Abstract
In mammals, molecular circadian rhythms are generated by autoregulatory transcriptional-translational feedback loops with PERIOD/CRYPTOCHROME containing complexes inhibiting the transcription of their own genes. Although the major circadian oscillator components seem to be identified, an increasing number of additional factors modulating core clock component functions are being discovered. In a systematic screen using short hairpin RNA in human clock reporter cells, we identified FBXL11 (also known as KDM2A), a histone-demethylase, whose gene dosage is crucial for a correct circadian period. Knockdown of FBXL11 leads to period shortening and overexpression to period lengthening. In addition, altering FBXL11 gene dosage modulates clock gene transcript levels, most prominently that of Nr1d1. FBXL11 exercises its role in the mammalian circadian clock by acting as a negative element on CLOCK/BMAL1 and RORα-induced transcription. It binds directly to the promoter regions of CLOCK/BMAL1-regulated genes via a CXXC-type zinc finger motif in a circadian phase-dependent manner; however, the histone-demethylase activity of FBXL11 is not required for transcriptional repression. Therefore, we propose FBXL11 as a novel component of the circadian clock that regulates the circadian gene expression by a so far unknown mechanism.
The environmental conditions on earth change on a daily basis due to the earth’s rotation around its axis. The lighting conditions and subsequently temperature, food availability, predators, and many other environmental factors oscillate with a 24-h period. Organisms have developed the so-called circadian clock to be able to anticipate such changes. Circadian clocks can be found in organisms ranging from bacteria to eukaryotes (e.g., in plants, vertebrates, and mammals, including humans). They are self-sustained cell-autonomous oscillators that are synchronized with the environment (entrained) and persist under constant conditions. The circadian clock in mammals is a hierarchical system with the suprachiasmatic nucleus (SCN) as the master clock in the hypothalamus, which is entrained by light via innervations from the eyes, and tissue-specific clocks in the various peripheral organs receiving time cues from the SCN via neuronal and humoral signals (Dibner et al., 2010). On a molecular level, the circadian oscillator consists of interlocked transcriptional and translational feedback loops. The positive factors CLOCK (Circadian Locomotor Output Cycles Kaput) and BMAL1 (Brain and Muscle ARNT-Like 1) (Gekakis et al., 1998) bind as heterodimers to E-box enhancer elements in the promoters of Period (Per) and Cryptochrome (Cry) genes to stimulate their transcription. The gene products (PERs and CRYs) then act as negative elements to inhibit their own transcription after a delay of several hours. By this mechanism, an oscillation in transcript and protein levels is generated (Buhr and Takahashi, 2013; Partch et al., 2014). For a correct circadian period, posttranscriptional and posttranslational events are crucial to generate a delay between transcription of the negative elements and their repressive action. An indication of the important role of posttranscriptional events is the fact that only a fraction of oscillating transcripts seems to be due to circadian de novo transcription (Koike et al., 2012). Furthermore, posttranslational modifications such as phosphorylation and dephosphorylation (Gallego and Virshup, 2007; Gao et al., 2013; Kwak et al., 2013; Reischl and Kramer, 2011; Sahar et al., 2010), clock protein ubiquitination (for a review, see Stojkovic et al., 2014) and degradation (Busino et al., 2007; Reischl et al., 2007; Siepka et al., 2007), subcellular localization (Vanselow et al., 2006), and epigenetic regulation (Eckel-Mahan and Sassone-Corsi, 2013) have been reported to play an important role for circadian rhythm generation. Although the major clock components are identified, a rising number of additional factors that modify circadian rhythms are being discovered (e.g., Maier et al., 2009; Zhang et al., 2009).
In a systematic short hairpin RNA (shRNA)–based screen for novel clock components, we identified FBXL11 (F-box and leucine-rich repeat protein 11, also known as lysine-specific demethylase 2A [KDM2A]) as a putative novel clock component. Here, we characterize the role of FBXL11 in the mammalian circadian clock. The knockdown of Fbxl11 resulted in short period oscillations in human U2-OS cells. Although the previous functions of FBXL11 have all been attributed to its histone-demethylase activity, we found that FBXL11 acts on the circadian clock by repressing CLOCK/BMAL1-mediated transcription in a demethylase-independent manner that requires binding of FBXL11 to DNA.
Materials and Methods
Plasmids
To express FBXL11, a human complementary DNA (cDNA) clone was acquired from imaGenes (accession number BC064360, Berlin, Germany) and the coding sequence was cloned into mammalian expression vectors (pLenti6 or pDEST26; Invitrogen, Carlsbad, CA). For the FBXL11 variants, the Fbxl11 coding sequence was mutated in pENTR (Invitrogen) and subsequently shuttled into the respective expression vectors. All mutations were verified by sequencing.
FBXL11-H212A and FBXL11-H305A: the histidine residues (either at position 212 or 305) were mutated to alanine using the site-directed mutagenesis kit from Stratagene (La Jolla, CA).
FBXL11-CXXC: the 2 cysteine residues at position 574 and 577 were mutated to alanine using the site-directed mutagenesis kit from Stratagene.
FBXL11-dF: the F-box (positions 889-936) was removed by a deletion of 345 nucleotides (amino acids 867-981).
FBXL11-PHD: the 2 cysteine residues at positions 620 and 623 were mutated to alanine using the site-directed mutagenesis kit from Stratagene (analogous to Koyama-Nasu et al., 2007).
FBXL11-LRR: to delete the C-terminal leucine-rich repeats (LRRs), the C-terminus (amino acids 1000-1162) was deleted.
Antibodies
The anti-FBXL11 antibodies used were from Abcam (ab27867 and ab31739; Abcam, Cambridge, UK), the anti–β-actin antibody was from Sigma (St. Louis, MO), the anti-V5 antibody was from Invitrogen, and the normal mouse IgG and all secondary antibodies were from Santa Cruz Biotechnology (Santa Cruz, CA).
shRNA
shRNA constructs were either purchased from Open Biosystems (GE Healthcare, Little Chalfont, United Kingdom) or generated using pGIPZ as a lentiviral backbone. The nonsilencing control construct used was from Open Biosystems. The sequences in Fbxl11 targeted by shRNA were as follows:
anti-fbxl11_1 ACATTCACAAAGTTCTCTG
anti-fbxl11_2 GCCCTCAGTGGCATCATCAAG
anti-fbxl11_3 GACGAGAAGCTGATACAGAAG
Virus generation, transduction, and selection were performed as described in Maier et al. (2009).
Tissue Culture
Cells were maintained in Dulbecco’s modified Eagle’s medium (DMEM) supplemented with 10% fetal bovine serum and antibiotics (100 µg/mL penicillin and streptomycin). For selection of transduced cells, either puromycin (10 µg/mL) or blasticidin (10 µg/mL) was added.
Luminescence Recording
Luminescence recording in reporter cells was performed either in a TopCount luminometer (PerkinElmer, Waltham, MA) or in a Lumicycle (Actimetrics, Wilmette, IL) as described earlier (Maier et al., 2009). Reporter cells were U2-OS cells stably expressing luciferase under the control of a Bmal1-promoter fragment (Maier et al., 2009) or a Period2-promoter fragment (Zhang et al., 2009). Knockdown of Fbxl11 was performed with pGIPZ-anti-fbxl11 (sequences see above) and the Open Biosystems pGIPZ nonsilencing vector as a control. Overexpression of FBXL11 was achieved by stably transfecting the reporter cells with pLenti6-FBXL11; pLenti6-GFP was used as an overexpression control. Cells were synchronized with a single dexamethasone pulse (1 µM) for 20 min prior to starting the luminescence recording. Raw data were detrended by dividing by the 24-h running average. Periods were estimated by fitting the cosine wave function using the ChronoStar software (Stefan Lorenzen, Bernhard-Nocht-Institut, Hamburg, Germany) as described in Maier et al. (2009).
For analyzing the effect of FBXL11 dosage (overexpression or knockdown of FBXL11), we transfected U2-OS cells with overexpression (pLenti6-FBXL11 and pLenti6-GFP) or knockdown constructs (pGIPZ-anti-fbxl11_1 and pGIPZ-shRNA-control) in varying ratios in a 96-well plate. The total virus load was kept equal in all experiments. The day after transfection, the medium was exchanged for antibiotic selection medium. After 5 to 10 days, the cells were synchronized and put into the TopCount luminometer for measurement for 7 days.
Cotransactivation Assays
Cotransactivation assays were performed in HEK293 cells as described previously in Vanselow et al. (2006), with a pGL3 plasmid containing 6 E-boxes as a reporter. The 2.0-kb Per1-promoter reporter construct is described in Sangoram et al. (1998), the 0.9-kb Bmal1-promoter reporter construct in Maier et al. (2009), and the 1.7-kb Nr1d1-promoter reporter construct in Gervois et al. (1999). The dual luciferase system (Promega, Madison, WI) was used for normalization.
One-Hybrid Assay
The 1-hybrid assay was performed essentially as the cotransactivation assay with pGL3 containing 6 E-boxes as a reporter. For the VP16 fusion protein, the Vp16 coding sequence was amplified from pMV2 (Sadowski, 1998) and cloned into the pDEST26 vector (Invitrogen) upstream of the att-integration sites used for shuttling different expression sequences. The result was an FBXL11 protein (wild-type or variant) with an N-terminal fusion to VP16.
RNA Extraction and Quantitative Polymerase Chain Reaction
RNA was extracted with the Pure Link RNA Mini Kit (Ambion, Life Technologies Corporation, Carlsbad, CA) according to the manufacturer’s protocol. Then, 2 µg RNA was reverse transcribed and used for quantitative polymerase chain reaction (PCR) in a CFX96 machine (Bio-Rad, Hercules, CA). For quantitative PCR, Quantitect primers (QIAGEN, Valencia, CA) were used except for Gapdh (hGAPDH_fwd TGCACCACCAACTGCTTAGC, hGAPDH_rev ACAGTCTTCTGGGTGGCAGTG) and Fbxl11 (hFBXL11_fwd ACCATCCCCATTACGAAGCC, hFBXL11_rev ACACCACACTCTCCTTGCAC). Quantitect primers were as follows: hBmal1, QT00011844; hClock, QT00054481; hCry1, QT00025067; hCry2, QT00094920; hNr1d1, QT00000413; hPer1, QT00069265; and hPer2, QT00011207.
Chromatin Immunoprecipitation
U2-OS BLH cells overexpressing either GFP or FBXL11 were grown in 15-cm dishes in DMEM containing the appropriate antibiotic for selection. When the cells reached confluency, the cells were synchronized with a single pulse of dexamethasone (1 µM) for 20 min. Plates were harvested 22 or 32 h after synchronization. Cells were washed twice with phosphate-buffered saline (PBS) and fixed with formaldehyde (1% in PBS) for 10 min at room temperature. Cells were again washed twice with PBS before they were scraped in 10 mL buffer 0 (100 mM Tris/HCl, 10 mM dithiothreitol [DTT], pH 9.4) and transferred to a 15-mL tube. After centrifugation at 2000 × g, 4 °C, for 10 min, the supernatant was discarded and the pellet resuspended in 1 mL buffer 1 (10 mM HEPES, 0.25% Triton-X-100, 10 mM EDTA, 0.5 mM EGTA, pH 6.5) and again pelleted by centrifugation. The supernatant was discarded and the pellet was resuspended in 1 mL buffer 2 (10 mM HEPES, 200 mM NaCl, 1 mM EDTA, 0.5 mM EGTA, pH 6.5) and pelleted by centrifugation. The supernatant was discarded and the pellet was resuspended in 0.5 mL lysis buffer (50 mM Tris/HCl, 1% sodium dodecyl sulfate [SDS], 10 mM EDTA, pH 8.1). The chromatin was then sonified (5 times for 15 sec), and insoluble particles were pelleted by centrifugation (13,000 rpm, 10 min). The chromatin supernatant was diluted 8-fold in chromatin dilution buffer (1.1% Triton-X-100, 2 mM EDTA, 20 mM Tris/HCl, 150 mM NaCl, pH 8.1) and used for immunoprecipitation. For this step, maximum-recovery tubes (Axygen, Union City, CA) were used. For the immunoprecipitation, 500 µL diluted chromatin was incubated with 1.5 to 3 µg antibody on a rotation wheel at 4 °C for 90 min. Then, 30 µL protein A-agarose (Santa Cruz Biotechnology) was added per reaction and incubated for another hour on the rotating wheel at 4 °C. The bead pellet was then subsequently washed with 500 µL TSEI (150 mM NaCl, 2 mM EDTA, 0.1% SDS, 1% Triton-X-100, 20 mM Tris/HCl, pH 7.5), TSEII (500 mM NaCl, 2 mM EDTA, 0.1% SDS, 1% Triton-X-100, 20 mM Tris/HCl, pH 7.5), TSEIII (1 mM EDTA, 0.5 Tergitol NP40, 0.5% Na-deoxycholate, 250 mM LiCl, 10 mM Tris/HCl, pH 7.5), and TSE IV (1 mM EDTA, 150 mM NaCl, 10 mM Tris/HCl, pH 7.5). The pellet was resuspended in 50 µL crosslink-reversal buffer (1% SDS, 150 mM NaCl, 2 mM EDTA, 20 mM Tris/HCl, pH 7.5) and incubated at 65 °C overnight to reverse the formaldehyde crosslink. The next day, the DNA was purified with the PCR Purification Kit (Invitec, Berlin, Germany) according to the manufacturer’s protocol and used for quantitative PCR afterward. The primers used for the quantitative PCR targeted the Gapdh-promoter (hGAPDH_TATAfw–GCGCCCCCGGTTTCT, hGAPDH_TATA re–CCTAGCCTCCCGGGTTTC) and the Nr1d1-promoter (hNR1D1 for–CCTTCTCTGGACTTTGCCCT, hNR1D1 rev–AAACCTTGCAAACGTGAGGG).
Results
FBXL11 Is Required for Normal Circadian Rhythms
To identify novel components of the mammalian circadian clock, especially those involved in the posttranslational regulation, we performed a systematic screen using a lentiviral shRNA library to knock down kinases, phosphatases, and F-box proteins of the human genome in human osteosarcoma cells (U2-OS) stably expressing luciferase under the control of a Bmal1-promoter fragment (Maier et al., 2009). One of the candidate genes we identified was Fbxl11 (Figure 1A). FBXL11 (also known as KDM2A, lysine-specific demethylase 2A) is a histone-demethylase that removes methyl groups from dimethylated (and, to a lesser extent, monomethylated) lysine 36 from histone H3 (Tsukada et al., 2006). Knockdown of Fbxl11 resulted in period shortening of up to 2 h depending on the shRNA construct, the dosage of the knockdown construct, and the reporter cell line (Figure 1 and Suppl. Figure S1). Period shortening upon Fbxl11 knockdown was observed with 3 independent shRNA constructs (Figure 1B) with different knockdown efficiencies (Figure 1C), indicating the effect is specific for Fbxl11 knockdown and not an off-target effect. In another experiment, the shRNA construct with the strongest knockdown efficiency resulted in the strongest period-shortening phenotype (Suppl. Fig. S1A,B). In addition, the period-shortening phenotype was dose dependent (Figure 1D). We could reproduce the period-shortening phenotype in U2-OS cells stably expressing luciferase under the control of a Period2-promoter fragment in a dose-dependent manner (Suppl. Figure S1A,B), indicating that the observed effect is not specific for the Bmal1-luciferase reporter system.
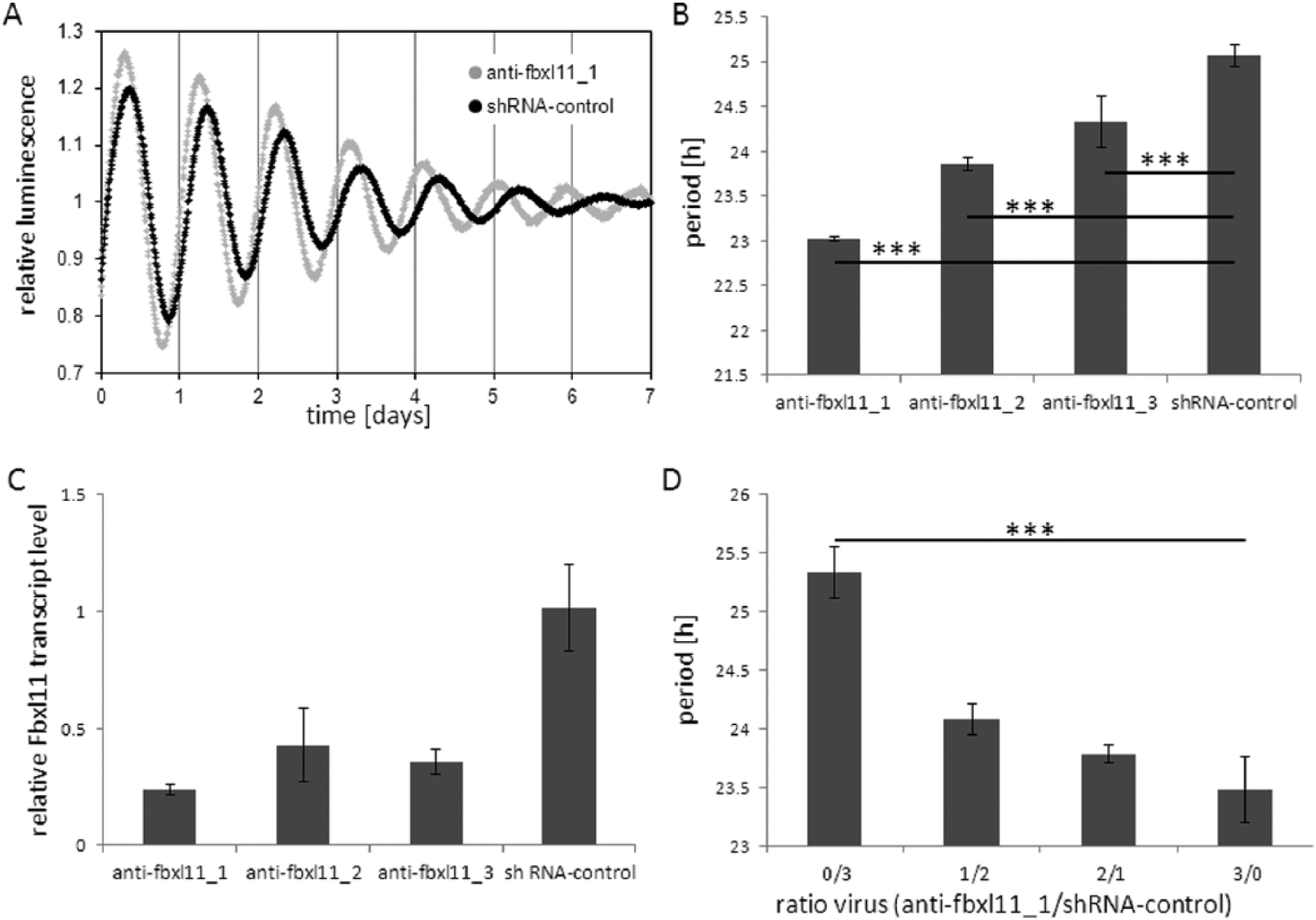
Knockdown of Fbxl11 shortens the circadian period. (A) The knockdown of Fbxl11 with short hairpin RNA (shRNA) (gray) leads to a shortened circadian period compared to an shRNA control (black) in human osteosarcoma cells (U2-OS) stably transfected with a Bmal1-luciferase reporter construct. Detrended representative time series are shown. (B) Period shortening upon Fbxl11 knockdown is dependent on the shRNA construct. p < 0.0001 (1-way analysis of variance), ***p < 0.0001 (Dunnett’s multiple-comparison post hoc test). (C) Knockdown efficiency of the 3 shRNA constructs used in B. (D) Period shortening is dose dependent. The data are highly significant (Kruskal-Wallis test, Dunn’s multiple-comparison post hoc test, ***p < 0.0001). Given is the mean ± SD (n = 4 for B and C, n = 16-20 for D).
To modify the level of FBXL11 in the opposite direction, we overexpressed FBXL11 in U2-OS reporter cells using a lentiviral vector. Overexpression of FBXL11 but not of GFP as a control led to lengthening of the circadian period in a dose-dependent manner (Figure 2A,B). Together, this indicates that the FBXL11 gene dosage is crucial for normal circadian rhythmicity.
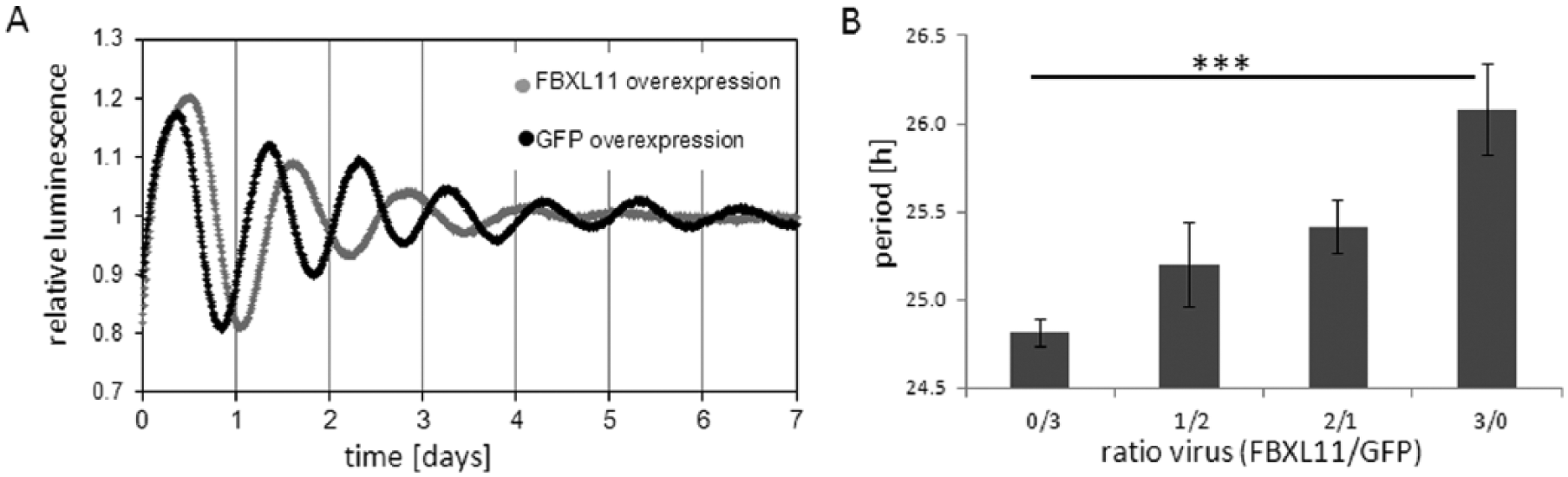
Overexpression of FBXL11 leads to a lengthening of the circadian period. (A) FBXL11 overexpression in U2-OS reporter cells lengthens the circadian period. Detrended representative time series for FBXL11 overexpression (gray) and control GFP overexpression (black) are depicted. (B) Period lengthening by FBXL11 overexpression is dose dependent as increasing ratios of FBXL11/GFP overexpression led to a gradual period lengthening. (Kruskal-Wallis test, Dunn’s multiple-comparison post hoc test, ***p = 0.006). Given is the mean ± SD (n = 5-6).
FBXL11 Levels Influence Endogenous Clock Gene Expression
To investigate how the level of FBXL11 in a cell modulates the endogenous circadian period, we analyzed clock gene transcript levels in synchronized and unsynchronized cells upon Fbxl11 knockdown and overexpression. In unsynchronized Fbxl11 knockdown cells, with a knockdown efficiency of 99 %, most of clock gene mRNA levels were unchanged or only slightly upregulated, except Nr1d1, which showed twice the amount of mRNA in Fbxl11 knockdown cells compared to cells transfected with a nonsilencing shRNA control. In cells overexpressing FBXL11 approximately 20-fold, only small alterations in the levels of clock gene mRNA could be observed. Genes that were upregulated upon Fbxl11 knockdown were downregulated upon FBXL11 overexpression, but these effects were only subtle (Figure 3A). To analyze clock gene expression in oscillating cells, U2-OS reporter cells with Fbxl11 knockdown or FBXL11 overexpression were synchronized with dexamethasone, and samples were taken every 4 h for 48 h and transcript levels of Per1, Bmal1, Dbp, and Nr1d1 were analyzed. We could detect only low-amplitude oscillations for transcripts of Per1, Bmal1, and Dbp, and thus it was difficult to assess a potential effect of Fbxl11 gene dosage on these rhythms (data not shown). This does not exclude an effect of the level of FBXL11 on the expression of these genes, but for further analysis, we focused on Nr1d1 expression, as both the amplitude of Nr1d1 transcript oscillation and the changes observed upon altering Fbxl11 gene dosage were substantial. Nr1d1 mRNA showed a clear circadian oscillation that was changed significantly upon knockdown or overexpression of FBXL11 with the direction of changes as expected from the steady-state experiments (increased levels of Nr1d1 mRNA in Fbxl11 knockdown and decreased levels of Nr1d1 mRNA in FBXL11 overexpression). When Fbxl11 levels were reduced (Figure 3B, left panel), the level of Nr1d1 transcript was elevated at the peak times, while Nr1d1 trough levels were unchanged, suggesting an effect on the transcription or accumulation of Nr1d1 mRNA. Overexpression of FBXL11 led to a reduced level of Nr1d1 mRNA at peak times (Figure 3B, right panel), while again trough levels were unaffected. Together, the results from steady-state and oscillating cells point toward a role of FBXL11 in regulating the expression of clock genes such as Nr1d1 either directly by inhibiting transcription or indirectly by modulating other clock components.
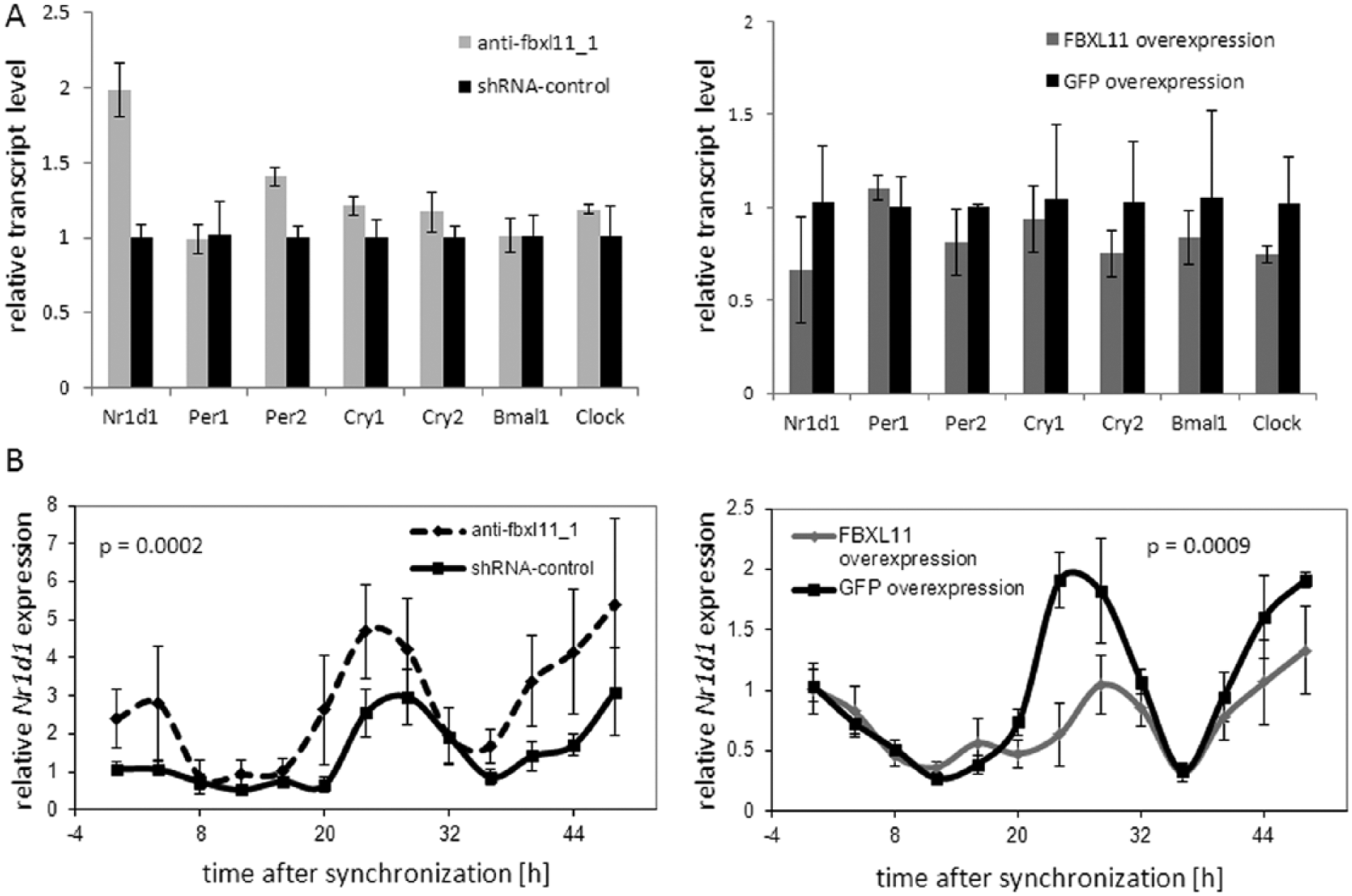
FBXL11 levels influence endogenous clock gene expression. (A) Knockdown or overexpression of FBXL11 leads to an altered amount of clock gene transcripts, in particular Nr1d1 in unsynchronized U2-OS Bmal1-promoter reporter cells. The Fbxl11 knockdown efficiency was 99% and FBXL11 overexpression was 20-fold. Given are the mean ± SD (n = 3). (B) Nr1d1 transcript levels are altered upon Fbxl11 knockdown (dashed line) or FBXL11 overexpression (gray line) in U2-OS reporter cells after dexamethasone synchronization with respect to a short hairpin RNA (shRNA) control (black, left panel) or GFP overexpression (black, right panel). Given are the mean ± SE (n = 3-6). Alterations in Nr1d1 transcript-level oscillations were significant for Fbxl11 knockdown (p = 0.0002, 2-way analysis of variance [ANOVA]) and FBXL11 overexpression (p = 0.0009, two-way ANOVA).
FBXL11 Can Inhibit CLOCK/BMAL1-Mediated Transcription
To analyze whether FBXL11 can influence the transcriptional activity of CLOCK/BMAL1, we performed cotransactivation assays in HEK 293 cells. To this end, a reporter construct–expressing luciferase under the control of an artificial promoter containing 6 E-boxes was used. When CLOCK and BMAL1 are cotransfected, transcription is increased strongly, while addition of wild-type FBXL11 reduces CLOCK/BMAL1-mediated transcription dramatically. Thus, FBXL11 can act as a repressor of CLOCK/BMAL1-mediated transcription (Figure 4A). FBXL11 is a protein composed of 1162 amino acids with a variety of functional domains (Figure 4B). At the N-terminus of FBXL11, there is a JmjC domain, known to be responsible for the histone-demethylase activity described for FBXL11. FBXL11 can demethylate mono- and dimethylated K36 of histone H3 (Klose et al., 2006). By mutating a single histidine to alanine (H305A or H212A), this activity can be abolished (Tsukada et al., 2006). FBXL11 also contains 2 zinc fingers, one of the CXXC type known to bind to DNA, and a PHD type. The F-box and LRRs at the C-terminus are additional functional domains. To analyze which domains are necessary for the repressor function, mutation and deletion variants of FBXL11 were generated and used in the CLOCK/BMAL1 cotransactivation assay (Figure 4A). The analysis of the FBXL11 variants revealed a prominent role for the CXXC-type zinc finger in repressing CLOCK/BMAL1-mediated transcription. By mutating the 2 cysteine residues of the CXXC motif to alanine residues, the repressor function of FBXL11 was completely abolished. The JmjC, F-box, and PHD domains did not affect the repressor activity, although the expression levels of the single FBXL11 variants were comparable (Suppl. Figure S2). There may be a role for the LRR region at the C-terminus, as a deletion of this region resulted in reduced repressor function of FBXL11. As LRRs are often involved in protein-protein interactions, one can speculate that another protein (probably a corepressor) is needed for FBXL11 to exhibit its full repressor function. The strong negative action of FBXL11 and the prominent role of the CXXC domain were also observed with a Per1-promoter fragment induced with CLOCK/BMAL1 and an Nr1d1- and Bmal1-promoter fragment induced by RORα (Suppl. Figure S3). This negative effect on transcription at different promoter fragments may be caused by direct, sequence-unspecific binding of FBXL11 to DNA since it is reported to bind unmethylated CpG islands in the promoter regions of mammalian genes (Blackledge et al., 2010). A direct interaction with CLOCK/BMAL1 or RORα is unlikely as both CLOCK/BMAL1- and RORα-induced transcription was repressed by FBXL11 (Figure 4 and Suppl. Figure S3), and a putative interaction site of FBXL11 is probably not able to interact with such diverse proteins as CLOCK/BMAL1 and RORα.
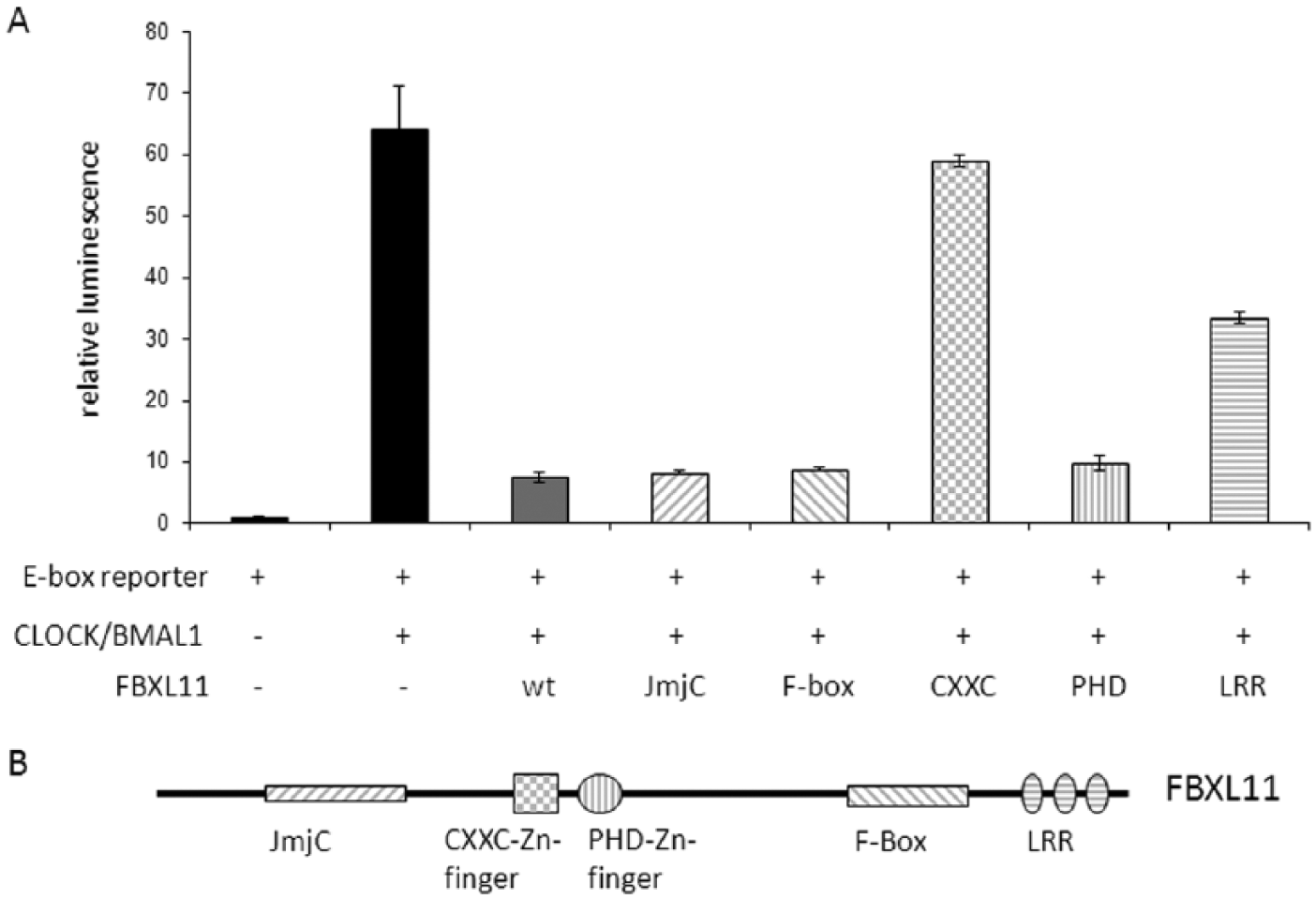
FBXL11 inhibits CLOCK/BMAL1 transactivation. (A) In a cotransactivation assay performed in HEK 293 cells with ectopically expressed CLOCK and BMAL1 as well as an artificial reporter containing 6 E-boxes upstream of the luciferase gene, FBXL11 acted negatively on CLOCK/BMAL1-mediated transactivation. Mutation and deletion analysis showed that the CXXC zinc finger is essential for this negative effect and that leucine-rich repeats (LRRs) at the C-terminus also contribute to FBXL11 repressor activity. For the normalization of luminescence, a dual luciferase system was used. Given is the mean ± SD (n = 3). (B) FBXL11 is a protein with many functional domains. It contains a histone demethylase (JmjC) domain, a CXXC- and a PHD-type zinc finger, an F-box domain, and C-terminal LRRs.
FBXL11 Binds DNA in a Time-of-Day Dependent Manner
From the previous experiments, we hypothesized that FBXL11 exercises its negative effect on CLOCK/BMAL1-mediated transcription via direct DNA binding. To test this, a mammalian 1-hybrid system was used: a fusion protein of FBXL11 and the viral transactivation domain VP16 was generated. As a reporter, again 6 E-boxes driven by the SV40-promoter upstream of the luciferase gene were used. When the fusion protein binds to DNA, the VP16 domain is positioned in the vicinity of the transcriptional start site, and transcription of luciferase is expected to be increased (Figure 5A). This is indeed the case for wild-type FBXL11 (Figure 5B) but not when the CXXC zinc finger is mutated, indicating a direct FBXL11 binding to DNA via its CXXC zinc finger motif.
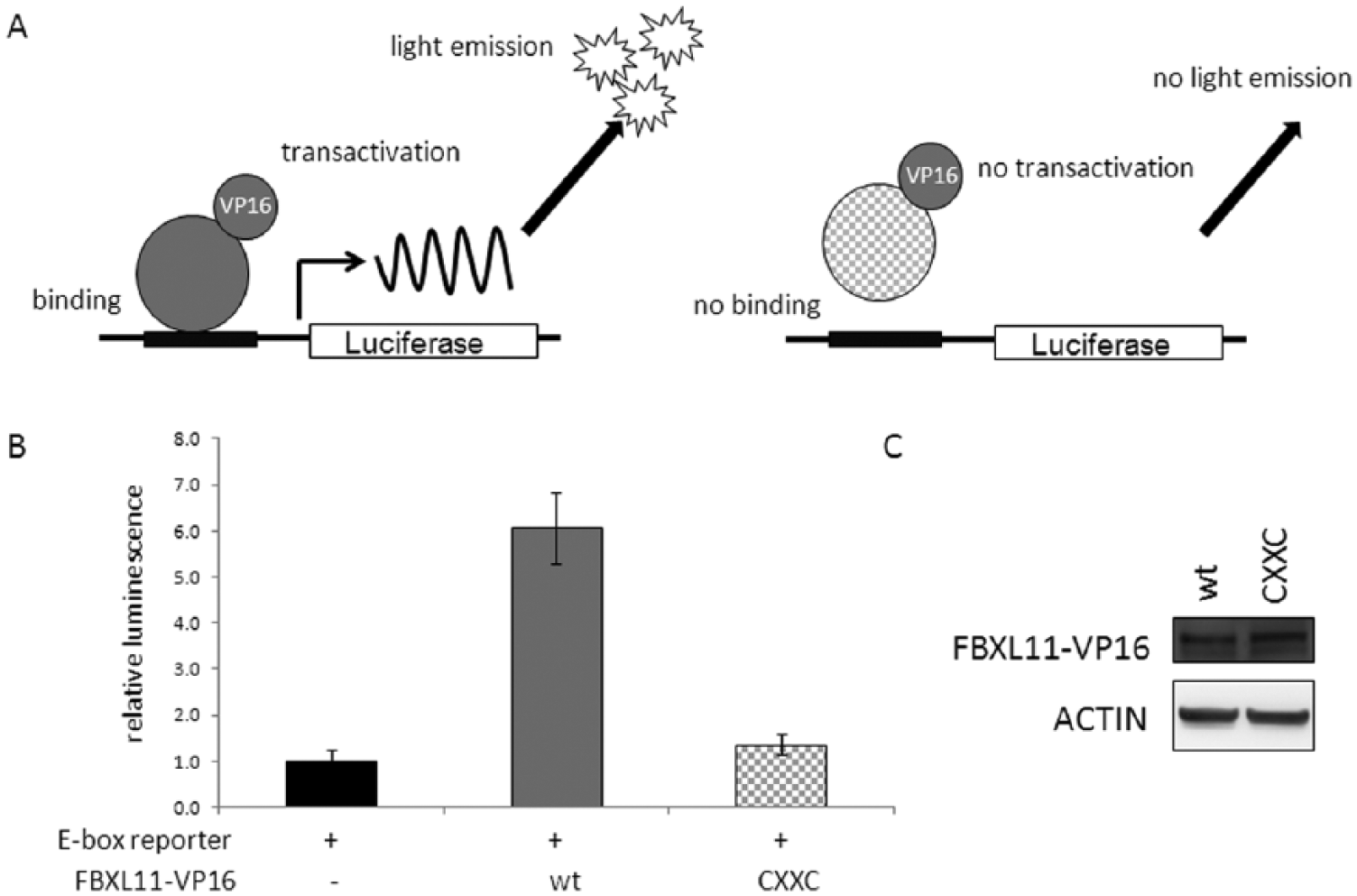
FBXL11 binds to DNA. (A) A 1-hybrid system was used to show the DNA-binding ability of FBXL11. FBXL11 wild-type or CXXC variant was fused to the viral VP16 transactivation domain. When the fusion protein bound to DNA via FBXL11, VP16 was positioned in the vicinity of the transcriptional start site and luciferase was transcribed. The reporter plasmid contained 6 E-boxes and a cytomegalovirus (CMV) promoter upstream of the luciferase gene. (B) FBXL11-VP16 wild-type was able to transactivate, whereas FBXL11-CXXC was not, indicating that the CXXC zinc finger is essential for FBXL11 binding to DNA. Given is the mean ± SD (n = 3).
To elucidate a putative mechanism for FBXL11 in the circadian system, we analyzed DNA binding of FBXL11 at different circadian phases in oscillating cells. To this end, we used synchronized U2-OS reporter cells stably overexpressing V5-tagged FBXL11 and performed chromatin immunoprecipitation to analyze FBXL11 binding at the Nr1d1-promoter 22 and 32 h after dexamethasone synchronization. These 2 time points represent the rising and declining phases of Nr1d1 transcript oscillation (see Figure 3B, right panel). We hypothesized that if the differential effects on Nr1d1 transcript level, which we detected at these time points, were indeed directly due to FBXL11, we should be able to see corresponding differences in FBXL11-DNA binding. The chromatin immunoprecipitation data showed FBXL11 binding to the Nr1d1-promoter to be lower when expression of Nr1d1 is rising (22 h), compared to when expression is declining (32 h) (Figure 6). Given the low number of data points (n = 4-5), the difference did not reach statistical significance. In contrast, binding to the Gapdh-promoter was equal at both time points, suggesting that FBXL11 modulates clock gene (at least Nr1d1) transcription via time-of-day dependent DNA binding.
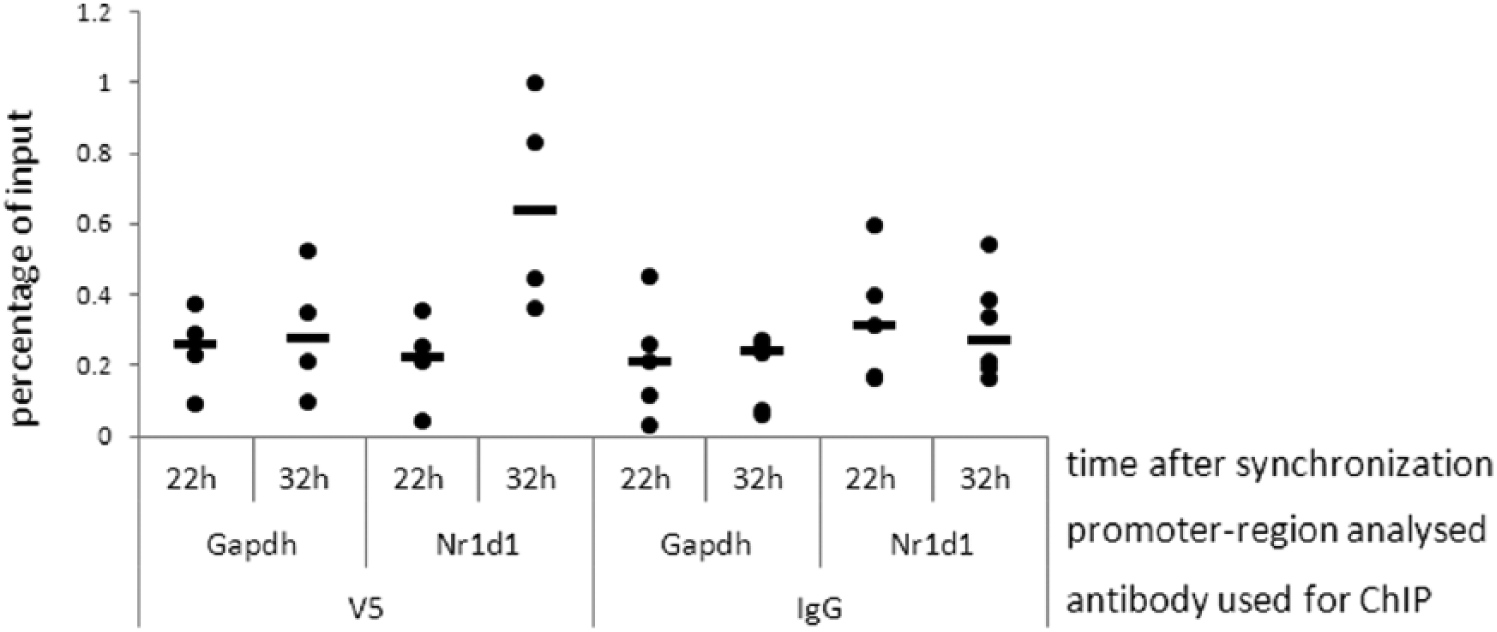
FBXL11 seems to bind to the Nr1d1-promoter in a circadian manner. U2-OS reporter cells overexpressing FBXL11-V5 were synchronized with dexamethasone and harvested 22 and 32 h after synchronization. Chromatin immunoprecipitation experiments were performed with anti-V5 antibody or normal mouse IgG as a control. Binding of FBXL11 to the Nr1d1- and Gapdh-promoter was analyzed with quantitative polymerase chain reaction. Given are the individual data points and the median (black bar). Statistical analysis: due to the low number of data points (n = 4-5), we were unable to determine a statistically significant difference in FBXL11 binding to the Nr1d1-promoter at 22 h and 32 h, but the data clearly show a trend toward increased binding of FBXL11 to the Nr1d1-promoter at 32 h.
Discussion
Here we identified FBXL11 as a novel negative element in the mammalian circadian clock. Depletion of Fbxl11 results in short-period oscillations, similar to a previously described histone-demethylase JMJD5 (Jones et al., 2010), which has the same demethylation specificity (H3K36me2) but is structurally different from FBXL11 and does not contain a CXXC-type Zn finger (Labbé et al., 2013). We therefore think the mechanism of action is different in the 2 histone-demethylases.
The observed short-period phenotype is typical for knockdown of a negative element in the circadian oscillator; for example, Cry1 knockout mice exhibit a shorter free-running period (van der Horst et al., 1999), and CIPC depletion (CLOCK-Interacting Protein, Circadian—a negative feedback regulator of the mammalian clock) causes period shortening (Zhao et al., 2007). In addition, using a mass-spectrometric approach to identify PER-interaction proteins, the group of C. Weitz (Harvard Medical School) and others described several novel members of the PER/CRY-inhibitory complex that cause shortened circadian periods when depleted: (1) PSF (polypyrimidine tract–binding protein-associated spicing factor) recruits SIN3A (SIN3 transcription regulator family member A) to the inhibitor complex, and depletion of both factors leads to period shortening (Duong et al., 2011). (2) The RNA helicases DDX5 (DEAD-box polypeptide 5) and DHX9 (DEAH-box protein 9) are part of the PER/CRY-inhibitory complex at the circadian time of inhibition (CT20), and depletion of either of the helicases shortens the circadian period (Padmanabhan et al., 2012). (3) The histone-methyltransferase HP1γ-Suv39h is also found in the PER-inhibitor complex. Depletion of Suv39h1 and Suv39h2 results in period shortening (Duong and Weitz, 2014). (4) Depletion of the RNA-binding protein NONO, which is part of the PER/CRY-inhibitory complex, shortens the circadian period (Brown et al., 2005).
Although the short-period phenotype for Fbxl11 depletion and the inhibitory action on CLOCK/BMAL1-mediated transcription put FBXL11 in the group of negative elements, it seems not to be part of the PER/CRY-inhibitory complex because (1) FBXL11 was not detected in a mass-spectrometric analysis of the PER2 repressor-complex (C. Weitz, Harvard Medical School, personal communication, 2010), and (2) a previous report indicates that FBXL11 does not exert its role in the circadian clock by protein-protein interaction with the core clock components—in coimmunoprecipitation experiments, no interaction could be detected between FBXL11 and the core clock components CRY1, CRY2, PER1, PER2, TIM1, BMAL1, CLOCK, and NR1D1 (Busino et al., 2007). (3) From our results, it is unlikely that FBXL11 is directly binding to the positive elements of the core clock and thereby inhibiting transcription, as the negative effect could be observed with both CLOCK/BMAL1-induced and RORα-induced transcription (see Suppl. Figure S3).
Therefore, we propose a mechanism of action for FBXL11 in the mammalian circadian system that is outside the PER/CRY-inhibitory complex and is also novel in the mode of action for FBXL11. So far, only functions of FBXL11 involving its demethylase activity have been described. It was reported that FBXL11 binds to unmethylated CpG islands in the promoter region of eukaryotic genes via a CXXC-type zinc finger motif acting there on histone methylation and thereby creating a different chromatin environment at unmethylated CpG islands to separate them from bulk chromatin (Blackledge et al., 2010). FBXL11 can repress the expression of ribosomal RNA (rRNA) in a JmjC-dependent manner by binding to the rRNA promoter and demethylating histone H3K36 (Tanaka et al., 2010). Apart from the action of FBXL11 on histone methylation, a role in demethylating and thereby repressing the activity of a nonhistone protein, nuclear factor (NF)–κB, is described (Lu et al., 2010). In contrast to this, the negative role of FBXL11 in the circadian system is not dependent on the histone-demethylase activity of FBXL11 as FBXL11-variants with nonfunctional JmjC domains are still able to repress CLOCK/BMAL1-mediated transcription. A homologue of FBXL11, FBXL10, which is also a functional histone-demethylase, can act in a JmjC-independent manner in repressing transcription of cJun. FBXL10 interacts with cJUN, binds to DNA (e.g., the cJun-promoter) via the CXXC domain, and represses transcription via interaction with SIN3A and a histone-deacetylase (Koyama-Nasu et al., 2007). Similar to FBXL10, FBXL11 binds to DNA via the CXXC domain and represses transcription in a JmjC-independent manner.
However, mechanistic details about the mode of FBXL11 action as well as putative interaction partners of FBXL11 are still unknown. As SIN3A is also implicated to have a role in regulating the circadian clock, and as knockdown of Sin3a with RNA interference led to moderate period shortening (Duong et al., 2011), we tested for an interaction between FBXL11 and SIN3A with coimmunoprecipitation but could not detect interaction with this method (data not shown). Another histone-demethylase, JARID1a, is known to be involved in circadian clock regulation. JARID1a forms a complex with CLOCK/BMAL1 and enhances transcription by inhibiting histone-deacetylase 1 in a JmjC-independent manner (DiTacchio et al., 2011). To test the hypothesis that FBXL11 exerts its negative function via the recruitment of a histone-deacetylase, we analyzed whether the effect of FBXL11 on CLOCK/BMAL1-mediated transcription in a cotransactivation assay could be abolished by the histone-deacetylase class I and II inhibitor trichostatin A. We found that the action of FBXL11 in this system is independent of the addition of trichostatin A (data not shown). Therefore, the mode of action of FBXL11 in the circadian system remains unclear. It is likely that FBXL11 binds to the Nr1d1-promoter (and possibly other circadian promoters) via its CXXC domain and acts as a repressor by recruiting a so far unknown factor. Binding of FBXL11 to the Nr1d1-promoter is rhythmic, but the expression of Fbxl11 in U2-OS cells or mouse liver does not oscillate in a circadian manner (data not shown). Therefore, one could assume DNA binding of FBXL11 as a hetero-dimer with the unknown factor and rhythmic abundance of the unknown factor. In favor of the “unknown-factor hypothesis” is the fact that deletion of the C-terminal LRR domain partly reduces FBXL11 repressor activity (see Figure 4A) and that LRRs are frequently involved in protein-protein interactions (Kobe and Kajava, 2001).
Footnotes
Acknowledgements
We thank Astrid Grudziecki for technical assistance, Ute Abraham for help with statistical analyses, Bert Maier for sharing unpublished data, Gerard Triquenaux (ENS Lyon, France) for supplying the Nr1d1 reporter construct, and Ivan Sadowski (University of British Columbia, Vancouver, Canada) for supplying a VP16-containing plasmid. This work was funded in part by the German Research Foundation (DFG, grant SFB 740/D2).
Conflict of Interest Statement
The author(s) have no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Notes
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
