Abstract
Introduction
Cervical squamous cell carcinoma (CESC) is the most common gynecological malignancy worldwide. Although the cancer susceptibility 18 (CASC18) gene was involved in the regulation of cancer biology, its specific role in CESC is not well characterized.
Methods
CASC18-related axis was predicted by bioinformatic analyses, and the competing endogenous RNA (ceRNA) interaction was further validated using quantitative real-time PCR, western blotting, RNA pulldown, and luciferase reporter assays. Transwell and wound healing assays were performed to verify the effect of CASC18 on SiHa and HeLa cell motility.
Results
We found that CASC18 was upregulated in CESC tissues. Moreover, interference with CASC18 attenuated NUAK1-mediated epithelial-mesenchymal transition (EMT) and thus suppressed cancer cell motility. Furthermore, the effects of CASC18 knockdown on CESC cells were partly rescued by transfection with the miR-5586-5p inhibitor. Additionally, our findings indicated that CASC18 acts as a ceRNA to enhance NUAK1 expression by sponging miR-5586-5p.
Conclusion
Our study showed a novel CASC18/miR-5586-5p/NUAK1 ceRNA axis that could regulate cell invasion and migration by modulating EMT in CESC. These findings suggest that CASC18 may potentially serve as a novel therapeutic target in CESC treatment.
Introduction
Cervical squamous cell carcinoma (CESC) is the most common gynecological tumor with the highest mortality rate. 1 Many patients are diagnosed with advanced metastatic lesions, which is one of the most critical barriers in the treatment of CESC. 2 Novel insights into the mechanisms underlying CESC invasion and migration are highly desired to improve the survival of these patients.
Recent studies have revealed that a large proportion of eukaryotic genomes are transcribed into non-coding RNA, including long non-coding RNA (lncRNA), whose transcripts are longer than 200 nt.3,4 Over 15,000 lncRNAs have been reported in human genome transcripts, 5 some of which play an important role in tumorigenesis and progression by regulating oncogenes and tumor suppressors. 6 For instance, roles of some non-coding RNAs, such as HOTAIR, MEG3, UCA1, and EWSAT1, during CESC development and progression were reported.7–10 However, only a small number of lncRNAs are well characterized. The competing endogenous RNA (ceRNA) hypothesis is attractive because of its potential to account for the function of some uncharacterized lncRNAs.11,12 As many lncRNAs are rich in miRNA response elements (MREs) via base complementation, they can act as microRNA (miRNA) sponges to competitively absorbing miRNAs and enhance the expression of target genes of the miRNA. 13 With the help of ceRNA prediction by bioinformatic approaches, the functions of lncRNAs will be purposively characterized after experimental verification. The lncRNA Cancer susceptibility candidate 18 (CASC18) is an RNA gene located in the 12q23.3 locus, which contains genes associated with cancer biology. 14 However, this gene was first reported to be a key regulator of neural cell differentiation and oocyte growth.14,15 In our previous study, we found that high CASC18 expression was associated with a high risk of occult lymph node metastasis in patients with squamous cell carcinoma of the tongue, 16 but the role of CASC18 in CESC remains unclear. As a key regulator of AMP-activated protein kinase (AMPK), NUAK1, also known as ARK5, is thought to be closely related to various diseases, including cancers. Upregulation of NUAK1 is associated with a high risk of metastasis and poor prognosis in numerous cancer types.17–20 Moreover, NUAK1 expression correlated with CASC18 expression in the TCGA-CESC dataset. Hence, the potential association between NUAK1 and CASC18 has drawn our attention.
In the present study, we found that CASC18 was upregulated in CESC and facilitated cell invasion and migration. Together with the prediction of shared miR-5586-5p binding sites within CASC18 and the NUAK1 3’-UTR, we hypothesized that CASC18 could facilitate CESC cell motility by antagonizing the inhibitory effect of miR-5586-5p on NUAK1 expression. Our results provide new insights into the molecular mechanism by which CESC promote metastasis, with the potential to identify novel therapeutic targets.
Methods
Tissues and cell line
Formalin-fixed and paraffin-embedded (FFPE) samples were collected from 32 patients with CESC with informed consent, including of matched cancerous and para-cancerous tissues and 24 additional cancerous tissues. All experimental protocols were approved by the ethics committee of Hunan Cancer Hospital (KYJJ-2020-222), Changsha, China. SiHa and HeLa, two CESC cell lines, were purchased from Shanghai GeneChem Co., Ltd. Cells were cultured in DMEM supplemented with 10% fetal bovine serum (FBS) (Gibco, Gaithersburg, MD) and penicillin/streptomycin (TransGen, Beijing, China) as routine and passaged every 2–3 days. A total of 5.0 × 104 cells were seeded in each well in a 24-well plate for 24 h culture. To knockdown CASC18 expression, siRNA (100 nM) transfection (si-CASC18#1: ACCTAGTCATACATCCTAA; si-CASC18#2: CCTTATCCCTGGATGGAAA, RiboBio, Guangzhou, China) was performed using Lipofectamine 2000 (Invitrogen, Carlsbad, CA, mixed in DMEM) for 48 h in cells. Transfection of miR-5586-5p mimic (50 nM, RiboBio, Guangzhou, China), miRNA mimic negative control (mimic-NC, 50 nM, RiboBio, Guangzhou, China), miR-5586-5p inhibitor (50 nM, RiboBio, Guangzhou, China), and miRNA inhibitor negative control (inhibitor-NC, 50 nM, RiboBio, Guangzhou, China) was used to overexpress or knockdown miR-5586-5p.
Bioinformatic analysis
The web tool KM-plotter (https://kmplot.com/) was used to estimate and visualize the overall survival ratio of patients with CESC in TCGA cohort with high and low CASC18 expression, divided by the best performing threshold as the cutoff. 21 The GEPIA 2.0 web tool (https://gepia2.cancer-pku.cn/) was used to identify the CASC18-related protein-coding genes, and pairs with Pearson correlation coefficient (PCC) value >0.5 (p < .05) were considered significantly correlated. 22 Binding miRNAs were predicted by LncBase v.2 (diana.e-ce.uth.gr/lncbasev2) for CASC18, and by miRWalk, 23 miRDB, 24 and TargetScan 25 for four CASC18-related protein-coding genes, and then intersected to obtain the final candidate miRNAs. The gene expression data of patients with CESC were obtained from the Cancer Genome Atlas (TCGA; cancergenome.nih.gov). Subsequently, the counts of miRNAs of interest were displayed as scatter plots, and the correlation between CASC18 and NUAK1 expression was visualized using GEPIA 2.0.
Fluorescence in situ hybridization
To determine the subcellular distribution of CASC18 in SiHa cells, fluorescence in situ hybridization (FISH) was performed using fluorogenic probes (RiboBio, Guangzhou, China) targeting CASC18, 18S, and U6 as described in our previous study. 16 Briefly, cells were firstly fixed in 4% formaldehyde for 10 min. After washing with PBS, the fixed cells were then treated with 0.5% Triton X-100 in PBS for 5 min at 4°C. The specimens were then incubated with blocking solution for 30 min at 37°C. The hybridization was performed in the dark at 37°C overnight, followed by washing with gradient sodium citrate buffer at 42°C. Finally, the nucleus was stained with DAPI, and FISH specimens were examined under a fluorescence microscope (Motic, BA410E, China) to capture fluorescent images with the same exposure setting and illumination.
Quantitative real-time PCR
For formalin-fixed paraffin-embedded (FFPE) specimens, at least four 10-µm scrolls were used to extract total RNA using RNeasy FFPE Kits (QIAGEN, Germantown, MD). For cells, TRIzol (Invitrogen, Carlsbad, CA) was used to purify total RNA from cell pellets. Reverse transcription of total RNA to cDNA was performed using the PrimeScriptTM RT-PCR kit (Takara, Dalian, China). Primer pairs for gene expression quantification were synthesized by RiboBio (Guangzhou, China) using the sequence (CASC18-F: 5’-TGTATAGTCTAGCCAAGTCC-3’, CASC18-R: 5’-ATTTCAGCCATCTTCAGTCCC-3’, β-actin-F: 5’-ACCCTGAAGTACCCCATCGAG-3’, β-actin-R: 5’-AGCACAGCCTGGATAGCAAC-3’) as described in our previous work. 16 qRT-PCR was performed to detect the expression levels of specific genes of interest using SYBR® Premix DimerEraser™ (Takara, Dalian, China) on a Roche LightCycler 480 II Real-Time PCR system (Roche, Basel, Switzerland). Finally, genes of interest were quantified and normalized to the expression of the β-actin internal control as 2−ΔΔCt. Differences between groups were compared by t test using SPSS 23.0, with p < .05 considered statistically significant.
Transwell assay
Cell invasion ability was assessed using a Transwell assay as described previously, with slight modifications. 26 An appropriate amount of Matrigel (BD Biosciences, San Jose, CA) was diluted and added to the upper chamber of the Transwell (Corning, NY, USA) for 4–6 h at 37°C. Cells in each group were suspended in a medium without serum at 2.0 × 105/well and added into the upper chambers. Then, 500 μL of medium plus 10% FBS was pipetted into each lower chamber. The cells were then cultured in Transwell plates for 24 h at 37°C in 5% CO2. After the removal of non-invading cells from the insert chamber, cells that had invaded to the lower surface were fixed and stained with crystal violet for 10 min. Finally, each experiment was repeated in at least triplicate, and stained cells from six randomly selected visual fields were counted under an optical microscope.
Wound healing assay
Cell migration was investigated by wound healing assay according to protocols as reported previously. 27 A sterile 200 µl pipette tip was used to scrape an injured area on the monolayer in a six-well culture dish when the cells reached 80% confluence. The culture was continued in a medium supplemented with 3% FBS and 1% penicillin/streptomycin (TransGen, Beijing, China). After the scratch was made, images were captured at 0 and 24 h, and analyzed using Image J (National Institutes of Health, Bethesda, MD, USA) software as described in our previous study. 16 Briefly, we measured the distances between the two edges of the scratched cells and the percentage of wound closure was calculated.
Western blot
The protein sediment was obtained by centrifugation at 15,000 g for 15 min, after lysing the harvested cells in each six-well plate well with radioimmunoprecipitation assay (RIPA) lysis buffer (Beyotime, Shanghai, China) at 4°C for 30 min. Protein concentration was measured by BCA Protein Assay Kit (Pierce, Rockford, IL). A total of 15 μg of protein extracts were then loaded into the gel and separated using 10% SDS-polyacrylamide gel electrophoresis (SDS–PAGE) according to the standard protocol. Subsequently, the proteins were transferred to PVDF membranes (Millipore, Billerica, MA), and the membranes were incubated in 5% fat-free milk for 30 min. Incubation with specific antibodies (ProteinTech Group Inc., Chicago, IL) against NUAK1 (22,723-1-AP, 1:2000 dilution), Vimentin (10,366-1-AP, 1:1500 dilution), E-cadherin (20,874-1-AP, 1:1000 dilution), and β-actin (20,536-1-AP; 1:2000 dilution) was carried out overnight at 4°C, followed by secondary antibody (HRP-conjugated Affinipure goat anti-rabbit IgG (H + L), SA00001-2, 1:2000 dilution) incubation at room temperature for 1 h. The signals from the proteins of interest were visualized using an enhanced chemiluminescence (ECL) reagent (Pierce, Rockford, IL).
Luciferase reporter assay
The sequence of wildtype NUAK1 3’-UTR (NUAK1-wt) was amplified by high-fidelity PCR with specific primers (cPrimer-NUAK1-3’UTR-F: 5'-CTCGAGCATTCCAGGGCGCC-3', cPrimer-NUAK1-3’UTR-R: 5'-TCTAGATTCAGTTCATTGCAATTTATTTAATTT-3', XhoI and XbaI restriction sites are underlined, Tm = 55°C). The seed sequence of miR-5586-5p binding site was replaced by its complementary sequence to form the miR-5586-5p binding site mutated 3’-UTR of NUAK1 (NUAK1-mut) using QuickMutation™Plus Site-Directed Mutagenesis Kit (D0208S, Beyotime, Shanghai, China) by PCR. The final amplified DNA fragments, NUAK1-wt and NUAK1-mut, were cleaved by XhoI and XbaI, and then ligated into the pmirGLO Dual-Luciferase miRNA Target Expression Vector (Promega, Madison, WI) to construct plasmids for the luciferase reporter assay, respectively. A total of 5.0 × 104 Siha cells were seeded in each well of a 24-well plate for 24 h culture. The vectors were then co-transfected with miR-5586-5p inhibitor (50 nM), inhibitor-NC (50 nM), mimic-NC (50 nM), miR-5586-5p mimic (50 nM), si-NC (100 nM), and si-CASC18#2 (100 nM) into SiHa cells using Lipofectamine 2000 for 48 h. A dual-luciferase reporter assay system (Promega, Madison, WI) was used to measure the relative luciferase activity in each group.
RNA pulldown assay
RNA pulldown assay was performed according to a previously described protocol. 16 A total of 1 × 106 SiHa cells were seeded in a 10 cm dish for 24 h culture. After transfection with si-CASC18#2 or si-NC for 48 h, SiHa cells were incubated with 1% paraformaldehyde solution in PBS for cross-linking, which was then blocked by adding 1.25 M glycine. Cell lysates were subsequently prepared using the lysis buffer (50 mM Tris-HCl PH 7.0, 10 mM EDTA, 1% SDS plus 200 U/mL RNase inhibitor (R0102, Beyotime, Shanghai, China)). Biotinylated probes (100 nmol, GenePharm, Shanghai, China) were incubated with 50 μL of M-280 streptavidin magnetic beads (Sigma-Aldrich, St Louis, MO) to generate probe-coated magnetic beads. Subsequently, the coating beads were incubated with cell lysates in 2 vol of hybridization buffer (50 mM Tris-HCl pH 7.0, 750 mM NaCl, 1 mM EDTA, 1% SDS, 15% formamide plus 200 U/mL RNase inhibitor (R0102, Beyotime, Shanghai, China)) at 4°C overnight, to capture CASC18 and RNA interacting with CASC18. After washing of beads at least six times with hybridization buffer, the RNA was purified using the TRIzol reagent according to manufacturer’s instructions. The abundance of miR-5586-5p was measured by using qRT-PCR.
Statistical analysis
Data were presented as mean ± standard deviation (SD) from at least three separate experiments. The t test was used for comparisons between the groups. The chi-square test was used for investigating the correlation between CASC18 expression and clinicopathological characteristics, and Yates’ correction for continuity was applied where necessary (1 ≤ expectation <5). The overall survival ratio of patients with CESC in each group was analyzed using the log-rank test and visualized using the Kaplan–Meier method. A p-value <.05 was considered statistically significant.
Results
Expression of CASC18 in cervical squamous cell carcinoma
To investigate the expression pattern of CASC18 in CESC, eight pairs of CESC and adjacent tissue samples were used to quantify CASC18 expression. The expression levels of CASC18 increased by 2.14 fold (p = .014) in CESC tissues compared to their adjacent normal tissues (Figure 1(a)). Notably, Kaplan–Meier plotter survival analysis showed that patients with higher CASC18 expression correlated with shorter overall survival (Figure 1(b)). To further investigate the association of CASC18 with CESC, the correlation between CASC18 expression and the clinicopathological characteristics of 42 patients with CESC was assessed. Our results indicated that high CACS18 expression was positively associated with distant metastasis (p < .05), suggesting a potential role of CASC18 in CESC metastasis (Table 1). Expression and prognostic value of CASC18. (a) Expression levels of CASC18 were compared in eight paired samples collected from the patients with CESC by qRT-PCR. (b) Overall survival of patients with CESC in TCGA population, grouped by low and high expression of CASC18, were visualized as Kaplan–Meier plots using log-rank test. Correlation between CASC18 expression and clinicopathologic characteristics of CESC patients. Bold values means statistically significant.

Selection of genes and miRNAs related to the CASC18 in cervical squamous cell carcinoma
FISH staining followed by qRT-PCR was performed to determine the subcellular localization of CASC18 in the SiHa cells. We found that CASC18 was localized in both the cytoplasm and nucleus (Figure 2(a)), suggesting that CASC18 might have the opportunity to at least partly function as a miRNA-sponge in the cytoplasm. To investigate the potential ceRNA axes regulated by CASC18 in CESC (Figure 2(b)), we obtained CASC18-related genes using GEPIA 2.0, with criteria of Pearson correlation coefficient ≥0.5 and p-value ≤.05, and four candidate genes, DEFB136, ZNF322, NUAK1, and RASSF6, were identified. Subsequently, miRNAs binding to CASC18 (score >0.8) were predicted using LncBase v.2, and those binding to CASC18-related genes were predicted using the miRWalk, TargetScan, and miRDB databases. The intersection of these miRNAs is miR-5586-5p, miR-5195-3p, and miR-4753-3p (Figure 2(c)). We then investigated the expression levels of these miRNAs in the TCGA-CESC subset and found that CESC barely expressed miR-4753-3p (Figure 2(d)). In addition, expression data for miR-5195-3p were absent from the database. Therefore, the CASC18/miR-5586-5p/NUAK1 axis was selected for subsequent studies, and the positive correlation (R = 0.54) between CASC18 and NUAK1 is shown in Figure 2(e). The expression levels of NUAK1 increased by 2.21 fold (p = .006) in CESC tissues compared to their adjacent normal tissues (Figure 2(f)). The same trend in overall survival was observable in patients with higher NUAK1 expression as those with higher CASC18 expression although the difference was not significant (p = 0.07) (Figure 2(g)). Screening of the potential ceRNA axis related to CASC18 in CESC. (a) The subcellular distribution of CASC18 was assessed by FISH in SiHa cells, with the reference of 18S and U6. (b) Flowchart for screening the potential ceRNA axes regulated by CASC18 in CESC. (c)The potential ceRNA axes regulated by CASC18 were investigated. Briefly, miRNAs binding to CASC18 were predicted using LncBase v.2, and those binding to CASC18-related genes, DEFB136, ZNF322, NUAK1, and RASSF6, were predicted using the miRWalk, TargetScan, and miRDB databases. Shared miRNAs are selected by taking intersection. (d) The counts of candidate miRNA expression in TCGA-CESC. (e) Scatter diagram shows a positive correlation between CASC18 and NUAK1 expression in CESC tissues. (f) Expression levels of NUAK1 were compared in eight paired samples collected from the patients with CESC by qRT-PCR. (g) Overall survival of patients with CESC in TCGA population, grouped by low and high expression of NUAK1, were visualized as Kaplan–Meier plots using log-rank test.
Knockdown of CASC18 inhibited cell invasion and migration via repressing NUAK1 and epithelial-mesenchymal transition
CASC18 knockdown was performed to investigate the biological function of CASC18 in CESC cells. Transfection with si-CASC18#1 and si-CASC18#2 decreased CASC18 expression by 71.3% and 81.0% in SiHa cells, and 59.7% and 64.7% in HeLa cells, respectively (Figure 3(a)). Transwell assays indicated that CASC18 knockdown inhibited cell invasion by 72.3% and 69.3% in SiHa and HeLa, respectively (Figure 3(b)). Similarly, CASC18 knockdown interference decreased cell migration (Figure 3(c)). Moreover, these changes in cell motility were partly rescued by the miR-5586-5p inhibitor, suggesting that this miRNA plays an essential role in this process. As epithelial-mesenchymal transition (EMT) is closely related to cell invasion and migration, as well as the role of NUAK1 in EMT regulation, we further assessed the changes in the expression levels of NUAK1 and EMT-related biomarkers using western blotting. We found that the expression of NUAK1 and the mesenchymal marker, Vimentin decreased with CASC18 knockdown, whereas that of the epithelial marker E-cadherin increased (Figure 3(d)). Similarly, transfection with the miR-5586-5p inhibitor attenuated the inhibitory effect of si-CASC18#2 on the expression of EMT-related proteins. These data indicated that the CASC18/miR-5586-5p/NUAK1 axis is important for the regulation of cervival cancer cell invasion and migration. Effect of CASC18 on cervical cancer cell invasion and migration. (a) qRT-qPCR was employed to detect the interference efficiency of siRNAs on CASC18 expression in cells. (b) Transwell assays were used to determine the invasion ability of each group of cells. (c) Wound healing assays were used to determine the migration ability of each group of cells. (d) Changes in NUAK1, Vimentin, and E-cadherin expression were determined using western blotting (cropped gels), with β-actin as the internal control. *, p < .05.
CASC18 acts as a miR-5586-5p sponge to regulate NUAK1 expression and epithelial-mesenchymal transition
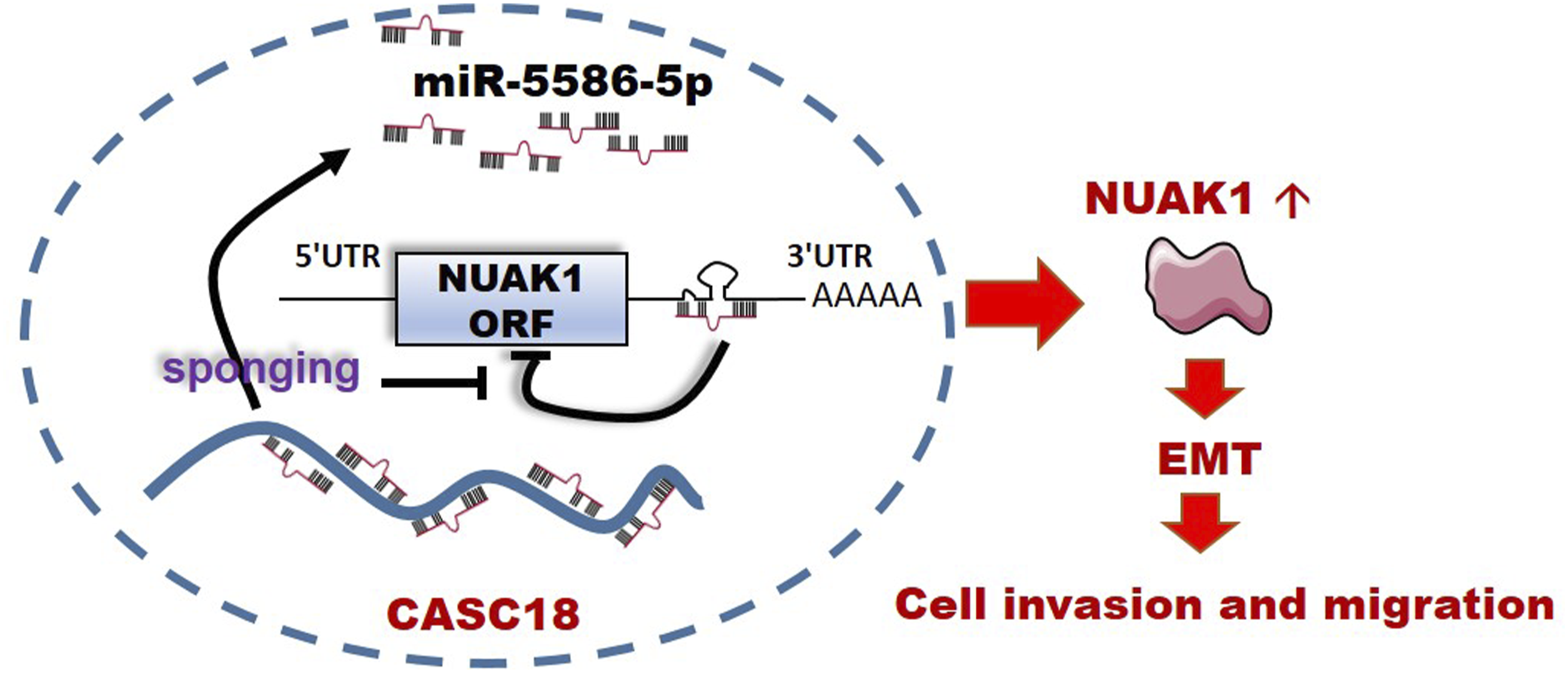
We first examined the effect of miR-5586-5p on NUAK1 expression. Our results showed that transfection of a gradient concentration of miR-5586-5p mimic reduced NUAK1 expression in SiHa cells, indicating the inhibitory effects of miR-5586-5p on NUAK1 expression (Figure 4(a)). Furthermore, we found miR-5586-5p mimic significantly decreased the expression of the reporter gene with NUAK1-wt 3’UTR, but had no effect on that with NUAK1-mut 3’UTR (Figure 4(b)). These results suggest that NUAK1 is a direct target of miR-5586-5p dependent on 3’-UTR binding. To further verify the interaction between CASC18, miR-5586-5p, and NUAK1, luciferase reporter and RNA pulldown assays were further performed in SiHa cells. When CASC18 RNAi was used, the expression of the reporter gene with NUAK1-wt 3’-UTR was significantly decreased. Conversely, it had no effect on the expression of the reporter gene with NUAK1-mut 3’-UTR (Figure 4(c)). Moreover, the miR-5586-5p inhibitor attenuated the inhibitory effect of si-CASC18#2 on the expression of reporter gene with NUAK1-wt 3’-UTR (Figure 4(c)), suggesting that the effect of CASC18 on NUAK1 expression depends on protecting miR-5586-5p from NUAK1 3’-UTR. Besides, the abundance of miR-5586-5p captured by the CASC18 probes decreased significantly when CASC18 was downregulated by RNAi (Figure 4(d)), further verifying the binding interaction between CASC18 and miR-5586-5p in SiHa cells. Therefore, our findings indicated that CASC18 promotes NUAK1 expression by sponging miR-5586-5p and facilitates CESC cell migration and invasion (Figure 5). CASC18 positively regulates NUAK1 by binding miR-5586-5p. (a) Effect of miR-5586-5p mimic on NUAK1 expression in SiHa cells. (b) Effect of miR-5586-5p mimic on luciferase activity of the reporter gene in SiHa cells. (c) Effect of miR-5586-5p inhibitor and si-CASC18#2 on luciferase activity of the reporter gene in SiHa cells. (d) Abundance of CASC18 and miR-5586-5p captured by CASC18 probes in different groups were quantified using an RNA pulldown assay followed by qRT-PCR. Data are presented as the mean ± standard deviation (SD). *, p < .05. Schematic diagram of the CASC18/miR-5586-5p/NUAK1 ceRNA axis. There are several miR-5586-5p binding sites within CASC18 transcript. In the cytoplasm of CESC cells, CASC18 sponged miR-5586-5p and prevented it from binding to the 3'-UTR region of NUAK1mRNA, thereby attenuating the inhibitory effect of miR-5586-5p on NUAK1 expression. Elevated levels of NUAK1 protein promoted EMT in CESC cells, which in turn promoted CESC cell migration and invasion.
Discussion
Cancer cell metastasis is one of the most important prognostic factors for patients with CESC. However, the underlying molecular mechanism remains to be fully elucidated. In the present study, we found CASC18 upregulation in CESC and investigated the role of the CASC18/miR-5586-5p/NUAK1 ceRNA axis in CESC cell metastasis.
Mehravar et al revealed that CASC18-D, a novel splice variant of the CASC18 gene, plays an important role in neural cell differentiation. 14 Another study has indicated the involvement of CASC18 in the regulation of oocyte growth. 15 As indicated by the full name of this gene, cancer susceptibility 18, CASC18 was located at the 12q23.3 locus, a potential cancer predisposition locus in the human genome, and flanked by APPL2,28,29 C12orf75 (OCC-1),30,31 and NUAK1, 32 all of which are well characterized as important regulators in cancers. However, the role of CASC18 in cancer biology has not been well characterized. Previously, we found that CASC18 promoted tongue squamous cell carcinoma cell invasion and migration, 16 consistent with the findings of the present study. As bioinformatic prediction indicated that there were shared miRNA-binding sites within CASC18 and the 3’-UTR of its neighboring gene NUAK1, we further verified the potential regulatory axis between these two genes. As an oncogene, NUAK1, also located at locus 12q23.3, has been reported to be involved in the tumorigenesis and metastasis of various cancers. 33 NUAK1 knockdown represses EMT and reduces cell mobility in ovarian cancer, another common gynecological cancer.34,35 Notably, EMT can provide cancer cells with an advantage in terms of cell mobility.36–39 Additionally, NUAK1 contributes to pancreatic cancer metastasis by enhancing MMP-2 and MMP-9 expression.40,41 All these studies support our findings that the CASC18/NUAK1 axis facilitates CESC cell invasion and migration by promoting EMT. Meanwhile, we inferred that CASC18 might have similar functions in squamous cell carcinoma originating from other tissue types, but through distinct signaling axes. MiR-5586-5p is another key regulator of the CASC18/NUAK1 pair. Although miR-5586-5p has not been well characterized, some studies have indicated that high miR-5586-5p expression predicts favorable outcomes in diffuse large B-cell lymphoma and pancreatic cancer.42,43 Similarly, the suppressive effect of CASC18 knockdown on CESC cell invasion and migration could be partly reversed by transfection with a miR-5586-5p inhibitor, indicating a protective effect of miR-5586-5p in patients with CESC. Although we provide some novel insights into the regulation of CESC cell mobility, there is some limitation in the present study. First, the roles of CASC18 still need to be further demonstrated in vivo. Second, the amount of clinical samples used in this study was small, and subsequent in-depth studies can be continued to detect the association between gene expression and clinicopathologic characteristics. In addition, clinical studies could be further designed to explore the clinical value of CASC18-related signaling axes as potential biomarkers.
Conclusions
Overall, we demonstrated a novel CASC18/miR-5586-5p/NUAK1 ceRNA axis and indicated its involvement in CESC cell invasion and migration. CASC18 enhanced NUAK1 expression through sponging miR-5586-5p and preventing it from binding to the 3'-UTR region of NUAK1 mRNA, which promoted EMT-mediated cell migration and invasion in CESC. Our findings provide new insights into the molecular mechanisms of CESC metastasis, which may provide fundamental clues for the development of novel diagnostic methods and drugs.
Supplemental Material
Cancer susceptibility 18 positively regulates NUAK Family Kinase 1 expression to promote migration and invasion via sponging of miR-5586-5p in cervical cancer cells
Cancer susceptibility 18 positively regulates NUAK Family Kinase 1 expression to promote migration and invasion via sponging of miR-5586-5p in cervical cancer cells by Jingrong Wang, Bocheng Zhang, Sha Gong, Ying Liu, Liang Yi and Ying Long in International Journal of Immunopathology and Pharmacology
Footnotes
Acknowledgements
We acknowledge TCGA database for providing their data for cancer research.
Authors’ contributions
Ylong, LY, and JW conceived and designed the study; JW, SG, LY, and BZ collected and prepared the samples; JW, BZ, LY, and SG carried out bioinformatic analyses; JW, BZ, and SG performed the experiments; JW, YLiu, and YLong wrote the manuscript. BZ, LY, and YLong revised the manuscript. All authors reviewed the manuscript and approved the final version.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The present study was supported in part by the National Natural Science Foundation of China (Grant no. 81703005 and 82273466), the Key Research and Development Projects of Hunan Province (grant number 2018SK2120 and 2018SK2121) and the Natural Science Foundation of Hunan Province (Grant no. 2022JJ40255 and 2023JJ60242).
Ethical statement
Trial registration
This study was not registered because it was an observational study, and the dose does not affect the treatment of participants at all.
Data Availability Statement
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
