Abstract
Introduction
Cancer is a serious health concern with high morbidity and mortality worldwide. 1 Despite developments in diagnosis and treatment strategies, the 5-years survival rate remains discouraging; therefore, improving prevention and early detection techniques may enhance the survival of patients with cancer.2,3 There is increasing evidence suggesting that the dysregulation of long non-coding RNAs (lncRNAs) is associated with the proliferation, invasion, chemoresistance, and metastasis of various tumours, and it modulates the progression of cancers through oncogenic or tumour-suppressive roles.4,5 Additionally, complex interactions between tumours and the immune microenvironment play an important role in predicting clinical outcomes and the therapeutic response of tumours.6–9 Cancer cells interact with their surrounding microenvironment during tumorigenesis and metastasis.10,11 Thus, revealing the complex relationship between the lncRNAs and TIME would be significant in understanding the tumour translational machinery.
The mechanisms of lncRNA regulation are numerous and complex. LncRNAs include eRNAs and competing endogenous RNAs (ceRNAs) of cancer. 11 Notably, eRNAs are transcribed from the enhancer and often converge at the activation of oncogenes or oncogene signalling pathways by regulating their target genes. 12 For example, Kallikrein Related Peptidase three eRNA can selectively enhance androgen receptor-regulated gene expression in prostate cancer. 13 TP53-induced eRNA was revealed to participate in the p53-dependent cell cycle, which restricts the cell cycle in many cellular processes 14 and might also influence drug responses in an intra- or cross-pathway manner. These findings reveal that eRNAs have important clinical utility functions in tumorigenesis. 15
Recently, immune-related mechanisms, which have played play an important role in cancer and immunotherapeutic strategies, have been under extensive investigation. 16 The latter investigations have focused on the targe components of TME or immune checkpoint genes, immunotherapy biomarkers of microsatellite instability (MSI), and tumor mutation burden (TMB). Wang et al. 17 reported that eRNA WAKMAR2 participates in immune responses in breast cancer. Ma et al. 18 reported that eRNAs LINC00987 is a functional and effective tumour suppressor and biomarker for assessing the TME and immunotherapy of lung adenocarcinoma . LINC02257 may be a multifaceted and significant immunotherapy-related eRNA in multiple cancer types, apart from glioma. 19 To the best of our knowledge, there is a lack of experimental evidence on immunotherapy-associated eRNAs, as well as their underlying functions and mechanisms, specifically in glioma.CRNDE is an elncRNA found on chromosome 16; it was activated early in colorectal cancer and later verified. 20 Emerging studies have identified CRNDE as a potential diagnostic biomarker and prognostic predictor for several cancer types.20,21 Currently, there is limited experimental evidence which has shown its role on the regulation of tumour microenvironment. Thus, it is necessary to further explore its expression and mechanism in pan cancer, which can provide a new reference for immunotherapy. In the present study, we combined data from different databases to investigate the role of CRNDE for tumour immunity as well as the prognosis of multiple cancers. The data implied that CRNDE was associated with IRX5, and modulated the TIME. Moreover, we conducted molecular biology verification in glioma to further confirm the role of CRNDE in vitro and in vivo.
Methods
Data sources and data processing
The clinical information and genomic matrix files from TCGA; (https://cancergenome.nih.gov) and GTEx (https://gtexportal.org) clinical sample databases were downloaded from the UCSC Xena browser website. The established RNA matrix file was merged with Ref-Seq and used for all subsequent RNA-seq expression analyses.2,22 Using this database, the expression and survival levels of all elncRNAs of interest in 33 tumours were obtained. The cancer cell line RNA-seq data of various types of cancer were downloaded from the Broad Institute Cancer Cell Line Encyclopedia (CCLE) (https://portals.broadinstitute.org/ccle). Validation datasets were downloaded from the Chinese Glioma Genome Atlas (CGGA) (https://www.cgga.org.cn/) and the GEO data sets (https://www.ncbi.nlm.nih.gov/geo/). Raw data were downloaded and analysed using the ‘limma’ package in R.
Pan-cancer prognostic analysis of CRNDE
The prognostic value of CRNDE was analysed using the ‘survival’ package in R. 23 The correlation between CRNDE expression and survival, including overall survival (OS), disease-specific survival (DSS), and progression-free interval (PFI), was performed by a univariate Cox regression analysis and the Kaplan-Meier survival analysis.
Analysis of tumour immune microenvironment
To evaluate the composition of immune cells in the tumour, we used immune econv, including TIMER, xCell, MCP-counter, CIBERSORT, EPIC, and quanTiseq to analyse the immune infiltration data based on CRNDE expression.24,25 Furthermore, the relationships between CRNDE expression and the tumor mutation burden (TMB) score, microsatellite instability (MSI) score, and expression of immune checkpoint genes in six types of cancer were analysed sequentially using Spearman’s correlation coefficients.
Nomogram establishment
Univariate and multivariate Cox regression models were established to evaluate clinical indicators (PRS type, histology, grade, age, chemostatus, IDH mutation, and 1p19q codeletion) and CRNDE-associated risk scores with the glioma patient operating system. In addition, the consistency between the actual survival and the nomogram was evaluated using calibration curves to predict survival probability. 26
Identify target genes
Gene correlation analysis was performed using GEPIA (https://gepia.cancer-pku.cn/). The co-expressed genes of CRNDE were obtained using TCGA and CGGA datasets (R > 0.4, p < 0.001). Heatmap and clustering data were constructed using roentgen packages.
Functional enrichment analysis
A GSEA was performed to determine the difference between the CRNDE-high and -low groups through the KEGG pathway databases from MsigDB using GSEA v.4.1.0. 27 The significance was defined as a p-value and FDR of <0.01.
Cell culture, CRNDE knockdown, and reverse transcription quantitative PCR
U87, LN229, and HS683 cells were cultured in a high sugar medium with 10% foetal bovine serum at 37°C and 5% CO2. To silence CRNDE expression, a specific smart RNA silencer (Guangzhou RiboBio Co, China), including siRNAs and small nucleolar RNAs (snoRNAs), was designed and synthesised against CRNDE. Total RNA content was extracted from U87, LN229, and HS683 cells using the TRIzol reagent. An iScript cDNA synthesis kit was used to synthesise cDNA for lncRNA and mRNA analysis. A RT-qPCR was then performed on an ABI 7500 PCR System with SYBR Green Real-time PCR Master Mix. 28 GAPDH was used as the reference gene. Sequences of the primers are shown in Supplementary Table 1.
MTT assay
Cell viability was monitored using a MTT assay. Transfected cells were seeded into 96-well plates. Each well was supplemented with 10 μl MTT (5 mg/mL) and incubated for 2 h. The cells were dissolved in dimethyl sulfoxide (DMSO, 100 μL). Absorbance was measured at a wavelength of 570 nm.
Invasion assay
Transwell chambers (Corning, USA) were diluted in a 1:8 ratio with Matrigel (BD, USA). The upper chamber contained 120 μL serum-free medium and was seeded at 5 × 104 cells, while the medium supplemented with 10% serum was added to the lower layer. 29 After transfection, the invasive cells were fixed with methanol, washed with PBS, stained with 0.1% crystal violet, and photographed using an inverted microscope (AMG, USA).
Wound healing assay
LN229, U87, and HS683 cells were incubated in 6-well plates. Cells were scraped with a 10-μL micropipette tip when the cell cultures reached 80% confluence. Analysis and photography of cells migration ability were performed using ImageJ software and an inverted microscope with a camera (AMG, USA).
Immunohistochemistry (IHC)
All clinical samples were collected from patients with primary glioma who underwent surgery at Nanyang Central Hospital. IHC was performed using paraffin-embedded sections (3 μm thick) of different grades of glioma and adjacent normal tissues. After dehydration with ethanol (70%, 90%, and 100%) and xylene, the stained sections were fixed. Endogenous peroxides were blocked with 0.3% H2O2 for 30 min. Subsequently, the sections were rinsed with PBS for 5 min. IRX5 and NRP1 primary antibodies were added after blocking in the blocking solution for 30 min. Sections were washed thrice with PBS, and the secondary antibody was added and incubated for 1 h at room temperature. Then, each well was washed three times with PBS, coloured for 5 min with DAB, and labelled with horseradish peroxidase for 30 min. DAPI was used to counterstain the nuclei.
Nissl staining
The sections were dehydrated for 1 h at room temperature with a mixture of chloroform and 100% ethanol (1:1) overnight. On the following day, nerves were rinsed twice with 95% and 100% ethanol for 15 min after which they were rinsed three times with distilled water. Nerves were stained at 37°C for 10 min and washed with distilled water. Subsequently, the samples were dehydrated with 95% ethanol for 5 min and 100% absolute ethyl alcohol for 10 min. Finally, neurons were randomly selected from each group for statistical analysis.
Statistical analysis
All computational and statistical analyses were performed using roentgen software v4.0.3 and Prism eight GraphPad (https://www.graphpad.com/scientific-software/prism/). The links between CRNDE expression and targets of interest, such as immune cell infiltration scores, TMB, and MSI, were assessed using the Spearman Correlation test. Data aer presented as means ± SD. A Student’s t-test was used for the analysis of gene expression data from the TCGA, GTEx, and CCLE databases. Pearson’s correlation test was used to assess the association among CRNDE, IRX5 and NRP1 in glioma. Statistical significance was defined as p < 0.05.
Results
Pan-cancer expression profiles of CRNDE
To investigate the expression of CRNDE from a pan-cancer perspective, we first described the expression level of CRNDE in multiple cancer types based on TCGA and GTEx databases. As shown in Figure 1(a), CRNDE was upregulated in 13 tumour types, whereas it was downregulated in two tumour types. To further validate the above results, we analysed four microarray sequencing datasets, GBM (GSE7696), KIRC (GSE53757), LIHC (GSE102079), and ACC (GSE33371), from the GEO database. As shown in Supplementary Figure 1, CRNDE was overexpressed in these four cancer types (p < 0.05). Thus, CRNDE expression in cancer patients in the GEO database was consistent with the results from TCGA database. Following that, we analysed the expression of CRNDE in 22 cancer cell lines from the CCLE dataset and found that CRNDE expression levels were overexpressed in different cancer cell lines and immune cell types (Figure 1(b), p < 0.001). Furthermore, the pan-cancer clinical relevance of CRNDE was analysed. Then, the association between CRNDE expression and age, sex, and race was also analysed. Patients in stage II had significantly higher CRNDE expression levels than patients in other stages (Figure 2(a), p < 0.001), patients older than 41 years had a significantly higher CRNDE expression level than younger patients (Figure 2(b), p.< 0.001), and female patients had higher CRNDE expression levels than male patients (Figure 2(c), p < 0.001). Meanwhile, racial disparity existed, and black patients had significantly higher CRNDE expression levels than Caucasian and Asian patients (Figure 2(d), p < 0.001). CRNDE expression in pan-cancer base on TCGA. Expression level of CRNDE across TCGA tumors compared to TCGA normal and GTEx data using GEPIA (a) and CCLE (b). It’s clear that CRNDE was notable upregulated in 11 cancers (red) and downregulated in two cancers (green). T: tumor; N: normal; n: number. Yaxis: transcript per million (log2 (TPM +1)). X axis: number of tumor and normal samples. *p < 0.05, **p < 0.01, ***p < 0.001. ACC: adrenocortical carcinoma; BLCA: bladder urothelial carcinoma; BRCA: breast invasive carcinoma; COAD: colon adenocarcinoma; DLBC: lymphoid neoplasm diffuse large B-cell lymphoma; ESCA: esophageal carcinoma; GBM: glioblastoma multiforme; HNSC: head and neck squamous cell carcinoma; KICH: kidney chromophobe; KIRC: kidney renal clear cell carcinoma; KIRP: kidney renal papillary cellcarcinoma; LAML: acute myeloid leukemia; LGG: brain lower grade glioma; LIHC: liver hepato cellular carcinoma; LUAD: lung adenocarcinoma; LUSC: lung squamous cell carcinoma; OV: ovarian serous cystadeno carcinoma; PAAD: pancreatic adenocarcinoma; PRAD: prostate adenocarcinoma; READ: rectum adenocarcinoma; SKCM: skin cutaneous melanoma; STAD: stomach adenocarcinoma; TGCT:testicular germ cell tumors; THCA: thyroid carcinoma; THYM: thymoma; UCEC: uterine corpus endometrial carcinoma; UCS: uterine carcinosarcoma. Relationship between CRNDE expression and clinical features in pan-cancer base on TCGA. Box-plots indicate CRNDE expression in different clinical features, inluding (a) stage, (b) age, (c) gender, and (d) race of pan-cancer from the TCGA dataset.

Pan-cancer prognostic significance of CRNDE expression
Using TCGA and GEO datasets, we investigated the link between CRNDE expression and pan-cancer prognosis. CRNDE had different prognostic values for different types of cancer. The high expression of CRNDE indicated a worse OS rate in patients with ACC (p < 0.01), GBM (p < 0.01), KIRC (p < 0.05), LGG (p < 0.001), THCA (p < 0.05), and LIHC (p < 0.05) and showed better clinical outcomes (OS) in LUAD (p < 0.001), and UCEC (p < 0.05) (Figure 3). Furthermore, as shown in Table 1, the Cox regression analysis indicated that CRNDE comprised a poor prognostic factor for OS in ACC (HR = 2.170, p = 0.001), GBM (HR = 1.288, p = 0.02), KICH (HR = 3.544, p = 0.002), KIRC (HR = 1.325, p = 0.002), LGG (HR = 1.905, p < 0.001), and THCA (HR = 2.838, p = 0.001). Meanwhile, CRNDE comprised a good prognostic factor for OS in KIRP (HR = 0.776, p = 0.039), LUAD (HR = 0.695, p < 0.001), THYM (HR = 0.315, p = 0.001), and UCEC (HR = 0.819, p = 0.044). The Cox regression analysis also indicated that high CRNDE expression was similarly linked to poor DSS in patients with ACC (HR = 2.123, p = 0.002), GBM (HR = 1.338, p = 0.014), KICH (HR = 2.138, p = 0.031), KIRC (HR = 1.253, p = 0.028), LGG (HR = 2.094, p < 0.001), and THCA (HR = 2.91, p = 0.006). Moreover, a high level of CRNDE expression was found to be a poor predictive factor for PFI with ACC (HR = 1.742, p = 0.002), LIHC (HR = 1.204, p = 0.02), and LGG (HR = 1.881, p < 0.001). Clinically, CRNDE was probably an independent prognostic factor for ACC, GBM, KICH, KIRC, KIRP, LGG, LUAD, THCA, and THYM. Kaplan-Meier Survival curves of CRNDE in pan-cancer base on TCGA. The Kaplan–Meier curves of OS according to CRNDE expression of (a) ACC, (b) GBM, (c) KIRC, (d) LGG, (e) LIHC, (f) LUAD, (g) THCA, (h)UCEC. Univariate Cox regression analysis of CRNDE.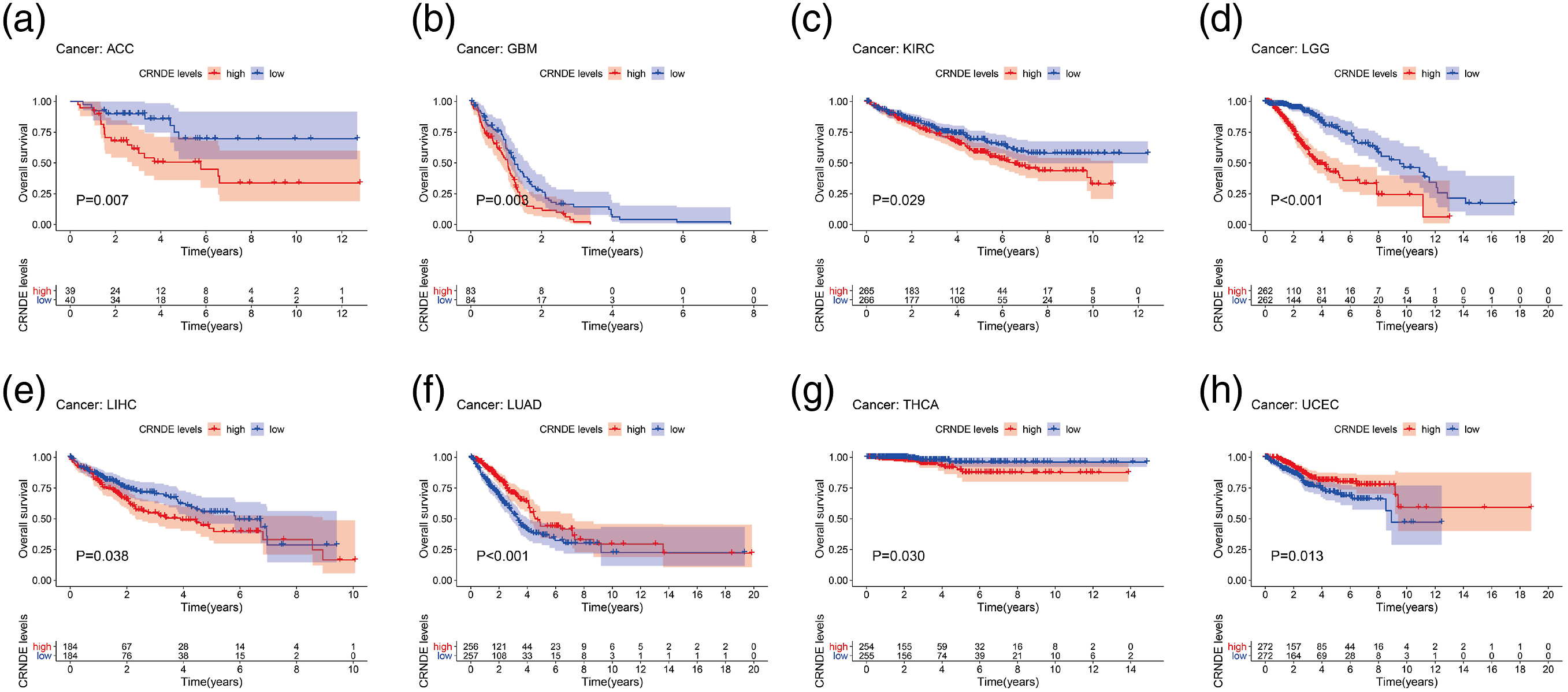

Correlation between CRNDE/IRX5 expression and TMB and MSI
Tumor mutation burden (TMB) and MSI are important determinants in the formation and progression of malignancies, and they are closely associated with tumour prognosis and immunotherapy response. Spearman correlation analysis demonstrated that expressions of CRNDE and its target gene IRX5 were significantly negatively associated with TMB in BRCA and ESCA. However, they were positively associated with TMB in UCEC, STAD, PAAD, KIRP, KIRC, and LGG. Meanwhile, CRNDE and IRX5 were positively associated with MSI in UCEC, STAD, SARC, LUSC, KIRC, and HNSC (p < 0.05, Figure 4(a)–(d)). The correlation between CRNDE/IRX5 and MSI and TMB in pan-cancer. Radar chart showing the correlation between CRNDE and (a) TMB and (b) MSI in 33 cancer types. Radar chart showing the correlation between IRX5 and (c) TMB and (d) MSI in 33 cancer types. Green and blue numbers represented the Spearman correlation coefficient. *p < 0.05, **p < 0.01, ***p < 0.001.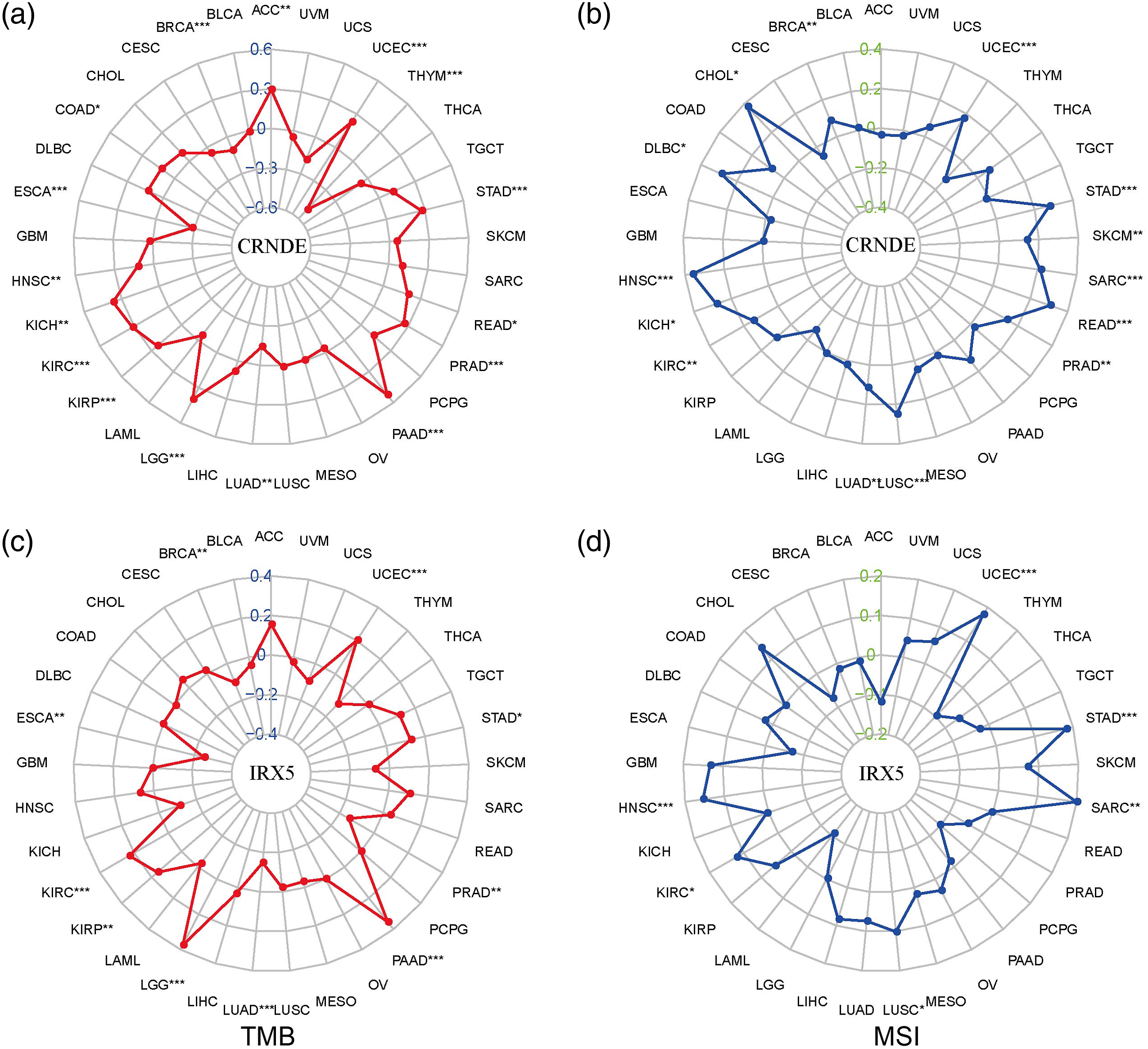
Correlations between CRNDE/IRX5 and tumour microenvironment
In the analysis of the immune infiltration landscape, the Pearson correlation coefficient was used as a measure of the association between CRNDE and IRX5. We found that CRNDE and IRX5 were positively correlated in the seven types of cancer that were associated with prognosis based on TCGA (R > 0.4, p < 0.05, Supplementary Table 2). For example, CRNDE was significantly associated with IRX5 in LGG (R = 0.893, p < 0.001), KIRP (R = 0.836, p < 0.001), and GBM (R = 0.747, p < 0.001). Moreover, CRNDE was also associated with 23 immune checkpoint markers in LGG, including BTLA, CD200, NRP1, TNFSF4, and CD276 (p < 0.05, Figure 5(a)). Interestingly, IRX5 was correlated with 33 immune checkpoint markers in LGG, such as BTLA, TNFRSF14, NRP1, LAIR1, and TNFSF4 (p < 0.05, Figure 5(b)). Next, to measure the relative quantity of tumour-infiltrating immune cells, we used immuneeconv (integrating the six latest algorithms: TIMER, xCell, MCP-counter, CIBERSORT, EPIC, and quanTiseq) to analyse immune infiltration data among seven cancer types (ACC, GBM, KICH, KIRC, KIRP, LGG, and THYM), according to the high expression of CRNDE, which is associated with poor prognosis. Notably, the expression levels of CRNDE and IRX5 were significantly associated with the expression of immune cells (Figure 6). Association of CRNDE and IRX5 gene expression with immune checkpoint markers expression in the seven cancer types that CRNDE associated prognosis. (a) Heat map shown the association of CRNDE expression with immune checkpoint markers expression. (b) Heatmap shown the association of IRX5 expression with immune checkpoint markers expression. For each pair, the top right triangle is colored to represent the p-value, and the bottom left triangle is colored to represent the correlation coefficient. *p < 0.05, **p < 0.01, ***p < 0.001. Association of CRNDE and IRX5 gene expression with immune cells in the seven cancer types that CRNDE associated prognosis. (a) Heat map shown the association of CRNDE expression with immune cells. (b) Heatmap shown the association of IRX5 expression with immune cells. *p < 0.05, **p < 0.01, ***p < 0.001, asterisks represent importance (*P), and different colors represent changes in correlation coefficients.

In response to the microenvironment, macrophages can exhibit different phenotypes(including M1 and M2 macrophages) in multiple stages of tumour progression. Interestingly, CRNDE expression was negatively associated with macrophages in GBM, according to the results of TIMER, XCELL, QUANTISEQ, and EPIC (Figure 6(a)). The results of XCELL and EPIC revealed that the expression of CRNDE was negatively correlated with macrophage M1 in GBM. In contrast, the expression of CRNDE was positively correlated with macrophages (M1 and M2) in LGG according to CIBERSORT, TIMER, XCELL, QUANTISEQ, and EPIC. CIBERSORT, XCELL, and QUANTISEQ showed a positive correlation between CRNDE and macrophage M2 in LGG. The result of IRX5 was consistent with that of CRNDE was observed in gliomas of different grades (p < 0.05, Figure 6(b)).
Validation of CRNDE and IRX5 in gliomas based on GEO and CGGA databases
Furthermore, we clarified the relationship between CRNDE and clinical features and prognosis. CRNDE expression was correlated with age, histological grade, chemo status, PRS type, histology, IDH mutation status, and 1p19q codeletion status of glioma patients based on CGGA (Figure 7(a)–(f), p < 0.001). The Kaplan-Meier analysis revealed that the high expression of CRNDE was related to poor prognosis (Figure 7(g)), and time-dependent ROC curve analysis showed that CRNDE was a good predictive marker for gliomas (Figure 7(h)). The results were further confirmed by data from the GEO database (p < 0.01, Figures 7(i) and (j)). To expand on the relationship between CRNDE, IRX5, and immune checkpoint markers, a total of 613 genes were found to be co-expressed with CRNDE based on TCGA database (R > 0.5, p < 0.001, Supplementary Table 3). A list of the top 20 positive correlation and top five negative correlation genes is shown in Supplementary Figure 2A. A circle chart of the correlation between CRNDE and the top five target genes is shown in Supplementary Figure 2B. Meanwhile, we observed that increased IRX5 expression was associated with poor prognosis in gliomas (p < 0.05, Supplementary Figure 2C–E). Clinical correlation analyses between CRNDE expression and prognosis in glioma. (a–f) The different expression of CRNDE in clinical features included (a) age, (b) grade, (c) Chemo status, (d) PRS type, (e) IDH mutation status and (f) 1p19q codeletion status. (g) Kaplan-Meier Survival curves of CRNDE based on CGGA dataset. (h) Time-dependent ROC curve depicting the predictive accuracy of the signature for OS in the entire dataset. (i and j) Validation of the prognosis of CRNDE in two GEO datasets.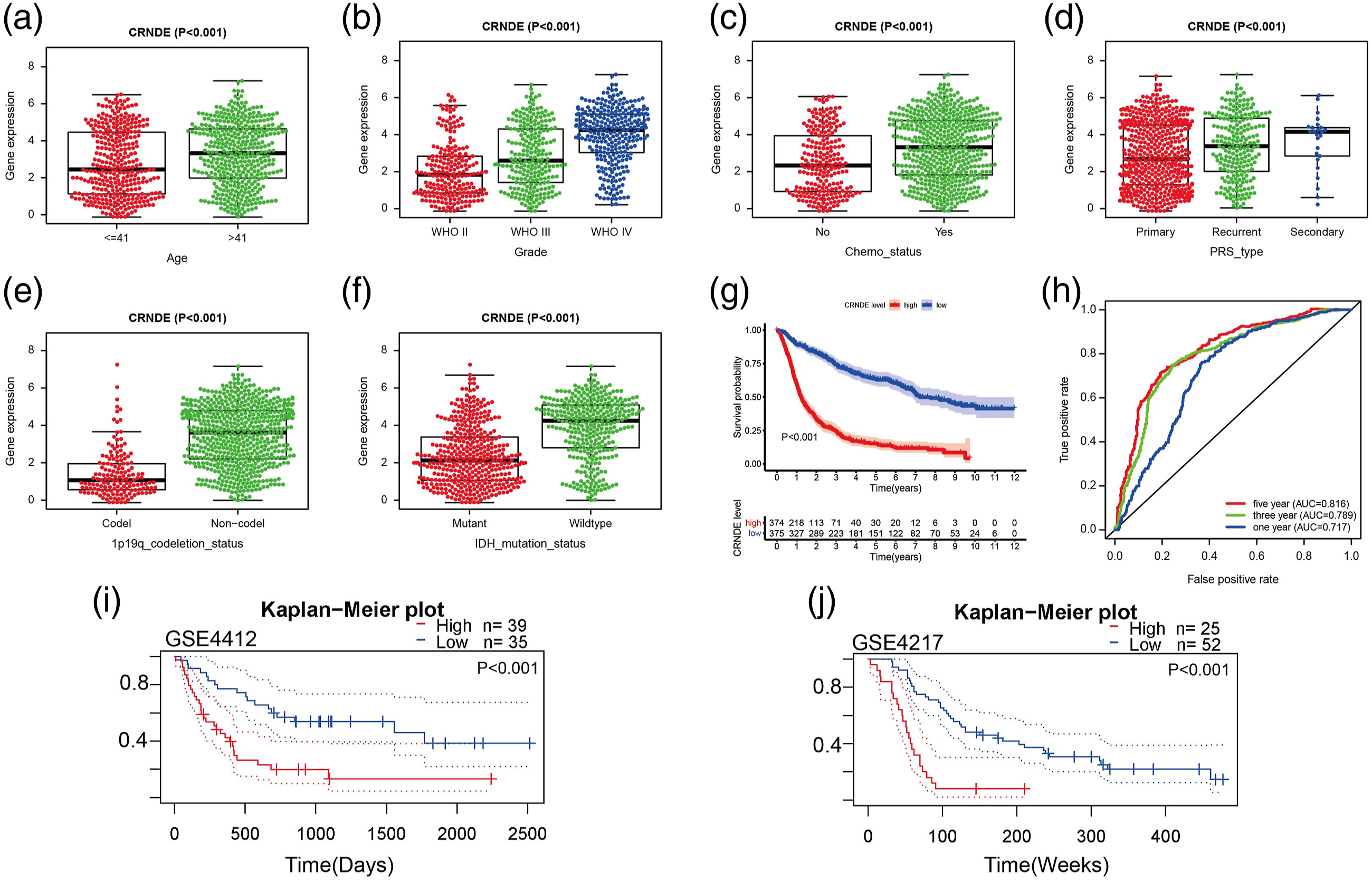
Establishment of a reliable nomogram for prediction of glioma prognosis
Univariate-Cox regression analysis uncovered a strong association of CRNDE, PRS type, histology, grade, age, chemostatus, IDH mutation, and 1p19q codeletion with glioma prognosis (Figure 8(a)). Among them, CRNDE, PRS type, grade, age, chemostatus, IDH mutation, and 1p19q codeletion served as independent prognostic indicators of glioma (Figure 8(b)). By integrating the above seven independent prognostic indicators, we used the nomogram survival to evaluate the survival prognosis of glioma patients (Figure 8(c)). In addition, we evaluated the nomogram prediction function using calibration curves. Our data showed that the 1-, 3-, and 5-years survival predicted by this nomogram was close to the actual survival (Figure 8(d)), suggesting the utility of our nomogram prediction. Establishment and validation of the nomogram for predicting OS of glioma patients. Clinical correlation analyses between CRNDE expression and prognosis in glioma base on CGGA: the univariate Cox analysis (a); the multivariate Cox analysis (b). Prognostic nomograms to predict the overall survival of glioma patients (c). Calibration plot shows the comparison between nomogram-predicted and actual 1-, 3-, 5-years OS (d).
Knockdown of CRNDE inhibited glioma cell proliferation, invasion, and migration and affected apoptosis
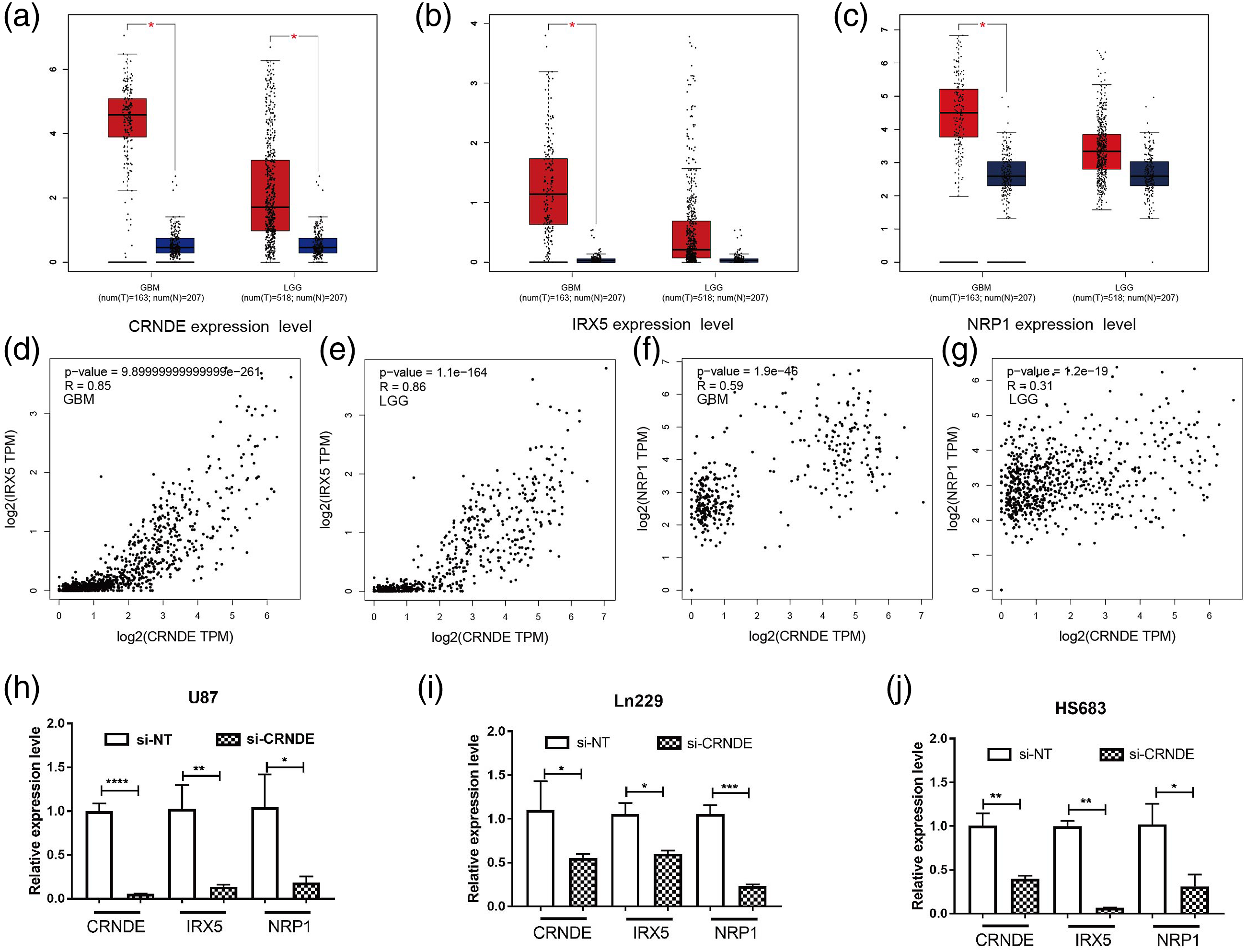
We further elucidated the anti-tumour role and mechanism of CRNDE in glioma cells. Additionally, MTT showed that the knockdown of CRNDE significantly inhibited glioma cell (LN229, HS683, and U87) proliferation (p < 0.05, Figure 9(a)–(c)). The outcome of the transwell assay indicated that CRNDE knockdown inhibited glioma cell migration (p < 0.05, Figures 9(d)–(f)). The wound healing assays revealed that the downregulation of CRNDE significantly decreased invasion (p < 0.01, Figures 9(g)–(i)). These in vitro results demonstrated that the knockdown of CRNDE suppressed glioma cell proliferation, migration, and invasion. Next, we compared CRNDE, IRX5, and NRP1 expression in glioma cells, which suggested significant differences in gene expression levels between cancerous and adjacent tissues (p < 0.05, Figure 10(a)–(c)). Interestingly, an investigation into the relationship between the immune checkpoint genes NRP1, CRNDE, and IRX5 in glioma cells revealed a significant association (R > 0.3, p < 0.001; Figures 10(d)–(g)). To further understand the correlation between CRNDE and IRX5 and immune checkpoint marker expression in glioma cells, we knocked down CRNDE (si-CRNDE) via transient transfection in U87, LN229, and HS683 cells. As expected, the expression level of the immune checkpoint marker NRP1 and the target mRNA IRX5 were notably reduced in U87, LN229, and HS683 cells (Figure 10(h)–(j), P < 0.05). Compared with para-carcinoma tissues, a significant upregulation of IRX5 and NRP1 mRNA expression in glioma samples of different grades was exhibited. Similar results were observed for the expression levels of IRX5 (green) and NRP1 (red) when examined using immunocytochemistry staining (Figure 11(a)–(c)). In addition, the apoptosis rate was examined using Nissl staining in vivo to confirm the role of CRNDE in glioma samples. The results demonstrated that the knockdown of CRNDE significantly altered the apoptosis rate when compared to the si-NC group (p < 0.0001, Figures 11(d) and (e)). Collectively, CRNDE knockdown not only inhibited glioma cell proliferation, invasion, and migration but also affected apoptosis. Knockdown of CRNDE inhibited glioma cell proliferation, invasion, and migration. (a–c) CRNDE knockdown inhibits proliferation in different grades glioma cells. (d–f) CRNDE knockdown inhibits invasion in different grades glioma cells. (g–i) CRNDE knockdown inhibits migration in different grades glioma cells. Data are shown as mean ± SD n = 3, *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001. The correlation of CRNDE and IRX5 and NRP1 in glioma. (a–c) The expression of (a) CRNDE, (b) IRX5 and (c) NRP1 in different grades glioma. (d–g) The correlationship of CRNDE and IRX5/NRP1 expression in different grades glioma. (h–j) Gene expression of CRNDE, IRX5 and NRP1 was determined by qPCR in (h) U87, (i) Ln229 and (j) HS683. Immunohistochemistry and nissl staining. (a) LGG and GBM Immunohistochemistry stained for IRX5. (b) LGG and GBM Immunohistochemistry stained for NRP1. (c) Graph quantifying immunofluorescent intensity using ImageJ. (d) Nissl staining. (e) Quantitative analysis of the number of Nissl-stained neurons.

Gene set enrichment analysis of CRNDE in gliomas
To predict the potential biological function of CRNDE in gliomas, we performed Gene Set Enrichment Analysis (GSEA) using TCGA GBM dataset. We listed the top 10 KEGG gene sets that had higher significance levels in the high-CRNDE expression group. Tumour-associated pathways such as the cell cycle, p53 signalling pathway, apoptosis, and pathways-in-cancer were investigated. Several immune-relative regulatory gene sets including primary immunodeficiency, ECM receptor interaction, antigen processing and presentation, lysosome, cytokine-cytokine-receptor-interaction, and focal adhesion were enriched in the high-CRNDE group (Supplementary Figure 3).
Discussion
Enhancer RNAs (eRNAs) are a subclass of non-coding RNAs that play a role in multiple crucial signalling pathways and are positively associated with the activity of their associated enhancers and regulated their target genes. They were reported to be involved in tumour growth, tumour metastasis, drug resistance, TIME regulation, angiogenesis, and cancer stem cell self-renewal.30,31 There is increasing evidence that eRNAs play a key role in tumour development and may be a potential target for tumour treatment.3,32,33 In the present study, we identified CRNDE as an important eRNA in the tumour immune system.
Certain studies have indicated that CRNDE is overexpressed in multiple human cancers. Specifically, aberrantly expressed CRNDE could regulate cancer chemoresistance and promote cancer cell growth, metastasis, and progression as a molecular ‘sponge’ or as a ceRNA to regulate miRNA.34–37 Thus, functional investigations are required to elucidate the underlying mechanisms of CRNDE. Notably, emerging evidence has shown that lncRNAs act as critical regulators of gene expression and they play fundamental roles in immune regulation.38,39 For instance, lncRNA inducing MHC-I and immunogenicity of tumour (LIMIT) comprises a cancer immunogenic lncRNA which may be targetable for cancer immunotherapy. LncRNA GATA3-AS1 facilitates tumour immune escape in triple-negative breast cancer L1. LncRNA SATB2-AS1 inhibits tumour metastasis and affects the tumour immune cell microenvironment in colorectal cancer by regulating SATB2. 40 Previous studies have revealed that CRNDE can serve as an important factor in immune cells and cancer cells. Ai et al. (2021) reported that CRNDE was obviously elevated and negatively correlated with IFN-γ production in tumour-infiltrating CD8 + tesla cells isolated from OSCC patients. Mechanistically, CRNDE specifically sponged miR-545–5p to induce T-cell immunoglobulin and mucin domain-3 (TIM-3), thus contributing to CD8 + T-cell exhaustion. Han et al. have revealed that lncRNA-CRNDE can promote the M2 polarization of macrophages through the high expression of CD163 to further promote tumour angiogenesis, which is one of the mechanisms by which CRNDE promotes liver cancer progression. The latter authors also reported that CRNDE could indirectly regulate tumour angiogenesis by promoting M2 polarisation of macrophages; this is also the mechanism of immune regulation in the microenvironment of hepatocellular carcinoma.41,42 However, the pan-cancer relationship between CRNDE and immune infiltration has not been revealed.
The central nervous system has a unique immune environment; however, it can still produce an effective antitumour immune response. Therefore, immunotherapy holds great potential for the treatment of gliomas. IC-targeted therapy has become a treatment for tumours with rapid progress and considerable curative effects. Immunotherapy methods and various CTLA-4- and PD-1-blocking drugs have been approved by drug regulatory agencies as clinical medications for non-small cell lung cancer, melanoma, and other diseases, but the application of ICIs in the treatment of gliomas remains in the pre-clinical trial phase.43,44 In addition to the selection of biomarkers for ICI treatment in patients with different types of gliomas in different stages and the development of a method combining inhibitors with other tumour treatment methods, further basic and clinical studies are needed. 44 Although the function of CRNDE in some tumours has been determined, CRNDE is still not fully understood from the perspective of pan-cancer tumour immunology. Our results link CRNDE to immune infiltration in gliomas, confirming that CRNDE is responsible for tumour immune infiltration regulation, which may form as a biomarker for glioma immunotherapies with immune checkpoint blockades, such as CTLA-4, PD-1, and PD-L1 inhibitors, providing new insights into tumour immunotherapy.
We initially conducted a comprehensive pan-cancer exploration of CRNDE expression and its association with patient clinical-pathological features using public databases (TCGA, GEO, and CGGA). Our research revealed that CRNDE expression was altered in various tumours and had different clinical outcomes (OS, DSS, and PFI) in different types of cancer. Additionally, we combined the direct targets of CRNDE IRX5 in our analysis. The relationship between both CRNDE and IRX5 and the immune microenvironment in 33 cancers were explored. The results demonstrated that both CRNDE and IRX5 were positively associated with MSI in patients with UCEC, STAD, SARC, LUSC, KIRC, and HNSC. Meanwhile, both CRNDE and IRX5 were positively associated with TMB in patients with UCEC, STAD, PAAD, KIRP, KIRC, and LGG. The results of the immune infiltration analysis showed that the low expression of CRNDE promoted the immune response of macrophage M1 and inhibited the immune response of macrophage M2 to inhibit tumour growth. The results of GSEA analysis also provided evidence to support the above statement. In addition, we concentrated on exploring the role of CRNDE and IRX5 in GBM and LGG by molecular biological methods. The high expression of CRNDE may have promoted glioma cell invasion and metastasis. Meanwhile, we knocked down CRNDE (si-CRNDE) via transient transfection in U87, LN229, and HS683 cells. As expected, the results suggested a possible interaction between both CRNDE and IRX5 and the immune checkpoint NRP1. The transwell assay confirmed that CRNDE promoted the invasion and migration of glioma cells. The immunohistochemistry results confirmed that IRX5 and NRP1 were highly expressed in glioma clinical tissues compared with para-carcinoma tissues. Moreover, Nissl staining confirmed a significant increase in Nissl+ neurons in si-CRNDE compared to si-NC, which found that si-CRNDE was able to inhibit the apoptosis of glioma cells and suggested the potential clinical utility of CRNDE in cancer therapy. We utilised multiple databases to analyse CRNDE and to explain the potential role of CRNDE in cancer. To the best of our knowledge, previous studies have not emphasised the independent pan-cancer prognostic capabilities of CRNDE, which were confirmed in our study. Immune checkpoint markers play an important role in tumours. However, previous research has failed to explore how CRNDE affects glioma immunity. Therefore, we not only analysed the relationship between CRNDE and its target gene IRX5 but also examined the immune signature genes related to CRNDE/IRX5 that are associated with prognosis. This research design for CRNDE and gliomas has, to the best of our knowledge, never been used or reported on before, by which it additionally provided a new perspective for future glioma research.
However, even though we explored and incorporated information from several databases and molecular biology experiments, there are still some limitations in the present study. First, the matrix processing method introduced biases into the measurement of immune cell levels. Second, we only evaluated the expression and prognosis of CRNDE and verified the expression through in vitro experiments, and the mechanism levels of CRNDE were not further clarified. Third, although CRNDE/IRX5 were found to be related to immune infiltration, as verified by RT-qPCR in glioma cells, experimental evidence for this conclusion has been insufficient. Thus, further laboratory validation is needed to verify the expression and function of CRNDE in vivo and in vitro.
Conclusions
Our comprehensive pan-cancer study of CRNDE broadened the clinical utility of eRNA, offering several avenues for further research on CRNDE/IRX5 expression, clinical relevance, target genes, and functions in tumorigenesis. Importantly, we set the stage for further investigations on the relationship between CRNDE/IRX5 and immune response. The latter may offer valuable new biomarkers for the immunotherapy of glioma and most human cancers. Future prospective research concentrating on CRNDE expression and the TIME would be useful to clarify the relationship between CRNDE/IRX5 and immune mechanisms.
Supplemental Material
Supplemental Material - CRNDE, an enhancer RNA of prognostic value in glioma, correlates with immune infiltration: A pan-cancer analysis
Supplemental Material for CRNDE, an enhancer RNA of prognostic value in glioma, correlates with immune infiltration: A pan-cancer analysis by Zheng Huo, LiShuo Guo, Wei Shao, Qian Ding, Yue Guo, and Qian Xu in European Journal of Inflammation
Supplemental Material
Supplemental Material - CRNDE, an enhancer RNA of prognostic value in glioma, correlates with immune infiltration: A pan-cancer analysis
Supplemental Material for CRNDE, an enhancer RNA of prognostic value in glioma, correlates with immune infiltration: A pan-cancer analysis by Zheng Huo, LiShuo Guo, Wei Shao, Qian Ding, Yue Guo, and Qian Xu in European Journal of Inflammation
Supplemental Material
Supplemental Material - CRNDE, an enhancer RNA of prognostic value in glioma, correlates with immune infiltration: A pan-cancer analysis
Supplemental Material for CRNDE, an enhancer RNA of prognostic value in glioma, correlates with immune infiltration: A pan-cancer analysis by Zheng Huo, LiShuo Guo, Wei Shao, Qian Ding, Yue Guo, and Qian Xu in European Journal of Inflammation
Supplemental Material
Supplemental Material - CRNDE, an enhancer RNA of prognostic value in glioma, correlates with immune infiltration: A pan-cancer analysis
Supplemental Material for CRNDE, an enhancer RNA of prognostic value in glioma, correlates with immune infiltration: A pan-cancer analysis by Zheng Huo, LiShuo Guo, Wei Shao, Qian Ding, Yue Guo, and Qian Xu in European Journal of Inflammation
Footnotes
Authors’ contributions
Q.X. and Z.H. conceived and designed the study. W.S, and Q.D. collected the data. L.S.G. and Z.H. analysed and interpreted the data. Y.G. and Q.D. verified the data. Q.X. and Z.H. performed all the statistical analysis. Q.X., W.S., Y.G., and Z.H. prepared figures and prepared the manuscript. All authors have read and approved the final manuscript.
Declaration of Conflicting Interests
The authors declare that there is no conflict of interest. The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was supported by the Natural Science Foundation of China (#81701185), Key Medical Science and Technology Program of Henan Province (#LHGJ20191463), and the specialised Science and Technology Key Project of Henan Province (#202102310406).
Ethical approval
This study and consent procedure were approved by Ethics Committee of Nanyang Central Hospital.
Data availability
TCGA pan-cancer datasets for this study can be downloaded at https://xenabrowser.net/datapages/, NCBI GEO (https://www.ncbi.nlm.nih.gov/gds), and CGGA (![]() ), accession numbers: GSE7696, GSE102079, GSE53757, GSE33371, GSE4412, and GSE4217. All the experiments were carried out in accordance with the Nanyang Central Hospital Ethics Committee’s guidelines and regulations. Institutional ethics committee approval was obtained for this study, and informed consent was obtained from all patients or their relatives.
), accession numbers: GSE7696, GSE102079, GSE53757, GSE33371, GSE4412, and GSE4217. All the experiments were carried out in accordance with the Nanyang Central Hospital Ethics Committee’s guidelines and regulations. Institutional ethics committee approval was obtained for this study, and informed consent was obtained from all patients or their relatives.
Supplemental Material
Supplemental material for this article is available online.
Appendix
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
