Abstract
Background
Kin17 is critical in regulating the proliferation and metastasis of tumors in various malignancies. However, the relationship between Kin17 expression, clinicopathologic features, and esophageal squamous cell carcinoma (ESCC) prognosis remains unclear.
Methods
The analysis of Kin17 messenger RNA (mRNA) expression involved the utilization of data from The Cancer Genome Atlas (TCGA) dataset through the platforms the University of ALabama at Birmingham CANcer data analysis portal (UALCAN) and the Gene Expression Omnibus (GEO). To determine the expression levels of Kin17 in tissues, immunohistochemistry was conducted. Using Pearson's chi-square test, the relationship between Kin17 expression and clinicopathological variables was evaluated. Cox proportional hazard models (both univariate and multivariate), receiver operating characteristic (ROC) curves, and Kaplan–Meier survival curves were used to analyze survival.
Results
In both the TCGA and GEO datasets, the mRNA level of Kin17 was greater in tumor tissues when compared to tumor-adjacent tissues (P < 0.001). Similarly, there was a significant expression of Kin17 (P < 0.0001) in ESCC tissues. Elevated Kin17 expression correlated significantly with increased Ki-67 levels (P < 0.001), advanced pathological tumor node metastasis stage (P = 0.01), and positive lymph node metastasis (P = 0.02). According to univariate and multivariate Cox models, high Kin17 expression was associated with poorer progression-free survival (PFS) (hazard ratio (HR): 1.990, 95% confidence interval (CI): (1.040–3.810)), and Kin17 was an independent prognostic variable for overall survival (OS) (HR: 2.321, 95% CI: (1.056–5.101)). ROC curve showed that the area under the curve for predicting PFS and OS using the combination of Kin17 and K-i67 was 0.7088 and 0.7031, respectively. High Kin17 expression was associated with unfavorable PFS (HR: 2.009, 95% CI: (1.059–3.811)) and OS (HR: 2.997, 95% CI: (1.488–6.040)).
Conclusions
Kin17 is abundantly expressed in ESCC tissues and is potentially useful for prognostic evaluation and as a target for therapeutic interventions in ESCC.
Keywords
Introduction
Esophageal cancer (ESCA) ranks seventh in the global incidence of malignant tumors and sixth in overall mortality. Approximately 70% of cases occur in males, with the incidence and mortality rates being two- to three-times higher in men than in women. China is the country with the highest incidence of ESCA worldwide, with the largest number of new cases, accounting for more than half of the global total.1–3 ESCA is mainly classified into two histological subtypes: esophageal adenocarcinoma and esophageal squamous cell carcinoma (ESCC). In China, 90% of ESCA cases are ESCC, with major risk factors including alcohol and tobacco use, as well as poor dietary habits such as consuming hot or hard foods, eating too quickly, and irregular eating patterns. Carcinogens from alcohol metabolism and tobacco, including aromatic amines, aldehydes, phenols, and nitrosamines, enter esophageal epithelial cells via the esophagus or systemic circulation, causing dysplasia and contributing to the development of ESCC.4,5
Unlike other cancers, early screening for ESCC is both difficult and unreliable. Typical symptoms of ESCC usually do not appear until the advanced stages of the disease. Factors such as tumor size, invasion of surrounding tissues and blood vessels, and lymph node metastasis can limit the treatment options for the tumor itself. 6 Although immunotherapy and targeted treatment have improved the prognosis of ESCC; nonetheless, the 5-year survival rate is still less than 20%. 7 Current pathological indicators, such as tumor location, size, and cell differentiation, rely heavily on the subjective experience of clinicians and pathologists, failing to provide precise individualized diagnosis. Meanwhile, serum tumor markers like carcinoembryonic antigen (CEA), cancer antigen (CA19-9), and squamous cell carcinoma (SCC) have limited specificity and are often insufficient for accurate therapeutic prediction in ESCC. With the growing application of targeted and immunotherapies, the potential of targeted gene therapies is increasingly recognized. Research is now focusing on identifying new molecular biomarkers for early detection and prediction of prognosis in ESCC.
Kin17 is a highly conserved protein that is widely distributed across mammalian tissues. It is usually expressed at very low levels in most tissues and organs, except for the heart, skeletal muscle, and testis. 8 It contains a tandem SH3 domain that binds to RNA and curved DNA at recombination hotspots in eukaryotic chromosomes.9,10 Kin17 is a key component in DNA replication, transcription, messenger RNA (mRNA) processing, cell cycle regulation, and the response to genotoxic stress.11,12 It is upregulated in response to DNA damage from ionizing radiation or UVC, depending on the functionality of the global genome repair system.13,14 As a DNA maintenance protein, Kin17 helps overcome replication perturbations caused by unrepaired lesions. 15 Studies have reported that Kin17 plays a critical role in cancer cell proliferation, invasion, and migration. It promotes cell proliferation in HepG2 and SMMC-7721 cells by upregulating cyclin D1 and p27, reduces migration and invasion in HeLa and SiHa cells by inhibiting the NF-κB-Snail pathway.16,17 However, the relationship between Kin17 and ESCC remains unclear.
Throughout our investigation, it was observed that Kin17 expression was upregulated in esophageal cancer tissues. We found an association between Kin17 expression and different clinicopathological parameters, and its impact on survival and prognosis in ESCC patients. Our data indicated that Kin17 is a crucial participant in ESCC progression and is potentially useful for prognostic evaluation and as a therapeutic target for ESCC.
Materials and methods
Kin17 mRNA expression analysis
The University of ALabama at Birmingham CANcer data analysis portal (UALCAN) is a biological analysis tool 18 that integrates data from diverse sources. Specifically, it incorporates RNA sequencing expression data from both tumor and normal samples obtained from The Cancer Genome Atlas (TCGA) database. All data are standardized to ensure consistency for subsequent analysis. UALCAN was employed to validate the differential expression of Kin17 between cancerous and normal tissues, as well as to evaluate individual cancer stages and nodal metastasis status in esophageal cancer. GSE23400 in the Gene Expression Omnibus (GEO) database retrieved Kin17 mRNA expression data from 53 pairs of ESCC patients’ tumor biopsy tissues and adjacent tissues, and compared them.
Clinical samples
Sixty-seven samples of ESCC were collected, including both adjacent normal tissue and tumor tissue specimens that were verified through pathological diagnosis. These samples were then preserved in paraffin for further analysis. The patients from whom these samples were obtained had undergone surgical procedures at the Affiliated Cancer Hospital of Guangzhou Medical University over a 1-year period from July 2016 to July 2017. Furthermore, 23 paraffin-embedded specimens from patients with benign esophageal squamous hyperplasia (ESH) were pathologically identified after surgical resection. The clinicopathological parameters of all patients are showed in Supplemental Table S1. This study was approved by the Institutional Research Ethics Committee of the Affiliated Cancer Hospital of Guangzhou Medical University (approval number: KY-2025004-01). All participants provided written informed consent before participation.
The International Union against Cancer and the United States Joint Commission on Cancer developed the standard for international tumor node metastasis (TNM) staging (8th edition), which was followed in 67 cases of postoperative TNM staging. The serum levels of tumor markers before surgery were based on the reference range established by the Department of Laboratory Medicine, the Affiliated Cancer Hospital of Guangzhou Medical University, CEA: (0.00–5.00) ng/mL, SCC: (0.00–2.70) ng/mL, CA19-9: (0.00–30.00) U/mL. For survival analysis, the date from surgery to death was used to define overall survival (OS), whereas the date from surgery to the first ESCC progression or death was used to determine progression-free survival (PFS). Final follow-up assessments were conducted on June 16, 2022.
Immunohistochemistry
Sections were baked for 30 min at 60°C, dewaxed, and rehydrated. Endogenous peroxidase was then inhibited for 10 min by applying 3% H2O2 after the pH 9.0 repair solution (97°C) was applied for 30 min, cooled at 26°C, and then circled. Following overnight incubation, primary antibody Kin17 (1:500 dilution, sc-32769, Santa Cruz Biotechnology, Inc.), Ki-67 (GM027, Shanghai Genetech Co., Ltd), and phosphate-buffered saline as a negative control, the slices were incubated with horseradish peroxidase (HRP)-labeled secondary antibody. Following visualization by immunostaining with diaminobenzidine, sections were counterstained with hematoxylin, dried, and mounted.
Immunohistochemical scoring
Two pathologists who were blinded to the clinical diagnosis scored the stained slides according to the procedure outlined by Merritt et al. 19 Histological images were captured under a light microscope with an objective magnification of 100× and 400×. By examining the proportion of positive cells and staining intensity, the expression of Kin17 was assessed. The evaluation criteria for staining positivity intensity were divided into four levels: no color was 0, light yellow was 1, yellow brown was 2, and brown was 3. The proportion of positive cells were calculated in multiple representative fields under high magnification. The proportion of positive cells and the staining intensity were multiplied to get an immunoreactivity score. The expression level of Ki-67 was assessed based solely on the proportion of positive cells, which was the more clinically relevant and commonly used metric. For further research, ESCC patients were divided into two groups based on the median values of immunohistochemistry (IHC) results: high and low expression. For Kin17, a value of ≥120 was defined as high expression, while <120 was defined as low expression. For Ki-67, an expression level of ≥50% was considered high expression, whereas <50% was considered low expression.
Statistical analysis
Statistical analyses were conducted utilizing GraphPad Prism (GraphPad Software, Inc.) and SPSS (IBM). For comparisons, Student's t-test is used for two groups, the Mann–Whitney U test for non-normally distributed data in two groups, and one-way analysis of variance for multiple groups. The correlation analysis of clinicopathological parameters and Kin17 expression was carried out using Pearson's chi-square test or Fisher's exact test. Factors contributing to ESCC prognosis were determined through univariate and multivariate Cox regression analyses. Combined Kin17 and Ki-67 to predict PFS and OS in ESCC patients were evaluated by ROC curve. Survival analysis was conducted using Kaplan–Meier survival curve, and differences between survival curves were assessed using a log-rank test. A significance level of P < 0.05 was considered statistically significant.
Results
Kin17 mRNA expression was associated with ESCC development
The result showed that Kin17 expression was significantly elevated in most cancer tissues, including ESCA (P < 0.001), when compared to normal specimens by using TCGA database (Figure 1(a)). Next, the GEO dataset GSE23400 was used to validate that Kin17 mRNA expression was higher in ESCC tissues than in normal tissues (Figure 1(d)) (P < 0.001). Further comparison revealed that Kin17 was highly expressed in nodal metastasis-positive and advanced stages ESCA patients than that in normal tissues, but no significant differences were observed between different stages or phases, as shown in Figure 1(b) and (c).
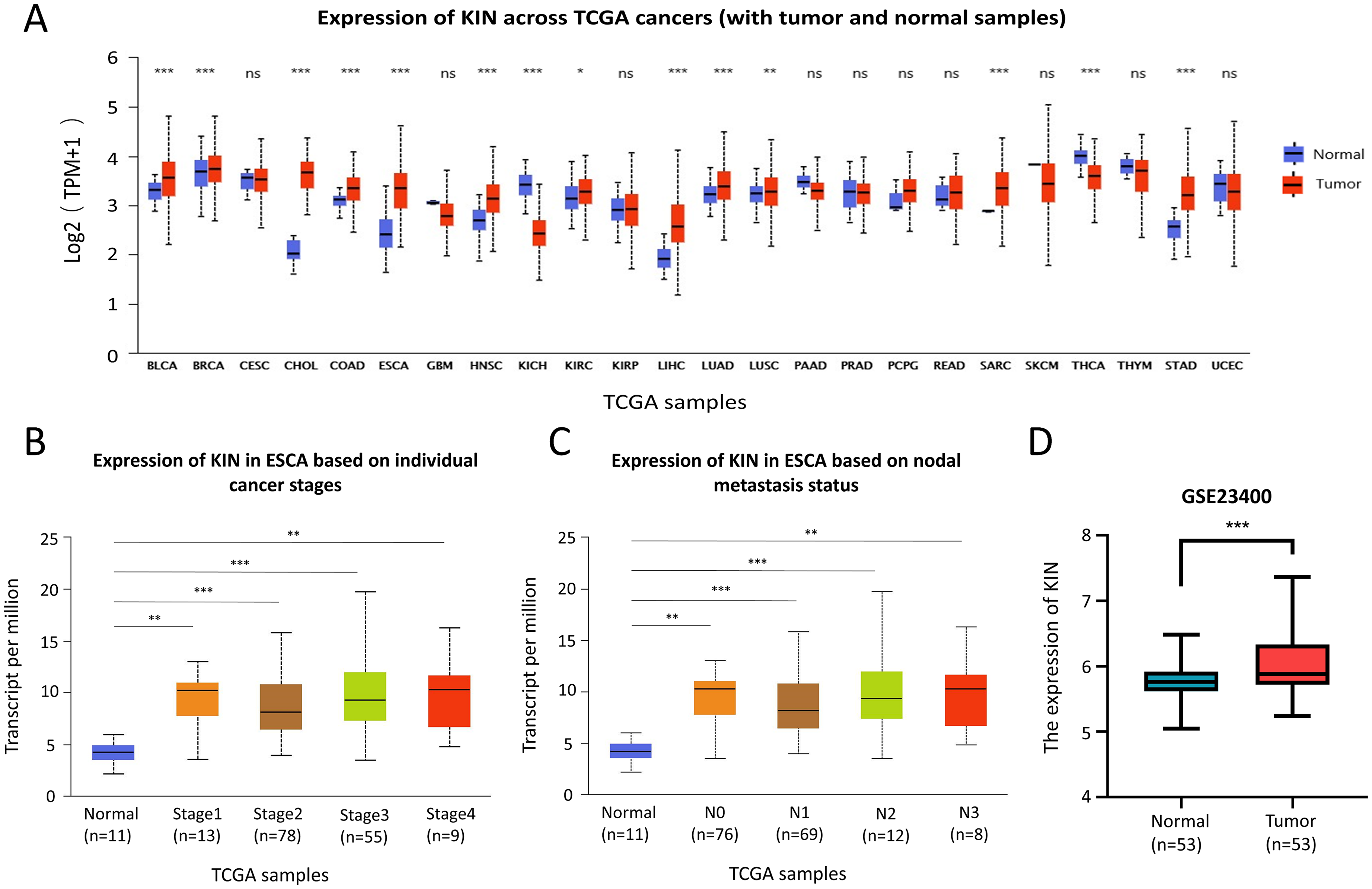
The mRNA expression profile of Kin17 in ESCA. (a) The expression of Kin17 in different human cancer tissues compared with normal tissues from the TCGA database. (b) Comparison of Kin17 expression in ESCA based on nodal metastasis. (c) Comparison of Kin17 expression in ESCA based on individual cancer stages. (d) The expression of Kin17 in ESCC tissue compared with the adjacent normal tissue in GEO datasets GSE23400. ***P < 0.001.
Kin17 expression was elevated in ESCC tissues
Immunohistochemical staining revealed that Kin17 showed strong nuclear staining in tumor cells and weak staining in some normal squamous epithelia of the corresponding tumor-adjacent tissues. However, in benign lesion tissues, strong positive staining was detected in a small area of squamous epithelium (Figure 2(a)). Based on IHC scores, Kin17 was upregulated in ESCC tissues (Figure 2(b) (P < 0.0001). Furthermore, ESCC patients with higher Ki-67 expression had significantly higher Kin17 expression (P < 0.001; Figure 2(c)).
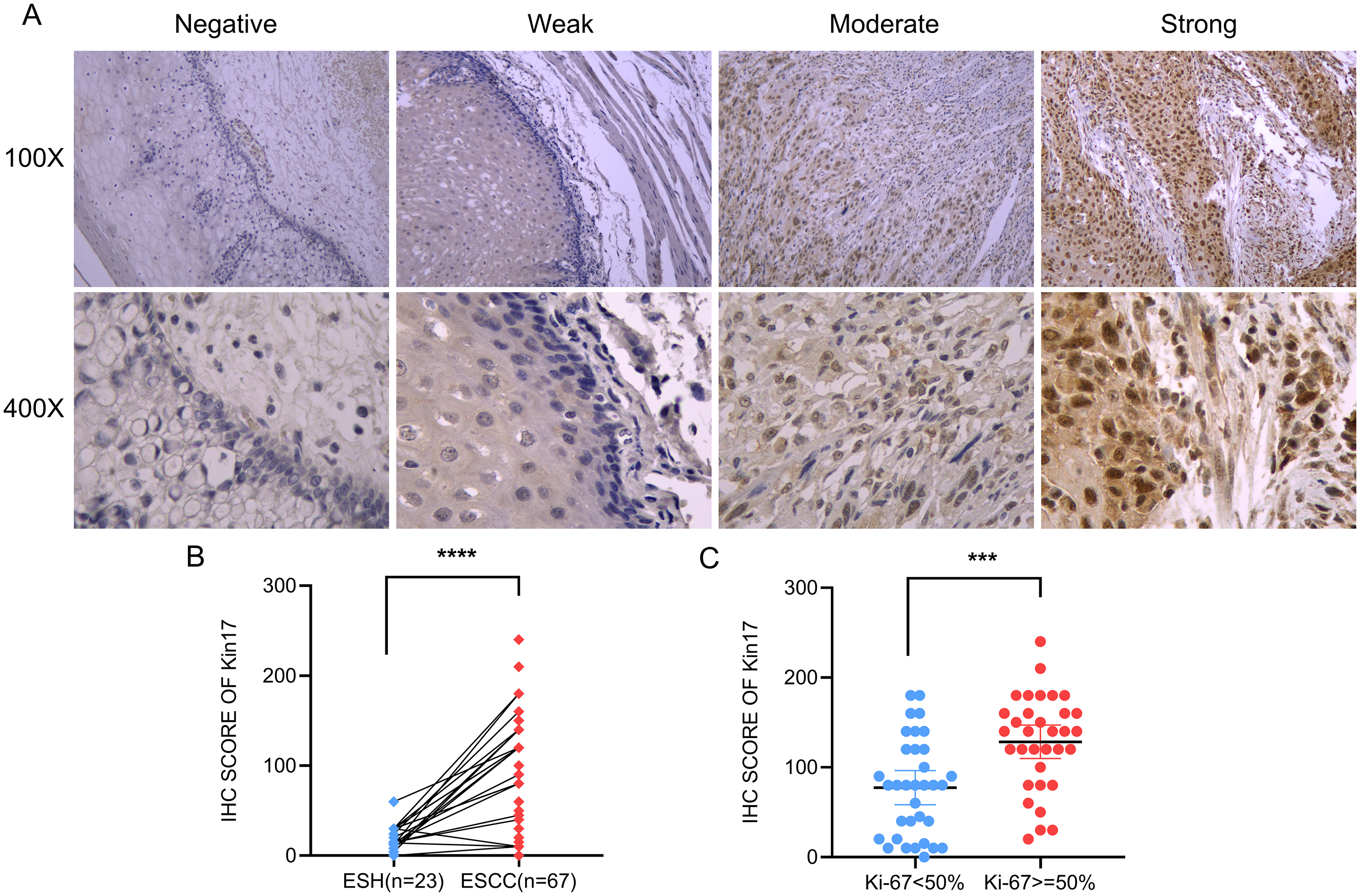
Representative images and comparison between groups of immunohistochemical staining for Kin17. (a) Representative images of IHC staining of Kin17 with negative, weak, moderate, and strong expression in tissues (100×, 400×). (b) The expression of Kin17 in ESCC tissues compared with ESH tissues based on IHC scores. (c) The comparison of Kin17 between groups at different Ki-67 expression levels based on IHC scores.
Correlation between Kin17 expression and clinical parameters
According to the IHC staining scores, 34 (50.7%) ESCC tissue samples had elevated Kin17 expression. The correlation results were showed in Table 1. Elevated Kin17 expression was correlated with advanced pathological tumor node metastasis (pTNM) stage (P = 0.01) and positive lymph node metastasis (P = 0.02) and significantly correlated with elevated Ki-67 expression levels(P < 0.001). Kin17 expression levels were not significantly correlated with other clinicopathological features.
Correlation between Kin17 and clinicopathological parameters in ESCC
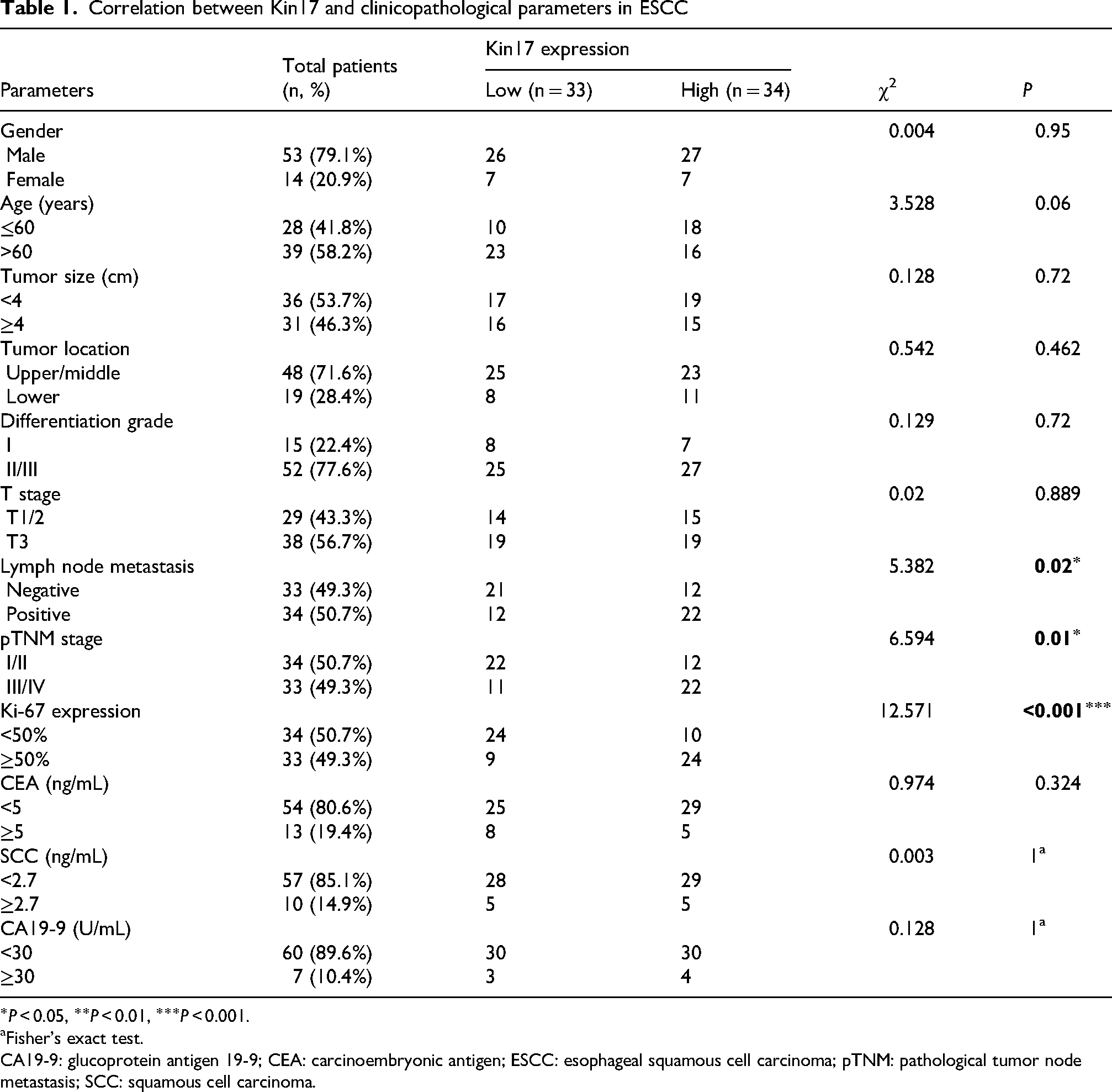
Fisher's exact test.
CA19-9: glucoprotein antigen 19-9; CEA: carcinoembryonic antigen; ESCC: esophageal squamous cell carcinoma; pTNM: pathological tumor node metastasis; SCC: squamous cell carcinoma.
Prognostic value of Kin17 in ESCC
Univariate analysis showed that a significant decrease in PFS was associated with high expression levels of Kin17 (HR: 1.990, 95% CI: (1.040–3.810)), Ki-67 (HR: 1.969, 95% CI: (1.028–3.773)), positive lymph node metastasis (HR: 2.271, 95% CI: (1.180–4.369)), and pTNM stage III/IV (HR: 2.563, 95% CI: (1.328–4.944)). Only pTNM stage III/IV (HR: 2.280, 95% CI: (1.142–4.554)) was an independent predictor for PFS (Supplemental Table S2). For OS, high Kin17 expression (HR: 3.039, 95% CI: (1.436–6.433)), high serum SCC expression levels (HR: 2.545, 95% CI: (1.139–5.687)), pTNM stage III/IV (HR: 2.160, 95% CI: (1.068–4.368)), and T3 stage (HR: 2.135, 95% CI: (1.033–4.410)) were correlated with decreased OS according to univariate analysis. Moreover, Kin17 expression (HR: 2.321, 95% CI: (1.056–5.101)) and serum SCC expression levels (HR: 2.768, 95% CI: (1.174–6.523)) were independent predictive factors for OS, in accordance with multivariate Cox models (Supplemental Table S3). The area under the ROC curve for predicting the survival outcomes of PFS and OS in ESCC patients by combining Kin17 and Ki-67 could reach 0.7088 and 0.7031 (Figure 3).
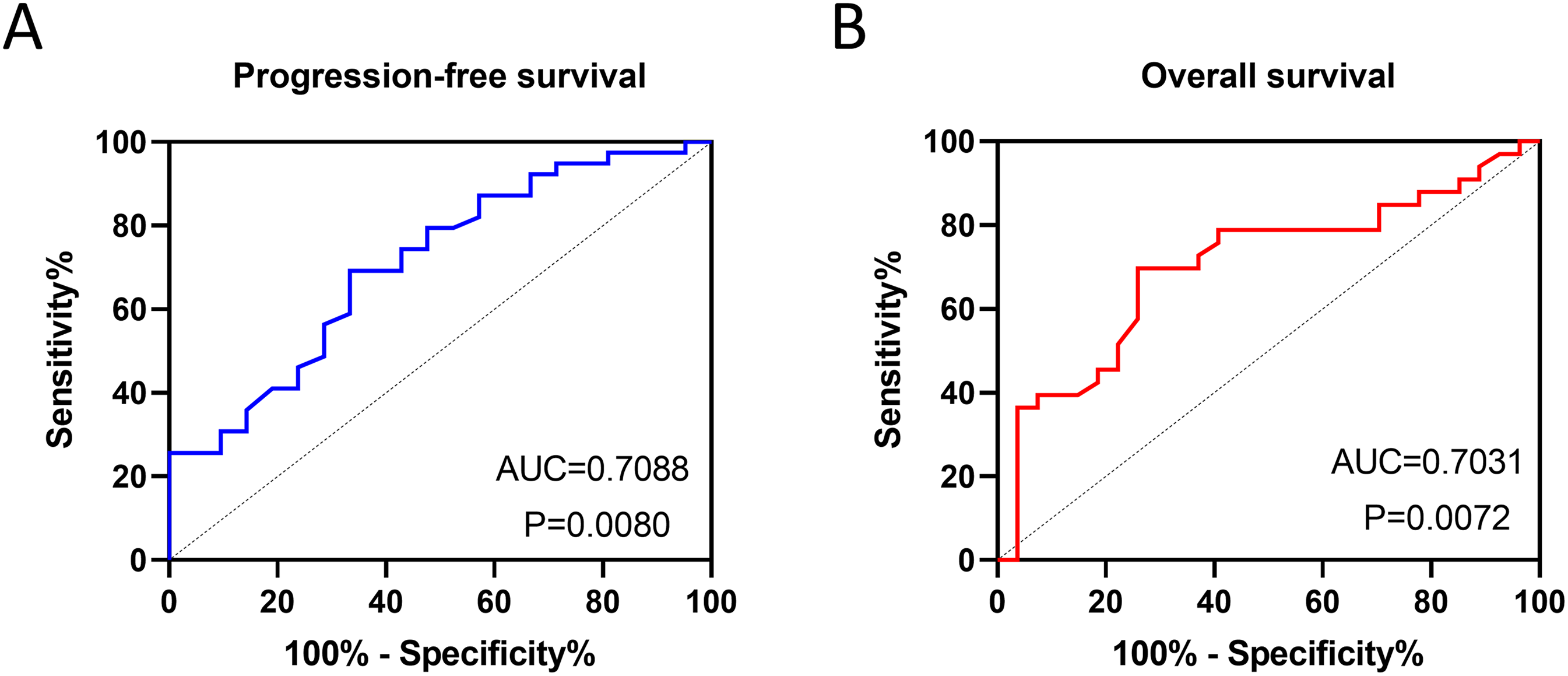
ROC curve in all ESCC patients to evaluate the accuracy of combining Kin17 and Ki-67 in predicting survival outcomes. (a) Progression-free survival. (b) Overall survival.
Survival outcome of Kin17 in ESCC
The results revealed that patients with high Kin17 expression had poor survival in terms of PFS (HR: 2.009, 95% CI: (1.059–3.811)) and OS (HR: 2.997, 95% CI: (1.488–6.040)) (Figure 4). The PFS and OS results for were analyzed by co-grouping Kin17 expression with different clinicopathological factors that influence survival outcomes. The survival outcomes of PFS and OS were consistent. Kaplan–Meier survival curves showed that patients who expressed high expression levels of Kin17 and had pTNM stage III/IV or positive lymph node metastasis had poor PFS (Supplemental Fig. S1(c) and (d)) and OS (Supplemental Fig. S2(c) and (d)). Conversely, patients in the T 1/2 stages with low Kin17 expression had favorable PFS (Supplemental Fig. S1(b)) and OS (Supplemental Fig. S2(b)). In addition, the result indicated that patients with low expression levels of Kin17 and SCC exhibited superior OS (Supplemental Fig. S2(a)), whereas PFS did not vary statistically between patients in the groups with high and low expression levels of Kin17 and Ki-67 (Supplemental Fig. S1(a)). Overall, Kin17 is critical for initiating and progressing ESCC.
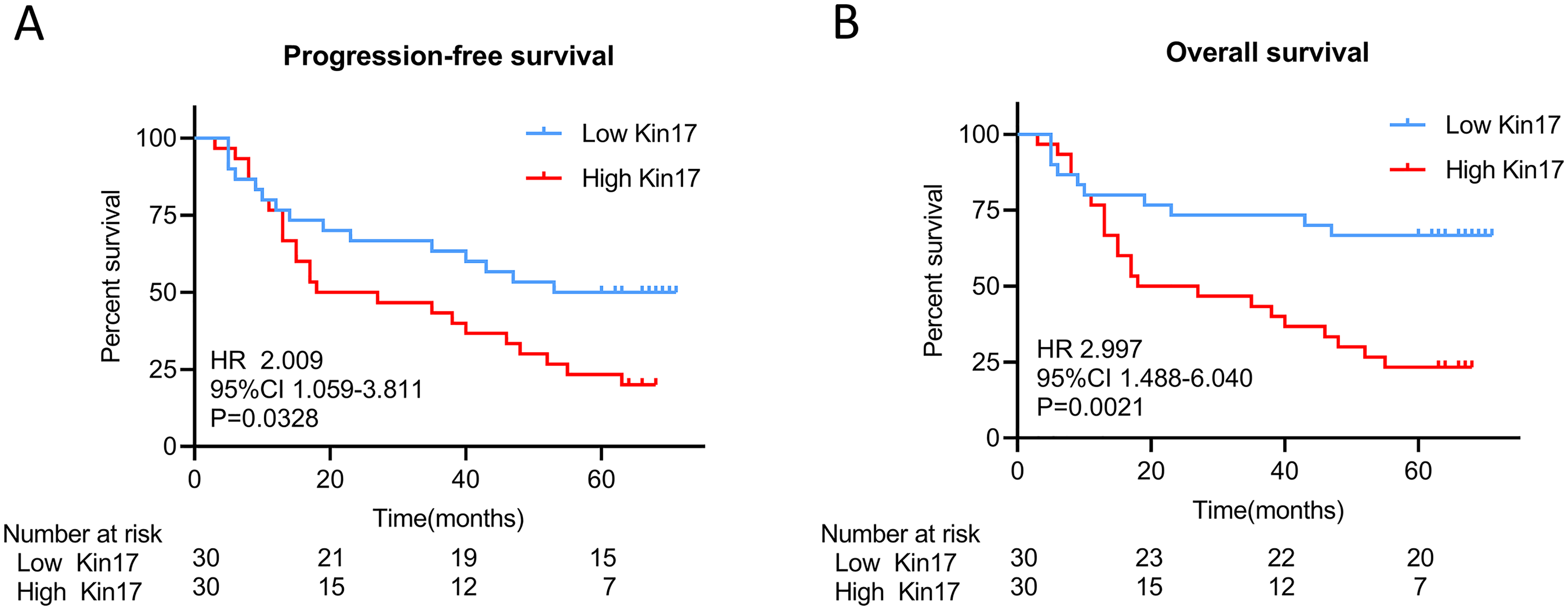
Kaplan–Meier survival curves in all ESCC patients grouped by high and low Kin17 expression. (a) Progression-free survival. (b) Overall survival.
Discussion
Patients with esophageal cancer have a poor prognosis because they lack distinct early symptoms and clear indicators. This is especially true when the disease is in its later stages and treatment is being administered.20,21 Thus, extending the life of patients with esophageal cancer greatly depends on the discovery of molecular markers for prognostic evaluation. 22 Recent studies have shown that Kin17 expression is elevated in various tumor types, suggesting that Kin17 expression could be used as a potential biomarker for different cancers.
According to Zeng et al., 23 invasive ductal carcinoma exhibits significantly higher Kin17 expression than benign breast disorders, through double label immunofluorescence analysis, Kin17 and Ki-67 in breast cancer tissues were mainly located in the nucleus, indicating that Kin17 and Ki-67 had the same subcellular localization in the nucleus. Here, TCGA, GEO, and IHC findings also indicated that Kin17 was upregulated in ESCC, suggesting high Kin17 expression was correlated with high Ki-67 expression. Although not as ideal as expected, the combined detection of Kin17 and Ki-67 expression still holds certain predictive value for the PFS and OS of ESCC patients and represents the optimal combination among various combinations examined in this study.
The Ki-67 protein, which encodes a 359KD non-histone nuclear protein, is a widely used cellular marker for proliferation. 24 All active stages of the cell cycle are characterized by the presence of Ki-67, which is strongly associated with mitosis. Ki-67 is commonly marker of cell proliferation in clinical practice and scientific experiments. Huang et al. 25 found strong correlations between Ki-67 expression and ESCC malignancy, pathological type, and lymph node metastasis. A noteworthy association between a high Ki-67 labeling index and poor OS in patients with esophageal cancer was further clarified by Jacobsen et al. 26
In non-small cell lung cancer, Zhang et al. 27 also found higher Kin17 expression than in benign pulmonary lesions, and it was correlated with tumor grading and lymph node metastasis status. This is consistent with the observation in ESCC. The elevation of Kin17 in ESCC tissues led us to postulate that Kin17 may be associated with the prognosis.
The univariate and multivariate Cox models indicated that high Kin17 expression was associated with poorer PFS and OS in ESCC patients, and it was an independent prognostic variable for OS. Kaplan–Meier survival curves showed that patients who expressed high levels of Kin17 and had pTNM stage III/IV or positive lymph node metastasis had poor survival outcomes.
According to Chen et al., 28 there is considerable upregulation of Kin17 expression in epithelial ovarian cancer tissues, resulting in lower 3- and 5-year OS rates, and it was an independent risk factor for OS. In hepatocellular carcinoma patients, it was found that Kin17 can affect the metastasis of cancer cells by regulating the expression of Smad2, p-Smad2, and epithelial-to-mesenchymal transition-related proteins in the TGF–β/Smad2 pathway, 29 Kin17 expression levels act as a prognostic factor, and patients with relatively higher Kin17 levels have poor OS and disease-free survival.
Furthermore, Kaplan–Meier survival curves showed that patients expressing low levels of Kin17 and SCC had better OS results. Tumor marker index (TMI) based on cyfra21-1 and SCC is an independent adverse prognostic factor, with high TMI patients having considerably poorer 5-year survival rates than low TMI patients, according to reports by Qiao et al. 30 and Yin et al. 31
However, this study was limited by the small sample size due to incomplete follow-up. In ESCC patients with high expression of Kin17, the impact on PFS and OS was not as significant as that observed in other types of tumors. Moreover, the predictive capacity of Kin17 combined with Ki-67 for PFS and OS in ESCC patients also did not reach the desired level. This limitation also affected the grouping of various tumor-related indicators and the resulting differences observed. Additionally, due to the lack of relevant molecular mechanism studies and functional analyses such as knockout or overexpression models, the mechanisms by which high expression of Kin17 contribute to poor prognosis in ESCC remain to be further elucidated.
In conclusion, Kin17 is highly expressed in ESCC tissues and is correlated with metastasis in ESCC patients. High Kin17 expression was associated with poor PFS and OS in ESCC patients. Therefore, Kin17 may be a promising prognostic biomarker for ESCC. Future work should expand the sample size, improve follow-up protocols, and employ rigorous experimental design to address existing gaps and provide stronger evidence for ESCC prognosis and targeted therapies.
Supplemental Material
sj-docx-1-jbm-10.1177_03936155251343636 - Supplemental material for High Kin17 expression is correlated with metastasis and prognosis of esophageal squamous cell carcinoma
Supplemental material, sj-docx-1-jbm-10.1177_03936155251343636 for High Kin17 expression is correlated with metastasis and prognosis of esophageal squamous cell carcinoma by Zhenkai Chen, Ruiqi Su, Liwen Jiang, Lina Yang, Xiaocong Lin, Qiyuan Huang, Kashif Rafiq Zahid, Yunen Lin and Tao Zeng in The International Journal of Biological Markers
Footnotes
Abbreviations
Acknowledgments
This research was funded by the Guangdong Basic and Applied Basic Research Foundation (grant No. 2023A1515010235), Start-up Fund for High-level Talents in Affiliated Hospital of Guangdong Medical University (grant No. 51301Z20200007), Medical Science and Technology Research Project of Guangdong Province (grant No. A2023168 and B2021180) and Discipline construction project of Guangdong Medical University (grant No. 4SG21266P and 4SG21276P). The funders had no role in study design, data collection and analysis, manuscript preparation or decision to publish.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was supported by the Guangdong Basic and Applied Basic Research Foundation (grant No. 2023A1515010235); Start-up Fund for High-level Talents in Affiliated Hospital of Guangdong Medical University (grant No. 51301Z20200007); Medical Science and Technology Research Project of Guangdong Province (grant No. A2023168 and B2021180); Discipline construction project of Guangdong Medical University (grant No. 4SG21266P and 4SG21276P).
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
