Abstract
Cryptochromes and photolyases belong to the same family of flavoproteins but, despite being structurally conserved, display distinct functions. Photolyases use visible light to repair ultraviolet-induced DNA damage. Cryptochromes, however, function as blue-light receptors, circadian photoreceptors, or repressors of the CLOCK/BMAL1 heterodimer, the transcription activator controlling the molecular circadian clock. Here, we present evidence that the functional divergence between cryptochromes and photolyases is not so univocal. Chrysodeixis chalcites nucleopolyhedrovirus possesses 2 photolyase-like genes: phr1 and phr2. We show that PHR1 and PHR2 are able to bind the CLOCK protein. Only for PHR2, however, the physical interaction with CLOCK represses CLOCK/BMAL1-driven transcription. This result shows that binding of photolyase per se is not sufficient to inhibit the CLOCK/BMAL1 heterodimer. PHR2, furthermore, affects the oscillation of immortalized mouse embryonic fibroblasts, suggesting that PHR2 can regulate the molecular circadian clock. These findings are relevant for further understanding the evolution of cryptochromes and photolyases as well as behavioral changes induced in insects by baculoviruses.
Photolyases are DNA damage repair enzymes that lesion-specifically and in a light-dependent manner remove cyclobutane pyrimidine dimers (CPD) or pyrimidine-(6,4)-pyrimidone photoproducts ((6-4)PP) from ultraviolet (UV)–exposed DNA in a process called photoreactivation (Müller and Carell, 2009; Eker et al., 2009). Photolyases, together with cryptochromes, belong to the cryptochrome/photolyase family (CPF) of flavoproteins. Cryptochromes were first identified as plant blue-light receptors (Cashmore et al., 1999). More recently, homologs of cryptochromes were identified in animals, where they act as circadian photoreceptor proteins (e.g., Drosophila and zebrafish) or core clock proteins (in mammals and Xenopus) (Kume et al., 1999; van der Horst et al., 1999; Cashmore, 2003). Additionally, cryptochromes were shown to be involved in magnetoreception in insects and birds (Gegear et al., 2010; Ritz et al., 2000). Interestingly, human CRY2 was recently shown to be involved in light-mediated magnetoreception in flies (Foley et al., 2011).
Cryptochromes and photolyases share a well-conserved core domain of about 500 amino acids, which binds 2 chromophores: flavin adenine dinucleotide (FAD) and either 5,10-methenyl-tetrahydrofolate (MTHF) or 8-hydroxy-7,8-didemethyl-5-deazariboflavin (8-HDF) (Fig. 1). Cryptochromes lack the nuclear localization signal-containing N-terminal extension characteristic for eukaryotic photolyases and are equipped with a unique C-terminal extension of variable length (Daiyasu et al., 2004; Eker et al., 2009). It has been proposed that the presence of different (species-specific) C-terminal extensions functionally separates cryptochromes from photolyases and that the unique character of these tails underlies the functional diversity among cryptochromes (Chaves et al., 2011b). While plant CRYs carry a long C-terminal tail, which is required for light function, Drosophila CRY has a short tail, which modulates the activity of the core. In the case of mammalian CRYs, it seems that function is dependent on a combined action between core and tail.
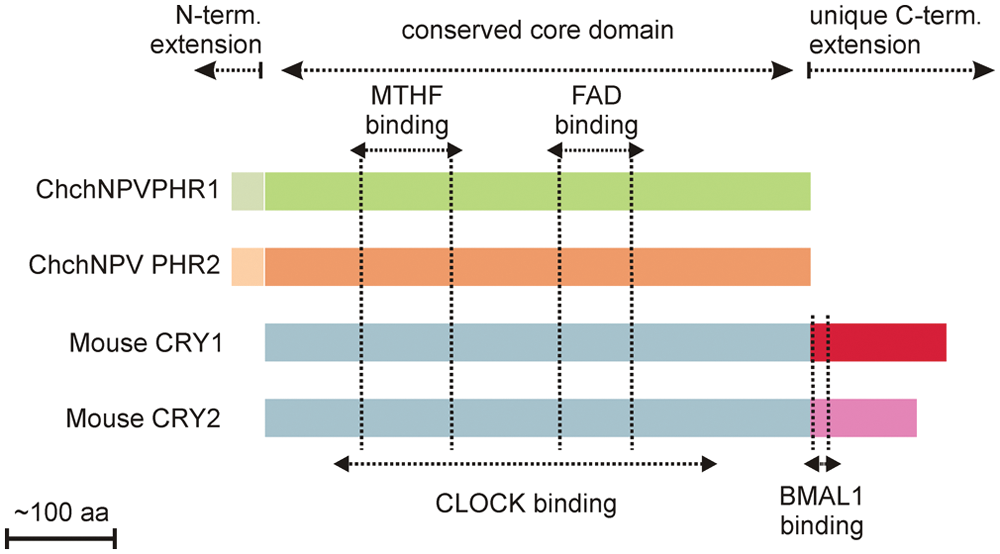
Comparison between PHRs and mammalian CRYs. Schematic representation of phr1, phr2, and mouse CRY1 and 2. Conserved and unique domains are indicated above; chromophore binding sites are indicated by vertical dotted lines. CLOCK and BMAL1 binding regions in mouse CRY are indicated below.
Mammalian cryptochromes (Cry1 and 2) are indispensable elements of the circadian clock (van der Horst et al., 1999) that allows most, if not all, organisms to anticipate to the environment by adjusting metabolism, physiology, and behavior to the momentum of the day. Circadian rhythms are generated by a self-sustained molecular clock, which has an intrinsic periodicity of approximately 24 hours and needs to be reset daily to keep pace with the day/night cycle as imposed by the rotation of the earth around its axis (Young and Kay, 2001). Although circadian clocks can respond to various cues (e.g., food availability, DNA damage), the main “zeitgeber” or cue is visible light (Dijk et al., 1995).
The circadian clock is based on an autoregulatory molecular oscillator in which a set of clock genes is periodically activated and repressed via interlocked positive and negative transcription/translation feedback loops (Reppert and Weaver, 2001; Shearman et al., 2000). In mammals, the main players in this molecular clockwork are the Clock, Bmal1, Rev-Erbα, Period (Per1 and 2), and Cryptochrome (Cry1 and 2) genes. The transcription activator heterodimer CLOCK/BMAL1 binds to E-box promoter elements (CACGTG) and turns on expression of the clock genes Cry, Per, and Rev-Erbα. Once synthesized, the CRY and PER proteins form complexes that inhibit CLOCK/BMAL1-driven transcription. In a second feedback loop, REV-ERBα inhibits the receptor tyrosine kinase–like orphan receptor (ROR)–driven transcription of Bmal1 (Preitner et al., 2002). In addition, posttranslational modification of clock proteins (e.g., phosphorylation, ubiquitylation, acetylation) controls the stability and subcellular localization of the clock proteins, thereby fine tuning the period length of the clock (Gallego and Virshup, 2007; Vanselow and Kramer, 2007). The circadian core oscillator is coupled to rhythmic output pathways via E-box– and ROR-containing clock-controlled genes (Reppert and Weaver, 2002; Takahashi et al., 2008).
The common ancestor of CPF members was likely a photolyase that made organisms more resistant against UV light (Cashmore et al., 1999). The photoreceptor function is developed later to help organisms sense day-night changes in order to avoid UV irradiation (Gehring and Rosbash, 2003). From phylogenetic analysis, it was deduced that animal cryptochromes are more closely related to (6-4)PP photolyases, whereas plant cryptochromes presumably evolved from CPD photolyases. This phylogenetic analysis, therefore, suggests that animal and plant cryptochromes arose by 2 independent evolutionary events (Cashmore et al., 1999). Members of the CPF are also encountered in various viruses and phylogenetically cluster with class II CPD photolyases (Eker et al., 2009). Baculovirus CPD photolyase–like genes were first described in Chrysodeixis chalcites nucleopoly-hedrovirus (ChchNPV) and were designated phr1 and phr2 (van Oers et al., 2004, 2005) (Fig. 1). These 2 photolyases belong to class II CPD photolyases (phylogeny in Eker et al., 2009; Biernat et al., 2011). PHR2 can rescue bacteria lacking a UV repair system, whereas PHR1 does not (van Oers et al., 2008). Further characterization indicated that the phr2 gene, in contrast to phr1, encodes a photolyase with DNA repair activity towards UV-induced CPDs.
The functional diversity and evolution of CPF members, however, are not understood entirely. The identification of photolyases with a dual function (i.e., circadian clock and DNA repair) suggests that these 2 functions are not mutually exclusive (Coesel et al., 2009; Heijde et al., 2010; Chaves et al., 2011a). Insects are known to change their behavior after baculovirus infection, but the underlying mechanisms are still not identified (Goulson, 1997). Identification of a possible clock regulatory function of PHR1 and/or PHR2 would take us a step further in understanding the baculovirus-induced behavioral changes. We investigated, therefore, whether PHR1 and PHR2 are able to interact with components of the mam-malian molecular clock and, consequently, have a transcription regulation function. We studied also the effect of baculoviral photolyases on the molecular oscillator and examined their possible function in the circadian clock.
Materials and Methods
Construction of Plasmids
Flag-tagged mouse Bmal1 and Clock expression constructs (further referred to as Bmal1-flag and Clock-flag) and the HA-mCry1 expression construct (all in pcDNA3) have been described previously (Chaves et al., 2006). To express ChchNPV phr1 and phr2 in mammalian cells, the complete coding region of both genes was PCR-amplified from genomic ChchNPV DNA using Phusion high-fidelity DNA polymerase (Finnzymes, Vantaa, Finland) and restriction site–flanked primers. The obtained HindIII-BamH1 phr1 and EcoRI-BamHI phr2 fragments were cloned into pcDNA3.1 (Invitrogen, Carlsbad, CA) to give pcDNA3.1-phr1 and pcDNA3.1-phr2. Essentially, the same PCR strategy was taken to generate constructs for the expression of C-terminally EGFP-tagged PHR, except that the reverse PCR primers were designed in such a way that the 3′ BamHI restriction site replaced the phr STOP codon. The obtained HindIII-BamH1 phr1 and EcoRI-BamHI phr2 fragments were first cloned into pJET (Fermentas, Vilnius, Lithuania) and subsequently recloned upstream of the egfp coding sequence of pEGFPN3 and pEGFPN1 (Clontech, Mountain View, CA), respectively. The pEGFPN3-phr1 and pEGFPN1-phr2 constructs are referred to as phr1-egfp and phr2-egfp.
Cells, Culture, and Transient Transfection Conditions
HEK 293T (human embryonic kidney) and NIH 3T3 (immortalized mouse embryonic fibroblast) cells were cultured at 37 °C and 5% CO2 in Dulbecco’s modified Eagle’s medium (DMEM)/Ham-F10 medium (1:1; Lonza, Basel, Switzerland) and supplemented with 10% fetal calf serum (Lonza), penicillin, and streptomycin. Transient expression of proteins was achieved by transfecting cells with the aforementioned constructs using FuGene reagent according to the manufacturer’s instructions (Roche, Basel, Switzerland).
Coimmunoprecipitation and Western Blot Analysis
For coimmunoprecipitation studies, HEK 293T cells transiently (co)expressing phr1-gfp or phr2-gfp, Bmal1-flag, and/or Clock-flag were lysed 24 hours after transfection in immunoprecipitation (IP) buffer, composed of 50 mM Tris-HCl, pH 7.5, 150 mM NaCl, 1% NP-40, and 1 mM EDTA, and supplemented with a complete protease inhibitor cocktail (Roche) and phosphatase inhibitors (Pierce, Rockford, IL) and sonicated. After centrifugation (20 minutes, 16,000g, 4 °C), a small amount of the lysate was stored (input), and the remainder was incubated with an anti-GFP antibody (Roche) for 3 hours at 4 °C under continuous rotation. After addition of G beads (GE Healthcare, Little Chalfont, UK), incubation was continued for another hour, after which the beads were washed twice with IP wash buffer (20 mM Tris-HCl, pH 7.5, 500 mM NaCl, 0.1% NP-40, and 0.05% sodium deoxycholate). Next, the beads and input samples were boiled in 2x Laemmli sample buffer and separated by SDS polyacrylamide gel electrophoresis in 4% to 12% gradient SDS-PAGE gels (Invitrogen). Subsequently, proteins were transferred to a nitrocellulose membrane and visualized using anti-Flag (1:1000 dilution; Sigma, St. Louis, MO) or anti-GFP (1:1000 dilution; Roche) primary antibodies. Horseradish peroxidase–conjugated anti-rabbit IgG (1:5000 dilution; Dako, Glostrup, Denmark) was used as a secondary antibody, and chemoluminescence was detected using ECL Western Blotting Reagents (GE Healthcare). HEK 293T cells were chosen because of their high efficiency of transfection. The use of NIH 3T3 cells does not allow a good detection of the proteins.
Immunofluorescence
HEK 293T cells were grown on cover slips in 12-well plates (105 cells/well) and (co)transfected with phr1-gfp, phr2, and/or flag-clock using Fugene reagent (Roche). After 24 hours, cells were fixed for 10 minutes with 4% paraformaldehyde in PBS. After washing 3 times with PBS, cells were permeabilized with 0.5% Triton X-100 (Sigma) in PBS for 5 minutes and washed with PBS. Subsequently, cells were blocked for 10 minutes in PBS, containing 1.5% BSA and 100 mM glycine, and incubated at room temperature with mouse anti-FLAG (1:1000 dilution; Sigma), or rabbit anti-PHR2 (1:500 dilution) (Xu et al., 2010) antibody was performed. As a secondary fluorescent, anti-rabbit antibody–Alexa 488 (1:1000 dilution; Invitrogen) or anti-mouse antibody–Alexa 594 (1:1000 dilution, Invitrogen) were used. Fluorescence images were taken with a Leica DM-RB microscope (Wetzlar, Germany) using band-pass filter cubes (L5 and TX2). Also, in this case, HEK 293T cells were chosen because of their high efficiency of transfection. Nevertheless, similar results were obtained with NIH 3T3 cells, but the amount of transfected cells was much lower. The experiment was repeated 3 times, and 150 cells were counted for each condition.
Luciferase Reporter Assay
The CLOCK/BMAL1 transcription assay was performed as previously described (Yamaguchi et al., 2000). HEK 293T cells were seeded in 24-well plates (5 × 104 cells/well) and cotransfected with the mPer2::luciferase reporter construct (1 ng), pRL-CMV–null Renilla luciferase (0.01 ng; used as an internal control), Clock (100 ng), Bmal1 (100 ng), and either mCry1 (100 ng; used as a positive control for inhibition of CLOCK/BMAL1 transcription), pcDNA3.1-phr1 (200 ng), or pcDNA3.1-phr2 (200 ng) expression construct, using Fugene (Roche). The total amount of 1.2 µg DNA per transfection was adjusted by adding pcDNA3 vector. Each transfection was performed in triplicate. After 24 hours, transcriptional activity was assessed with the Dual-Luciferase 10 Reporter Assay (Promega, Fitchburg, WI) by measuring the ratio of firefly activity to Renilla luciferase activity. For statistical analysis, a 2-tailed t test was used. Also, in this case, HEK 293T cells were chosen because of their high efficiency of transfection. However, similar results were obtained with NIH 3T3 cells (Suppl. Fig. S1).
Real-Time Luminometry
For real-time bioluminescence monitoring of circadian core oscillator performance, NIH 3T3 cells were cotransfected with the mPer2::luciferase reporter construct and either pcDNA3.1-phr1, pcDNA3.1-phr1-egfp, or pcDNA3.1-phr2. As a control, we used pcDNA3.1 (empty vector) or pEGFP. Two days after transfection, cells were clock-synchronized by replacing the medium with 2 mL fresh medium containing forskolin (30 µM), luciferin (100 µM), HEPES (25 nM), 10% fetal calf serum, and antibiotics. Dishes were sealed with a glass cover slip and parafilm and placed in a LumiCycle 32-channel automated luminometer (Actimetrics, Wilmette, IL) in a dry, temperature-controlled incubator at 37 °C. In a parallel series of experiments, the Bmal1::luciferase reporter was used. Real-time bioluminescence recording (60-second measurements at 10-minute intervals) and data processing have been described before (Oklejewicz et al., 2008) using the Actimetrics LumiCycle Analysis program. Amplitudes were calculated both for baseline subtracted data and raw data (by comparing peak and trough values). The choice of NIH 3T3 cells relies on the oscillatory capacity of these cells. HEK 293T cells, on the other hand, cannot be synchronized and make it impossible to study circadian clock oscillations.
Results
PHR1 and PHR2 Can Physically Interact with CLOCK
Taking into account the structural similarity between cryptochromes and photolyases as well as our recent observation that Potorous tridactylus CPD photolyase can replace CRY proteins in the repression of CLOCK/BMAL1-driven transcription (Chaves et al., 2011a), the first step in studying whether baculovirus photolyases may potentially exhibit a circadian clock function was to determine whether PHR1 and/or PHR2 can physically interact with CLOCK and BMAL1, the positive elements of the mammalian transcription/translation feedback loop. To this end, we performed pull-down experiments using HEK 293T cells transiently expressing phr1-egfp or phr2-egfp in combination with either Flag-Clock or Flag-Bmal1. As shown in Figure 2A, PHR1 failed to coprecipitate with BMAL1, whereas, interestingly, the protein could be pulled down together with the CLOCK protein. In the absence of the CLOCK protein, PHR1 could not be precipitated with anti-FLAG antibodies, showing that the coprecipitation of PHR1 is not an aspecific feature of the antibody used. Similar results were obtained for PHR2 (Fig. 2B). From these data, we conclude that the PHR1 and PHR2 proteins can physically interact with CLOCK.
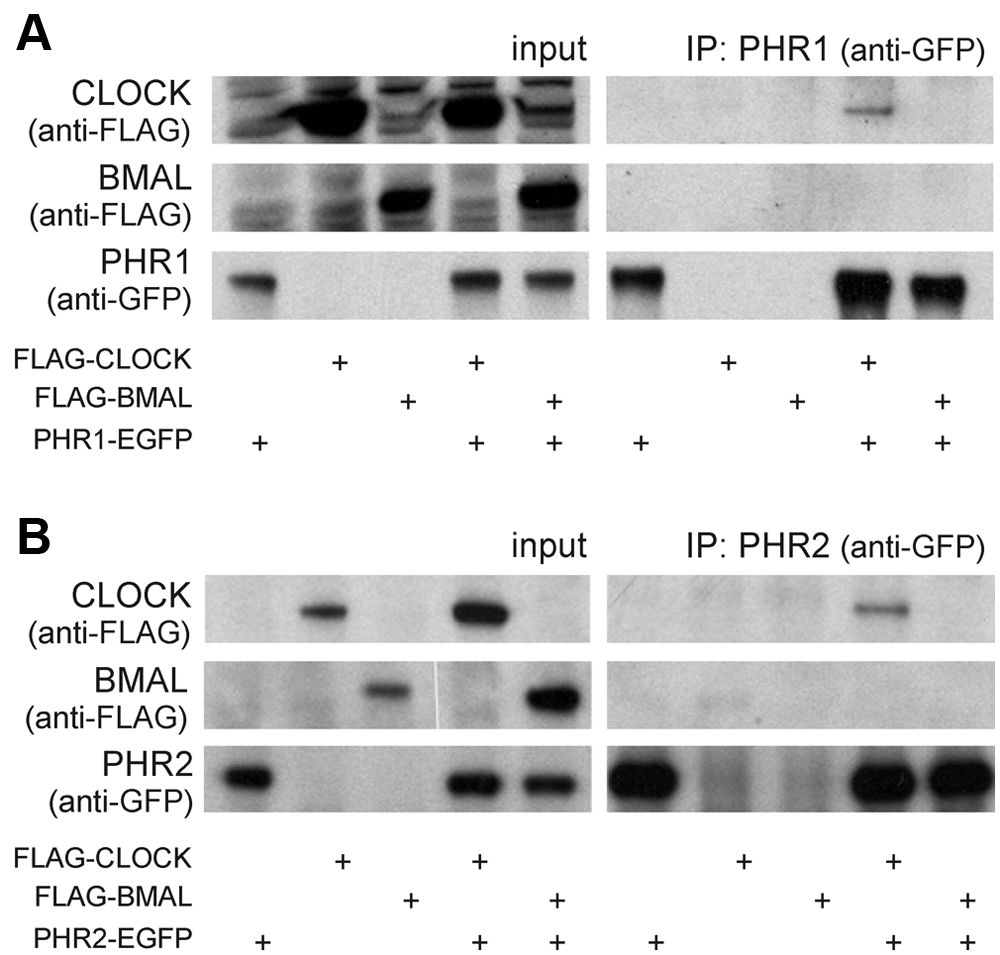
PHR1-EGFP and PHR2-EGFP interact with FLAG-CLOCK. HEK 293T cells were transiently transfected with either phr1-egfp (A) or phr2-egfp (B) and cotransfected with empty vector, Flag-Clock, or Flag-Bmal1 (indicated by plus symbols below). Anti-GFP antibodies were used to immunoprecipitate PHR1-EGFP and PHR2-EGFP. Left panels: Immunoblot analysis of total cell lysates, confirming the presence of the various transiently expressed proteins. Right panels (IP): Immunoblot analysis of precipitated PHR1-EGFP and PHR2-EGFP (anti-GFP antibodies) and CLOCK and BMAL1 (anti-FLAG antibodies).
PHR1 and PHR2 Colocalize with CLOCK
Overexpressed FLAG-CLOCK mainly localizes in the cytoplasm, but when coexpressed with CRY1, it completely accumulates in the nucleus (Chaves et al., 2006). To investigate the consequences of the interaction between CLOCK and the PHR proteins on the subcellular localization of CLOCK, we performed an immunofluorescence assay in HEK 293T cells transiently expressing these proteins (Fig. 3A and 3B). As might be expected on the basis of the predominantly nuclear localization of baculoviral photolyases in insect Sf9 cells and their strong affinity to condensed DNA (Xu et al., 2010), PHR1 and PHR2 were also found mainly in the nucleus (Fig. 3A). When CLOCK was coexpressed with PHR1 or PHR2, the subcellular localization of CLOCK shifted from cytoplasmic to nuclear (Fig. 3B). This finding not only further illustrates that baculoviral PHR proteins can interact with CLOCK but also suggests that they may affect the performance of the circadian core oscillator.
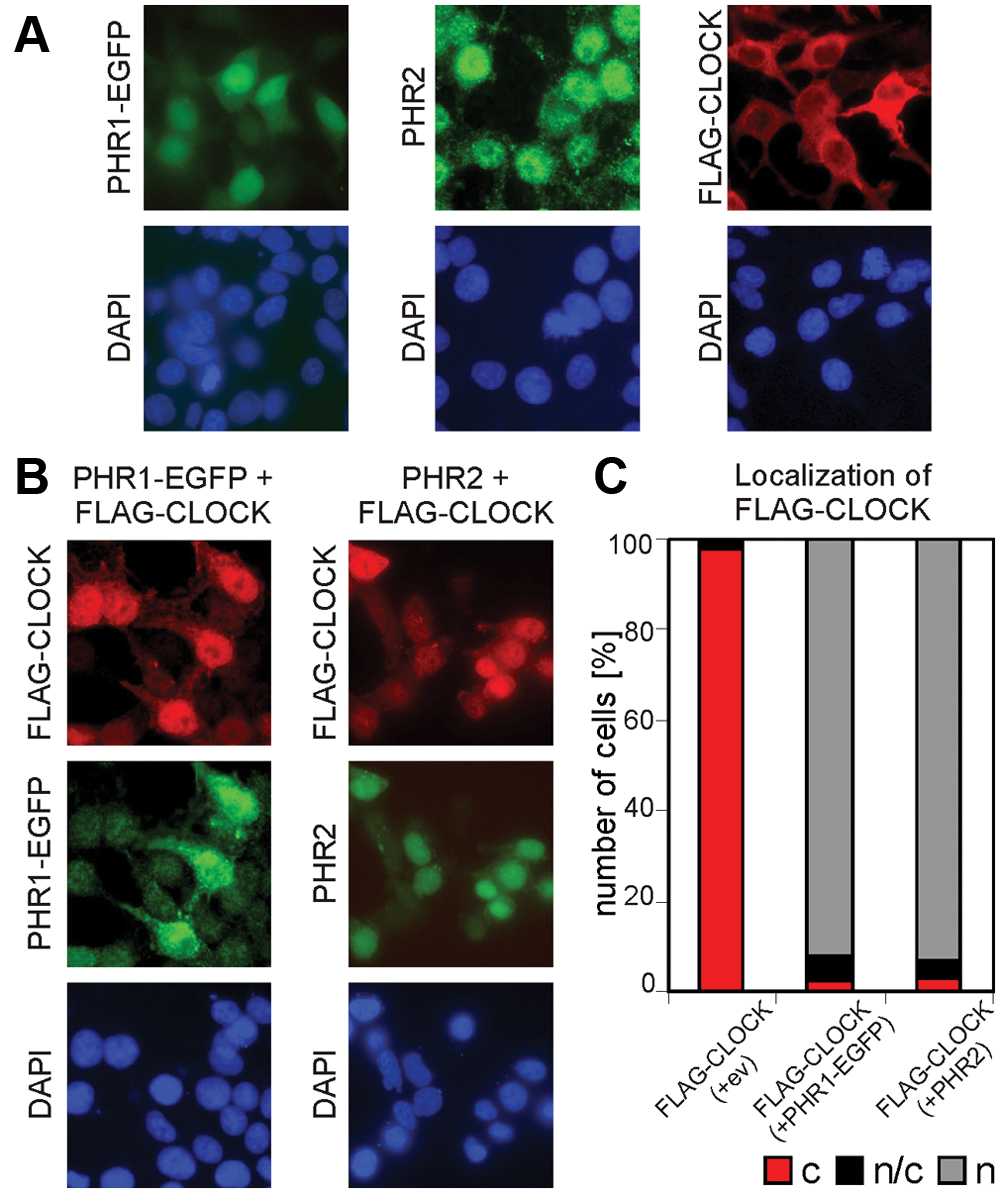
Subcellular localization of PHR1-EGFP, PHR2, and FLAG-CLOCK and nuclear transport of FLAG-CLOCK. Immunofluorescence of HEK 293T cells transiently single-transfected with phr1-egfp, phr2, and Flag-Clock (A) or with combinations of photolyase and Flag-Clock (B). DAPI nuclear staining in blue. (C) Quantification of the cellular localization of the transiently expressed proteins. This is the average of 3 experiments, and 150 cells per experiment were counted per condition.
PHR2 Acts as an Inhibitor of CLOCK/BMAL1-Driven Transcription
In order to test whether the interaction of PHR1 and/or PHR2 with CLOCK affects the transcription activation properties of the CLOCK/BMAL1 heterodimer, we performed an mPer2::luciferase reporter gene–based transcription inhibition assay in HEK 293T cells (similar results were obtained in NIH 3T3 cells) (Suppl. Fig. S1). The coexpression of the reporter gene with Clock and Bmal1 caused an increase in the transcription of the reporter gene compared to mPer2::luciferase alone (Fig. 4). As expected, coexpression of mCRY1 (100 ng) drastically inhibited CLOCK/BMAL1-driven transcription. When mCRY1 was replaced by PHR1 (200 ng) or PHR2 (200 ng), only the latter protein was capable of consistently and significantly (p < 0.01) reducing the transcriptional activity of the CLOCK/BMAL1 heterodimer (Fig. 4). In all conditions, the total amount of plasmid DNA was equalized with the empty vector. The difference in the amount of plasmid used for mCRY1 and either phr1 or 2 relies on the fact that mCRY1 is a more efficient repressor than the photolyases. When varying amounts of plasmid were used, we observed a dose-dependent effect on the repression for mCRY1 and PHR2 (Suppl. Fig. S1). This finding suggests that despite the binding of PHR1 to CLOCK, this interaction is not sufficient to inhibit CLOCK/BMAL1.
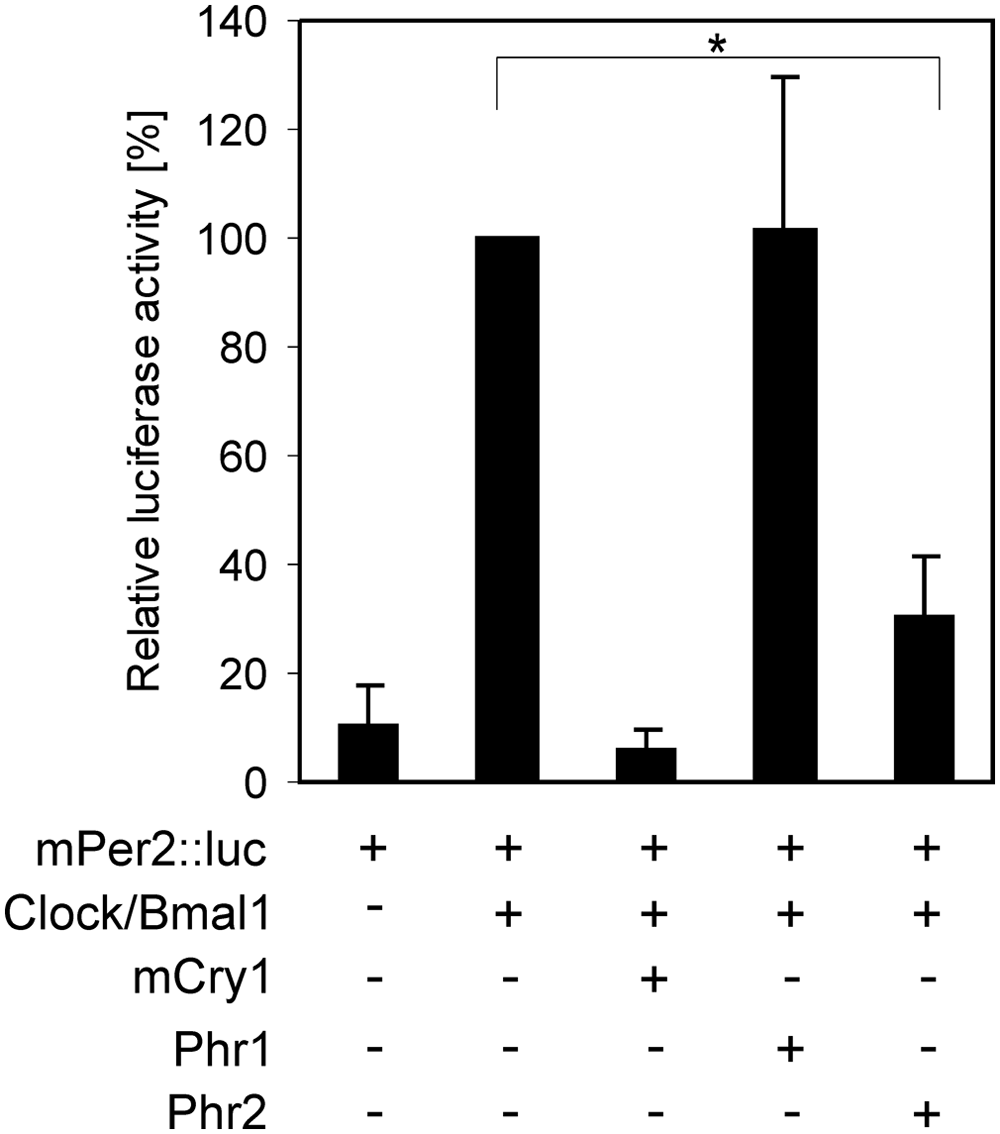
Effect of PHR on CLOCK/BMAL1-driven transcription. Relative luciferase activity in HEK 293T cells. In all reactions, cells were transfected with the reporter constructs Per2::luciferase (luminescence read-out) and PRL (as internal control). Cells were cotransfected with Clock and Bmal1 and Clock and Bmal1 with either mCry1 (100 ng), phr1 (200 ng), or phr2 (200 ng), as indicated under the graphic. The luciferase activity in samples containing Clock/Bmal1 was set to 100%. Error bars (±SEM) were calculated from 3 independent experiments. p values lower than 0.05 are indicated with asterisks.
PHR2 Dampens the Oscillation of Synchronized NIH 3T3 Cells
To obtain insight into the physiological consequences of overexpression of PHR1 or PHR2 on the molecular clock, we next investigated the core oscillator performance in forskolin-synchronized NIH 3T3 cells transiently coexpressing the Bmal1::Luc reporter plasmid (Fig. 5). This was done in combination with either phr1-egfp or phr2 expression vectors, using the empty vector (ev) as control. As an additional control, we also monitored oscillations in cells transiently expressing the mCRY1 protein, which, as expected, drastically reduced the amplitude of Bmal1 rhythms (Fig. 5A). The amount of plasmid DNA transfected for each of the tested constructs was 750 ng, which is at least 3-fold above the necessary amount to obtain repression of CLOCK/BMAL1 (Fig. 4). The overexpression of PHR2 resulted in a significant decrease of approximately 40% (amplitude of 60% ± 12%; p = 0.02) in the amplitude of oscillations (Fig. 5C), whereas overexpression of PHR1-EGFP did not significantly influence the bioluminescence rhythm (amplitude of 85% ± 20%; p = 0.2) (Fig. 4B). Similar results were obtained using an untagged PHR1 (Suppl. Fig. S2). These findings are in agreement with our observation that PHR2, but not PHR1, is able to inhibit CLOCK/BMAL1-driven transcription (Fig. 4).
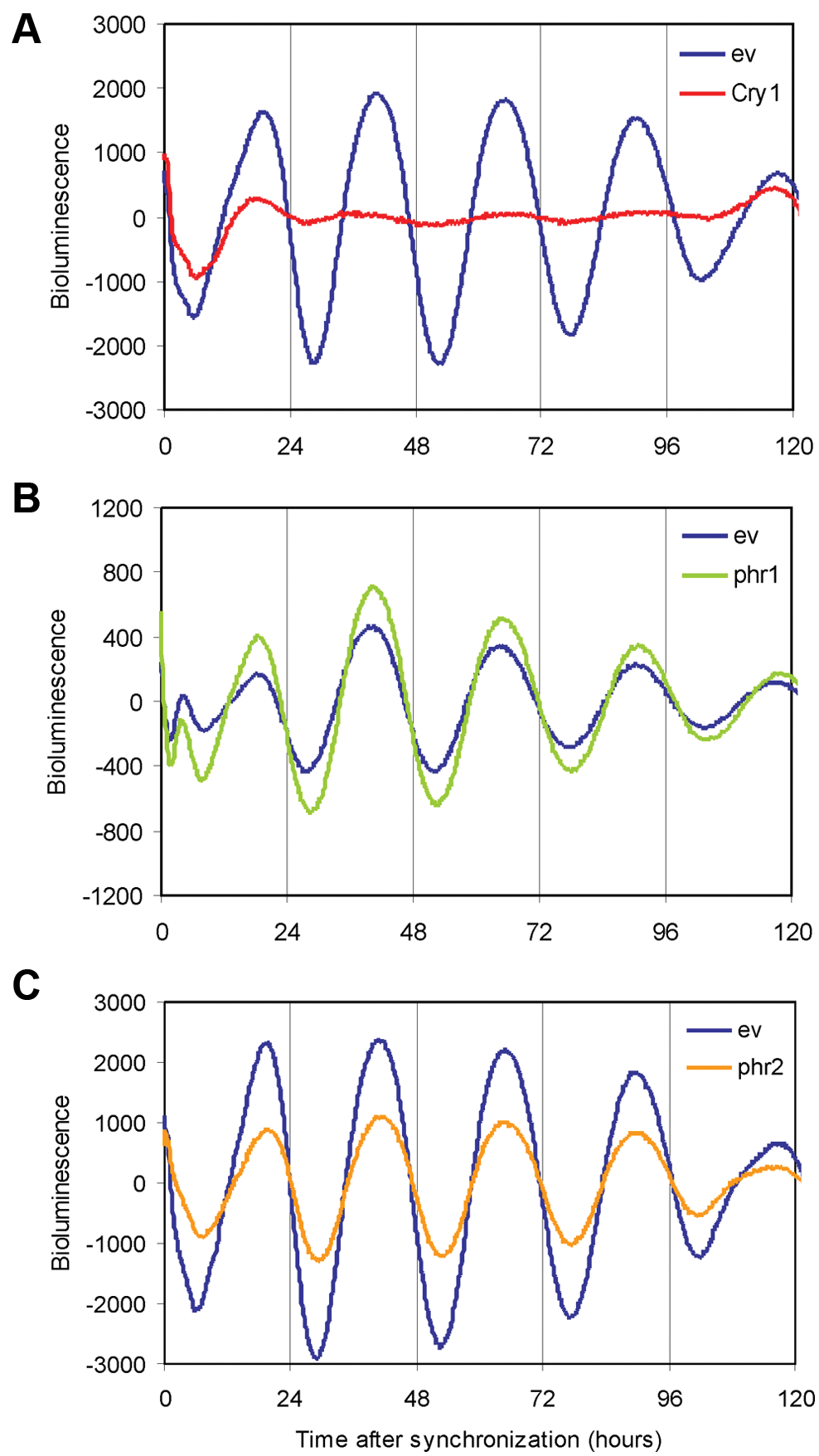
Effect of PHR on the circadian oscillation of synchronized NIH 3T3 cells. Representative examples of bioluminescence rhythms in synchronized NIH 3T3 cells (n = 4). Bmal1::luciferase (250 ng) was used as a reporter construct to follow oscillation in real time. The control oscillation was monitored by cotransfection with the empty vector (ev) (blue). Oscillations of cells cotransfected with mCry1 are depicted in red (A), with phr1-egfp in green (B) and with phr2 in orange (C). In each case, 750 ng of plasmid was transfected. The values in the y-axis correspond to baseline subtracted bioluminescence values.
Discussion
In the present study, we demonstrate that the Chrysodeixis chalcites nucleopolyhedrovirus PHR1 and PHR2 proteins resemble mammalian cryptochrome proteins in that they can interact with CLOCK, a protein that is characterized as an integral component of the transcription/translation feedback loop of the mammalian circadian clock. Cryptochromes act in the molecular oscillator as repressors of the CLOCK/BMAL1 complex by interacting with CLOCK via the core domain and with BMAL1 via the predicted coiled-coil domain of CRY (Chaves et al., 2006). As a consequence, the fact that the photolyases studied here bind CLOCK, but not BMAL1, is highly understandable, as the predicted coiled-coil domain, required for interaction with BMAL1, is absent in the baculovirus and other PHR proteins (Fig. 1). Only PHR2, however, is capable of inhibiting CLOCK/BMAL1-driven transcription, indicating that interaction with CLOCK by itself is not sufficient to inhibit CLOCK/BMAL1-driven transcription. We find, moreover, that overexpressed PHR2 (but not PHR1) affects the molecular oscillation in mammalian cells by decreasing the amplitude of the oscillation in cultured fibroblasts, which is likely due to its capacity to efficiently inhibit CLOCK/BMAL1-driven transcription. This finding is remarkable, bearing in mind the evolutionary distance between mammals and baculoviruses.
The phr2 gene encodes, in contrast to phr1, an active photolyase, which can rescue photolyase-deficient Escherichia coli and has CPD photolyase activity in vitro (van Oers et al., 2008). Sequence analysis shows that PHR1 differs from other class II CPD photolyases. More importantly, these discrepancies are found in conserved regions of the sequence (van Oers et al., 2008). In light of this observation, it is not surprising that the nonfunctional photolyase PHR1 does not affect the molecular clock. For photoreactivation and for repression of the CLOCK/BMAL1 heterodimer, most likely, the same subdomains in the photolyase-like core domain are required. A remaining question is why ChchNPV would be equipped with 2 photolyase-like genes, especially if one of these, phr1, does not seem to have any known function. Further studies are needed to reveal in more detail the properties of PHR1 in ChchNPV and its homologs in other plusiine-infecting baculoviruses (Xu et al., 2010) to come to a better understanding of PHR1 functions.
Recently, a clock repression function has been reported for Phaeodactylum tricornutum (diatom) and Ostreococcus tauri (green alga) (6-4)PP photolyases (Coesel et al., 2009; Heijde et al., 2010). It has been proposed that this duality of function of photolyases is encountered in the lower specialized organisms, which is supported by our analyses of the baculovirus PHR proteins. The taxonomic position of the organism, however, may not be an appropriate criterion to exclude a circadian role of photolyases, as the Potoruos tridactylus CPD photolyase also affects CLOCK/BMAL1-driven transcription (Chaves et al., 2011a). The functional subdivision of cryptochromes, that is, (circadian) photoreception, core circadian clock, and a role in magnetoreception, is also not entirely understood, and it is becoming evident that the same CRY can display more than one function. Additionally, the lack of a light-driven function for mammalian CRY remains controversial. The recent report by Reppert and coworkers (Foley et al., 2011) shows that human CRY2 can also mediate light-dependent magnetoreception in flies, despite the fact that mammalian CRYs have a “dark” function. It is accepted that the evolution of cryptochromes is the result of point mutations in a photolyase ancestor gene, combined (in most organisms) with the acquisition of a C-terminal extension (Chaves et al., 2006; Zhu et al., 2003; Balland et al., 2009). Domain analyses of PHR1 and PHR2 and in vivo studies with mutant ChchNPVs will further pinpoint differences between the viral PHRs and provide insight into the function and evolution of baculovirus photolyases.
Although over 50 baculovirus genomes have been sequenced so far (Harrison, 2009), only few possess a photolyase gene. From an ecological point of view, carrying an active photolyase enzyme would be highly beneficial for the insect virus. It is known that upon infection by a baculovirus, the infected larvae become hypermobile and often climb up to the top of the plant canopy, allowing efficacious spreading of the progeny virus over the foliage and highly facilitating transmission of the virus within the insect population (Goulson, 1997). As a consequence, however, the virus is more exposed to UV light, which could lead to its rapid inactivation (Sun et al., 2004). To survive such conditions, some insect viruses are equipped with a photolyase with DNA repair activity towards UV-induced lesions. This is also found for the entomopox virus AMEV (Nalcacioglu et al., 2010).
Here, we have shown that the role of photolyases in virus-insect host interaction may be much broader then previously thought and may also encompass a circadian clock–related function. It is tempting to speculate about the ecological benefit for baculoviruses to contain a photolyase with a potential clock-related function. Insects were among the first organisms where a master clock was identified (Helfrich-Förster et al., 1998), and there is a striking similarity between the oscillators in mammals and insects. It is crucial to emphasize that both CLOCK and BMAL1 have their orthologs in insects: CLOCK and CYCLE, respectively (Rosbach, 2009). We hypothesize that the circadian role of PHR2 (as identified using the mammalian clock system) is not accidental and that this protein might also affect the circadian clock of the insect host. Possibly, the photolyase encoded by the viral genome not only protects against UV light but also might have a role in virus-induced behavioral changes by influencing the host’s circadian clock. This question is still open, and in vivo studies with phr1 and phr2 knockout ChchNPVs in Chrysodeixis chalcytes larvae are needed to reveal the whole set of functions of the encoded PHR proteins.
Footnotes
Acknowledgements
The authors thank Dr. Kazuhiro Yagita (Osaka University Graduate School of Medicine, Japan) for kindly providing them with the mPer2::luciferase, Bmal1-flag, and Clock-flag expression constructs. The Bmal1::luciferase construct was a generous gift of Dr. Ueli Schibler (Geneva University, Switzerland). This work was supported in part by grants from the Netherlands Organization for Scientific Research (NWO): CW ECHO 700.57.012 to GTJvdH, and ALW2PJ/06023 and MEERVOUD-836.05.070 to MMvO.
The author(s) have no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Notes
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
