Abstract
The transcriptional circadian clock network is tuned into a 24-h oscillator by numerous posttranslational modifications on the proteins encoded by clock genes, differentially influencing their subcellular localization or activity. Clock proteins in any circadian organism are subject to posttranslational regulation, and many of the key enzymes, notably kinases and phosphatases, are functionally conserved between the clocks of mammals, fungi, and plants. We now establish sumoylation, the posttranslational modification of target proteins by the covalent attachment of the small ubiquitin-like modifier protein SUMO, as a novel mechanism regulating key clock properties in the model plant Arabidopsis. Using 2 different approaches, we show that mutant plant lines with decreased or increased levels of global sumoylation exhibit shortened or lengthened circadian period, respectively. One known functional role of sumoylation is to protect the proteome from temperature stress. The circadian clock is characterized by temperature compensation, meaning that proper timekeeping is ensured over the full range of physiologically relevant temperatures. Interestingly, we observed that the period defects in sumoylation mutant plants are strongly differential across temperature. Increased global sumoylation leads to undercompensation of the clock against temperature and decreased sumoylation to overcompensation, implying that sumoylation buffers the plant clock system against differential ambient temperature.
Introduction
Posttranslational modification instantly alters the characteristics of target proteins and provides organisms with a rapid means to respond to changes in their surrounding environment without having to realign transcription or translation to that change. Circadian timekeeping is dependent on posttranscriptional and posttranslational regulation of clock gene products to tune the transcriptional system into a 24-h oscillator (Gallego and Virshup, 2007; Mehra et al., 2009; Seo and Mas, 2014).
Like in other higher clades of circadian organisms, the most abundant posttranslational modification identified to date in the plant circadian clock is phosphorylation (Hsu and Harmer, 2013; Choudhary et al., 2015). Phosphorylation can alter the stability of clock proteins and their function (Fujiwara et al., 2008). Interestingly, the role of posttranslational modifications in timekeeping appears to be well-conserved. For example, Casein Kinase (CK) 1 and 2 are involved in circadian regulation in mammals and fungi (Lee et al., 2009; Querfurth et al., 2011). Both kinases were recently implicated in timekeeping in the simplistic proto-clock system of the green algae Ostreococcus tauri (O’Neill et al., 2011; van Ooijen et al., 2013a, 2013b; Le Bihan et al., 2015). In plants, phosphorylation of CCA1 by CK 2 affects its binding to target promoters (Daniel et al., 2004; Portolés and Más, 2010).
Increasing evidence implicates several additional posttranslational modifications in timekeeping in various model organisms (van Ooijen and Millar, 2012). For example, sumoylation functions as a regulator of the mammalian clock through dynamic modification of BMAL1 (Cardone et al., 2005; Lee et al., 2008). Recent proteomic data sets suggest that sumoylation in plants often co-regulates the function of target proteins with phosphorylation (Nukarinen et al., 2017). Sumoylation is the modification of proteins by the Small Ubiquitin–related Modifier (SUMO), a reversible and highly dynamic modification that is conserved throughout the eukaryote lineage (Melchior, 2000). SUMO is a ~10-kDa protein that structurally resembles ubiquitin and requires maturation by SUMO proteases before attachment to a target protein (Ha and Kim, 2008). SUMO is attached covalently to a lysine residue in the substrate protein, in an adenosine triphosphate–dependent reaction involving an activating enzyme (E1), a conjugating enzyme (E2), and a ligase (E3; Fig. 1; Gareau and Lima, 2010). While E1 and E2 are vital enzymes (Saracco et al., 2007), E3 is biochemically not strictly required for the formation of a covalent bond between SUMO and substrate (Desterro et al., 1999). However, in vitro and in vivo studies show that it does facilitate and enhance binding (Kagey et al., 2003; Miura et al., 2009; Okada et al., 2009). Sumoylation is reversible by deconjugation of SUMO from a substrate protein, by the same SUMO proteases required for maturing SUMO (Chosed et al., 2006; Colby et al., 2006; Gareau and Lima, 2010).
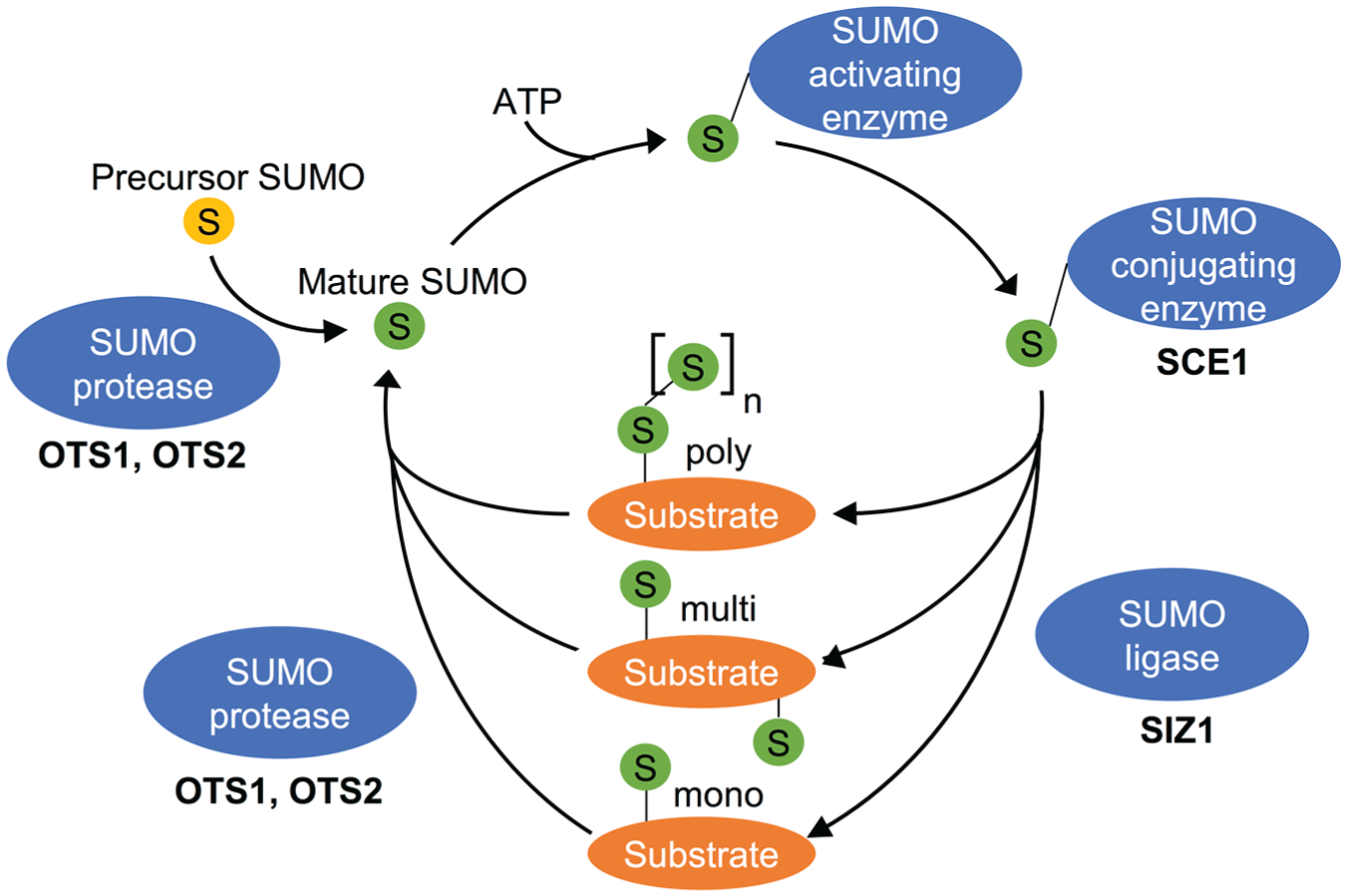
The sumoylation pathway. (A) The precursor SUMO is C-terminally cleaved by SUMO proteases. Mature SUMO is bound by SUMO activating enzyme (SAE) and subsequently transferred to SUMO conjugating enzyme (SCE). Ligation to a lysine residue can be facilitated by SUMO ligases. One or more lysine residues on a target protein can be sumoylated by a single or a chain of SUMO. Modification by SUMO can be reversed by SUMO proteases. The SUMO machinery enzymes investigated in this study are indicated in bold.
Abiotic stress causes substantial increases in the level of sumoylation and the overall number of sumoylated proteins (Kurepa et al., 2003; Saracco et al., 2007; van den Burg et al., 2010; Miller et al., 2010). Upon heat stress, SUMO conjugation rapidly increases, reducing the overall level of free SUMO. The modification is only transient, and sumoylation levels return to normal a few hours after heat shock (Kurepa et al., 2003; Saracco et al., 2007). Increased sumoylation in response to stress is thought to be an important mechanism by which cells can reprogram transcription to cope with a changed environment (Yoo et al., 2006; Tomanov et al., 2013). In plants, many abiotic stress responses are gated by the circadian clock (reviewed in Spoel and van Ooijen, 2014). Plants respond to milder changes in temperature, leading to altered growth and development in a process called thermomorphogenesis (Quint et al., 2016). Remarkably, several of the proteins involved in thermomorphogenesis have been identified as sumoylation targets (Conti et al., 2014; Sadanandom et al., 2015; Tan et al., 2015; Lin et al., 2016), indicating that SUMO plays a role not only in regulating response to temperature extremes but also within the temperature range not regarded as “stressful.”
The circadian clock is characterized by temperature compensation, which means that the free-running period of the clock remains stable over a wide range of physiologically relevant temperatures (Pittendrigh, 1954; Kondo et al., 1993; Somers et al., 1998; O’Neill and Reddy, 2011). Temperature compensation is a defining feature of the circadian clock, and although the underlying mechanism(s) are not fully elucidated, it is clear that it involves transcriptional, posttranscriptional, and posttranslational regulation. In plants, temperature compensation has been shown to involve the clock genes CCA1, LHY, RVE8, PRR7, PRR9, and GI, covering all expression phases of a full 24-h cycle (Gould et al., 2006; Salomé et al., 2010; Rawat et al., 2011). In addition, several genes involved in light sensing and photoperiodic responses are involved in temperature compensation (Edwards, 2005; Edwards et al., 2006; Ito et al., 2012; Gould et al., 2013; Nagel et al., 2014; Edwards et al., 2015). At the posttranslational level, phosphorylation and, in particular, CK2 activity are known regulators of timekeeping and temperature compensation (Sugano et al., 1998; Daniel et al., 2004; Baker et al., 2009; Lee et al., 2009; van Ooijen et al., 2013a). Inhibition of phosphorylation through mutations or chemical treatment compromises temperature compensation in plants, mammals, and fungi (Mehra et al., 2009; Portolés and Más, 2010; Zhou et al., 2015). In this article, we report our investigations into a possible role of sumoylation in circadian timekeeping and, notably, given the known roles of sumoylation in temperature responses, the potential link between sumoylation and temperature compensation of circadian rhythms.
Materials And Methods
Plant Lines, Genotyping, and Growth Conditions
The Columbia-0 plant lines ots1 ots2 (Conti et al., 2008), siz1-2 (Miura et al., 2005), sum1-1 (Saracco et al., 2007), SUM1-OX, and sum1-1 35S:amiR-SUM2 (van den Burg et al., 2010) were described previously. The reporter line CCA1pro:LUC, in Col-0 background, was kindly provided by Karen Halliday (University of Edinburgh).
CCA1pro:LUC Col-0 was crossed with ots1 ots2 and siz1 to give CCA1pro:LUC ots1 ots2 and CCA1pro:LUC siz1. F1-F3 progeny were genotyped by polymerase chain reaction to confirm T-DNA insertions (SIZ1 LP 5′-gagctgaagcatctggttttg-3′ and RP 5′-cacgacagatgaagcattgtg-3′ [Miura et al., 2007], OTS1 RP 5′-cgacaagaagtggtttagacc-3′ [Conti et al., 2008], and LP 5′-gatgatgcaaggaggctagtg-3′, OTS2 LP 5′-gcttcttccggtttaaaccac-3′ and RP 5′-tttttcttctggcgactcatg-3′, and SALK Lba1 5′-tggttcacgtagtgggccatcg-3′ [Alonso et al., 2003]) and presence of transgene (CCA1prom 5′-tccatttccgtagcttctggtctc-3′, LUCrev 5′-gccttatgcagttgctctcc-3′). Soil-grown plants were grown under long day conditions (16 h light/8 h dark) at 22 °C using 60- to 80-µmol × m–2 × s–1 white light-emitting diode (LED) tube light (Impact, T8). Seedlings were grown on half-strength Murashige and Skoog media without vitamins and sucrose at 21 °C in 100 µmol × m–2 × s–1 white tube light (Phillips, Alto II).
Luminescent Imaging
The protoplast transfection vector system for rapid circadian imaging was provided by David Somers (Kim and Somers, 2010), and experiments were carried out at 20 °C under constant blue (470 nm) and red (630 nm) LED light as described previously (Hansen and van Ooijen, 2016). For imaging of seedlings, 5- to 6-day-old seedlings were sprayed with luciferin (5 mM luciferin [Biosynth AG], 10 mM Tris pH 8, 0.01% Triton), transferred to constant light conditions (50 µmol × m–2 × s–1 630 nm red LED light, and 25 µmol × m–2 × s–1 470 nm blue LED light) at 15 °C, 21 °C, or 27 °C and imaged every 2 h with a CCD camera (Hamamatsu). Images were analyzed with MetaMorph software. All circadian parameters described in this article were determined using nonlinear regression analysis in GraphPad Prism, similar to what is published in Putker et al. (2017): A centered fifth-order polynomial fit is performed to remove trend and create a residuals plot of the data on which a circadian damped cosine wave is fitted to determine period length using the user-defined equation: Y = (m*X) + amplitude*exp(–k*X)*cos(((2*π*(X-phase))/period)) where X is time, Y is signal, k is the decay constant (such that 1/k is the half-life), which is constrained to >0. Initial values are chosen automatically (m: 1; phase: 1; period: 24). The range of time that is analyzed for both fits is set based on visual inspection of raw data and usually excludes the first part of any time series where traces can be erratic, as well as the last part that is often dampened. A sample data set analyzed as above can be obtained as a Prism file from the corresponding author on request.
Results
Sumoylation Tunes Circadian Rhythms in Plants
Besides the several SUMO isoforms itself, the sumoylation machinery involves the enzymatic activities of a SUMO activating enzyme, a SUMO conjugating enzyme, SUMO ligases, and specific SUMO proteases (Fig. 1). We subjected existing Arabidopsis lines defective in these activities to a cell-based clock assay to rapidly determine if sumoylation could be important for timekeeping. Leaf mesophyll cells were isolated from adult wild-type and mutant plants, and these so-called protoplasts were transfected with a construct carrying the clock-regulated CCA1 promoter driving expression of the firefly luciferase protein (CCA1pro:LUC) for longitudinal imaging of circadian rhythms. This assay provides a rapid and reliable means of assessing clock defects (Kim and Somers, 2010; Hansen and van Ooijen, 2016) in any line where the mutant phenotype does not affect protoplast viability. Most mutant or transgenic lines tested exhibited no or only a minor period difference to wild-type plants (Fig. 2A, B), presumably reflecting the known functional redundancy between isoforms and enzymes within the sumoylation machinery (Saracco et al., 2007) and/or the well-documented capacity of the circadian clock network to buffer against perturbations (Zhang and Kay, 2010). However, a double mutant of the 2 isoforms of SUMO protease OVERLY TOLERANT TO SALT (ots1 ots2) exhibited a marked lengthening in circadian rhythms. The ots1 ots2 mutant exhibits an increased level of overall sumoylation (Conti et al., 2008), suggesting that sumoylation might be a delay mechanism within the clock network. To verify that result, the ots1 ots2 line was crossed to the stably transgenic, rhythmically luminescent clock marker line CCA1pro:LUC. Indeed, a comparison of the resulting luminescent signal reveals that increased global sumoylation affects circadian period length (Fig. 3A). We hypothesized that if increased overall sumoylation slows circadian rhythms, decreased global sumoylation might have the opposite effect. Decreased global sumoylation is observed in the SUMO ligase mutant siz1 (Miura et al., 2005), which could not reliably be tested in the protoplast assay because of its dwarfed phenotype that affects protoplast viability. The siz1 line was therefore crossed to the CCA1pro:LUC line to analyze clock defects on a whole-plant level. Circadian rhythms in the resulting line revealed that reduced overall levels of sumoylation are indeed associated with a short circadian period (Fig. 3B), supporting the notion that SUMO ligase SIZ1 and SUMO proteases OTS1/OTS2 exhibit opposing effects in tuning the circadian clock in plants.
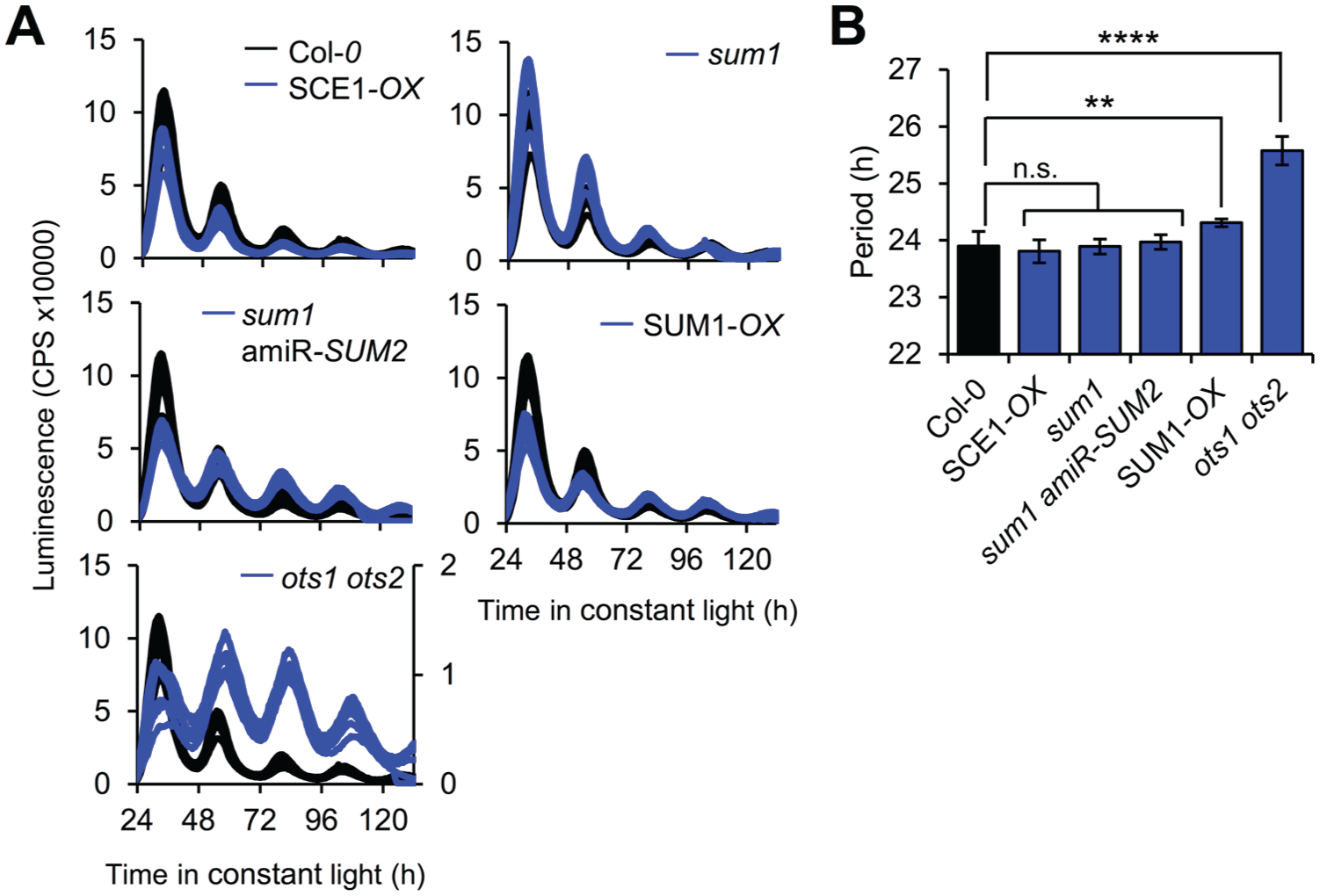
Sumoylation affects timekeeping in plants. (A) Transient expression of CCA1pro:LUC in protoplasts from the indicated mutant backgrounds imaged in constant light at 20 °C. SUMO isoforms are indicated (SUM1, SUM2); SCE1 is Sumo Conjugating Enzyme 1; OTS1/2 is Overly Tolerant to Salt homolog 1 or 2; amiR stands for artificial micro-RNA. The ots1 ots2 data are scaled to the right y-axis. (B) Circadian period as calculated from the traces in panel A (mean value ± SEM, n = 7 biological replicates for all genotypes except ots1 ots2 [n = 6]). Significant differences from Col-0 were determined by ordinary 1-way analysis of variance with Dunnett’s multiple-comparisons test. p-value >0.05 (ns), <0.01 (**), <0.0001 (****). Result has been replicated in 3 independent biological replicate experiments.
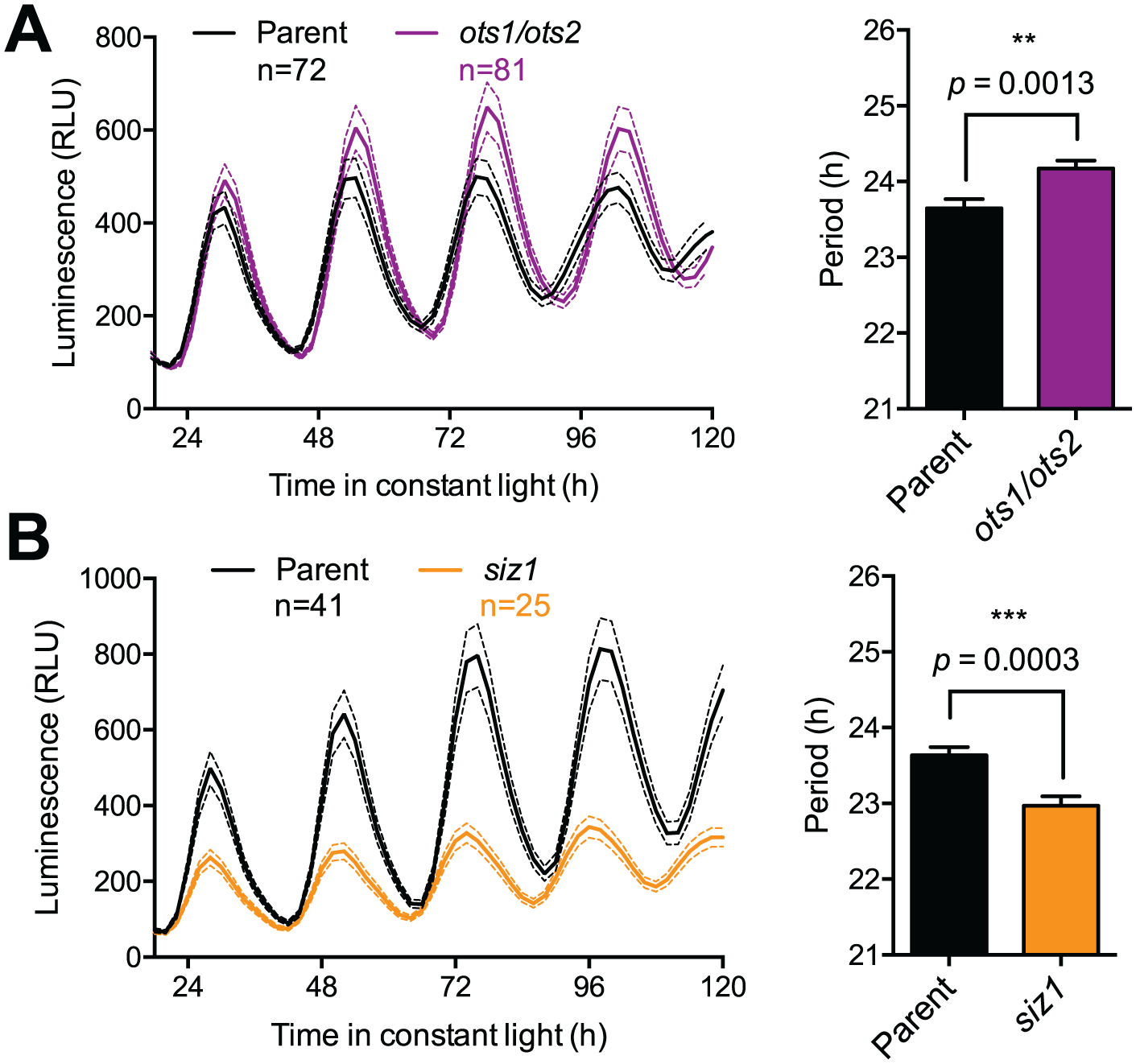
SUMO protease and ligase activities have an opposing effect on circadian period. Circadian gene expression as reported by CCA1pro:LUC was analyzed in the Col-0 background (black traces) compared with the ots1 ots2 (A) or siz1 (B) mutant backgrounds at 21 °C. Resulting luminescent traces (left) revealed significant changes to the period (right) of circadian rhythms (mean value ± SEM of the indicated number of biological replicates; comparisons between mutant data and wild-type using unpaired t tests; p values as indicated). Result has been replicated in 3 independent biological replicate experiments.
Temperature Compensation Is Affected by Sumoylation
Given the general role of sumoylation in temperature responses, the effects of siz1 and ots1 ots2 on circadian period were analyzed at moderate cold and warm conditions. Seedlings of CCA1pro:LUC siz1, CCA1pro:LUC ots1 ots2, and the parent line CCA1pro:LUC Col-0 were grown at 21 °C for 5 to 6 days under light-dark cycles and then transferred to luminescent imaging conditions of constant light at either 15 °C or 27 °C. Remarkably, while a long-period circadian phenotype was very evident in the CCA1pro:LUC ots1 ots2 at 15 °C (Fig. 4A), there was no significant difference in circadian period at 27 °C (Fig. 4A). This result suggests that desumoylation is required for proper temperature compensation at colder temperatures. To test whether the opposite is true for SUMO conjugation, CCA1pro:LUC expression in the siz1 background was compared with the parent line at 15 and 27 °C. Astonishingly, we observed no significant period difference at 15 °C whereas a strong period-shortening effect was observed at 27 °C (Fig. 4B). This result shows that in contrast to SUMO protease activity, SUMO ligase activity is especially important for circadian timekeeping at higher temperatures.
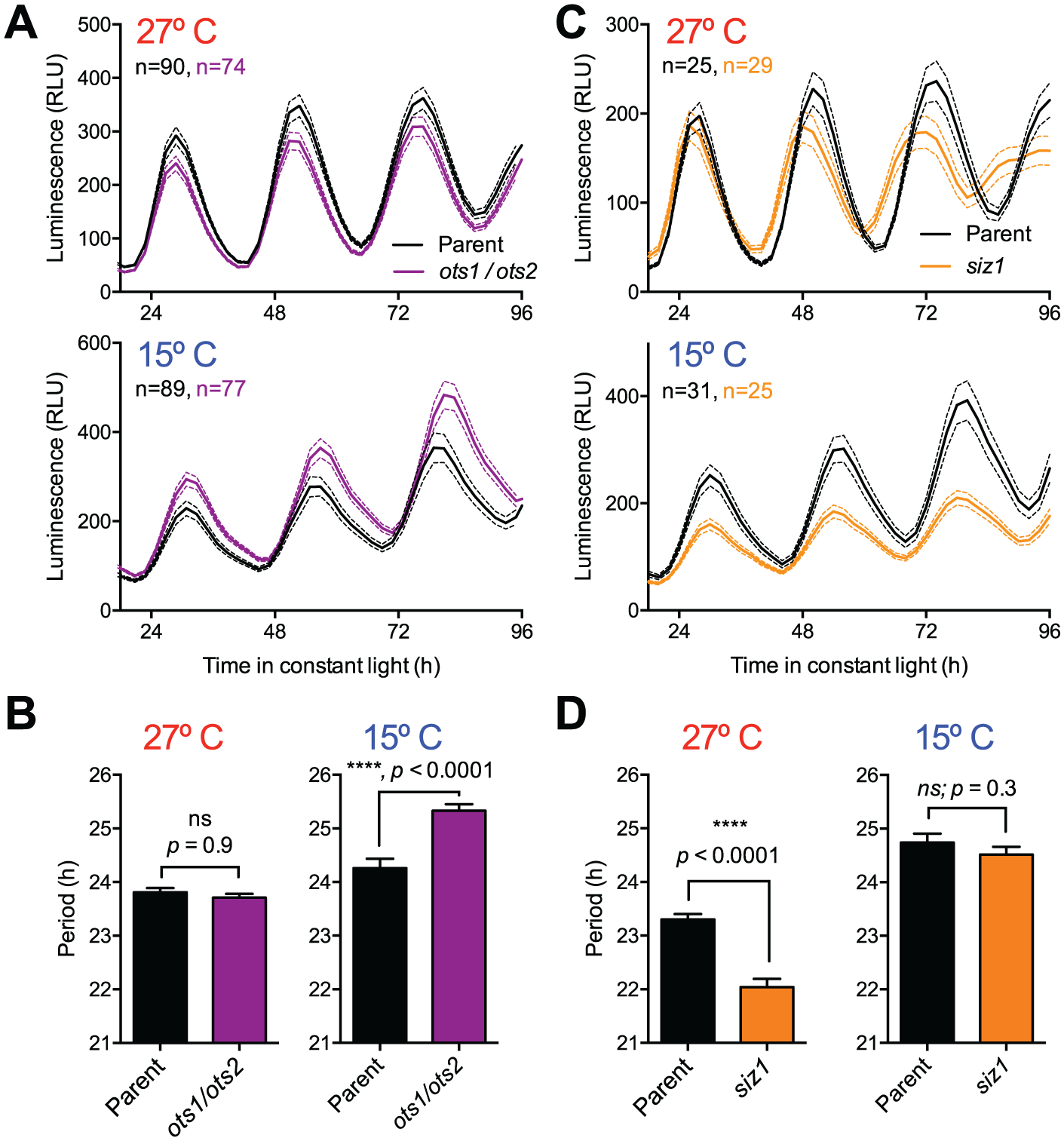
Circadian rhythms in ots1 ots2 and siz1 at high and low temperatures. Seedlings of CCA1pro:LUC ots1 ots2 (A, B) and CCA1pro:LUC siz1 (C, D) compared with the parent line CCA1pro:LUC Col-0 imaged at 27 °C (top) or 15 °C (bottom) in constant light over 4 days. Luminescent traces revealed significant changes to the period of circadian rhythms (mean value ± SEM of the indicated number of biological replicates; comparisons between mutant data and wild-type using unpaired t tests; p values as indicated). Result has been replicated in 2 independent biological replicate experiments.
The effects of reduced SUMO ligase and protease activity are better visualized if the quantified period differences are plotted relative to those observed in wild-type plants (Fig. 5). Biochemical reactions proceed faster at higher temperatures, except those that are temperature compensated, such as the circadian system. The clock in the siz1 background is undercompensated at higher temperatures (i.e., speeds up with temperature increase), indicating that SUMO ligase activity is required for temperature compensation at high temperatures. The clock in the ots1 ots2 background is undercompensated at the lower temperatures (i.e., slows down with decreased temperature), indicating that SUMO protease activity is required for temperature compensation at low temperatures.
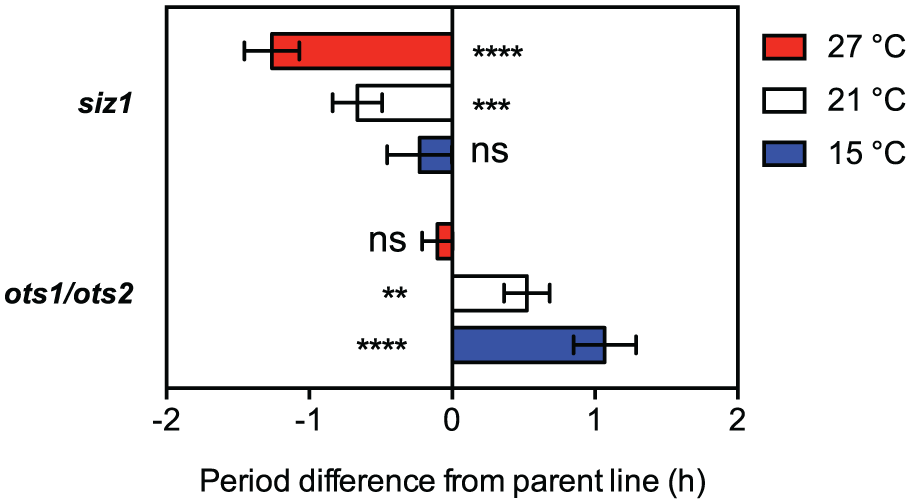
Differential effects on the circadian period of mutants of the SUMO machinery across a range of physiologically relevant temperatures. CCA1pro:LUC ots1 ots2 and CCA1pro:LUC siz1 period difference from parent line at 27 °C, 21 °C, and 15 °C, from traces in Figure 3 and 4. Mean of difference in period is provided ±SEM; significant differences were determined by t tests between the mutant and the parent line for each temperature. p-value >0.05 (ns), <0.01 (**), <0.001 (***), <0.0001 (****). Result has been replicated in 2 (15 and 27 °C) or 3 (21 °C) independent biological replicate experiments.
Discussion
The data in Figures 1 to 3 imply that sumoylation contributes to the tuning of circadian period, differentially over moderate temperature ranges. Combined, these data suggest sumoylation is a delay mechanism under normal lab conditions of ~21 °C. To compensate clocks against temperature, SIZ1 activity (i.e., SUMO ligase activity; Miura et al., 2005) negates the speeding that higher temperatures would have, whereas OTS1/2 activity (i.e., SUMO protease activity; Conti et al., 2008) negates the slowing effect of decreased temperature. It is important to note that although reported differences from controls in this study are relatively small, the true effect of these enzymes on the clock and temperature compensation might well be greater, as the several additional SUMO ligases and proteases that have been identified might balance mutant phenotypes. However, no significant period changes were observed upon mutation of both predominant SUMO isoforms nor upon overexpression of the only known conjugating enzyme (Fig. 2) in a protoplast assay system at 21 °C. The authors deem it likely that future studies using stably transgenic clock marker lines in additional mutant backgrounds will reveal clock phenotypes in these mutants that, too, are dependent on environmental conditions: the data in Figure 5 suggest that had the original screen been carried out at 27 or 15 °C, the ots1/ots2 or siz1 mutant backgrounds, respectively, might not have displayed their phenotype. A second important note is that temperature might not be the only environmental factor that influences the overt phenotypes of sumoylation mutants. In the supplementary files of a previous study (Jin et al., 2008), the effects of the siz1 mutation on circadian rhythms in red light versus blue light were compared. The authors of that study concluded that there were no changes in circadian period length, although no statistical analyses were reported. Performing these analyses using the published period estimates, standard deviations, and replicate numbers (Suppl. Fig. S1), we find period differences, while biologically small, that are statistically significant for some genotypes and crucially depend on light conditions. Raw data are provided only for the red light experiment, but based on the published period estimates, a slight period shortening was observed at 21 °C under blue light, not dissimilar to results presented here. Our current results suggest that this effect might have been more pronounced if experiments were carried out at slightly lower temperatures. Unfortunately, both sets of results are difficult to compare, because in the current study, results were gathered under red and blue light. It is, however, an interesting avenue for further study to elucidate if the phenotype of the siz1 mutation and other sumoylation mutants depend on light spectrum as well as temperature.
It is not likely that the effects of sumoylation mutants on the clock are mediated via differential sumoylation of a single target. Indeed, a small number of clock-relevant proteins has already been reported to be sumoylated (Miller et al., 2010; López-Torrejón et al., 2013; Lin et al., 2016). To obtain a general idea of clock-relevant targets that might be subject to sumoylation, we compared the identified SUMO targets from a proteome-wide screen (Miller et al., 2010) with known circadian-expressed transcripts (Covington et al., 2008). From the 297 known SUMO targets for which transcript expression profiles were available, 43% (127) are encoded by a circadian expressed transcript (Fig. 6A; Suppl. Table S1), which is more than the ~30% that would be expected by chance. A gene ontology (GO) analysis (Mi et al., 2017) of the 127 SUMO targets that have circadian transcripts returned the protein classes “transcription factor” and “nucleic acid binding” as the most frequent (Fig. 6B). A GO-term enrichment analysis comparing the 127 targets to the set of circadian transcripts returned helicase activity, negative regulation of transcription (DNA templated), negative regulation of gene expression, and chromatin and chromosome organization as some of the most enriched terms. Not surprisingly, cellular compartments within the nucleus were enriched, including nucleolus, chromosome, and nucleoplasm (Suppl. Table S2-S4). Interestingly, histone H2B is part of the overlapping data set. H2B is conserved in the eukaryote linage, and sumoylation was observed not only in plants but also in yeast and humans (Nathan et al., 2006; Vertegaal et al., 2006). Modification of H2B by ubiquitin E3 ligases is required for proper timekeeping in both Arabidopsis and mammals (Himanen et al., 2012; Tamayo et al., 2015). Mining existing data sets could point to more proteins on which regulatory sumoylation is functionally conserved or to proteins with a conserved function in timekeeping. However, not every clock-relevant protein is encoded by a rhythmically expressed transcript, nor does every rhythmically expressed transcript have a product that is involved in timekeeping. Detailed biochemical experimental studies will now be necessary to identify the target proteins through which sumoylation affects timekeeping and, more importantly, to characterize the effect that sumoylation has on the biochemical properties of these target clock proteins.
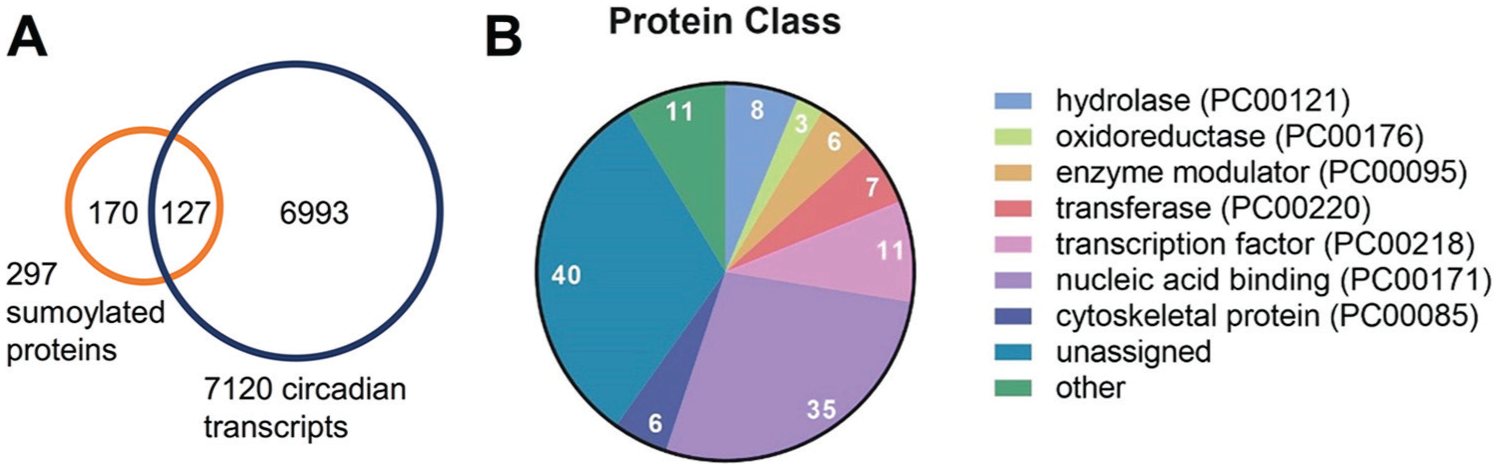
Overlap between sumoylation targets and circadian transcripts. (A) Venn diagram of overlap between sumoylated proteins identified by Miller et al. (2010) and all circadian transcripts in the dataset from Covington et al. (2008). (B) Protein classes associated with the overlapping data set assigned using Panther. Panther protein class IDs indicated in brackets.
Footnotes
Acknowledgements
The authors would like to thank Sarah Hodge for technical assistance with the Hamamatsu cameras, Michael Skelly and Gary Loake for the SCE1-OX line, and Karen Halliday for providing the circadian reporter line. This research was supported by the Royal Society research grants RS120372 and RS140275 and by the Royal Society University Research Fellowship UF110173 (all awarded to GvO).
Conflict of Interest Statement
The author(s) have no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Notes
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
