Abstract
Keywords
Background
Renal cell carcinoma (RCC), including clear cell RCC (ccRCC), papillary RCC (pRCC), and chromophobe RCC (chRCC), is the most common solid lesion in the kidney, accounting for approximately 90% of kidney malignancies and 3% of all cancers. 1 In 2022, 79,000 new kidney cancer cases and 13,920 kidney cancer-related deaths had been estimated for the United States. 2 Among these three subtypes, ccRCC is the main RCC tissue subtype, accounting for 75% of all RCC diagnoses, while also being the most invasive, with a high metastasis risk and recurrence rate. 3 Studies have shown that 30% of patients with ccRCC present with tumor cell metastasis at the time of diagnosis, 4 while 25%–40% of patients exhibit metastasis after receiving treatment. 5 Conventional chemotherapy and radiotherapy are ineffective in patients with metastatic ccRCC. 6 In recent years, immunotherapy has greatly contributed toward the treatment of cancer. Immune checkpoint inhibitors monotherapy or immune-based combinations are associated with improved survival irrespective of ECOG performance status (PS) 0 or 1. 7 In the pooled overall survival (OS), HR of men treated with single agent immunotherapy was found to be higher than that in women. In other cases, a superior benefit was observed in female patients treated with the immuno-oncology combinations compared to that observed in men. 8 However, for metastatic RCC, which is insensitive to first-line immunotherapy, cabozantinib has been shown to be effective in patients with primary refractory mRCC. 9 Although research has primarily focused on the mechanisms underlying cancer development and progression, the etiology and carcinogenesis of ccCC remain unclear. 5 Therefore, it is important to understand the molecular mechanisms of ccRCC, in conjunction with a pressing need to explore novel prognostic predictors and therapeutic targets for ccRCC.
In the clinical context, biomarkers refer to any measurable molecular indicator for cancer risk, occurrence, or patient outcome. Biomarker testing in cancer involves the profiling of tumor or body fluids to detect changes in DNA, RNA, proteins, or other biomolecules that can provide information related to diagnosis, prognosis, precision medicine/guiding cancer treatment, predicting drug response, or cancer monitoring.10,11 In RCC, biomarkers can be used to predict response and help guide which immunotherapy regimen to use. 12 Activating transcription factor 3 (ATF3) is a stress-induced transcription factor that is central in the regulation of metabolism, immunity, and tumorigenesis, while also being a hub of the cellular adaptive response network. Multiple extracellular signals, such as endoplasmic reticulum stress (ER), cytokines, chemokines, and lipopolysaccharides, are associated with ATF3 induction. 13 Numerous studies have demonstrated ATF3′ role as a regulator of prostate, 14 breast, 15 gastric and colon, 16 lung, 17 and liver 18 cancers. Pelzer et al. observed an upregulation of ATF3 in prostate cancer in vivo and in vitro after androgen stimulation. 19 The overexpression of microRNA-301a-3p promotes cell invasion and proliferation by targeting runt-related transcription factor 3 in prostate cancer. 20 Rohini et al. showed that overexpression of miR-590-3p decreased proliferation and increased the apoptosis in breast cancer cells. 21 In contrast, ATF3 facilitates death receptor 5 (DR5) induction and apoptotic cell death in human p53-deficient colorectal cancer cells after zerumbone or celecoxib treatment. 22 Furthermore, previous works have indicated that saleramide triggers ER stress in human non-small cell lung cancer (NSCLC) cells, which regulates the induction of the ATF4-ATF3-c/EBP homologous protein (CHOP) axis, leading to DR5-dependent apoptosis. 23 These observations suggest that ATF3 can act as an oncogene or a tumor suppressor, however, its role of in ccRCC has been underexplored. A recent study reported that ATF3 inhibits growth and metastasis of ccRCC by deactivating the EGFR/AKT/GSK3β/β-catenin signaling pathway. 24 However, the upstream regulatory mechanisms of ATF3 remain unclear.
Here, we analyzed ATF3 expression in ccRCC as well as its potential correlation with patient survival. Moreover, we explored the miRNA and long non-coding RNA (lncRNA)-associated regulation of ATF3. Our findings showed that ATF3 acts as a suppressor of ccRCC by regulating the apoptotic signaling pathway in response to ER stress.
Materials and methods
UALCAN
The UALCAN (https://ualcan.path.uab.edu/) web resource was used to investigate ATF3 expression and the association between ATF3 and various clinicopathological parameters (cancer stage, nodes metastasis, tumor grade, sex, age, and subtypes) of ccRCC, as well as the survival analysis for ATF3 in pan-cancer. 24
The human protein atlas
The Human Protein Atlas (HPA; https://www.proteinatlas.org/) was used to explore ATF3 protein expression and perform survival analysis for ATF3 in ccRCC. 24
Gene expression profiling interactive analysis
Gene Expression Profiling Interactive Analysis (
Analysis of ATF3-Interacting genes and proteins
The GeneMANIA database (https://www.genemania.org/) was used to construct an ATF3 gene interaction network, while we similarly queried the STRING online database (https://string-db.org/) to build a protein–protein interaction (PPI) network for ATF3.
Gene ontology term and Kyoto Encyclopedia of Genes and Genomes Pathway Enrichment Analyses
The DAVID (https://david.ncifcrf.gov/summary.jsp) online analytical tool was used for Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment and Gene Ontology (GO) analyses including Biological Process (BP), Cellular Component (CC), and Molecular Function (MF) analyses for genes interacting with ATF3. 25
Tumor Immune Estimation Resource (TIMER)
Tumor Immune Estimation Resource (TIMER; https://cistrome.shinyapps.io/timer/) is a web server for the comprehensive analysis of tumor-infiltrating immune cells. We used TIMER to analyze the correlation between ATF3 expression in pan-cancer and the correlation between ATF3 and key genes regulating ER stress in KIRC. TIMER was also used to investigate the expression of key genes that regulate ER stress in KIRC. 25
Kaplan–Meier plotter database analysis
The prognostic value of ATF3 in KIRC was analyzed using the Kaplan–Meier plotter (https://kmplot.com/). Patient samples were divided into two groups according to the median expression (high and low), and OS was analyzed using risk ratios (h) with 95% confidence intervals (95% CI) and log-rank p values. The prognostic value of GAS5, DANCR, and RASSF8-AS1 in KIRC was estimated using the Kaplan–Meier plotter.
Candidate miRNA prediction
Upstream-binding miRNAs of ATF3 were predicted using several target gene prediction programs, including PITA, RNA22, miRmap, microT, miRanda, PicTar, and TargetScan. Only predicted miRNAs that commonly appeared in more than two of the programs used were included in subsequent analyses. These predicted miRNAs were regarded as candidate miRNAs for ATF3 expression.
starBase database analysis
We utilized starBase (https://starbase.sysu.edu.cn/) to perform expression correlation analyses for miRNA-ATF3, lncRNA-hsa-miR-221-3p, or lncRNA-ATF3 in KIRC. The expression and prognostic values of hsa-miR-221-3p and has-miR-378a-3p in KIRC were analyzed by the database. In addition, starBase was used to predict candidate lncRNAs that could bind to hsa-miR-221-3p.
Statistical analysis
The statistical analysis in this study was automatically conducted by starBase, at the end of each query. A p-value <.05 or log rank p value <.05 was considered as a statistically significant result.
Results
The expression and prognostic values of ATF3 in ccRCC
Using the TIMER online database, we explored the mRNA expression of ATF3 in human pan-cancers. We observed decreased expression of ATF3 in the following cancer types, when compared with the corresponding normal tissues: Bladder cancer (BLCA), breast cancer (BRCA), kidney chromophobe (KICH), KIRC kidney renal papillary cell carcinoma (KIRP), liver hepatocellular carcinoma (LIHC), lung adenocarcinoma (LUAD), lung squamous cell carcinoma (LUSC), prostate adenocarcinoma (PRAD), rectal cancer (READ), thyroid cancer (THCA), and uterine corpus endometrial carcinoma (UCEC) (Figure 1(a)). The protein expression of ATF3 was further investigated in ccRCC using the HPA database, which revealed that ATF3 protein levels were lower in ccRCC than in normal tissues (Figure 1(b)). In addition, the ATF3 expression in ccRCC samples and adjacent normal tissues was further analyzed using the data obtained directly from TCGA database, which showed that ATF3 expression was significantly decreased in ccRCC tissues (Figure 1(c)). A marked decrease of ATF3 expression in ccRCC was also observed in tumor samples relative to that of normal samples (Figure 1(d)). We also observed that mRNA levels of ATF3 were consistently lower in KIRC than in normal tissues in the UALCAN database (Figure 1(e)). Our results of the survival analysis indicated no statistical significance of ATF3 in predicting the prognosis of patients with BLCA, BRCA, KICH, KIRP, LICH, LUAD, LUSC, PRAD, READ, THCA, or UCEC using the UALCAN database (Figure S1(a–k)). Nevertheless, patients with ccRCC, who presented with downregulated ATF3 expression exhibited poor OS according to the Kaplan–Meier plotter (Figure 2(a)), UALCAN (Figure 2(b)), and HPA databases (Figure 2(c)). These findings illustrate that ATF3 expression is decreased in ccRCC and is significantly associated with the prognosis of patients with ccRCC. Expression of ATF3 in ccRCC. (a) The mRNA of ATF3 expression in different types of cancer was investigated with the TIMER database. (b) The protein of ATF3 expression in ccRCC was examined by using the HPA database. (c) TCGA database and statistical analyses of ATF3 expression in 72 pairs of ccRCC tissues and adjacent normal tissues. (d) Analysis of ATF3 expression in ccRCC and normal tissues in the TCGA database. (e) The mRNA of ATF3 expression in ccRCC was examined by using the UALCAN database. **p value < 0.01; ***p value < 0.001; ****p value < 0.0001. Survival curve evaluating the prognostic value of ATF3 for ccRCC. (a) Survival curves using the Kaplan–Meier plotter is shown for OS. (b) Survival curves using the UALCAN database is shown for OS. (c) Survival curves using the HPA database is shown for OS.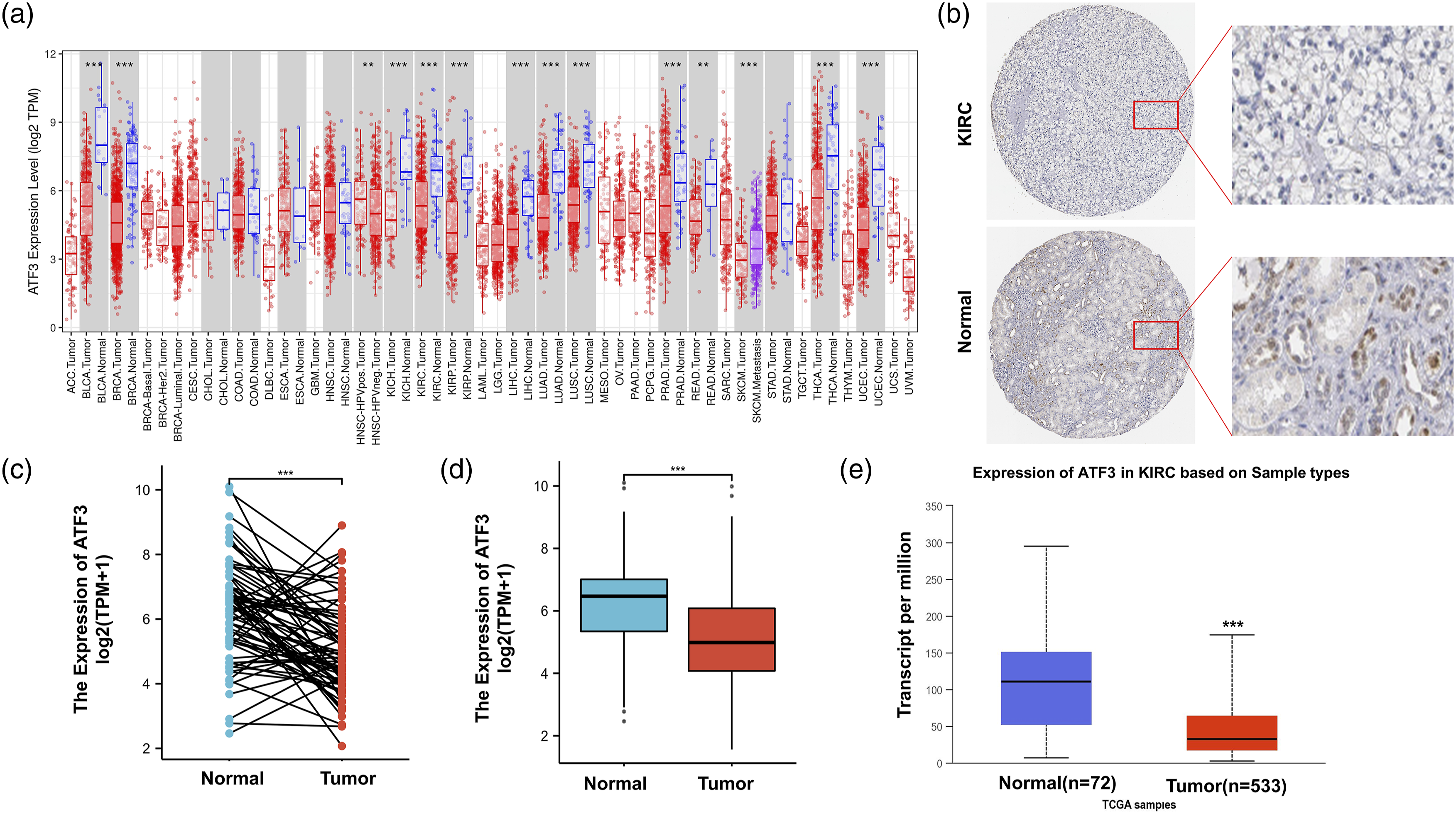

ATF3 expression and clinical parameters of patients with ccRCC
We used the UALCAN database to investigate the expression of ATF3 according to different clinical parameters among the patient groups. Based on cancer stage, ATF3 expression was significantly decreased in patients at stages 1, 2, 3, and 4 compared with that of the corresponding normal controls (Figure 3(a)). In regard to node metastasis status, ATF3 expression was lower in patients with ccRCC classified as N0 and N1 (Figure 3(b)). Downregulation of ATF3 expression was also observed in patients with grade 2, 3, and 4 ccRCC compared with that of normal controls (Figure 3(c)). In terms of sex, ATF3 expression was significantly lower in ccRCC samples from male and female patients compared to that of their sex-matched controls (Figure 3(d)). Moreover, ATF3 levels were significantly downregulated in patients with ccRCC across different age groups (Figure 3(e)). Finally, based on ccRCC subtypes, ATF3 levels were significantly downregulated in patients with the ccA and ccB subtypes (Figure 3(f)). Taken together, these results further corroborate that ATF3 expression is closely related to ccRCC progression and metastasis. Box plots evaluating ATF3 expression among different groups of patients based on clinical parameters using the UALCAN database. Analysis is shown for cancer stage (a), nodes metastasis (b), tumor grade (c), gender (d), age (e), subtypes (f). *p < 0.05, **p < 0.01, ***p < 0.001, ****p value < 0.0001.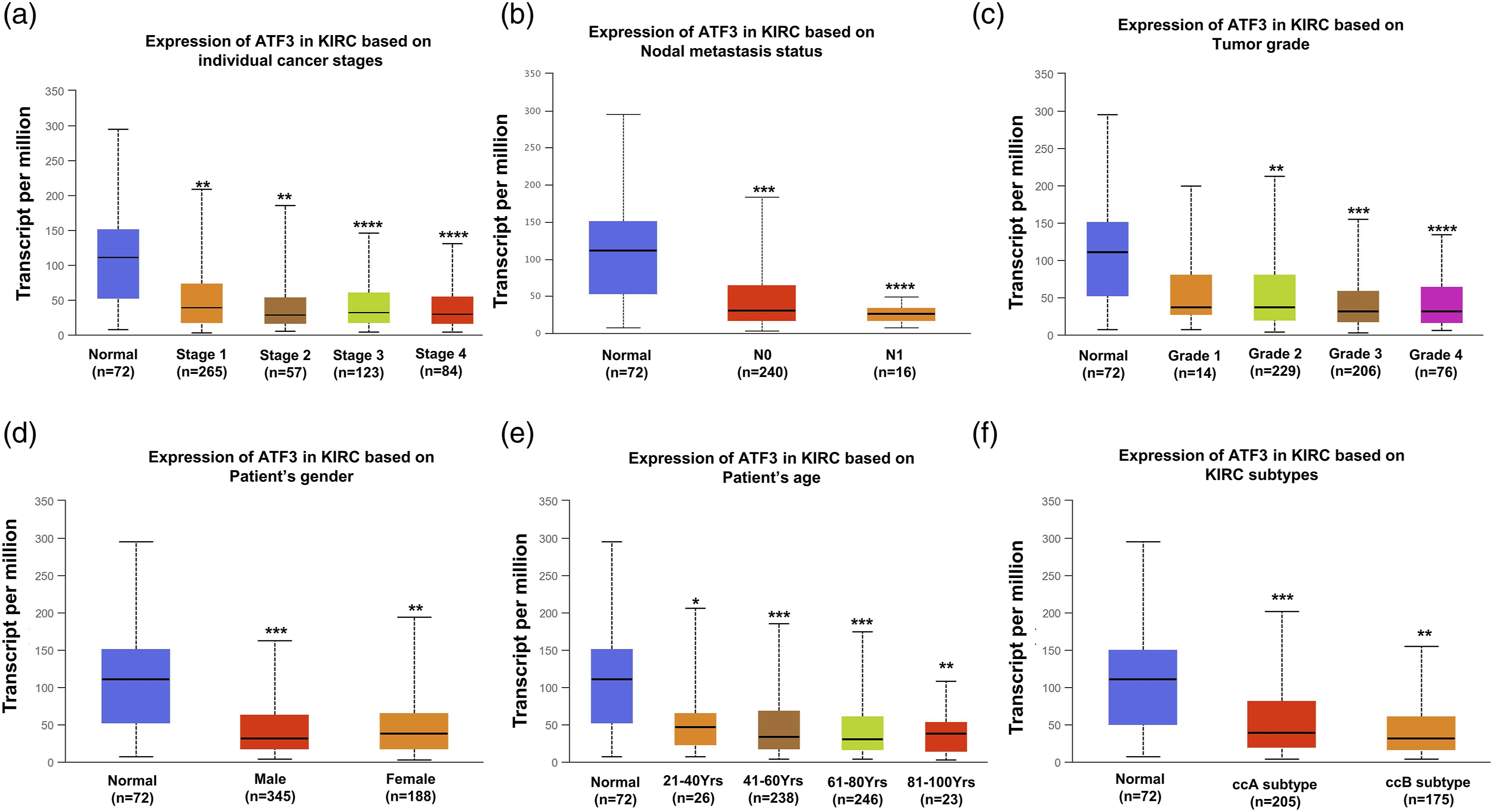
Prediction and analysis of upstream miRNAs of ATF3
Non-coding RNAs have been known to gene expression. To examine whether ATF3 expression is modulated by ncRNAs, we first predicted the upstream miRNAs that could potentially bind to ATF3 and identified 28 candidate docking miRNAs (Figure 4(a)). Expression correlation analysis showed that ATF3 expression inversely correlated with that of hsa-miR-221-3p and has-miR-378a-3p to a significant extent, while positively correlated with that of has-miR-494-3p in ccRCC (Figure 4(b)). Based on the mechanism of action of miRNA in regulation of target gene expression, we expected a negative correlation between miRNAs and ATF3 expression. Therefore, we assessed the expression and prognostic value of hsa-miR-221-3p and has-miR-378a-3p in ccRCC using the starBase database. The results showed that while hsa-miR-221-3p levels were distinctly elevated in ccRCC tissues, those of has-miR-378a-3p were decreased, compared to the respective expression levels of each miRNA in normal tissues (Figure 4(c)). Survival analysis indicated that patients with ccRCC, who also presented with upregulated hsa-miR-221-3p expression, exhibited poor OS, but no statistical significance was observed for has-miR-378a-3p in predicting the prognosis of patients with ccRCC (Figure 4(d)). These findings suggest that hsa-miR-221-3p is the most potent regulatory miRNA of ATF3 in ccRCC. Identification of hsa-miR-221-3p as a potential upstream miRNA of ATF3 in ccRCC. (a) The expression correlation between predicted miRNAs and ATF3 in ccRCC analyzed by starBase database. (b) The miRNA-ATF3 regulatory network established by Cytoscape software. (c) The expression of hsa-miR- 221-3p and hsa-miR-378a-3p in ccRCC and control normal samples determined by starBase database. (d) The prognostic value of hsa-miR-221-3p and hsa-miR-378a-3p in KIRC assessed by starBase database.
Prediction and analysis of upstream lncRNAs of hsa-miR-221-3p
Correlation analysis between lncRNA and hsa-miR-221-3p in ccRCC.

Correlation analysis between lncRNA and ATF3 in ccRCC.


Expression analysis and survival analysis for upstream lncRNAs of hsa-miR-221-3p in ccRCC. (a–c) The expression of PAXIP1-AS2, AP000766.1, and OIP5-AS1 in ccRCC and control normal samples determined by starBase database. (d–f) The OS analysis for PAXIP1-AS2, AP000766.1, and OIP5-AS1 in ccRCC by using the starBase database.
Identification of ATF3-interacting genes and proteins and GO and KEGG pathway analysis
Using the GeneMANIA and STRING online databases, we identified 20 genes and proteins that interacted with ATF3 (Figure 6(a), (b)). We further assessed the role of ATF3 in ccRCC carcinogenesis, while conducting GO and KEGG enrichment analyses to identify ATF3-interacting genes. The GO analysis showed that ATF3 and its associated genes were involved in ER stress-related biological processes (Figure 6(a), (c)) (Table S2). The KEGG analysis further revealed that ATF3 and its related genes were involved in the IL-17, tumor necrosis factor, apoptosis, and MAPK signaling pathways (Figure 6(d)) (Table S3). These results strongly underscore ATF3’s involvement in the regulation of the apoptosis signaling pathway in response to ER stress in ccRCC. (a) The gene–gene interaction network of ATF3 was constructed using GeneMANIA database. (b) The PPI network of ATF3 was generated using STRING. (c, d) GO and KEGG enrichment analysis for ATF3.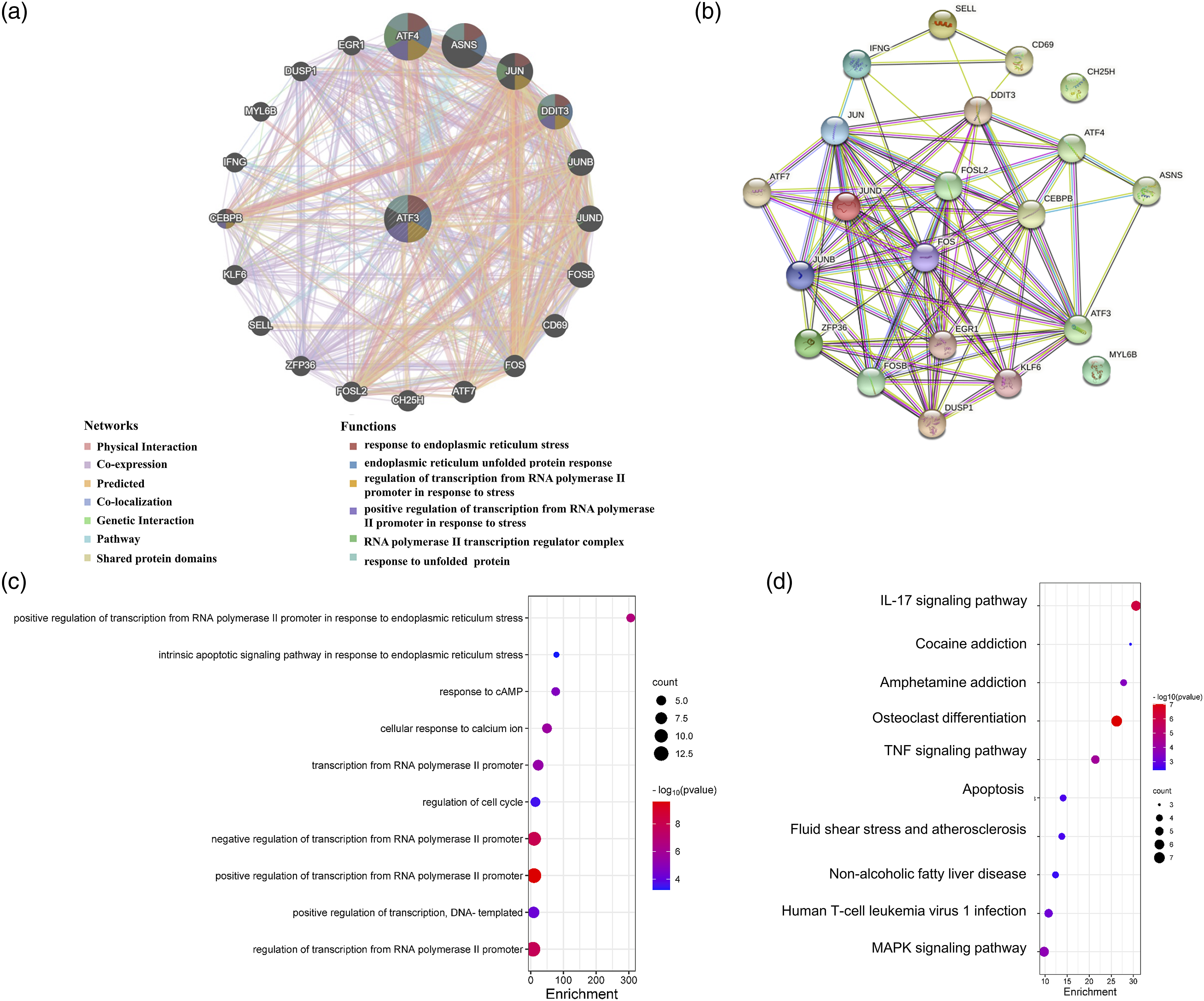
Relationship between ATF3 and ER stress in ccRCC
To verify whether ATF3 is closely related to ER stress, we explored the correlation between ATF3 and key genes that regulate ER stress using the TIMER database. Our results showed that ATF3 expression closely mirrored that of SERP1, SERP2, IRE1α (ERN1), ATF6, and PERK (EIF2AK3) (Figure 7(a)). Moreover, the expression levels of SERP1, SERP2, and ATF6 were significantly decreased in ccRCC. The IRE1α and PERK expression in ccRCC was similar to that of normal tissues (Figure 7(b)). These observations suggest that the anticancer effect of ATF3 may be mediated through regulation of the apoptotic signaling pathway in response to ER stress in ccRCC. Correlation analysis between ATF3 and genes of ER-stress and the expression of genes of ERstress in ccRCC determined by TIMER database. (a) Correlation analysis among ATF3 and SERP1, SERP2, ATF6 and EIF2AK3. (b) The expression of SERP1, SERP2, ATF6 and EIF2AK3 in ccRCC.
Discussion
ccRCC is a complex and heterogeneous disease with different clinical features, while exhibiting a mortality rate of 30%–40% compared to bladder and prostate cancer, with a corresponding approximately 20%. 26 Despite the increase in the total incidence off ccRCC over the past three decades, especially in developed countries, the associated mortality rate has decreased rapidly, owing to early diagnosis and therapy. 27 However, irrespective of the progress in disease control and survival, locally advanced disease and distant metastases continue to occur in many patients. 28 Therefore, elucidating the molecular mechanisms of ccRCC carcinogenesis may provide key clues for developing effective therapeutic strategies and identifying promising prognostic biomarkers. In this regard, ATF3, which is mostly used as a marker of nerve injury that is significantly altered in animal models of osteoarthritis,29,30 has been increasingly supported by mounting. Accumulating evidence has demonstrated that ATF3 plays key roles in the initiation and progression of multiple human cancers. However, knowledge of the role of ATF3 in ccRCC remains inadequate and requires further investigation.
In the present work, we demonstrated that ATF3 expression was markedly decreased in ccRCC compared to that in normal lung tissues, at both the protein and mRNA levels. Moreover, patients with ccRCC and downregulated ATF3 expression exhibited poor OS. Downregulation of ATF3 expression was observed in patients with grade 2, 3, 4, and nods metastasis ccRCC patients compared with that of normal controls. Gao et al. demonstrated that ATF3 suppresses the growth and metastasis of ccRCC. 24 This report, together with our analytical results, indicates that ATF3 acts as a suppressor of ccRCC.
It has been shown that ncRNAs, including miRNAs, lncRNAs, and cyclic RNA (circRNAs), participate in the regulation of gene expression through a ceRNA mechanism.31–33 Overexpression of the clustered mRNAs hsa-miR-221 and hsa-miR-222 has been reported to directly lead to increased growth and tumorigenesis in oral carcinoma cells. 34 Here, we identified 28 miRNAs, with hsa-miR-221-3p, expression found to be significantly negatively correlated with that of ATF3. Moreover, hsa-miR-221-3p was enriched in ccRCC relative to normal lung tissues and correlated with the poor OS patients with ccRCC. Previous studies have shown that miR-221 and miR-222 play key roles in modulating clinical outcomes in patients with both solid and hematological tumor malignancies.35–37 These findings, combined with our data, suggest that hsa-miR-221-3p might be the most potential regulatory miRNA of ATF3 in ccRCC.
The lncRNAs play significant roles in the regulation of various biological functions, including epigenetic and posttranscriptional regulation. 38 To identify the upstream regulatory lncRNAs of the hsa-miR-221-3p/ATF3 axis, we predicted 64 lncRNAs associated with hsa-miR-221-3p using the starBase database. By conducting expression, survival, and correlation analyses, the lncRNAs PAXIP1-AS2 and OIP5-AS1 were returned as potential upstream regulators of the hsa-miR-221-3p/ATF3 axis in ccRCC. The PAXIP1-AS2 is an antisense RNA with no known function. A recent study indicated that PAXIP1-AS2 expression led to reduced amounts of the RAD18 protein and DNA polymerase η, and subsequently resulted in decreased translesion DNA synthesis (TLS). As such, PAXIP1-AS2 was considered to be a TLS regulator. 39 Regarding, OIP5-AS1, this lncRNA reportedly functions as an oncogene in multiple malignancies,40–42 but it also plays a protective role in esophageal squamous carcinoma cells 43 and esophageal carcinoma. 44 Taken together, the PAXIP1-AS2 and OIP5-AS1/hsa-miR-221-3p/ATF3 axis were identified as potential regulatory pathways in ccRCC.
The ER is involved in protein synthesis, folding and modification, as well as lipid synthesis and calcium storage. The ER stress is induced when intracellular homeostasis is disrupted, and the demand on ER function exceeds its capacity. 45 The ER stress is a double-edged sword. On one side, when environment disruptions are relatively mild, ER stress can reverse such effects. However, when cells are unable to restore homeostasis, intense and chronic ER stress leads to apoptosis.45,46 Numerous studies have documented that ER stress contributes to tumorigenesis virus cancer types, including glioblastomas and carcinomas of the breast, stomach, colon, esophagus, lung, prostate, pancreas, and liver. 47 However, in the context of RCC, ER stress can significantly inhibit tumor progression. 48 Our results strongly suggest that ATF3 is involved in the regulation of the apoptotic signaling pathway in response to ER stress, in ccRCC. In addition, ATF3 expression positively correlated with that of ER stress-related genes, such as SERP1, SERP2, IRE1α (ERN1), ATF6, and PERK (EIF2AK3). Furthermore, the expression of SERP1, SERP2, and ATF6 was significantly decreased in ccRCC. In the present work, we observed that promoting ER stress could suppress RCC progression. 49 Moreover, another study reported that the expression of ATF6 and PERK, but not of IRE1α, was downregulated in patients with advanced ccRCC tumor stage and grade. This finding demonstrated that ER stress was suppressed in such patients. 50 The collective observations from previous works, further confirm our conclusion that the anticancer effect of ATF3 is likely mediated through the regulation of the apoptotic signaling pathway in response to ER stress in ccRCC.
Nevertheless, our study had certain limitations. First, our current findings are based solely on the in silico analysis of publicly available data. Moreover, funding restrictions meant that clinical data supporting the diagnostic value of ATF3 in ccRCC are lacking. In future studies, we will be incorporating a follow-up experimental verification process of computational results. Overall, our study provides insights that may contribute to the formation of a new theoretical basis for research in ccRCC.
Conclusions
Our in silico analysis findings showed that expression of ATF3 decreased in ccRCC, and negatively correlated with the poor prognosis in such patients, while the PAXIP1-AS2 and OIP5-AS1/hsa-miR-221-3p/ATF3 axes were identified as potential regulatory pathways in ccRCC. Our results suggest that anticancer effect of ATF3 in ccRCC may be mediated through the regulation of the apoptotic signaling pathway in response to ER stress (Figure 8). The model of PAXIP1-AS2 and OIP5-AS1/hsa-miR-221-3p/ATF3 axis in carcinogenesis of ccRCC.
Supplemental Material
Supplemental Material - Activating transcription factor 3 is a new biomarker correlation with renal clear cell carcinoma progression
Supplemental Material for Activating transcription factor 3 is a new biomarker correlation with renal clear cell carcinoma progression by Zhicong Yang, Yongwang Hou, Jingqi Li, Dandan Xu, Zhichao Yang, and Xinsheng Wang in International Journal of Immunopathology and Pharmacology
Footnotes
Acknowledgements
We thank The First Affiliated Hospital of Hebei North University for their approval of the study.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by the Hebei Health Commission Scientific Research Foundation Project (20220596).
Supplemental Material
Supplemental material for this article is available online.
Appendix
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
