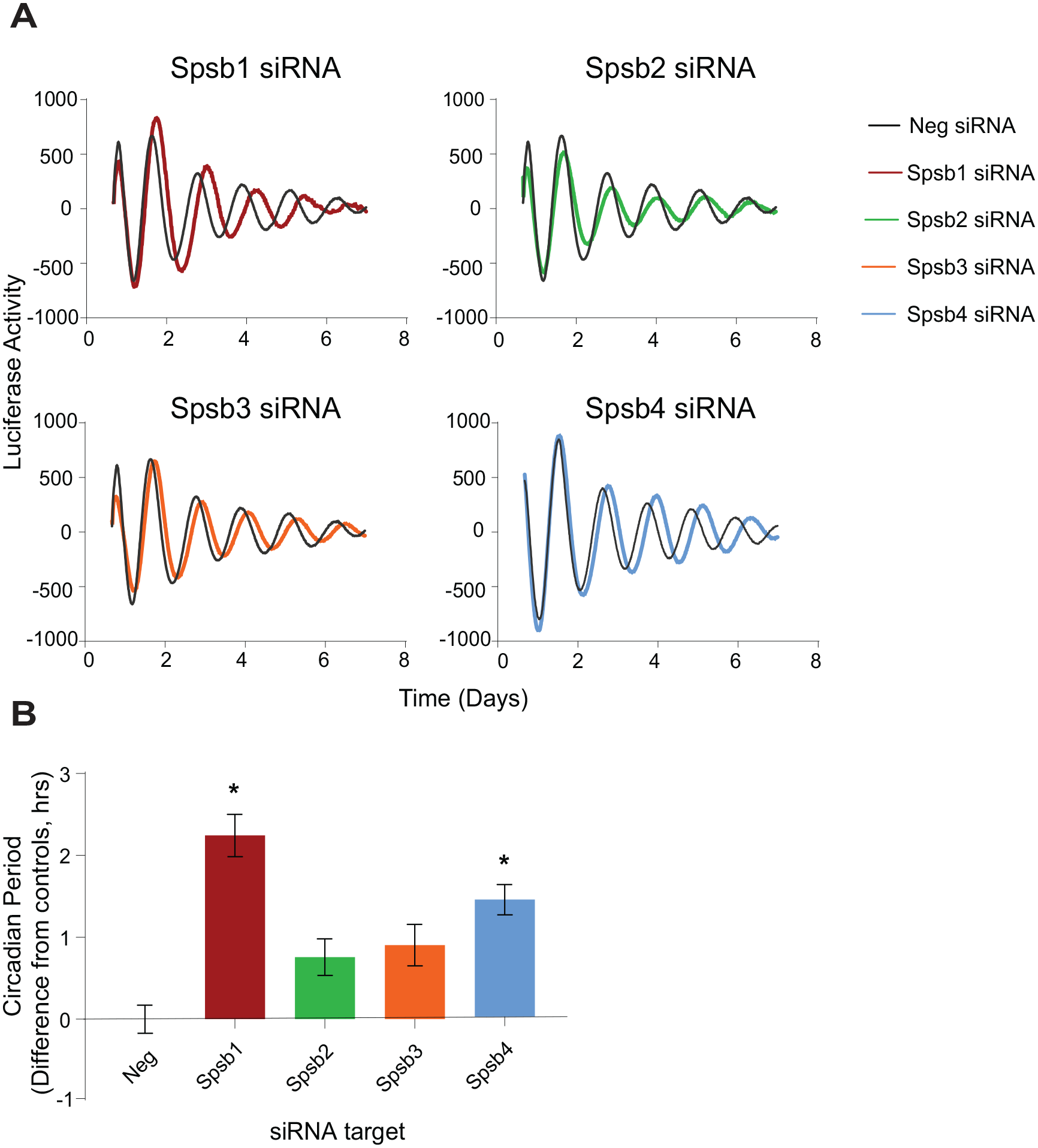
Spsb1 and Spsb4, but not Spsb2/Spsb3, knockdown lengthens circadian period. (A) Average bioluminescence rhythms produced by Bmal1-luc U2OS cells transfected with 10 pmol of the indicated small interfering RNAs (mean, n = 3-4 cultures from a typical experiment). (B) Circadian period data from each experiment were normalized to the average period of the negative controls in each trial (2-4 independent trials for each cDNA) are shown (mean ± SEM, n = 9-12 cultures for each). *p < 0.0001 versus negative controls, analysis of variance, Tukey’s multiple comparisons test.
We next determined if there is possible redundancy among Spsb members. For example, it is possible that we did not see an effect of Spsb2 knockdown because there was sufficient Spsb3 to compensate for its loss. However, knocking down both Spsb2 and Spsb3 together had no additional effect on period, lengthening it by only ~0.6 h compared with controls and similar to effects of knocking down each individually (p > 0.05 compared with control and single knockdown periods; Suppl. Fig. S6). Thus, similar to their relative inability to degrade RevErbα, SPSB2 and SPSB3 appear dispensable for normal clock function.
In contrast, knocking down Spsb1 and Spsb4 together in the same cells produced an approximately additive effect on period (Fig. 4). In these experiments (using 9 pmol of each siRNA, compared with 10 pmol used for Fig. 3), individual Spsb1 knockdown lengthened period by 1.6 ± 0.1 h and Spsb4 knockdown lengthened period by 1.0 ± 0.1 h, but knocking down both Spsb1 and Spsb4 in the same cultures lengthened period by 3.0 ± 0.2 h, relative to controls (Fig. 4). Taken together, these data suggest that Spsb1 and Spsb4, but not Spsb2 or Spsb3, are partially redundant regulators of circadian oscillator function.
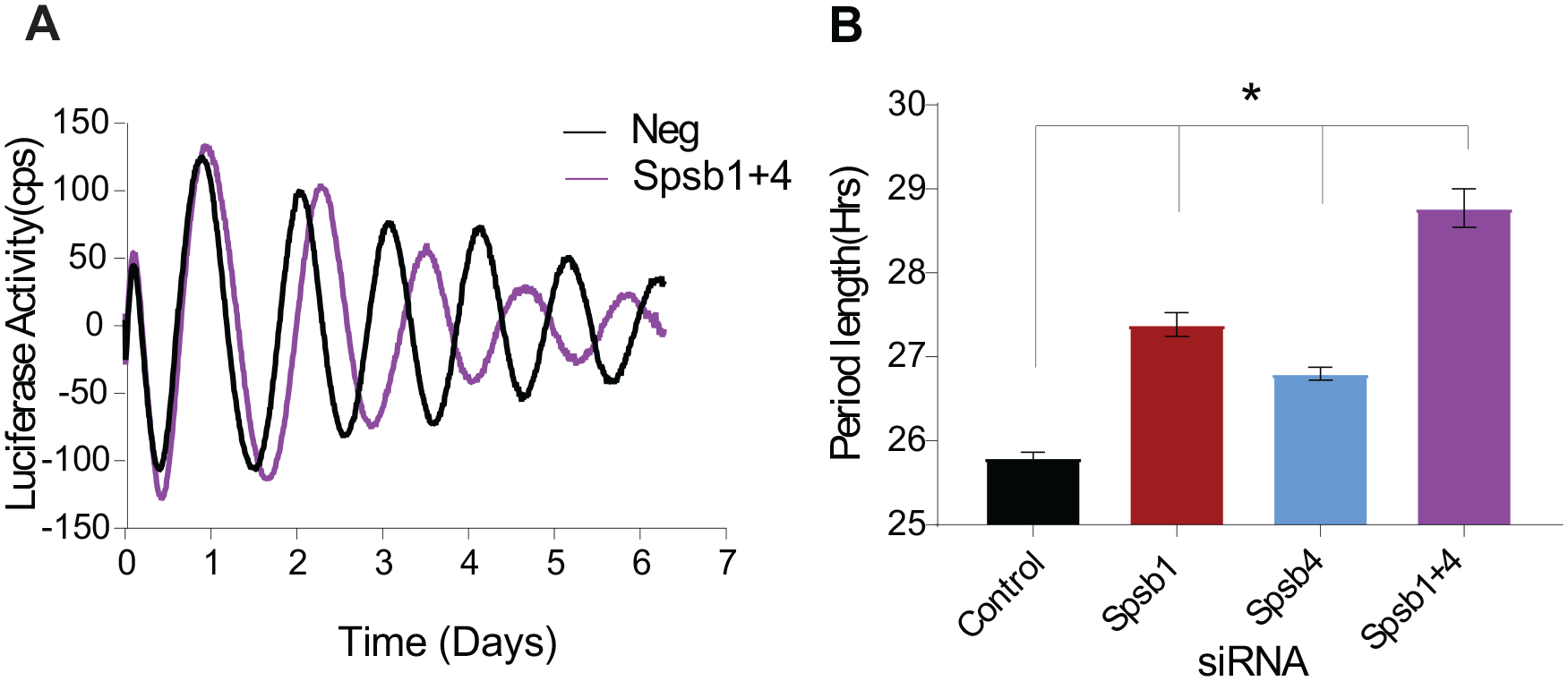
Spsb1 and Spsb4 double knockdown has additive effects on circadian period. (A) Representative bioluminescence rhythms produced by Bmal1-Luc cells transfected with the indicated small interfering RNAs (siRNAs; 9 pmol of Spsb1 and Spsb4 siRNAs each, filling in with negative control siRNAs to ensure molar equivalence). (B) Average period lengths combined from several experiments (mean ± SEM, n = 14-15 cultures per siRNA, from 4 independent trials) *p < 0.0001 versus negative controls and Spsb1 or Spsb4 versus Spsb1+4, analysis of variance, Tukey’s multiple comparisons test.
Given their specific roles in regulating RevErbα stability and the previous notion that changes in RevErbα dynamics can regulate period (Ukai-Tadenuma et al., 2011), we next determined if Spsb1/4 depletion altered the circadian patterns of endogenous RevErbα abundance. We did this in Bmal1-luc U2OS cells transfected with an equal amount of Spsb1 and Spsb4, or negative control, siRNAs and synchronized with 50% horse serum 48 h later. Since RevErbα levels are rhythmic with a peak at ~22 h post-synchronization (DeBruyne et al., 2015), we harvested cultured cells at 2-h intervals starting at 18 h after synchronization to detect changes that may affect overall rhythmic accumulation or degradation of RevErbα. In cells transfected with negative control siRNAs, RevErbα protein levels showed a strong oscillation, peaking at about ~24 h post-synchronization and falling to a trough about 12 to 14 h later (Fig. 5A, B). In Spsb1/Spsb4–depleted cells, RevErbα levels oscillated, but its peak levels were ~50% higher than controls. This elevation in abundance extended the duration in which RevErbα protein levels were higher than the half-maximal levels in negative controls by 3 to 4 h (Fig. 5B), a time frame consistent with the ~3-h lengthening in period (Fig. 4). Importantly, the siRNA-mediated knockdown persisted throughout the duration of the experiment (Fig. 5C). Moreover, the increase in RevErbα protein levels is not due to an increase in RevErbα gene expression (Fig. 5D), consistent with the role of SPSB1 and SPSB4 as posttranslational regulators of RevErbα stability.
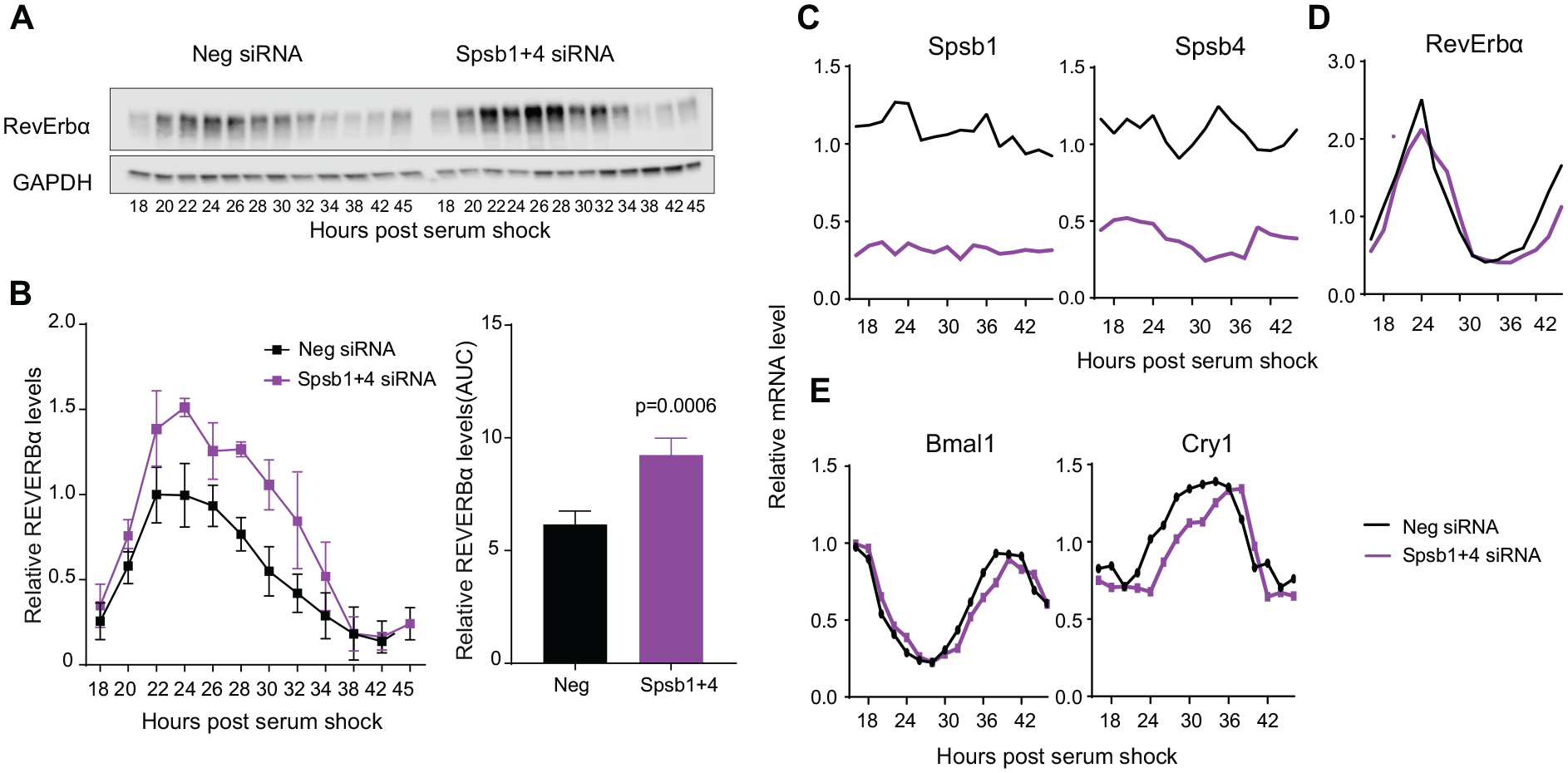
Spsb1 and Spsb4 depletion slows RevErbα degradation. (A) Representative western blot of RevErbα protein abundance rhythms in U2OS cells transfected with a control (Neg) small interfering RNA (siRNA) or a combination of Spsb1 and Spsb4 siRNAs. Cells were synchronized with 50% horse serum 48 h after transfection and collected at the indicated times. (B) Left: Quantification of RevErbα protein abundance from 4 experiments (mean ± SEM, n = 4). Right: Bar graph of the areas under the curve on the left. Data are represented as mean ± SEM, determined for n = 4 independent trials separately. The p value shown is that from a Student’s t test. (C) Spsb1 and Spsb4, (D) RevErbα, (E) Bmal1 and Cry1 mRNA profiles of synchronized control or Spsb1 and 4–depleted U2OS cells collected every 2 h. Data are plotted relative to the average of the negative siRNA samples.
We also determined if these effects on endogenous RevErbα protein levels translated to altered expression of its target genes Bmal1 and Cry1. We predicted that we would see evidence of a prolonged repression in the expression of these genes that matched the RevErbα profile in Spsb1/4-depleted cells. Indeed, the mRNA expression profile of Cry1 in the Spsb1/4-depleted cells was delayed compared with their negative controls, but there was only a very subtle effect in the timing of Bmal1 expression (Fig. 5E). This differential effect is likely due to the phase difference between the Bmal1 and Cry1 expression profiles in relation to the RevErbα protein abundance rhythm: Cry1 is expressed earlier than Bmal1 and thus is likely more sensitive to the effects of manipulating RevErbα stability (Suppl. Fig. S7). For instance, the most robust effect of Spsb1/4 depletion on RevErbα levels (time 24-32) highly corresponds to the bathyphase (trough) in Bmal1 expression but substantially overlaps in time with the increase in Cry1 expression. Thus, the change in the circadian RevErbα abundance profile does correlate well with changes in Cry1 expression. Since RevErbα regulation of Cry1 expression can determine circadian period (Ukai-Tadenuma et al., 2011), the most parsimonious interpretation of our data collectively is that SPSB1 and SPSB4 are redundant regulators of circadian clock function via their role in determining RevErbα stability. Future studies aimed at mutating the precise sites on RevErbα itself that impair its regulation by Spsb1/4 are necessary to formally test this hypothesis. Nonetheless, our data suggest that SPSB1/4 modulation of RevErbα stability (but not RevErbβ) could be another entryway for manipulating overall clock function.
Discussion
The UPS plays a critical role in regulating many cellular processes necessary for cell survival. Defects in this system can result in pathogenesis of many human diseases (Predmore et al., 2010; Johnson, 2015; Tramutola et al., 2016). It is especially essential in processes such as the circadian system, whose timing of feedback loops is dictated by the time-dependent degradation of its components. In the circadian clock, the role of the UPS in determining protein half-life is critical for proteins such as PER1/2, CRY1/2, and RevErbα, with a daily rhythm in abundance (Siepka et al., 2007; Stojkovic et al., 2014). There has also been high interest in E3 ligases as therapeutic targets because of their ability to confer substrate specificity (Bulatov et al., 2018). However, there is still much to learn in this area, as substrates have been identified for a fraction of the ~600 mammalian genes encoding apparent E3 ligases (Li et al., 2008), and E3 ligases are known for an even smaller fraction of degraded proteins.
Adding to this complexity is that individual protein substrates can be targeted by multiple E3 ligases. Remarkably, in addition to SPSB1/4, four other E3 ligases, Arf-bp1 and Pam (Yin et al., 2010), Siah2 (DeBruyne et al., 2015), and FBXW7 (Zhao et al., 2016) have also been shown to regulate RevErbα stability/degradation. Although the effects/roles of each of these E3 ligases have not been directly compared, it is hard to imagine that they are all redundant with each other, as removing each has detectable consequences on RevErbα levels (Spsb1/4, Arf-bp1/Pam; Yin et al., 2010), cycling (Siah2; DeBruyne et al., 2015), and/or function (Fbxw7; Zhao et al., 2016). Moreover, depleting Spsb1 or 4 (or both) or Siah2 lengthens period without notable differences in rhythm amplitudes (DeBruyne et al., 2015), whereas removing Arf-bp1/Pam and Fbxw7 has the opposite effect: it reduces rhythm amplitudes without altering circadian period (Yin et al., 2010; Zhao et al., 2016). The contrasting roles of these E3 ligases in modulating either clock amplitude or period also indicate that they likely have distinct functions in regulating RevErbα stability and the circadian clock. Indeed, the Spsb family members display tissue specificity in rhythmic expression (Kleiber and Singh, 2009; Zhang et al., 2014), providing one potential avenue for separating functions. In addition, SPSB1 and SPSB4 appear to target only RevErbα, not RevErbβ, which is at least distinct from Siah2, which regulates both paralogs (DeBruyne et al., 2015). Similarly, FBXW7 was also found to selectively interact with and degrade RevErbα but not RevErbβ, owing to the exclusive presence of a highly conserved sequence present in RevErbα (Zhao et al., 2016). Overall, we predict that these RevErbα E3 ligases likely regulate its stability in a context-specific, or possibly a target gene promotor–specific manner. These possibilities, however, have yet to be explored.
In addition, it is also possible that having multiple E3 ligases targeting the same protein substrates is essential to fully ensure its proteolysis but in a very highly regulated manner. Several studies have suggested that the time-dependent degradation of RevErbα may be critical in regulating its overall function as a transcriptional repressor. Although we cannot discount the role of other factors in causing the period lengthening observed in U2OS cells, various lines of evidence suggest that RevErbα stability is the most likely mechanism. For example, constitutive RevErbα overexpression leads to a constitutive inhibition of Bmal1 transcription, arresting clock function, and altering the ability of RevErbα to repress the Cry1 promoter that regulates circadian period (Ukai-Tadenuma et al., 2011). Moreover, disrupting RevErbα function impairs the amplitude of the circadian clock (Zhang et al., 2009; Zhao et al., 2016), and removing both RevErbα and RevErbβ ablates rhythmicity (Bugge et al., 2012; Cho et al., 2012). These findings argue that the precisely timed regulation of RevErbα’s appearance and its disappearance via degradation are critical control mechanisms governing RevErbα’s function. Emerging in vivo studies support this notion as removing either RevErbα or proteins that regulate its stability results in physiological consequences in a host of tissues, including the liver (Duez and Staels, 2008; Le Martelot et al., 2009; Bugge et al., 2012), brown adipose (Gerhart-Hines et al., 2013), white adipose (Jager et al., 2016), muscle (Woldt et al., 2013), and brain (Jager et al., 2014). However, it remains to be determined whether disrupting RevErbα stability will always have the same consequence. Nonetheless, the multiplicity in RevErbα E3 ligases and their potential differential roles suggests the exciting possibility that each ligase may provide a unique opportunity to manipulate RevErbα to achieve different physiological outcomes.
Supplemental Material
Supplemental_Figure_1 – Supplemental material for The E3 Ligases Spsb1 and Spsb4 Regulate RevErbα Degradation and Circadian Period
Supplemental material, Supplemental_Figure_1 for The E3 Ligases Spsb1 and Spsb4 Regulate RevErbα Degradation and Circadian Period by Tsedey Mekbib, Ting-Chung Suen, Aisha Rollins-Hairston and Jason P. DeBruyne in Journal of Biological Rhythms
Supplemental Material
Supplemental_Figure_2 – Supplemental material for The E3 Ligases Spsb1 and Spsb4 Regulate RevErbα Degradation and Circadian Period
Supplemental material, Supplemental_Figure_2 for The E3 Ligases Spsb1 and Spsb4 Regulate RevErbα Degradation and Circadian Period by Tsedey Mekbib, Ting-Chung Suen, Aisha Rollins-Hairston and Jason P. DeBruyne in Journal of Biological Rhythms
Supplemental Material
Supplemental_Figure_3 – Supplemental material for The E3 Ligases Spsb1 and Spsb4 Regulate RevErbα Degradation and Circadian Period
Supplemental material, Supplemental_Figure_3 for The E3 Ligases Spsb1 and Spsb4 Regulate RevErbα Degradation and Circadian Period by Tsedey Mekbib, Ting-Chung Suen, Aisha Rollins-Hairston and Jason P. DeBruyne in Journal of Biological Rhythms
Supplemental Material
Supplemental_Figure_4 – Supplemental material for The E3 Ligases Spsb1 and Spsb4 Regulate RevErbα Degradation and Circadian Period
Supplemental material, Supplemental_Figure_4 for The E3 Ligases Spsb1 and Spsb4 Regulate RevErbα Degradation and Circadian Period by Tsedey Mekbib, Ting-Chung Suen, Aisha Rollins-Hairston and Jason P. DeBruyne in Journal of Biological Rhythms
Supplemental Material
Supplemental_Figure_5 – Supplemental material for The E3 Ligases Spsb1 and Spsb4 Regulate RevErbα Degradation and Circadian Period
Supplemental material, Supplemental_Figure_5 for The E3 Ligases Spsb1 and Spsb4 Regulate RevErbα Degradation and Circadian Period by Tsedey Mekbib, Ting-Chung Suen, Aisha Rollins-Hairston and Jason P. DeBruyne in Journal of Biological Rhythms
Supplemental Material
Supplemental_Figure_6 – Supplemental material for The E3 Ligases Spsb1 and Spsb4 Regulate RevErbα Degradation and Circadian Period
Supplemental material, Supplemental_Figure_6 for The E3 Ligases Spsb1 and Spsb4 Regulate RevErbα Degradation and Circadian Period by Tsedey Mekbib, Ting-Chung Suen, Aisha Rollins-Hairston and Jason P. DeBruyne in Journal of Biological Rhythms
Supplemental Material
Supplemental_Figure_7 – Supplemental material for The E3 Ligases Spsb1 and Spsb4 Regulate RevErbα Degradation and Circadian Period
Supplemental material, Supplemental_Figure_7 for The E3 Ligases Spsb1 and Spsb4 Regulate RevErbα Degradation and Circadian Period by Tsedey Mekbib, Ting-Chung Suen, Aisha Rollins-Hairston and Jason P. DeBruyne in Journal of Biological Rhythms
Supplemental Material
Supplemental_Figure_Legends_1 – Supplemental material for The E3 Ligases Spsb1 and Spsb4 Regulate RevErbα Degradation and Circadian Period
Supplemental material, Supplemental_Figure_Legends_1 for The E3 Ligases Spsb1 and Spsb4 Regulate RevErbα Degradation and Circadian Period by Tsedey Mekbib, Ting-Chung Suen, Aisha Rollins-Hairston and Jason P. DeBruyne in Journal of Biological Rhythms
Footnotes
Acknowledgements
This work was funded by the National Institutes of Health (NIH) National Institute of General Medical Sciences grants 1SC1GM109861 and 1R35GM127044 to J.P.D., as well as in part by NIH National Institute on Minority Health and Health Disparities grants 8G12MD007602, 8U54MD00758, 1G20RR031196, S21MD000101, and C06RR18386 and NIH National Institute of Neurological Disorders and Stroke grant U54 NS083932.
Conflict of Interest Statement
The authors report no conflicts of interest. The authors alone are responsible for the content and writing of the paper.
Notes
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
