Abstract
Coagulase-negative staphylococci (CoNS) belong to saprophytic microbiota on the skin and mucous membranes of warm-blooded animals and humans, but are also isolated from foodstuffs such as meat, cheese, and milk. In other circumstances, some CoNS can act as pathogens. Thus the presence of CoNS may not be an immediate danger to public health, but can become a risk factor. In particular antibiotic-resistant genes could be transferred to other potentially pathogenic microorganisms. Furthermore, CoNS are known to be strong biofilm producers and this is also a risk factor for public health.
The aim of the present work was to determine the genotypic and phenotypic profiles of 106 CoNS belonging to four different species isolated from five different Italian cheeses for the presence of some adhesion and virulence features. In order to verify a possible correlation between the formation of biofilm and staphylococcal virulence factors, we checked the presence of adhesin genes by PCR and we investigated the ability of these strains to make biofilm at different temperatures. Furthermore, in some conditions, we analyzed surface proteins and autolytic pattern of selected strains. In conclusion, we checked the presence of norA and mecA genes responsible for fluoroquinolones and methicillin resistance, respectively. We found resistant genes in a proportion of the food isolates in amounts of 9.4% (mecA) and 5.7% (norA). These data support the importance to continuously examine the microbiota not only for the creation of a database but also to safeguard public health.
Keywords
Introduction
Coagulase negative staphylococci (CoNS) are normal inhabitants of humans and other organisms and are mainly isolated as commensal strains from skin and from mucous membranes. Usually most of the species are harmless, however since 1970, CoNS have been regarded as etiologic agents of various infections. 1 The pathogenesis of CoNS is mainly attributed to a combination of extracellular factors and properties such as adherence and biofilm formation. 2 Furthermore CoNS are frequently isolated from dairy products and from the surface of various types of cheese, either as contaminant species or as useful species for the determination of the flavor or development of the organoleptic properties.3–5
The microbial diversity of the cheese depends on both the milk microbiota and on practices of production. 6 Over 400 species have been detected in raw milk: Gram-negative and Gram-positive bacteria, lactic acid bacteria, yeasts, and molds. This biodiversity decreases in the core of the cheese where lactic acid bacteria are the prevalent species but it persists on the surface where numerous species are detected. The difference between cheeses is due particularly to changes in the dynamics of the microbial population used in the production. Cheese microbial population still remains difficult to control due to their complex dynamics and to their interactions.7,8 A broader knowledge of the microbiota composition could make possible a better selection of specific organoleptic properties or to prevent quality defect, spoilage, or undesirable accumulation of pathogens and/or toxins.
The balance between a commensal behavior versus a pathogenic behavior, apart from a permissive host, probably relies on the potency of the armamentarium of virulence for a datum strain.
Many different studies on the pathogenesis of CoNS, in particular on S. epidermidis, indicated a key role of biofilm formation in a successful colonization of foreign bodies.9,10 After initial adhesion on biotic or inert surfaces, staphylococci under appropriate conditions can expand as multilayered communities; in this form they are more protected from environmental changes. Complex biofilm architecture results in a low metabolism of cells, decreased transcription and translation, and a shift from aerobic production of energy to fermentation, resulting in a non-aggressive and protected mode of growth typical of persister cells; persisters are less sensitive to antibiotics and the host immune defence.11,12 One of the first factors described mediating biofilm accumulation was the polysaccharide intercellular adhesin PIA, synthesized by icaADBC encoded enzymes. 13 Nevertheless, strains that are able to produce a protein-dependent biofilm were also identified.11,14–17 For example, in S. epidermidis clinical isolates the associated accumulation protein Aap was identified as a polysaccharide-independent intercellular adhesin mediating biofilm formation. 15 Apart from Aap, CoNS produce various other microbial surface proteins that recognize adhesive matrix molecules of eukaryotic cells (MSCRAMMS), as well as other adhesive proteins, which enable bacteria to bind different surfaces moieties. Among them fibrinogen, fibronectin, and vitronectin adhesins (Fnb, EmbP, AtlE) and biofilm associated proteins (Bap) were identified.10,18,19
The food chain can be also considered an important source for antibiotic-resistant transmission. In fact, CoNS that are usually overlooked as food-borne bacteria are emerging as important opportunistic pathogens that may be highly resistant to antimicrobial drugs. 20 The antibiotic resistance may be transferred from CoNS to commensal or other pathogenic bacteria. 21 CoNS can be resistant to methicillin, mainly due to the expression of the mecA gene coding for penicillin binding protein 2a (PBP2a), a transpeptidase with a low affinity for betalactams. Furthermore the SCCmec elements could be transferred from CoNS to S. aureus, 22 representing an additional risk factor for the spreading of antibiotic resistance in the communities. Methicillin resistance is often accompanied by resistance to other antimicrobial agents such as fluoroquinolones. 23 In S. aureus nor family genes coding for efflux pumps provide resistance to fluoroquinolones. The NorA protein of S. aureus confers resistance to a variety of hydrophilic fluoroquinolones.
The aim of the present work was to determine the genotypic profiles of adhesion and virulence genes and the adhesive behaviors at different temperatures of 106 CoNS strains isolated from five different Italian cheeses. We investigated the presence of genes associated to adhesion and biofilm formation such as icaA, embP, aap, and atlE and we checked the presence of genes (norA and mecA) associated with antibiotic resistance (fluoroquinolones, methicillin). Biofilm formation was performed at different temperatures corresponding to different steps for the process of cheese-making and to the human body temperature. Finally we analyzed surface protein and autolytic pattern of selected strains at different temperatures.
Materials and methods
Bacterial strains and culture conditions
A total of 106 CoNS were isolated from the surface of the Italian cheeses Casera Valtellina, Scimudin, Gorgonzola, Taleggio, and Formaggio di Fossa (Table 1). Gorgonzola and Taleggio cheeses were purchased, while Casera Valtellina, Scimudin, and Formaggio di Fossa were supplied by Cooperlat-Fattorie Italy. All isolates were identified as reported by Fontana and co-workers. 8 All strains were also confirmed in their coagulase phenotype by using coagulase test as reported by Essers and Radebold. 24 Staphylococcus saprophyticus DSM20229, Staphylococcus equorum CIP103502, and Staphylococcus sciuri DSM20352 were used as reference strains.
Distribution of CoNS isolated from the surface of five Italian cheeses divided for bacterial species and type of cheese.

CoNS were routinely grown in Tryptic Soy broth (TSB, Oxoid, UK) under vigorous agitation at 37°C for 24 h. As solid medium Tryptic Soy Agar (TSA, Oxoid, UK) was used. Methicillin-resistant strains were tested on Mannitol Salt Agar (MSA, Oxoid, UK) at 37°C for 24 or 48 h. All strains were stored at −80°C in cryovials with 15% of glycerol in TSB.
PCR amplification and sequence analysis
Primers and PCR conditions are summarized in Table 2. DNA preparation and PCR were performed as follows. A single colony picked on TSA plate was resuspended in 100 μL of TE buffer (Tris-HCl 10 mM pH 8, EDTA 10 mM pH 8) and boiled for 10 min. The mixture was then centrifuged at 13,000 rpm at 4°C to eliminate bacterial debris. A total of 5 μL of supernatant containing bacterial DNA partially purified was used for PCR amplifications. Amplicons were analyzed by gel electrophoresis.
Primers used for PCR-based detection of staphylococcal factors involved in adhesion and antibiotic resistance. For adhesion genes was investigated the presence of icaA, embp, aap, and atlE, respectively. For the antibiotic resistant the presence of norA and mecA genes were analyzed, respectively.

PCR products were purified by using GenElute Gel Extraction Kit (Sigma-Aldrich, USA). Sequencing was carried out by PRIMM biotech (Italy). To confirm the specificity of amplicons, sequences obtained were employed for similarity searching in the available databases by using BLAST algorithm. 25
Susceptibility testing
Bacterial strains positive to mecA and norA genes were further investigated by susceptibility test on the corresponding antibiotics. Antimicrobial susceptibility was determined by test disks (Oxoid, UK) at the following concentration: 10 μg for methicillin, 5 μg for oxacillin, 30 μg for cefoxitin, and 5 μg for ofloxacin. The procedure was performed according to the Clinical and Laboratory Standards Institute (CLSI). Plates were then incubated at 37°C for 24 h. Methicillin resistance was also evaluated at 37°C for 48 h.
Quantification of biofilm formation
Quantification of in vitro biofilm production was based on method previously reported. 26 Briefly, the wells of a sterile 96-well flat-bottomed polystyrene plate were filled with 200 µL of TSB supplemented with 0.25% glucose. A 100-fold dilution of overnight bacterial cultures was added into each well. The plates were incubated aerobically for 24 h at 37°C or 20°C, and for 72 h at 8°C, without agitation. After rinsing with PBS, adhered cells were stained with 0.1% crystal violet, rinsed twice with double-distilled water, and thoroughly dried. The dye bound to adherent cells was solubilized with 20% (v/v) glacial acetic acid and 80% (v/v) ethanol. The OD of each well was measured at 590 nm. Each data point is composed of four independent measures.
Surface protein extraction and processing
Surface proteins were extracted as previously reported. 27 A total of 50 mL of each bacterial culture (OD600 = 0.6) were centrifuged and pellets were washed twice in PBS and then suspended with 500 µL PBS containing 1% SDS. Samples were incubated at 37°C for 15 min and after centrifugation the supernatants were collected and used for SDS-PAGE and zymogram analyses. The protein content in the samples was determined by the Bradford procedure. 28
SDS-PAGE and zymogram
SDS-PAGE was carried out by standard methods with an SDS–polyacrylamide separating gel (10% acrylamide, pH 8.8) and constant voltage at room temperature. Following electrophoresis proteins were stained with Coomassie Brilliant Blue (Bio-Rad). Renaturing SDS-PAGE was performed according to the methods of Artini and co-workers. 29 SDS-polyacrylamide separating gel (10% acrylamide, pH 8.8) containing 0.2% (wt/v) lyophilized Micrococcus luteus cells provided by Sigma, was used to detect the lytic activities. After electrophoresis, the gels were soaked (twice, 15 min) in distilled water at room temperature. The gels were then transferred into the renaturing buffer (50 mM Tris-HCl pH 8.0 containing 1% Triton X-100) and shaken at 60 rpm for 2 h at 37°C to allow renaturation. The renatured autolysins appeared as clear translucent bands on opaque background. For each experiment, two gels were prepared from the same stock solution and run in the same apparatus at the same time. The presence of M. luteus cells in the gel did not evidence differences in the migration of the standards.
Results
Planktonic growth and adhesive behavior at different temperatures
A total of 106 isolates from five different cheeses belonging to four species, were used in this study. Among them, S. equorum (45 strains), S. saprophyticus (33 strains), S. vitulinus (25 strains), and S. caprae (2 strains). In the analysis three reference strains described in the Materials and methods section were also included.
The 106 CoNS isolates and the three reference strains were preliminarily screened for the ability to growth in planktonic form at 20°C and 37°C (Figure 1). The first temperature chosen corresponded to that used during the thermic treatment of cheese production (Table 3), while the second one corresponded to the temperature of the potential host (human). All strains were then analyzed for the ability to form biofilm at the same temperatures. Of total strains, 32% (34/106) resulted no biofilm producers. The remaining 72 cheese staphylococcal isolates ranged from weak biofilm producers (OD590 <0.25), intermediate (0.25 < OD590 <0.5), to strong biofilm producers (OD590 >0.5) at 20°C and 37°C.

Planktonic growth curves of eight selected CoNS isolates at three different temperatures. (a) 37°C; (b) 20°C; (c) 8°C.
Cheese-making chart. The table indicates the range of temperature and humidity required during the different steps of the production.

On the basis of their behavior at 37°C and 20°C, eight isolates out of 106 (UC8192, UC8424, UC8390, UC8405, UC8193, UC8337, UC8203A, UC8322) were further selected to assess sessile and planktonic growth also at 8°C (Figures 1 and 2 ). The rationale of this choice was the selection of strongest biofilm producers within each bacterial species. Growth curves of staphylococcal strains analyzed here showed that duplication times at 20°C and 37°C were comparable, while at 8°C it was significantly higher. In particular, two out of 8 strains, UC8193 and UC8203A, did not reach the optical density required for further analyses at 8°C. The planktonic optimal growth temperature did not necessarily corresponded to the optimal growth temperature for the sessile condition. In fact, the duplication time at 37°C was always shorter than the duplication time at 20°C, whereas in some isolates (UC8390 and UC8337) the biofilm produced at 20°C was more abundant than the biofilm formed at 37°C (Figure 2). Moreover, four strains (UC8192, UC8424, UC8193, and UC8322) were also strong biofilm producers at 8°C. Furthermore, strain UC8322 was stronger biofilm producer at 8°C rather than at 37°C or 20°C. The best biofilm producer at all tested temperatures was UC8192 strain.

Biofilm production of eight selected staphylococcal isolates at three different temperatures (37°C, 20°C, and 8°C). Data are reported as absorbance at 590 nm after coloration with 0.1% crystal violet. Results are representative of three independent experiments.
Detection of adhesion related genes
PCR technique was used to evaluate the presence of different genes involved in microbial adhesion such as icaA involved in PIA synthesis, embP coding for fibronectin adhesin, aap coding for the intracellular adhesin, and atlE involved in the first phase of biofilm formation.
The presence of the icaA gene was demonstrated by amplification of the corresponding fragment in 21 (19.8%) of 106 isolates. All 106 CoNS isolates resulted positive for gene aap, coding for accumulation-associated protein, the additional factor that mediates biofilm formation. Twenty cheese isolates resulted positive for embP encoding for a fibronectin adhesion (18.8%). The AtlE gene was only found in seven strains (6.6%) out of 106. In Table 4 the frequency of adhesive genes in No-biofilm and Biofilm producers strains, respectively, are reported. We observed that atlE gene was absent in all no-producing biofilm strains.
Association between biofilm production and gene frequency. The isolates and the three reference strains were divided in producers or non-producers biofilm on the basis of results obtained by crystal violet assay. Two groups were then linked with the presence of adhesion and antibiotic resistant genes.

Antibiotic resistance genes detection and susceptibility tests
The presence of norA gene involved in quinolone resistance and of mecA gene involved in methicillin resistance were demonstrated by amplification of the corresponding fragment (Table 5). A considerably high proportion (9.43%) of 106 CoNS tested was resistant to methicillin, oxacillin, and cephalosporin. The mecA gene was detected in six out of 25 S. vitulinus isolates (24%) and in four out of 33 S. saprophyticus (12%) isolates. All genetic mecA-positive were phenotypic methicillin and oxacillin resistant as determined by disk diffusion method (Table 5). No S. equorum or S. caprae mecA positive strains were identified. All CoNS strains were also tested for the presence of the norA gene. Six CoNS tested strains (5.7%) were norA-positive. The norA gene was detected in two out of 45 S. equorum isolates, three out of 37 S. saprophyticus isolates, and only in one out of 25 S. vitulinus isolates. All genetic norA-positive were also phenotypic ofloxacin resistant as determined by disk diffusion method. No strain carrying both mecA and norA genes was identified. Table 4 shows the frequency of mecA and norA genes in no-biofilm and biofilm producers strains, respectively. We observed that mecA gene was absent in no-producer biofilm strains.
In vitro antibiotic susceptibility according to CLSI.

ND, not determined; R, resistant; S, sensitive.
Antimicrobial susceptibility was determined by test disks on bacterial strains resulted positive to mecA and norA genes by PCR. In the table only bacteria positive to PCR analysis are shown.
Surface protein and autolytic profiles
Figure 3 shows surface protein and autolytic patterns of the selected eight isolates by SDS-PAGE and zimography. These analyses did not reveal dramatic changes among different temperatures. However the analysis of the zymograms of bacteria grown at 20°C and 37°C showed evident differences in some strains. In fact, it was observed that the protein band of apparent molecular weight around 125 KDa is no longer visible for the strains UC8192 and UC8322 grown at 20°C compared with the same strains grown at 37°C. On the contrary, it can be observed different bands appeared for the strain UC8203A grown at 20°C, in particular a band of apparent molecular weight around 90 KDa was evident. In strain UC8424 the band of about 250 KDa disappeared during growth at 8°C. Nevertheless the strain UC8405 did not show differences in autolytic profiles at three different temperatures. In other strains the autolytic profile was not detectable in tested conditions. Furthermore, these patterns did not correlate either with the species or with the source of the isolate.
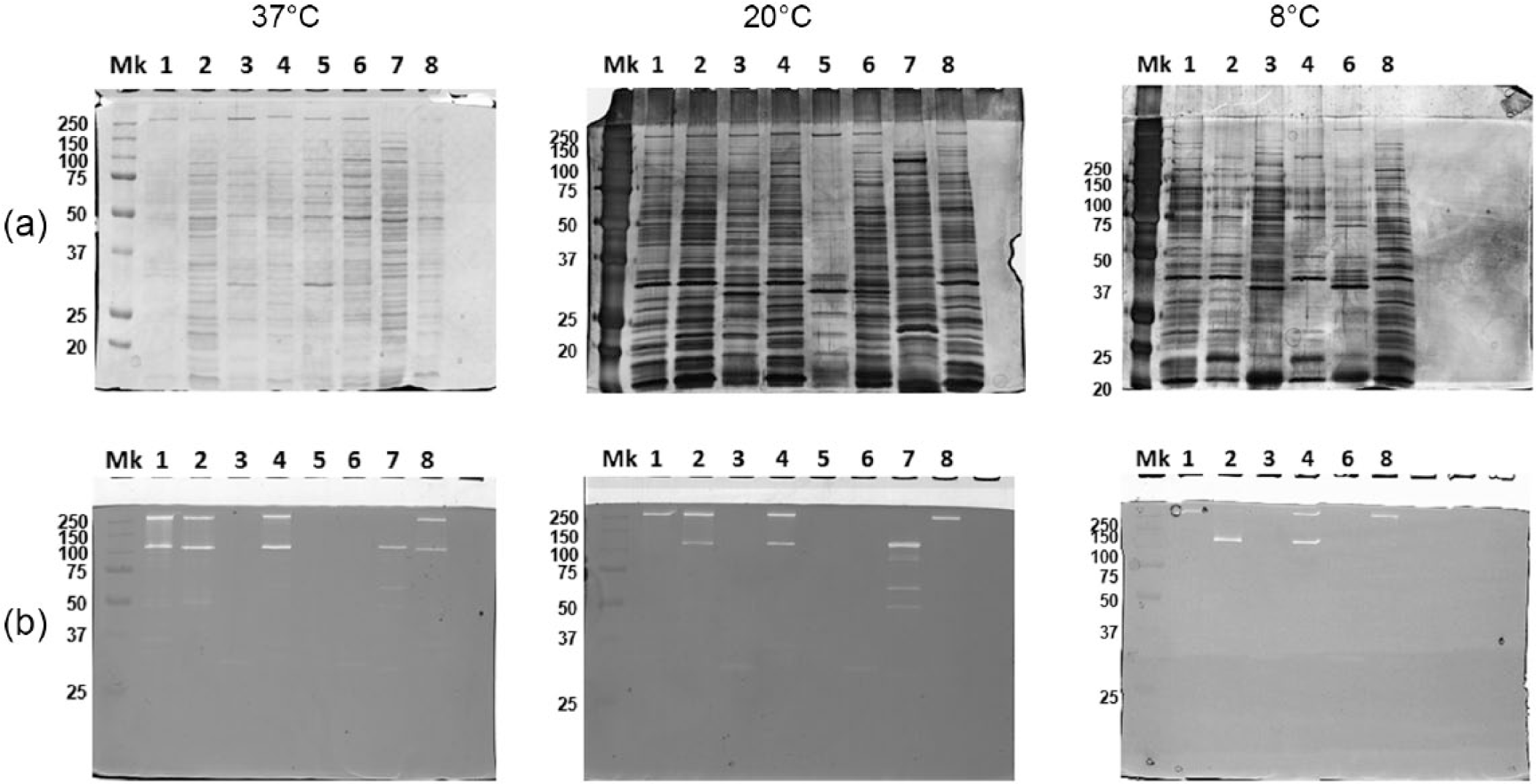
Surface protein pattern (a) and surface autolytic pattern (b) of eight isolates analyzed at different temperature (37°C, 20°C, and 8°C). Mk (molecular weight marker Biorad).
Discussion
In this paper we analyzed the ability to form biofilms of 106 strains of CoNS isolates from five types of traditional cheeses. We analyzed the genotypic profile and looked for a possible correlation between biofilm formation and virulence factors such as resistance to antibiotics and adhesive features. CoNS isolates were identified in four species belonging to: S. equorum (44/106), S. saprophyticus (36/106), S. vitulinus (25/106), and S. caprae (2/106). The analysis of the biofilm did not show correlation between its production and the type of cheese where the bacteria were isolated. Of the strains analyzed 32% were not biofilm producers.
In this study, we analyzed the presence of the norA and mecA genes because it has been shown that strains of methicillin-resistant staphylococci may be resistant to other antibiotics as penicillins, penems, carbapenems, and cephalosporins. 30 Our data showed that the mecA and norA genes frequency is quite high. The frequency for the mecA gene is 9.4% and for the norA gene 5.7%. If the microorganisms are divided into biofilm producers and biofilm non-producers, it can be noticed that any species producing bacterial biofilms harbors mecA gene and only one strain carries the norA gene (S. saprophyticus UC8389). Despite an increase in the frequency of genes mecA (13%) and norA (7%) in the biofilm producers, the correlation between the presence of resistance genes and biofilm remains doubtful. mecA resistance in staphylococci is often accompanied by resistance to other antimicrobial agents such as fluoroquinolones, but in our study none of the 106 food isolates possess a resistance to both methicillin or ofloxacin. mecA gene was found only in S. saprophyticus and S. vitulinus while norA gene was found in at least one strain of all species analyzed.
Analysis on the incidence of antibiotic resistances revealed that pathogens more frequently acquire resistances and multiple resistances as a result of continuous antibiotics use. 21 Cabreras-Contreras and co-workers 31 observed a correlation between antibiotic resistance and biofilm production for clinical methicillin-resistant S. epidermidis (MRSE). Furthermore Pozzi and co-workers 32 observed that excision of the SCCmec element in the methicillin-resistant S. aureus (MRSA) strain BH1CC resulted not only in oxacillin susceptibility but also in reduced biofilm production. For these observations it is important to verify the frequency of resistance genes and control the progress of antibiotic resistant bacteria. By analyzing the frequency of atlE with biofilm production it can be observed that this gene is absent in non-producing biofilm strains and it is 10% present in others. In biofilm producers an increase in the frequency of the icaA gene passing from 14% to 25% can be also observed. It is not surprising that all the isolates were aap-positive. In fact, as reported in literature, the positivity to aap in S. epidermidis is greater than 94%. 33 The isolate S. vitulinus UC8417, icaA-positive, was a strong biofilm producer both at 20°C and at 37°C, moreover it was also found oxacillin resistant. The isolate S. vitulinus UC8424, even though it was an icaA-negative strain, it was a strong biofilm producer at 8°C, at 20°C, and at 37°C; this strain also was oxacillin resistant. The isolate S. vitulinus UC8152, icaA-positive and embP-positive, was instead a weak biofilm producer both at 20°C and at 37°C. So the presence of icaA together with another proteinaceous adhesin is not sufficient for high biofilm production. The three isolates described above, considering their phenotype together with the genotype, are potential pathogens. Among 106 strains, 15 icaA-negative isolates were strong biofilm producers at least at two temperatures, among them the isolate S. equorum UC8388, which is also embP and atlE-positive, it is resistant to ofloxacin and, for this reason it could be considered another example of potential pathogen. Three S. saprophyticus isolates (UC8131, UC8191, and UC8327), three S. equorum isolates (UC8189, UC8202, and UC8353), and one S. vitulinus (UC8192) were strong biofilm producers at both two temperatures with only aap-positivity. In one case, S. caprae UC8322, grown on Scimudin, the biofilm formed at 8°C is almost three-fold stronger than the biofilm formed at 37°C and more than 13-fold stronger than the biofilm formed at 20°C.
In conclusion, the presence of CoNS may not be an immediate risk to public health, however, it may be a risk factor as antibiotic resistance genes may be transferred to other microorganisms some of which may be pathogenic. 3 In the same way the presence of CoNS that produce biofilm can be a risk factor. Therefore it is important to continue to monitor the microbiota not only for the creation of a database but also to safeguard public health.
Footnotes
Acknowledgements
The authors thank Cristina Avanzolini and Marco Vitale for technical support.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This work was supported by Grants from the Grandi Progetti Strategici (MIUR D.M. 697/2007) “Ruoli dei Biofilm microbici per la qualità e sicurezza dei prodotti caseari”, and MISE GU 205 del 2/9/2008 Nuove Tecnologie per il Made in Italy “La shelf-life nella filiera del latte: sviluppo di tecnologie innovative per il miglioramento delle caratteristiche di qualità, sicurezza e conservazione dei prodotti lattiero-caseari Made in Italy”.
