Abstract
Currently, no glucocorticoid dose prediction model is available for clinical practice. This study aimed to utilise machine learning techniques to develop and validate personalised dosage models. Participants were patients with SLE who were registered at Nanfang Hospital and received prednisone. Univariate analysis was used to confirm the feature variables. Subsequently, the random forest (RF) algorithm was utilised to interpolate the absent values of the feature variables. Finally, we assessed the prediction capabilities of 11 machine learning and deep-learning algorithms (Logistic, SVM, RF, Adaboost, Bagging, XGBoost, LightGBM, CatBoost, MLP, and TabNet). Finally, a confusion matrix was used to validate the three regimens. In total, 129 patients met the inclusion criteria. The XGBoost algorithm was selected as the preferred method because of its superior performance, achieving an accuracy of 0.81. The factors exhibiting the highest correlation with the prednisone dose were CYP3A4 (rs4646437), albumin (ALB), haemoglobin (HGB), anti-double-stranded DNA antibodies (Anti-dsDNA), erythrocyte sedimentation rate (ESR), age, and HLA-DQA1 (rs2187668). Based on validation, the precision and recall rates for low-dose prednisone (⩾5 mg but <7.5 mg/d) were 100% and 40% respectively. Similarly, for medium-dose prednisone (⩾7.5 mg but <30 mg/d), the accuracy and recall rates were 88% and 88%, and for high-dose prednisone (⩾30 mg but ⩽100 mg/d), the accuracy and recall rates were 62% and 100% respectively. A robust machine learning model was developed to accurately predict prednisone dosage by integrating the identified genetic and clinical factors.
Keywords
Introduction
Systemic lupus erythematosus (SLE) is an autoimmune inflammatory connective tissue disease affecting multiple organs. The global incidence of SLE is estimated to range from 1.5 to 11 per 100,000 individuals per year. 1 Glucocorticoids (GCs) continue to serve as a fundamental component in the management of SLE, 2 Despite the availability of immunosuppressive agents, 3 up to one third of patients never discontinue GCs. 4 GCs have pros and cons. GCs remain statistically linked to the occurrence of severe infections and delayed cardiovascular morbidity in SLE.5,6 In addition, GCs exert deleterious effects on bone resorption 7 and even induce bone necrosis. 8 GCs are an independent predictor of decreased quality of life in SLE. 9 The harmful effects of glucocorticoids on chronic organ damage have recently been validated in recent studies. 10 Excessive immune activation caused by GCs may result in long-term inflammation. GCs encourage the growth of harmful T helper cells and lead to tissue damage by causing inflammation in the nerves and intestine. 11 The absence of GC therapy may be associated with the absence of chronic harm accumulation. 12 Based on the uncertainty of benefits versus risks, the optimal management of GCs in SLE is recognised as a significant contemporary challenge. 13
The efficacy of a drug response is influenced by various factors, including pharmacogenomics and patient-specific attributes such as sex, age, and concurrent medication. Pharmacogenomics plays a pivotal role in personalised medicine. 14 Therefore, it is imperative to further investigate and validate the synthesis of these potential factors and their application in devising prednisone dose adjustment strategies. Machine learning is a promising avenue for integrating and leveraging data from patients 15 and exhibits superior generalisation and accuracy compared with traditional models.16–18 Notably, recent advancements in algorithmic principles have led to the development of more sophisticated algorithms such as eXtreme Gradient Boosting (XGBoost), light gradient boosting machine (LightGBM), Categorical Boosting (CatBoost), Gradient Boosting Decision Tree (GBDT), and TabNet, which have received significant recognition within the algorithmic community.19–23 Recently, a surge in the application of machine- and deep-learning methodologies in practical applications has been observed. A notable example is the development of a predictive model for tacrolimus blood concentration in patients with autoimmune disorders. 24 In addition, CatBoost was used to develop a pioneering model for sodium valproate dose prediction. 25 Numerous studies have demonstrated the advantages of machine-learning algorithms over traditional statistical techniques.
Despite the widespread utilisation of GCs, very few small-scale clinical trials26–29 and observational studies30–33 have been conducted to ascertain the optimal form, dosage, or administration regimen. Currently, there is no GCs dose prediction model that can provide guidance for clinical practice. In the Systemic Lupus International Collaborating Clinics Inception Cohort, 2 the use of GCs was primarily based on physicians’ clinical practices rather than individual patient characteristics. To optimise the management of GCs, our study aimed to employ machine learning techniques to construct personalised dosage models for GCs.
Materials and methods
Study design and participants
This prospective observational study was conducted at NanFang Hospital, affiliated with Southern Medical University between 2017 and 2023, and enrolled adult patients with SLE diagnosed based on the 1997 American College of Rheumatology classification criteria. Every 3 months, the daily prednisone dosage and laboratory parameters were recorded, and SLE disease activity was assessed. During the one-year follow-up period, rheumatologists made adjustments to the GC therapy based on the daily prednisone dose and disease remission status. The study protocol was approved by the Ethics Committee of Nanfang Hospital, Southern Medical University (NFEC-2021-320) and all patients provided written informed consent.
The inclusion criteria for this study were: (1) initial doses of ⩾5 mg/day at the baseline visit; (2) ability to complete 1 year of follow-up. The exclusion criteria were as follows: (1) initiation of GCs prior to study enrolment, (2) irregular medication, (3) intricate and severe coexisting medical conditions and a projected lifespan shorter than the duration of the study. Patients who met the inclusion criteria and none of the exclusion criteria underwent polygenic drug testing. They were then closely monitored and followed up for 1 year, during which relevant indicators of efficacy and safety were systematically collected and documented. Finally, 129 patients who received daily doses of prednisone were included in this study. The patient enrolment process is illustrated in Figure 1.
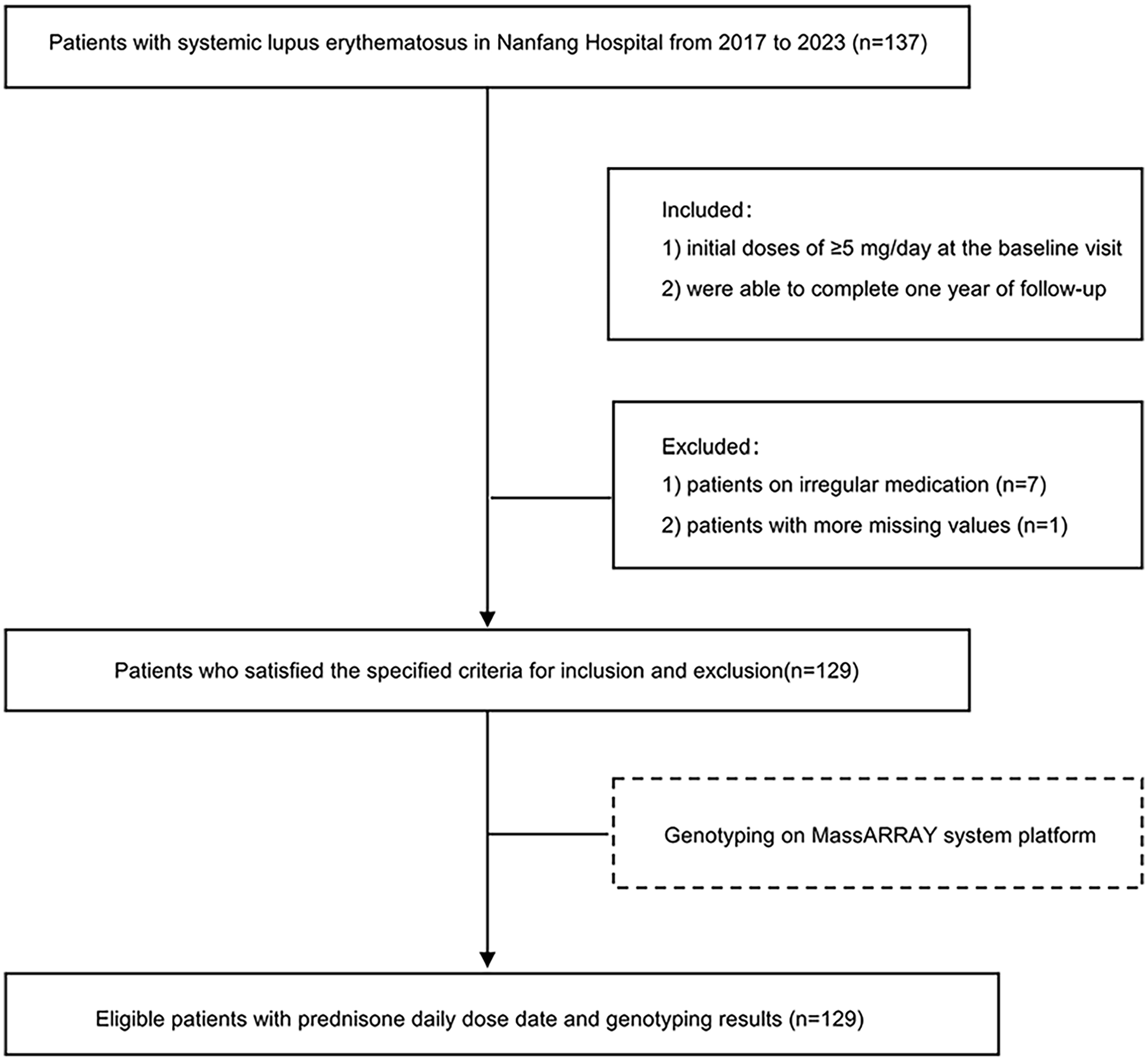
Enrolment of patients.
Patients and clinical assessment
The demographic characteristics, inspection information and combination of medication were recorded at baseline. Further, information collected included erythrocyte sedimentation rate (ESR), C-reactive protein (CRP), anti-nuclear antibodies (ANA), anti-double-stranded DNA antibodies (Anti-dsDNA), Sm antibodies (Anti-Sm), RNP antibodies (Anti-RNP), alanine aminotransferase (ALT), albumin (ALB), serum creatinine (Scr), uric acid (UA), white blood cell count (WBC), blood platelet count (PLT), haemoglobin (HGB), Ca, K, and glucose (Glu), total cholesterol (TC), triglyceride (TG), low density lipoprotein (LDL), high density lipoprotein (HDL). Data on disease activity according to the SLE disease activity index (SLEDAI) were collected. When SLEDAI ⩽ 4, the prednisone dose is appropriate and the treatment is effective.
GCs use
The average daily oral prednisone doses between visits were recorded. The oral average daily prednisone doses were categorised into three categories based on clinical expertise31,34: low (⩾5 mg but <7.5 mg/d), medium (⩾7.5 mg but <30 mg/d), and high (⩾30 mg but ⩽100 mg/d).
Genotyping
Drug-related target genes were identified by conducting a comprehensive search of the PharmGKB, Clinical Pharmacogenetics Implementation Consortium (CPIC), and PubMed databases to identify medications commonly used for the treatment of SLE. A total of 45 single nucleotide polymorphisms (SNPs) in the target genes (Supplemental Table S1) were assessed. The primer design was completed and validated by Simcere Diagnostics. Venous blood samples were collected from the participants and genomic DNA was isolated. The amplified samples were analysed using a computer-based assay specifically using the MassARRAY system platform. The specimens were collected on a microchip and analysed. Specimens were obtained from the microchip and subsequently subjected to scanning. The resulting scan data were processed using Typer4.0 software. Time-flight mass spectrometry was used to determine the mutation status of individual target genes by quantifying the nucleotide count at a specific mutation site. A peak was classified as a mutation if it was in close proximity to the base of the mass spectrum peak.
Variable selection and model establishment
The modelling process is illustrated in Figure 2. To ensure that the study design for the three-category problem was robust and to enhance the reliability of the results, we selected Cohen’s w to calculate the required sample size. Classification variables such as sex, medication combination, and genetic type were encoded using heat as the sole method. To address extreme imbalances, classification variables with a single category comprising more than 90% of the total should be removed. In addition, variables with more than 20% missing data were excluded. To ascertain the pertinent factors associated with prednisone dosage, feature screening was performed using univariate differential analysis. It is imperative to train both linear and nonlinear models to ensure data precision. Next, the implementation of single-factor feature screening was performed. It was necessary to interpolate the missing values of the seven variables to estimate the dosage for patients diagnosed with SLE. The random forest method (RF) proved effective in managing the complex relationships among numerous characteristics, thus facilitating the interpolation of missing values using RF. Statistical analysis of the modelling data after interpolation is presented in Supplemental Table S2.

Workflow of data processing and model establishment.
The dataset was divided into a training cohort and a validation cohort in an 8:2 ratio, with the objective variable ‘prednisone daily dose’ and seven selected variables serving as covariates. Ten algorithms were constructed to predict the prednisone dose in patients with SLE. The training set was further divided into six segments, with five segments assigned for training and the sixth segment designated for verification. This process was repeated six times with each iteration involving the selection of a unique verification set. Finally, the performance of the model was evaluated by computing the mean of six validation outcomes. The selection grid search technique was used to identify the most favourable combination within the model’s parameter space, resulting in the highest performance. Six-fold cross-validation was performed to evaluate the model performance and ascertain the optimal parameter combination for each scenario. The specifics of the model and validation optimisation parameters are presented in Supplemental Tables S3 and S4.
Statistical analysis
Patient characteristics are reported as median (IQR), depending on the degree of similarity to a normal distribution. Categorical variables were analysed, and the results included both absolute and relative frequencies. The Mann–Whitney U test was used to perform univariate analysis of continuous variables. The chi-squared test was used to assess categorical data when the sample size was ⩾ 5, whereas Fisher’s exact test was used when the sample size was <5. A confusion matrix was generated using Sklearn software. The machine learning and deep learning algorithms were executed on a Windows 10 operating system utilising an Intel® Core™ i5-10400F CPU @ 2.90 GHz with 12 CPUs and 512 GB of RAM. Data analysis was performed using Python 3.8.8 and IBM SPSS Statistics 22. The significance threshold was defined as P < 0.05 (two-sided).
Results
Baseline information
In total, 129 patients diagnosed with SLE were enrolled in this study. Baseline characteristics and genotyping information are shown in Table 1 and Supplemental Table S5. The median age was 35 years (IQR 18–71), and 89.15% were female. Hydroxychloroquine (HCQ) was the most commonly used combination medication (80.62%). In addition, each patient was administered calcium supplements and proton pump inhibitors. In addition, the dataset provides information on the missing percentages for each variable. Regarding genotyping, only five SNPs (SLC19A1 rs1051266, ATIC rs4673993, ABCB1 rs2032582, CYP2D6 rs1065852, and ABCB1 rs1045642) had missing data with a miss rate of <2%. There remaining SNPs had no missing data.
Demographic clinical characteristics.

The use of medication corresponds to ‘1’, non-use of medicines corresponds to ‘0’. Continuous variables were presented as ‘mean (interquartile range, IQR)’, while categorical variables were presented as ‘frequency (percentage, %)’.
ESR: erythrocyte sedimentation rate; CRP: C-reactive protein; ANA: anti-nuclear antibodies; Anti-dsDNA: anti-double-stranded DNA antibodies; Anti-Sm: Sm antibodies; Anti-RNP: RNP antibodies; ALT: alanine aminotransferase; ALB: albumin; Scr: serum creatinine; UA: uric acid; WBC: white blood cell count; PLT: blood platelet count; HGB: hemoglobin; Glu: glucose; TC: total cholesterol; TG: triglyceride; LDL: low density lipoprotein; HDL: high-density lipoprotein; SLEDAI: systemic lupus erythematosus disease activity index; Cyc: cyclophosphamide; MMF: mycophenolate mofetil; HCQ: hydroxychloroquine.
Changes in daily prednisone dose and disease activity
After 12 months, among these patients, 13 (10.08%) received a daily prednisone dose of 5–7.4 mg, 69 (53.49%) received 7.5–29 mg, and 47 (36.43%) received 30–100 mg (Figure 3a). According to SLEDAI (Figure 3b), 129 patients had SLEDAI ⩽ 4 and had stable disease control as assessed by a rheumatologist.

Changes in daily prednisone dose and disease activity. (a) Percentage of patients on daily prednisone dose at each visit. (b) Percentage of patients on SLEDAI at each visit. Data labels (numbers) refer to absolute values.
Univariate analysis
Statistically significant associations (P < 0.05) between the univariate and prednisone daily dose variables were identified (Supplemental Table S6). Following a screening procedure, the seven variables in Table 2 were deemed significant and included in the subsequent processing and modelling stages.
List of variables significantly related to prednisone dose.
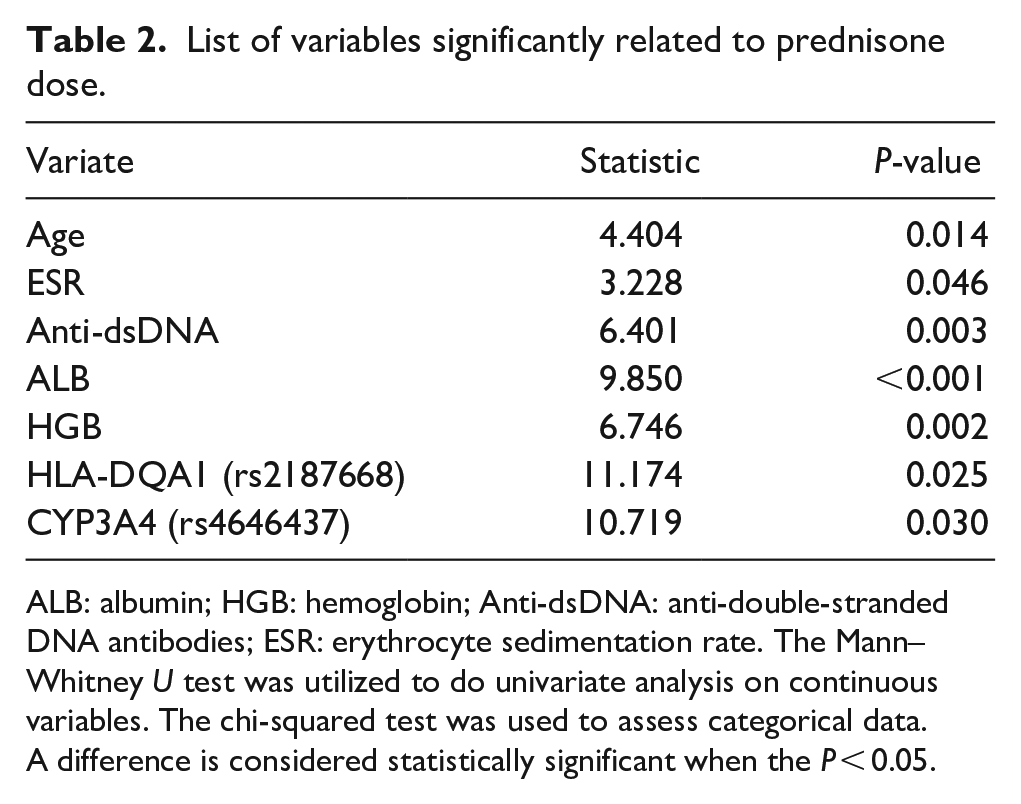
ALB: albumin; HGB: hemoglobin; Anti-dsDNA: anti-double-stranded DNA antibodies; ESR: erythrocyte sedimentation rate. The Mann–Whitney U test was utilized to do univariate analysis on continuous variables. The chi-squared test was used to assess categorical data.
A difference is considered statistically significant when the P < 0.05.
Model performance
Ten algorithms were employed to establish the models, each of which was fitted with the optimal parameters for the training set. The performance of each model was assessed using various evaluation indicators, resulting in four model results: true negative (TN), false negative (FN), true positive (TP), and false positive (FP). Accuracy is frequently used to assess the overall performance of a model by predicting the proportion of correctly classified samples. The accuracy specifically pertains to the ratio of truly positive samples to all samples predicted to be positive. The recall denotes the accurate proportion of all positive samples predicted by the model. The F1 score is the harmonic mean of the accuracy and recall. The formulae are as follows:




Table 3 presents the prediction performances of the nine models. XGBoost had Precision = 1, Recall = 0.4, and F1_score = 0.57 for predicting the low-dose prednisone (⩾5 mg but <7.5 mg/d); Precision = 0.88, Recall = 0.88, and F1_score = 0.88 for predicting the medium-dose prednisone (⩾7.5 mg but <30 mg/d); Precision = 0.62, Recall = 1, and F1_score = 0.77 for predicting the high-dose prednisone (⩾30 mg but ⩽100 mg/d); and Accuracy = 0.81 for the whole XGBoost model, surpassing other algorithms and exhibiting superior overall performance. Consequently, XGBoost was used to predict the daily dosage of prednisone and to evaluate the influence of various factors on dose prediction.
Prediction performance of different algorithms.

Dose Regimen of the daily prednisone dose of ⩾5 mg but <7.5 mg/d corresponds to ‘0’, regimen of the daily prednison dose of ⩾ 7.5 mg but <30 mg/d corresponds to ‘1’ and regimen of the daily prednison dose of ⩾30 but ⩽100 mg/d corresponds to ‘2’.
SVM: support vector machines; RF: random forest; Adaboost: adaptive boosting; XGBoost: eXtreme gradient boosting; LightGBM: light gradient boosting machine; CatBoost: categorical boosting; MLP: multi-layer perceptron.
Therefore, the significance scores of the seven selected variables were calculated and ranked using XGBoost (Table 4 and Figure 4). Notably, the importance score of CYP3A4 rs4646437 (0.311) was remarkably higher than that of the other variables, followed by ALB (0.182) and HGB (0.154). The four subsequent variables, Anti-dsDNA, ESR, Age, HLA-DQA1 (rs2187668), had importance scores of 0.104, 0.097, 0.097, and 0.057, respectively, indicating their significance in the model. A higher importance score signified a greater impact of the variable on the prediction of daily prednisone dose.
Variable importance score.

ALB: albumin; HGB: hemoglobin; Anti-dsDNA: anti-double-stranded DNA antibodies; ESR: erythrocyte sedimentation rate.

Ranking of variable importance.
The validation cohort comprised 26 patients, among whom five patients received a low daily dose (⩾5 mg but <7.5 mg/d), 16 patients received a medium dose (⩾7.5 mg but <30 mg/d), and five patients were prescribed a high dose (⩾30 mg but ⩽100 mg/d). The recommended prednisone dose for patients was determined using a confusion matrix derived from the XGBoost prediction model (Figure 5). The model exhibited a high level of accuracy in recommending a low daily dose regimen for two patients, achieving a precision rate of 100% and a recall rate of 40%. Similarly, the model accurately recommended a medium-dose regimen for 16 patients, with only two patients receiving an incorrect dose. This resulted in precision and recall rates of 88% and 88%, respectively. Furthermore, the model accurately recommended a high-dose regimen for eight patients; however, three patients were prescribed an incorrect dose. The precision rate for this category was 62% and the recall rate reached 100%.

Confusing matrix diagram.
Discussion
In this study, we developed a predictive model to determine the daily dose of prednisone. This involved evaluating three specific dose regimens (⩾5 mg but <7.5 mg/d, ⩾7.5 mg but <30 mg/d, and ⩾30 mg but ⩽100 mg/d). To achieve this, we used a leading-edge deep learning technique that yielded a prediction model with a commendable performance level (accuracy = 0.81). Subsequently, we ranked the variables that exhibited a strong correlation with daily prednisone dose using an importance score. The importance is ranked as follows: CYP3A4 (rs4646437) > ALB > HGB > anti-dsDNA > ESR > HLA-DQA1 (rs2187668). Finally, a confusion matrix was used for model validation, which revealed that the low-dose prednisone regimen exhibited 100% precision and 40% recall rate. Similarly, the medium-dose regimen demonstrated a precision of 88% and recall rate of 88%, whereas the high-dose regimen had a precision of 62% and recall rate of 100%. Our findings indicate that the XGBoost model successfully integrated individual features, enabling their conversion into a strategy for optimising the daily dosage of prednisone in the treatment of patients with SLE. The XGBoost algorithm is a nonparametric estimation ensemble algorithm constructed on decision trees. The influence of the correlation between independent variables on the model is negligible. In cases where the sample size and dataset dimensions are substantial, the XGBoost algorithm exhibits superior performance compared to the logistic regression algorithm. When considering empirical evidence, machine learning and deep learning methodologies possess indisputable advantages over conventional modelling techniques. These advantages include the ability of machine-learning and deep-learning models to handle intricate, high-dimensional, and interactive variables that traditional models cannot accommodate.
Numerous factors have been documented to influence the expression of metabolic enzymes and transporters, thereby affecting the pharmacokinetics of GCs. Cytochromes play a crucial role in the detoxification and maintenance of steroidal compounds. A significant proportion (78%) of hepatic clearance drugs undergo oxidative metabolism mediated by cytochrome P450 in families 1, 2, and 3. Among these, CYP3A4/5 (37%) exhibit the highest contribution. 35 Notably, CYP3A4 is involved in the 6f-hydroxylation of dexamethasone, contributing to the negative feedback loop of dexamethasone autoregulation. 36 Furthermore, prednisone stimulates CYP3A4. 37 The activity of the CYP3A4 enzyme is enhanced by the regulation of CAR receptor protein production, which is involved in transcription, 38 resulting in an improvement in the metabolic activity of CYP3A4 substrates such as drugs used in combination with prednisone. Schirmer et al. 39 discovered that the T > C variation in the CYP3A4 (rs4646437) gene, located on intron 7, influences the expression level of CYP3A4 protein in the liver and the activity of the enzyme. According to our findings, which are supported by previous research, the SNP in CYP3A4 (rs4646437) is the principal determinant of daily prednisone dosage.
An additional genetic variation, specifically HLA-DQA1 (rs2187668), was identified. The HLA-DQA1 genotype (rs2187668) may play a role in the development of cutaneous lupus erythematosus by abnormally affecting antigen presentation. 40 Bullich et al. 41 provide empirical support for the involvement of HLA-DQA1 (rs2187668) in predicting the efficacy of immunosuppressant drugs. In conjunction with our findings, it has been postulated that alterations in these two SNPs could potentially affect prednisone metabolism and therapeutic outcomes. Our study represents a pioneering effort to incorporate a patient’s genetic composition into the management of GC dosing, thereby facilitating the precise administration of GCs, akin to the accurate delivery of immunosuppressive medications such as tacrolimus.
SLE exhibits a distinctly higher prevalence in particular sex and age cohorts, with women in their reproductive years having an elevated vulnerability. 42 We acknowledge that age is a crucial predictive factor in the dosage model. Furthermore, our findings suggest that specific biochemical markers, namely ALB, HGB, anti-dsDNA, and ESR, have considerable significance in the developed model for predicting the daily prednisone dosage. In particular, ALB correlates with the likelihood of mortality from all causes and infections in individuals with severe SLE-related illnesses. 43 Patients with SLE demonstrate a significant decrease in ALB levels compared to healthy individuals. Furthermore, a negative correlation was observed between disease activity and serum ALB levels. 44 Altered ESR has been consistently observed in acute SLE cases. 45 The measurement of anti-dsDNA is a critical factor in evaluating SLE disease activity, particularly in cases involving lupus nephritis. 46 In a personalised medication model for prednisone, ALB, HGB, anti-dsDNA, and ESR serve as notable predictors that can assist clinicians in appropriately adjusting prednisone dosage regimens.
In addition to the factors mentioned above, prognostic indicators and relapse variables include 24-hour proteinuria, urine sediment, and complement C3 and C4 levels, as well as clinical symptoms such as lupus nephritis, neurologic issues, vasculitis, haematological abnormalities, and refractory hypertension. However, insufficient clinical data on these variables hindered our ability to conduct further enquiries. It is crucial to include relevant markers as clinical variables in future investigations to estimate GC dosage.
Patients differ in age, sex, underlying health status, and genetic background, all of which can influence disease activity and treatment response. For example, young patients may tolerate the disease better and show relatively mild symptoms owing to better physical status, even when some indicators are abnormal; however, the condition of older patients or patients with other underlying diseases, even if the same index is abnormal, may be more serious. Patients with SLE usually require long-term drug therapy with glucocorticoids and immunosuppressants. These drugs are effective in managing the disease but may also affect the metrics in the model, leading to a change in the relationship between the indicators and disease severity.
The developed model will help physicians make preliminary assessments of patient sensitivity to glucocorticoids. Kandane-Rathnayake et al. 47 discovered that the dose of GCs was significantly higher in disease with higher activity. Higher doses of GC treatment often indicate low responsiveness, which means that these patients face adverse effects from high-dose GCs; therefore, physicians must consider whether to adjust the treatment strategy and use biological agents instead of GCs in these patients.
This study has some limitations. First, it only included participants of Han Chinese descent. Variations in GC pharmacokinetics according to race may result in a reduced predictive power of our model in patients of other ethnicities. We used a limited sample size from a single medical centre, highlighting the need for a larger samples encompassing several medical centres in further studies. Incorporating additional data will enhance the performance of the model. In addition, the exclusion of partial information on combination medications was attributed to an inadequate number of samples in our prediction model. Consequently, the inclusion of additional ethnic groups and combination regimens is necessary to enhance discerning factors and broaden the applicability of the model.
Conclusion
We developed a GC dose prediction model using machine learning and deep learning methodologies, and subsequently evaluated multiple algorithms. XGBoost exhibited superior performance as a dose prediction model. To facilitate implementation in clinical settings, the model requires further research based on empirical data and potential expansion to incorporate novel variables. This study contributes to a more precise model for the management of GC dosage in patients with SLE.
Supplemental Material
sj-docx-1-iji-10.1177_03946320251331791 – Supplemental material for Construction and evaluation of glucocorticoid dose prediction model based on genetic and clinical characteristics of patients with systemic lupus erythematosus
Supplemental material, sj-docx-1-iji-10.1177_03946320251331791 for Construction and evaluation of glucocorticoid dose prediction model based on genetic and clinical characteristics of patients with systemic lupus erythematosus by Xin Luo, Jinjun Zhao, Danfeng Zou, Xiaoning Luo, Meida Fan, Hongling Hu, Ping Zheng, Yilei Li, Renfei Xia and Liqian Mo in International Journal of Immunopathology and Pharmacology
Footnotes
Acknowledgements
We express our gratitude to Ze Yu and Jinyuan Zhang for their contribution in conducting data analysis.
Author contributions
All authors participated in the study’s design, data collection, analysis, and critical revision for key intellectual content. XL wrote the manuscript and contributed to data curation. JJZ contributed to Methodology and review. MDF and XNL contributed to data curation and investigation. HLH contributed to data analysis. PZ and YLL contributed to project administration. RFX and LQM contributed to study design. All authors read and approved the final manuscript. Additionally, all authors gave their approval for the final version to be published.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by grants from the Wu Jieping Medical Foundation (320.6750.2020-04-1); Guangdong Basic and Applied Basic Research Foundation (2019A1515110148); Medical Research Foundation of Guangdong Province (C2022073); and Hospital Pharmaceutical Research Fund of Guangdong Provincial Hospital Association (YXKY202208).
Ethicals approval
The studies involving human participants were reviewed and approved by the Ethics Committee of NanFang Hospital, Southern Medical University (NFEC-2021-320). The patients provided their written informed consent to participate in this study.
Informed consent
Written informed consent was obtained from all subjects before the study*.
Data availability
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
