Abstract
Introduction
Renal cell carcinomas are derived from the renal tubular epithelium, 80% of which are ccRCC. Although its malignancy is low, its morbidity and mortality have been increasing in recent years. 1 Moreover, 20–30% of patients who undergo radical nephrectomy still develop distant organ metastases or focal recurrence during follow-up. 2 Metastasis is the leading cause of death in ccRCC patients. With the continuous in-depth study of molecular biology, studies on the pathogenesis of ccRCC have been conducted; however, the specific molecular mechanism of the occurrence and development of ccRCC remains unclear. Therefore, actively identifying new molecular biomarkers are of great help in improving ccRCC prognosis.
TJs are junctional complexes that mediate cell-to-cell adhesion in epithelial and endothelial cellular sheets 3 and play crucial roles in cell polarity, adhesion, and permeability. 4 TJs mainly comprises two types of proteins, one of which includes structural membrane proteins, including occludin, CLDNs, and junctional adhesion molecules. 5 Among them, CLDNs are the skeleton proteins of TJs, and their function is the most important because no paracellular barrier can form without CLDNs. 6 To date, 26 (human) or 27 (rodent) CLDNs have been produced in epithelial and endothelial cells in a tissue-specific manner. 7 CLDNs are ubiquitous in humans; however, many studies have shown that CLDNs’ expressions were altered in many types of cancers. Whether upregulated or downregulated, the structure and function of TJs are often abnormal in several cancers. 8 The direct effect of this abnormal expression is the destruction of the composition ratio of closely connected components, which causes changes in the structure and function of closely connected components. Studies have confirmed that the disruption of cell polarity is an early manifestation of tumour invasion and metastasis. 9 Moreover, the abnormal expression of CLDNs is considered a mechanism underlying the malignant progression of cancer. 10 The differential expressions and functions of CLDNs may be associated with their high tissue and cell specificity. CLDNs can be used as markers for tumour diagnosis and as independent indicators of prognosis and tumour recurrence. 11 For example, high expression of CLDN6 is associated with worse overall survival (OS), disease-specific survival, and progression-free interval in adrenocortical carcinoma and bladder urothelial carcinoma. 12 CLDN10 expression is associated with poor prognosis in ovarian cancer. 13 In patients with advanced gastric cancer with high expression of CLDN18.2, the combination of CLDN18.2 antibody IMAB 362 and chemotherapeutic drugs can significantly improve patient survival. 14 Antibody-coupled agents targeting CLDN6 can effectively kill hepatocellular carcinoma cells. 15 However, the tissue-specific mechanism of CLDN expression changes in tumours remains unclear, which may be related to precise molecular regulation.
CLDNs are closely associated with the occurrence, development, and metastasis of several malignant tumours. However, no studies have reported the expression of CLDNs in ccRCC and their influence on the occurrence and development of ccRCC. In this study, we aimed to analyze the expression and clinical relevance of CLDNs in ccRCC using an online public database. Our results suggest that CLDNs may have complex and unique functions and may be potential therapeutic targets for future clinical interventions in ccRCC.
Materials and methods
Gene expression profiling interactive analysis
The GEPIA database (http://gepia.cancer-pku.cn/index.html) is based on TCGA and GTEx is a database for the gene expression analysis of 9736 tumours and 8587 normal tissues. 16 We sequentially analyzed the differences in the expression of 24 CLDN family members between ccRCC and normal kidney tissues. The thresholds were as follows: |Log2 FC|cutoff = 1, p < .01.
UALCAN
The UALCAN (http://ualcan.path.uab.edu) is a cancer gene chip database and integrated data mining platform, that includes TCGA genome data of 31 types of cancers. 17 The results of the GEPIA database were validated against those of the UALCAN database. Promoter methylation, clinicopathological parameters, and survival analyses were performed. In the UALCAN analyses, the threshold was set at p < .05.
Human protein atlas
Human protein atlas (https://www.proteinatlas.org) is a website that analyzes the expression levels of human proteins from protein and RNA levels in various human tissues and organs based on immunohistochemistry and RNA sequencing methods. 18 In this study, we obtained immunohistochemical images using this tool to directly compare the protein expression of different CLDN family members in healthy human and ccRCC tissues.
The Kaplan-Meier plotter
The Kaplan Meier plotter (http://kmplot.com/analysis/) is capable to assess the correlation between the expression of all genes (mRNA, miRNA, protein) and survival in 30k+ samples from 21 tumor types. 19 The prognostic value of CLDNs in ccRCC patients was also analyzed. Statistical significance was set at p < .05.
cBioPortal
The cBioPortal (http://www.cbioportal.org/) is a comprehensive web resource that can be used to explore multidimensional cancer genomic data. 20 We investigated the genetic variation of the CLND family members in ccRCC using the
cBioPortal for Cancer Genomics. Twenty-four genes were queried in 512 ccRCC samples (TCGA, pan-cancer atlas).
String-DB database
The String-DB database (https://string-db.org) is used to study the interaction between biological genes or proteins. It combines known and predictable protein functions to form a biological database of the interactions between proteins. 21 Proteins that interact with CLDNs upstream and downstream can be analyzed using this database. PPI enrichment p-value <.0001, minimum required interaction score: medium confidence (.400).
GeneMANIA
GeneMANIA (http://www.genemania.org) is a resource-rich website that contains gene information, analyzes gene lists, and prioritizes genes for functional assays with highly accurate prediction algorithms. 22 This was used to determine the predictive value of CLDNs.
Metascape
Metascape (https://metascape.org/gp/index.html#/main/step1) is an online functional annotation website, integrated with more than 40 biological information databases. 23 Metascape not only contains enrichment analyses of biological pathways, structural analyses of protein interaction networks and abundant gene annotation capabilities but also presents the results in an easy-to-understand, high-quality graphic language.
Tumor immune estimation resource
TIMER (https://cistrome.shinyapps.io/timer/) associates tumour-infiltrating immune cells with gene expression levels, gene mutations, and other data by analyzing the expression profile data of tumuor samples in TCGA. 24 We examined the association between the expressions of CLDN7, 8, 10, 11, 14, 16, and 19 and the level of immune invasion in ccRCC.
Statistical analysis
The expression difference of CLDNs between ccRCC and normal renal tissue was analyzed by GEPIA database. For normally distributed measurement data, paired t test was used for comparison between paired samples, independent t test was used for comparison between independent samples, and one-way variance was used for comparison between multiple groups. Non-normal distribution measurement data were compared between the two groups using non-parametric test method. Pearson correlation coefficient was used to analyze the correlation of GEPIA data. Kaplan-Meier plotter database was used to draw survival curve by Kaplan-Meier method, and comparison between groups was performed by Log-rank test. p < .05 was considered statistically significant.
Results
Differential expressions of CLDNs in ccRCC patients
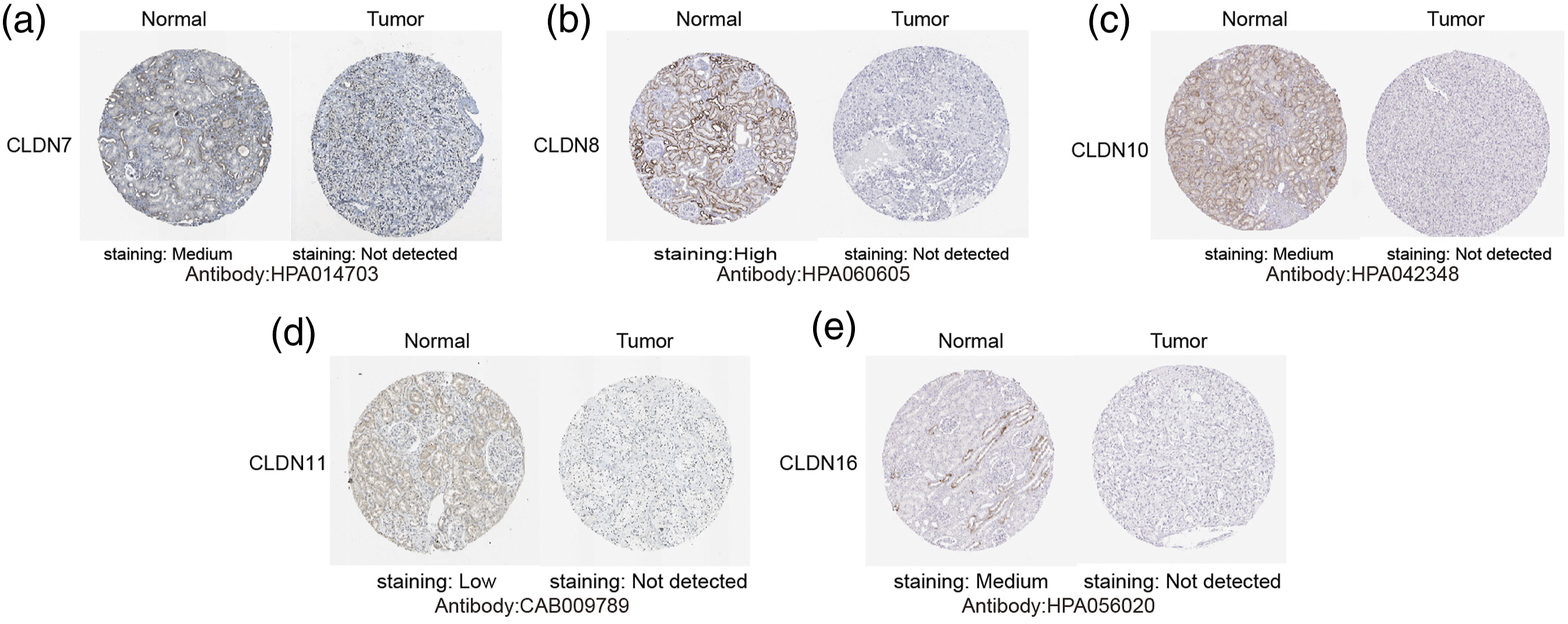
To explore the distinct expression of CLDNs in ccRCC patients, mRNA expression was analyzed using the GEPIA database, which was used to analyze the mRNA expression in The Cancer Genome Atlas (TCGA) and Genotype-Tissue by Peking University Expression (GTEx) samples. As depicted in Figure 1, seven genes exhibited low expression in the ccRCC samples compared with that in normal kidney tissue samples and included CLDN7, 8, 10, 11, 14, 16, and 19. There were no significant differences in the expression of other genes. The UALCAN database was used to verify the above results. As shown in Figure 2, the mRNA levels of CLDN7, 8, 10, 11, 14, 16, and 19 in ccRCC tissues were significantly lower than those in normal kidney tissues. These results are consistent with those from the GEPIA database. Next, we used HPA to explore the protein expression of different CLDNs in the ccRCC tissues. Furthermore, HPA revealed low protein expression of CLDN7, 8, 10, 11, and 16 in ccRCC tissues, while medium and high protein expression was observed in normal kidney tissues (Figure 3). No data related to the protein expression of CLDN14 and CLDN19 in ccRCC were found in the HPA. Moreover, the protein and transcription levels were consistent. The mRNA expressions of CLDNs in ccRCC (GEPIA). p < .01. The mRNA expressions of CLDNs in ccRCC (UALCAN). (a)–(g) represents CLDN7, 8, 10, 11, 14, 16, and 19. ***p < .001. The protein expressions of CLDNs in ccRCC (HPA). (a)–(e) represents CLDN7, 8, 10, 11, and 16.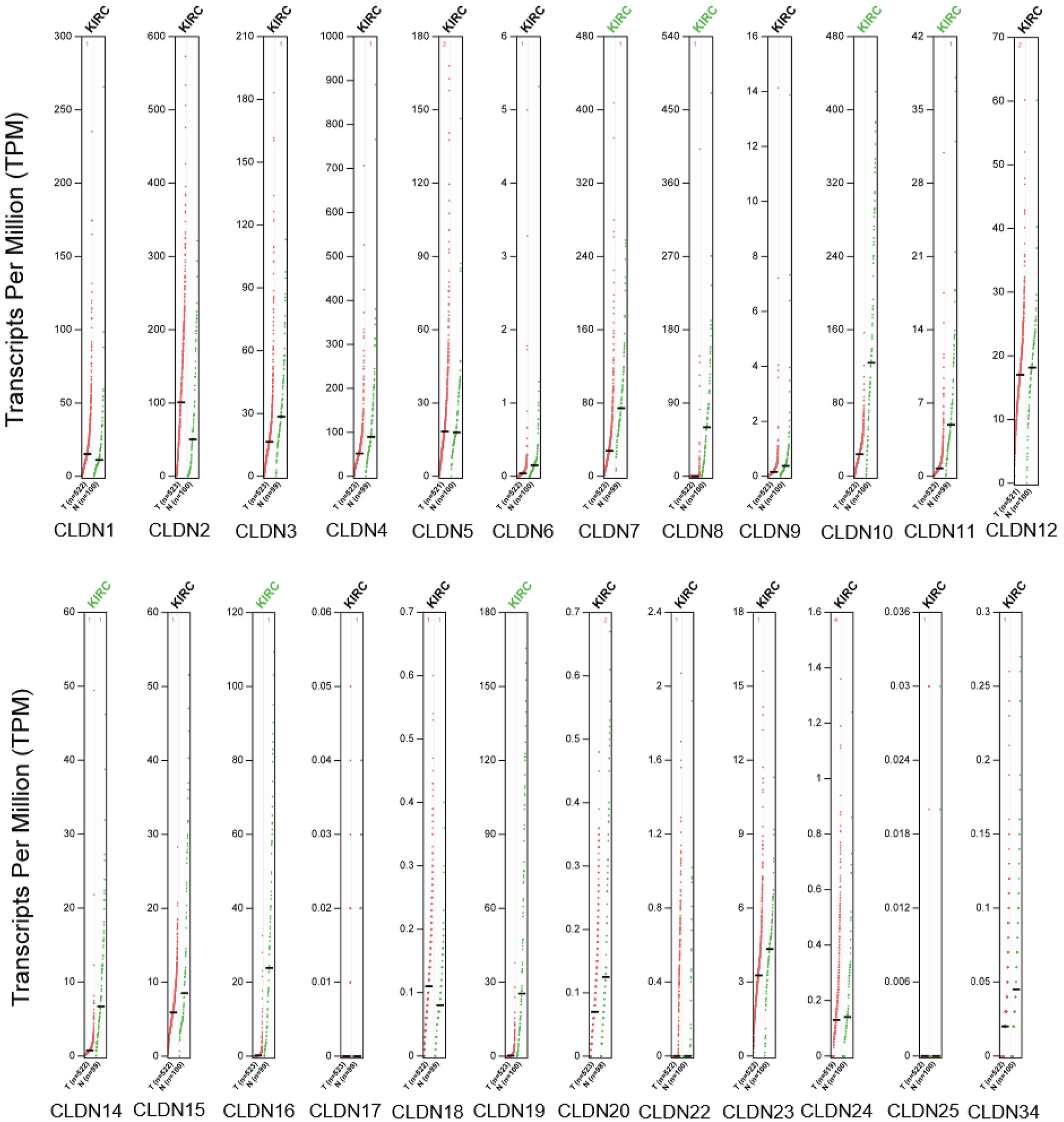

Promoter methylation status of CLDNs in ccRCC patients
Moreover, given the important role of promoter methylation in the process of renal tumours, we used the UALCAN to investigate the promoter methylation levels of CLDNs in ccRCC and found that the promoters of CLDN7, 8, 10, 11, 14, 16, and 19 were hypermethylated in ccRCC tissues, compared to those in normal kidney tissues (Figure 4). Promoter methylation status of CLDNs in ccRCC (UALCAN). (a)–(g) represents CLDN7, 8, 10, 11, 14, 16, and 19. *p < .05, ***p < .001.
Relationship of mRNA levels of CLDNs with clinicopathological parameters in ccRCC patients
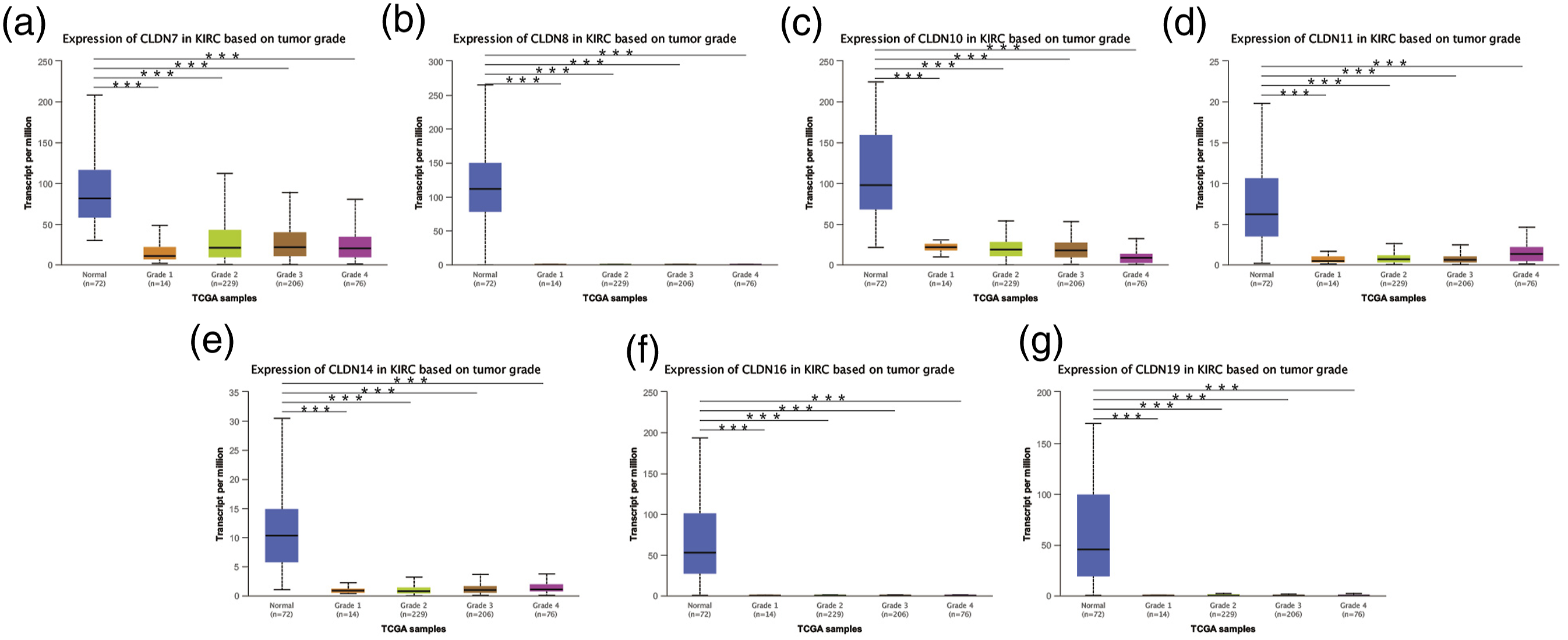
After mRNA expression and promoter methylation were analyzed, we explored the relationship between mRNA levels of CLDNs and clinicopathological parameters in ccRCC patients using UALCAN, which included the patients’ cancer stages and tumour grades. The mRNA expression of CLDNs was significantly correlated with the cancer stage. As the tumours progressed, patients with more advanced cancer stages tended to express lower mRNA expressions of CLDN7, 8, 10, 11, 14, 16, and 19 (Figure 5). Furthermore, as shown in Figure 6, the low expression of CLDN7, 8, 10, 11, 14, 16, and 19 significantly correlated with advanced tumour grades. In conclusion, the mRNA expressions of CLDN7, 8, 10, 11, 14, 16, and 19 were significantly associated with clinicopathological parameters in ccRCC patients. Correlation between CLDNs expressions and cancer stages in ccRCC patients (UALCAN). (a)–(g) represents CLDN7, 8, 10, 11, 14, 16, and 19. ***p < .001. Correlation between CLDNs expression and tumour grades in ccRCC patients (UALCAN). (a)–(g) represents CLDN7, 8, 10, 11, 14, 16, and 19. ***p < .001.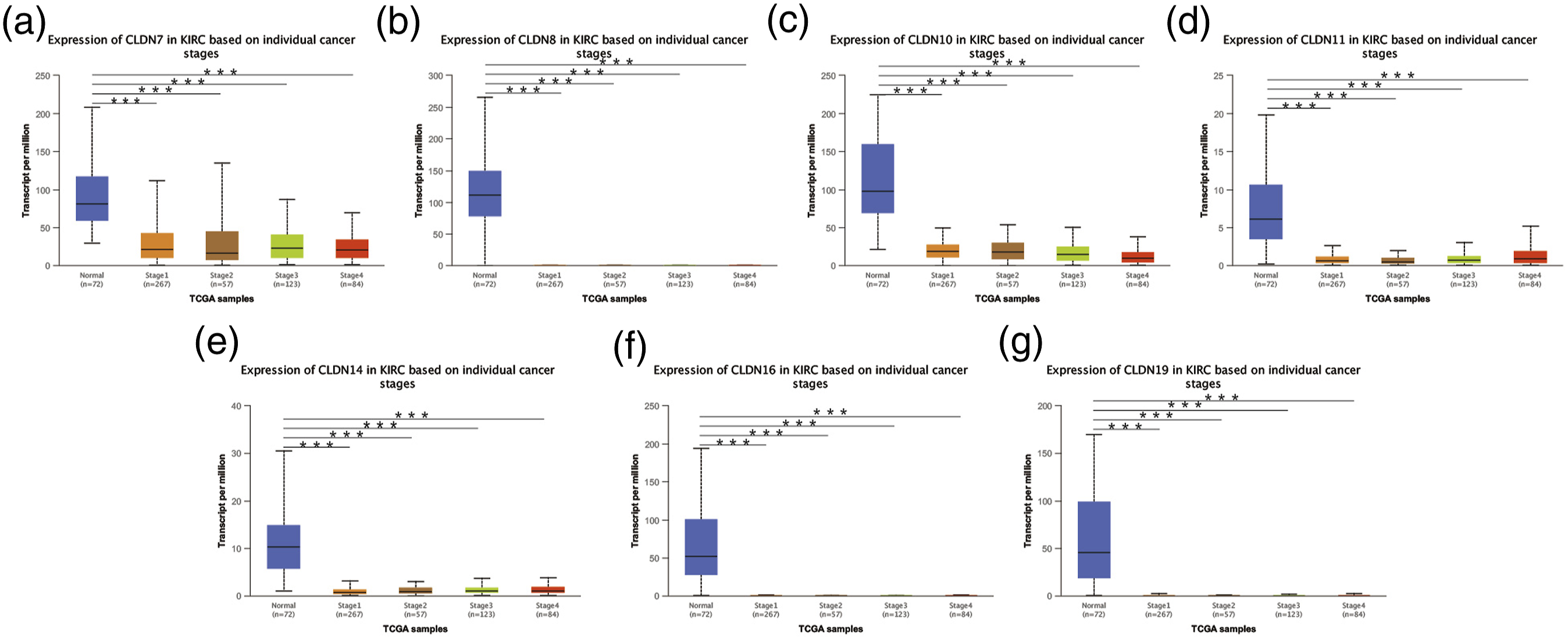
Prognostic value of the mRNA expressions of CLDNs in ccRCC patients
Furthermore, we analyzed the prognostic value of CLDNs in patients with ccRCC using the Kaplan-Meier plotter. As is shown in Figure 7, low CLDN7 (HR = 0.71, p = .074), CLDN8 (HR = 1.37, p = .037), CLDN10 (HR = 0.4, p < .0001), CLDN11 (HR = 2.07, p < .0001), CLDN14 (HR = 1.83, p = .0003), CLDN16 (HR = 1.35, p = .05), and CLDN19 (HR = 0.81, p = .25) mRNA expressions were correlated with shorter OS in ccRCC patients. To verify the prognostic value of CLDNs in ccRCC patients, UALCAN was utilized. As shown in Figure 8, low CLDN7 (p = .038), CLDN8 (p = .014), CLDN10 (p = .00014), CLDN11 (p = .012), CLDN14 (p = .015), CLDN16 (p = .77), and CLDN19 (p = .24) mRNA expressions were correlated with shorter OS in ccRCC patients. A statistically significant difference was considered when the p-value was less than .05. Combining the analysis of the two databases, we found that ccRCC cases with lower CLDN7, 8, and 10 mRNA expression levels showed a shorter OS. The differences were statistically significant. The prognostic values of CLDNs in ccRCC patients in the OS curve (Kaplan-Meier Plotter). (a)–(g) represents CLDN7, 8, 10, 11, 14, 16, and 19. The prognostic value of CLDNs in ccRCC patients in the OS curve (UALCAN). (a)–(g) represents CLDN7, 8, 10, 11, 14, 16, and 19.

Genetic alteration analyses of CLDNs in ccRCC patients
We investigated the genetic variation of the CLDN family members in ccRCC using the cBioPortal for Cancer Genomics. Twenty-four genes were queried in 512 ccRCC samples (TCGA, Pan-Cancer Atlas). Figure 9(a) shows the alteration frequency of genetic variation in ccRCC. As is shown in Figure 9(b), queried genes were altered in 244 (48%) queried samples. In addition, CLDN7, 8, 10, 11, 14, 16, and 19 were altered in 4%, 1.2%, 4%, 4%, 1.4%, 5%, and 4% of the queried ccRCC samples, respectively. Previous studies have shown that the CLDNs are dysregulated in a variety of tumours and are involved in diagnosis, tumorigenesis, and prognosis.
25
Thus, the CLDNs are worthy of further research on ccRCC. The genetic variation of the CLDNs in ccRCC (cBioPortal). (a) The alteration frequency of the CLDNs in ccRCC. (b) The Oncoprint of the CLDNs in ccRCC.
Protein-protein interactions comprehensive analysis of CLDNs in ccRCC patients
Functional interactions between proteins are necessary for the molecular mechanisms and metabolism of malignancies. Therefore, we conducted a PPI network analysis of the differentially expressed CLDNs using the String-DB database to explore the potential interaction proteins among them. As expected, several nodes (17) and edges (87) were identified in the PPI network (Figure 10(a)). CLDN23, CLDN12, TJP1, OCLN, and CD44 interacted with the differentially expressed CLDNs. They are mainly involved in biological processes, such as tight junction-specific obliteration of the intercellular space, cellular barriers, and tight connections between cells. The GeneMANIA results also revealed that the functions of the differentially expressed CLDNs and their associated molecules (such as CLDN24, CLDN15, CDY1, CLDN20, CLDN22, and CLDN23) were primarily related to tight junction organization, tight junction, apical junction assembly (Figure 10(b)). PPI network of different expressions of CLDNs. ((a) String-DB (b) GeneMANIA).
Enrichment analysis of related genes and pathways of CLDNs in ccRCC patients
By searching for genes related to CLDN7, 8, 10, 11, 14, 16, and 19 in the UALCAN database, 4203 genes were found to be positively correlated with them. The lowest and highest Pearson correlation coefficients were 0.3 and 0.89, respectively. After the list of these genes was input into Metascape (Figure 11), the results showed that the main pathways involved in these genes included epithelial cell differentiation, regulation of cell adhesion, cell morphogenesis, and cell-cell junction organization. Summary of CLDN-related genes enrichment pathways (Metascape). Bar graph of enriched terms across input gene lists, coloured by p-values.
Immune cell infifiltration of CLDNs in ccRCC patients
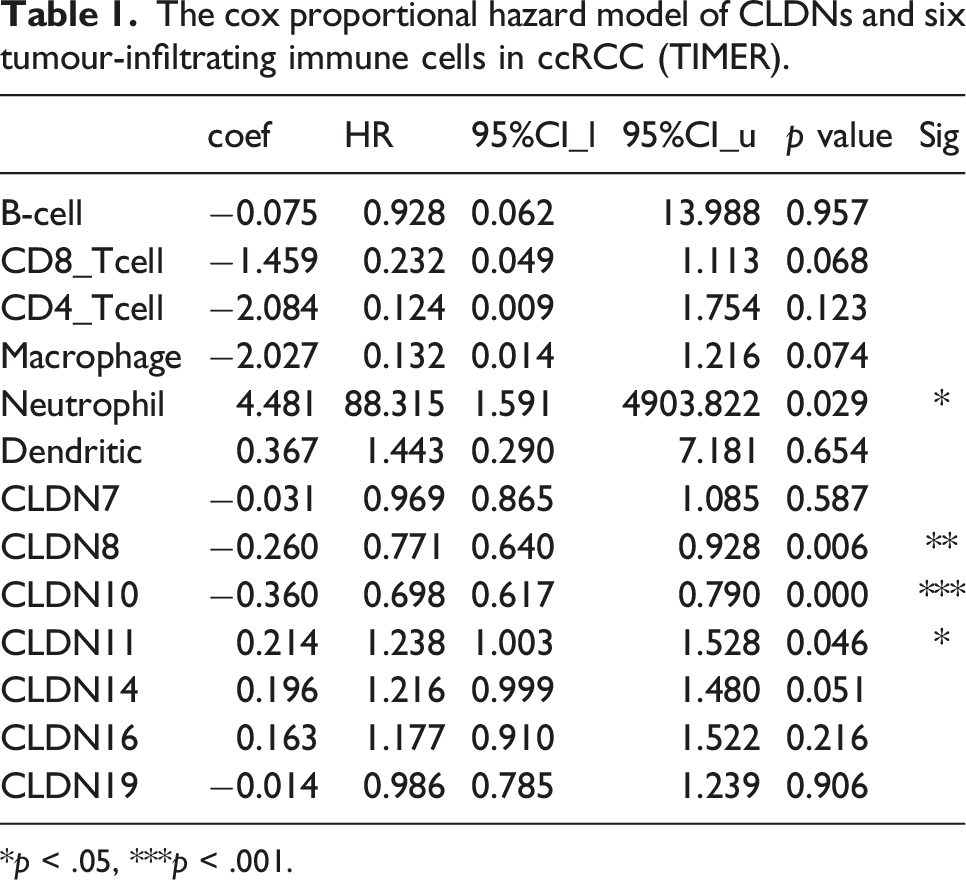
Immune infiltration in the tumour microenvironment (TME) plays a key role in tumour development. Therefore, we analyzed the association between CLDN expression levels and immune cell infiltration using the TIMER database. CLDN7 and CLDN10 expressions were positively correlated with B-cell infiltration (Figure 12(a) and (c)). CLDN8 expression was positively correlated with the infiltration of B cells, CD8+ T cells, macrophages, neutrophils, and dendritic cells (Figure 12(b)). CLDN11 expression was positively correlated with the infiltration of CD4+ T cells, macrophages, neutrophils, and dendritic cells (Figure 12(d)). CLDN14 expression was positively correlated with the infiltration of B cells and dendritic cells (Figure 12(e)). CLDN16 expression was positively correlated with the infiltration of B cells, CD4+ T cells, macrophages, neutrophils, and dendritic cells (Figure 12(f)). CLDN19 expression was positively correlated with the infiltration of CD4+ T cells and dendritic cells (Figure 12(g)). Moreover, we used the Cox proportional hazard model and found that neutrophils and CLDN8, 10, and 11 were significantly correlated with the clinical outcome of patients with ccRCC after adjustment for confounding factors (Table 1). Correlations between differentially expressed CLDNs and immune cell infiltration (TIMER). (a)–(g) represents CLDN7, 8, 10, 11, 14, 16, and 19. The cox proportional hazard model of CLDNs and six tumour-infiltrating immune cells in ccRCC (TIMER). *p < .05, ***p < .001.
Discussion
Renal cell carcinoma (RCC) is one of the most common tumours of the urinary system, accounting for 3–5% of adult malignant tumours in recent years. 26 ccRCC accounts for up to 80% of all kidney cancers. 27 Despite continuous cancer treatment, the prognosis of patients with advanced and metastatic ccRCC remains poor, and the mortality rates continue to increase. 28 Approximately 20–30% of patients with ccRCC have already experienced local or systemic metastasis when clinically diagnosed.1,29 Under the existing conditions, the diagnosis and treatment of ccRCC are still one of the most difficult problems. Therefore, it is particularly important to explore the genes related to the development of ccRCC and discover new molecular markers suitable for clinical application.
CLDNs are the main components of tight junctions in epithelial cells and have functions, such as maintaining cell junctions, polarity, and signal transduction, thus regulating cell proliferation, apoptosis, migration, and invasion.30,31 Moreover, the dysregulation of CLDN expression is closely related to the occurrence and development of many malignant tumours. CLDN2 inhibits ccRCC progression by inhibiting YAP activation. 32 In bladder cancer, loss of CLDN11 may be associated with proliferation, recurrence, and invasiveness. 33 Reduced CLDN12 expression predicts poor prognosis in cervical cancer. 34 CLDN15 is a novel diagnostic marker of malignant pleural mesothelioma. 35 However, the expression levels of the CLDN family members in ccRCC and the affection of their differential expression on ccRCC development remain unknown.
In this study, we found seven CLDN family members with low expressions in ccRCC tissues compared with that in normal kidney tissue samples, namely CLDN7, 8, 10, 11, 14, 16, and 19. They were significantly associated with the cancer stages and tumour grades in patients with ccRCC. With the development of tumours, their expression levels decrease, indicating that they play roles as tumour suppressor genes. It has been reported that CLDN8 could serve as an independent prognostic factor in ccRCC, in which it suppresses 786–O proliferation, migration, and invasion through the EMT and AKT pathways. 36 This finding is consistent with the results of our analysis. The Kaplan-Meier Plotter database was used to analyze the effects of CLDN7, 8, 10, 11, 14, 16, and 19 on the survival rate of patients with ccRCC. The results indicated that the OS rate of patients with ccRCC with low CLDN8, 10, 11, and 14 expressions were significantly reduced. These results were verified by the UALCAN database, suggesting that patients with low CLDN7, 8, 10, 11, and 14 expressions had significantly lower OS rates. CLDN16 and CLDN19 also showed similar characteristics in patients with ccRCC. Although their expressions in tumour tissues are lower than that in normal tissues, it needs to be further verified whether CLDN16 and CLDN19 play inhibitory roles in ccRCC, similar to other CLDN family members.
As we all know, methylation is an important modification of protein and nucleic acid, regulating gene expression and shut down, and is one of the most important contents of epigenetics. Studies have shown that abnormal methylation leads to instability in gene expression and transcription, which is an important reason for the occurrence and development of tumours. 37 Epigenetic modifications, such as methylation of CLDNs have been reported to be promising prognostic markers in various cancers.38,39 Therefore, the methylation levels of CLDN7, 8, 10, 11, 14, 16, and 19 were analyzed using the UALCAN database, and the results showed that their methylation levels were significantly increased in ccRCC. Furthermore, studies have shown that the downregulation of CLDN7 induced by promoter hypermethylation is associated with progression and poor prognosis in human ccRCC. 40 Related studies have shown that DNA hypermethylation is an important factor in the decreased expression of CLDN10 in ccRCC. 41 Our findings are consistent with these results.
Cancer is currently regarded as a disease characterized by uncontrolled cellular growth caused primarily by genetic alterations that mainly occur in a set of cancer driver genes, conferring transformed cells with certain selective advantages over neighbouring cells. 42 For example, mutations in CLDN1, 5, 14, 16, and 19 have been reported to cause neonatal ichthyosis, sclerosing cholangitis, non-syndromic deafness, familial hypomagnesemia, and other symptoms in humans. 6 We investigated the genetic variation of the CLDN family in ccRCC using the cBioPortal for Cancer Genomics. Twenty-four genes were queried in 512 ccRCC samples (TCGA, Pan-Cancer Atlas), and these queried genes were altered in 244 (48%) samples. However, whether these genetic alterations are the drivers of ccRCC, whether they contribute to tumorigenesis, and the exact tumorigenesis mechanism remain to be determined. CLDNs expression in cancer is regulated by factors other than genetic alterations.
Next, using String-DB database mining analysis, CLDN7, 8, 10, 11, 14, 16, and 19 interacted with CLDN24, CLDN15, CDY1, CLDN20, CLDN22, and CLDN23. They are mainly involved in biological processes, such as tight junction-specific obliteration of the intercellular space, cellular barriers, and tight connections between cells. GeneMANIA results also revealed that the functions of the differentially expressed CLDNs and their associated molecules (such as CLDN24, CLDN15, CDY1, CLDN20, CLDN22, and CLDN23) were primarily related to tight junction organization, tight junction, and apical junction assembly. Pathway enrichment analyses of interacting proteins using the Metascape database showed that CLDN7, 8, 10, 11, 14, 16, and 19 were mainly involved in epithelial cell differentiation, cell morphogenesis, and cell-cell junction organization. Disruption of CLND strands allows foreign molecules and microorganisms to enter tissues through the paracellular space, causing inflammation, which is the most common predisposing factor for cancer. 43 Moreover, the disruption of CLDN strands also allows growth factor infiltration into the mucosa to promote neoplastic transformation and growth. 44 These results indicated that the downregulated expression of CLDN7, 8, 10, 11, 14, 16, and 19 in ccRCC directly damages and affects the function of tightly coupled structures.
TME mainly includes stromal and immune cells, and the degree of infiltration of immune cells is highly correlated with the occurrence, development, and immunotherapy response of tumours. 45 In this study, TIMER analysis revealed that the expression of CLDNs may be significantly correlated with the infiltration of six immune cell types, indicating that CLDNs are closely related to tumour immune cell infiltration and are also involved in the pathogenesis of ccRCC. Moreover, CLDN7, 8, 10, 11, 14, 16, and 19 were down-regulated in ccRCC. This abnormal expression may play a pathogenic role by increasing tumor immune cell infiltration, but the specific mechanism remains unclear. Our study provides a theoretical basis for the design of novel immunotherapies.
However, our experiment was only at the data analysis level and should be verified experimentally to provide sufficient evidence for the conclusions of this paper. Only one TCGA dataset is used in this paper, and the data sources are very limited. The work in this paper lacks validation on external datasets. And a lot of the work in this paper is done based on the online website, without the use of personalized algorithms and codes. The analysis in this paper is too conventional and lacks innovation. Overall, the depth of this work is not enough.
In summary, we systematically analyzed the expression and prognostic value of CLDNs in ccRCC. Our findings suggest that CLDN7, 8, and 10 could be used as abiological markers for prognosis evaluation and malignancy analysis of ccRCC.
Conclusions
In this study, reliable bioinformatics analyses revealed that CLDN7, 8, 10, 11, 14, 16, and 19 were expressed in low levels in ccRCC tissues, which may play roles as tumour suppressor genes in tumour progressions. Moreover, the low expressions of CLDN8, 10, 11, and 14 are associated with poor prognosis of patients and can be used as potential biomarkers for the diagnostic and prognostic assessment of patients with ccRCC, providing clues and a theoretical basis for subsequent studies on ccRCC.
Footnotes
Acknowledgements
We acknowledge TCGA and GEO databases for providing their platforms and contributors for uploading their meaningful datasets.
Author contributions
QHD and XQW conceived and designed the experiments; JJX performed the data analyses; SMZ and MXH supervised the whole experimental work and revised the manuscript. QHD wrote the paper. All authors read and approved the manuscript.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Ethical approval
TCGA and GEO belong to public databases. The patients involved in the database have obtained ethical approval. Users can download relevant data for free for research and publish relevant articles. Our study is based on open source data.
