Abstract
Background
Hepatobiliary cancers present a heterogeneous group of diseases, and molecular residual disease (MRD) evaluation based on circulating tumor DNA (ctDNA) is anticipated to offer greater sensitivity in monitoring disease progression than glycoprotein-based tumor markers such as alpha-fetoprotein or carbohydrate antigen 19-9 (CA199).
Methods
The panels for MRD surveillance were customized for each patient based on their specific genetic mutation characteristics. The changes in ctDNA mean variant allele frequencies (mVAF) and single nucleotide variants (SNVs) were analyzed from baseline to post-operative and between post-operative measurements.
Results
A unique tumor-informed whole-exome sequencing (WES) assay revealed significant variations in gene mutations between individuals. Among 63 cases, a total of 1952 SNVs were detected in tumor tissue from 63 patients using WES; only 6 loci (0.3%) were shared by at least 2 patients, indicating that over 95% of the 20–40 loci screened were unique to individual patients . Only 17 gene alterations were common to at least 2 patients, suggesting that alterations vary widely between individuals. The mVAF and the number of SNVs in ctDNA at baseline was dramatically higher than in first post-operative MRD (MRD1). The mVAF clearance was observed in three patients, whose ctDNA was positive at MRD1 but subsequently became negative at the second post-operative MRD (MRD2). Patients exhibiting vascular invasion demonstrated a significant increase in mVAF levels and SNV numbers. Furthermore, we revealed that mVAF levels were significantly associated with clinicopathologic characteristics, including gender, age, tumor subtype, stage, metastasis, vascular invasion, hepatitis B, liver cirrhosis, and tumor differentiation. Importantly, we have shown that the detection of an MRD-guided medication regimen modification is crucial to achieve clinical complete remission.
Conclusions
This study provided data supporting the use of a more reliable assay for MRD analysis in hepatobiliary cancers based on a tumor-informed assay. Dynamic monitoring of post-operative MRD is important for assessing disease progression, risk of recurrence, and response to treatment.
Introduction
Hepatobiliary cancers are a diverse group of diseases with morbidity and mortality depending on its stage of presentation. 1 Alpha-fetoprotein (AFP) and carbohydrate antigen (CA199) are commonly used glycoprotein-based tumor markers for screening and prognostic purposes. However, their sensitivity and specificity are limited because not all cancers produce these glycoproteins. Circulating tumor DNA (ctDNA) is a promising non-invasive biomarker that can be measured and quantified in liquid biopsies to identify molecular residual disease (MRD) and assess disease progression as well as response to treatment. 2 In solid tumors, MRD refers to the small number of tumor cells that persist following radical cancer treatment. ctDNA consists of DNA fragments released during the apoptosis, necrosis, and rupture of tumor cells. 3 By monitoring ctDNA levels, it is possible to obtain the level of microscopic residual lesions, providing guidance for more precise individualized post-operative treatment, which is widely and intensively used in breast, bowel, and lung cancers.4–7 Pre-operative plasma MRD testing predicts early recurrence of hepatocellular carcinoma (HCC).8,9 HCC patients with detectable somatic mutations in plasma ctDNA after surgery had worse disease-free survival than those without mutations. 10 Therefore, ctDNA is currently mostly utilized to assess whether patients with MRD detected by ctDNA following surgery and/or adjuvant therapy, may benefit from additional or intensive systemic therapy to mitigate the risk of recurrence.11–13 However, further research is needed to explore the potential risk of preventing disease progression and to evaluate post-operative outcomes.
Although fresh and archived biopsy material has traditionally been used for genomic profiling of solid tumor, this approach is hampered by several restrictions, such as limited availability, low repeatability, and high risk of failure. 14 In addition, when surgical tissue or biopsies are unobtainable from patients, timely detection and prompt treatment of the condition may become unfeasible. As a tool for real-time monitoring of cancer patients, liquid biopsy methods are gaining increasing attention. Recent technological advances in ctDNA detection and characterization can facilitate treatment tailoring in accordance with real-time monitoring of tumor evolution. 15 Multiple platforms have achieved exciting analytical performance in detecting MRD, encompassing both tumor-informed (personalized panel) and tumor-agnostic ctDNA assays.16–18 Tumor-agnostic ctDNA assay is a blood-based test that does not rely on a priori on tumor profiling as another assay for ctDNA detection method.19–21 Tumor-informed ctDNA assays designed for MRD detection depend on initial genomic profiling of tumor tissue to identify patient-specific alterations, so that these precise alterations can be detected in ctDNA.22,23 Prior knowledge of tumor-specific mutations in “tumor-informed” approaches can increase the sensitivity of ctDNA testing, perhaps determining which alterations are tumor-derived and which have arisen from non-tumor origins with potential false positives including clonal hematopoiesis of certain potential, and then improving specificity. However, to date, the application of tumor-informed ctDNA assays for MRD assessment has been relatively limited in hepatobiliary cancers.
In the current study, we aimed to assess the applicability and feasibility of a tumor-informed ctDNA assay for evaluating MRD in hepatobiliary cancers. A unique tumor-informed ctDNA assay was customized to investigate the correlation among baseline, postoperative ctDNA VAF, single-nucleotide variant (SNV) numbers and clinical parameters, and the changes in ctDNA VAF and SNV numbers from baseline to post-operative and among post-operative measurements were analyzed. ctDNA VAF and SNV number changes also be attempted to determine the application of predicting disease progression, post-operative efficacy, and post-operative recurrence.
Materials and methods
Patients’ enrollment and sample collection
A total of 63 hepatobiliary patients were enrolled in our study, including 31 patients with HCC, 29 patients with biliary tract cancer (BTC), and the other 3 with mixed tumor types. Tumor tissue samples were collected firstly and > 20% tumor cells were detected by whole exome sequencing (WES). While blood leukocytes were taken as controls to exclude germline and to confirm that all detected mutations were of tumor origin. Baseline blood samples were collected from 22 patients 7 days prior to surgery, and post-operation blood samples for ctDNA detection were obtained from 58 patients. Patient clinical data, including personal and family history, tumor subtype, and stage, were collected. This study was approved by the Ethical Committee of Eastern Hepatobiliary Surgery Hospital, Second Military Medical University (Shanghai, China) with reference number of EHBHKY2023-K002-P001 and written consents were received from all participating patients.
Tumor-informed assay
Genomic DNA was collected through using QIAamp DNA FFPE Tissue Kit and QIAamp DNA Blood Midi Kit (Qiagen, Hilden, Germany). WES was conducted with a mean depth of 500x on tumor biopsies and matched peripheral-blood mononuclear cell samples. A self-developed commercial method was used to screen for somatic SNV loci for monitoring MRD. Performance evaluation was carried out using standard references comprising two commercially available standards consisting of five positive sites each at VAF of 0.1% and 1%, alongside a wild-type sample. Additionally, a set of single nucleotide polymorphism references was generated by amalgamating DNA from three distinct samples (160 sites). To establish reference standards for our assay, we blended genomic DNA (gDNA) samples devoid of the SNV loci of interest to create VAFs of 0%, 0.005%, 0.01%, 0.02%, 0.05%, and 0.1%. For generating circulating free DNA (cfDNA) references, we titrated a clinical cfDNA mixture from five patients with mononucleosomal HEK293 DNA to create VAFs ranging from 0.005 to 0.1% across 20–40 positive sites. The OriCleverClover algorithm continuously tested and optimized the filter conditions, while the OriSelector algorithm was continuously optimized. To optimize tumor notification assays, we used normal tissues as controls. Using WES and self-developed optimization algorithms, we selected 20–40 ctDNA monitoring SNV sites with a VAF of ≥ 0.02% to ensure a sensitivity and specificity of ≥ 99%.
ctDNA-MRD detection
The screening of 20–40 SNVs from the aforementioned tumor tissues was performed using a Multiplex polymerase chain reaction (PCR) assay by NEB Next Ultra II Q5 Master Mix. A library was prepared using VAHTS® Universal DNA Library Prep Kit for Illumina V3 and sequenced on the NovaSeq 6000 system. A sample was considered MRD-positive if more than two mutations were detected within the SNV pool encompassing all mutations, and MRD-negative if zero or 1 mutation was detected. The distinct VAFs observed among various mutations present in ctDNA indicated differing levels of clonality among the mutations.
Statistical analysis
R statistical software version 3.4.3 package (R Foundation for Statistical Computing) was used for all analyses. Regression analysis was performed to analyze the differences between ctDNA and clinical parameters. For normally distributed continuous variables, the Student's t-tests was conducted to compare differences between two groups, whereas for non-continuous variables, the F-test was used for statistical comparisons. Continuous variables were presented as mean, and p-value less than 0.05 was regarded as statistically significant.
Results
Patient demographics and clinicopathologic characteristics
The clinicopathologic characteristics of the patients are summarized in Supplementary Table 1. The cohort in this study comprised predominantly male patients (n = 46, 73.02%) compared to women (n = 17, 26.98%). The average age was 57.5 years, ranging from 30 to 74 years. Of all patients, 22 (34.92%) were in stage I/II and 37 (58.73%) were in stage III/IV. Fifteen patients (23.81%) had a history of alcohol, and 10 (15.87%) had a history of smoking. All 63 patients underwent surgery or puncture, and baseline blood samples were collected within 7 days before surgery. Baseline ctDNA was detected on 22 patients, 20 of whom tested positive and 2 tested negative. Fifty-eight patients had at least one MRD monitoring after surgery: 2 patients were monitored five times, 1 patient was monitored four times, 8 patients were monitored three times, 12 patients were monitored two times, and 35 patients were monitored one time (Supplementary Table 1). Seventeen patients underwent both baseline and postoperative MRD monitoring. Regardless of baseline, we analyzed the dynamic changes in ctDNA levels among 23 patients who had at least two postoperative tests (12 two times, 8 three times, 1 four times, and 2 five times).
Tumor tissues and ctDNA genomic landscape of hepatobiliary patients
Tumor tissues from 63 patients were sequenced using WES testing, and several SNVs were selected for ctDNA-MRD detection (Figure 1 and Supplementary Table 2). Among 63 cases, the most frequently altered genes were TP53 (65.1%), TERT (31.7%%), TTN (30.2%), CDKN2A (23.8%), OBSCN (23.8%), ARID1A (20.6%), TSC2 (20.6%), AGO2 (19%), MUC16 (19%), and SLC45A4 (19%) (Figure 1(a)). A total of 1952 SNVs were detected in tumor tissues of 63 patients using WES, with only 6 loci (0.3%) being common to at least 2 patients (Supplementary Table 3), indicating that more than 95% of the 20–40 loci screened were independently possessed by each patient. MRD surveillance sites vary widely among individuals, which meant that personalized customization was necessary. The WES screened 30 clonal somatic mutations for the design of a personalized, tumor-informed ctDNA-MRD assay; the most frequently altered genes (only SNVs were observed) are shown in Figure 1(b). Additionally, we presented the high-frequency mutated genes detected in ctDNA post-surgery (Figure 1(c)), most of which were not associated with drug responsiveness and tumor prognosis, except for TP53, PIK3CA, and MAP2K4, which were not included in the commercial next generation sequencing (NGS) panel. As shown in Figure 1(d) and Supplementary Table 4, only 17 gene alterations were observed to be shared by at least 2 patients, suggesting that alterations vary widely among individuals. This heterogeneity is well known in hepatobiliary tumors, which exhibit both spatial and temporal heterogeneity. In addition to confirming that there were great individual differences in gene variation characteristics, this study was expected to improve the accuracy of MRD detection by obtaining gene variation characteristics of each patient with a tissue priority method, and then customize the panel for MRD monitoring according to the characteristics of each patient.
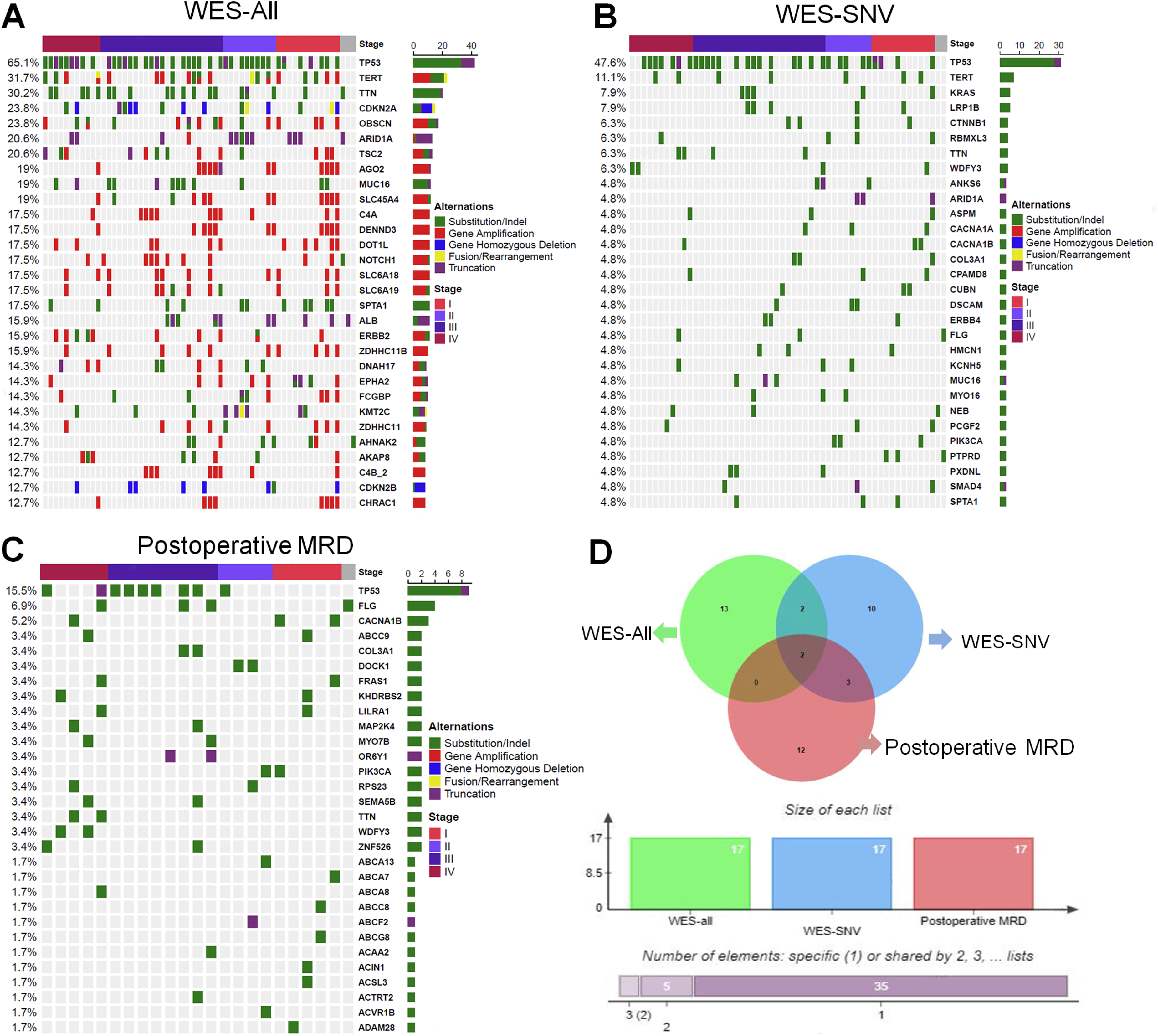
Genomic landscape of hepatobiliary patients. (a) The genomic alteration profiles from WES testing for all patients. (b) The WES custom genomic alteration profiles for SNV mutation site. (c) The ctDNA genomic alteration profiles. (d) According to 17 genes with MRD sample size ≥ 2, the differences between WES-all group, WES customized SNV mutation sites and postoperative MRD detection among the three groups were analyzed.
Association of changes in VAF levels from baseline to postoperative MRD
Seventeen patients, comprising 2 with gallbladder cancer, 13 with HCC, and 2 with intrahepatic cholangiocarcinoma, underwent both baseline and postoperative MRD monitoring. We analyzed the dynamic changes of ctDNA in these patients from pre-operative to post-operative. Supplementary Table 5 provides the clinicopathologic characteristics of these patients, with the majority (64.71%) having undergone a radical operation. At baseline, 16 patients tested positive for ctDNA and 1 was negative. However, the first post-operative MRD (MRD1) testing revealed that 7 patients were positive and 10 were negative, with a median post-operative sampling time of 60 (12–754) days. The mVAF in baseline ctDNA was 35.55%, whereas in post-operative MRD1 it was 10.16%, and the mean numbers of SNV were 24 and 8, respectively. We found that the mVAF and the numbers of SNV in ctDNA at baseline were dramatically higher than in post-operative MRD1 (P < 0.0001, Figure 2(a) and (b)).
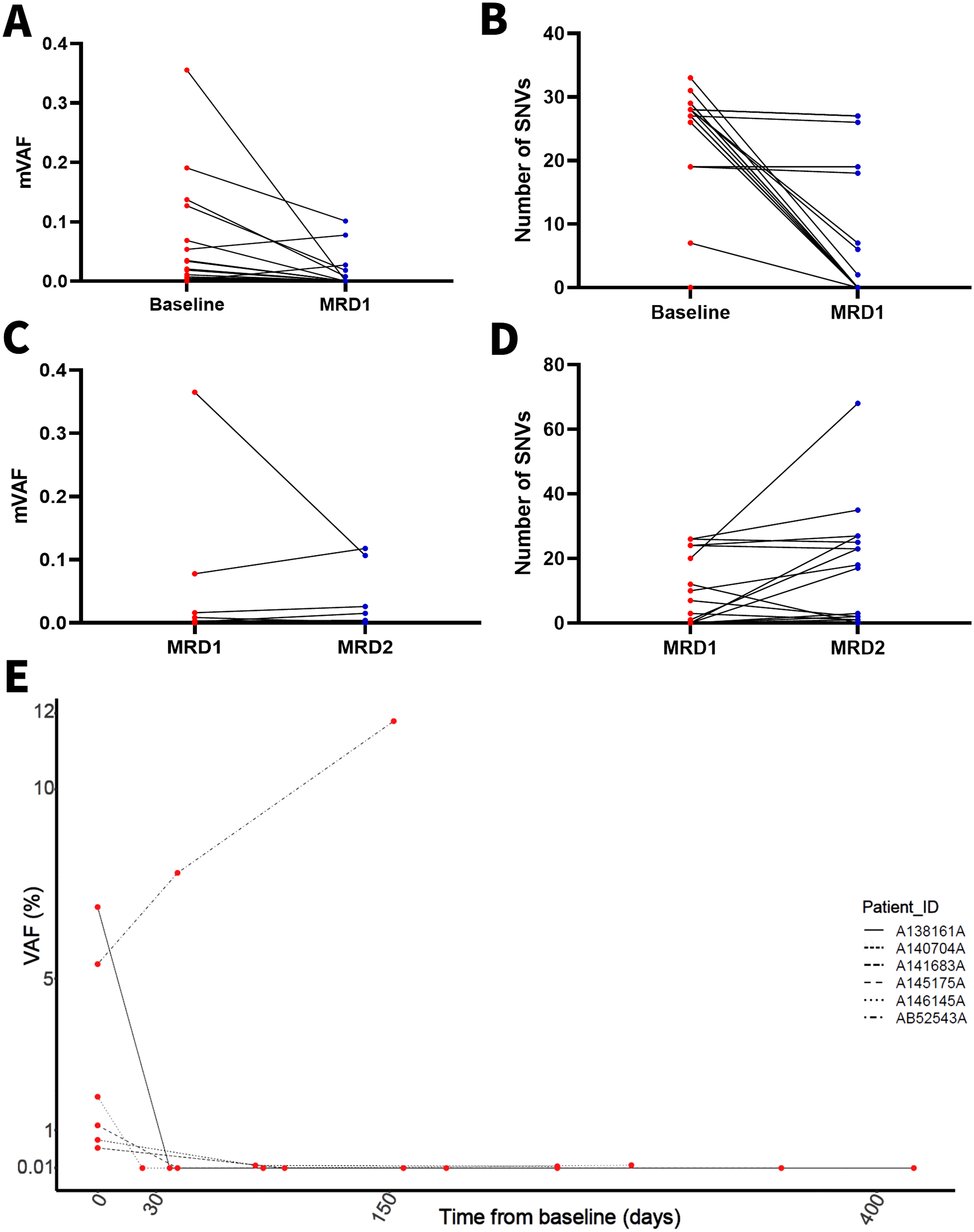
Analysis of ctDNA dynamic. (a) The mVAF change was determined from baseline to postoperative MRD1. (b) The number of SNVs change was determined from baseline to postoperative MRD1. (c) The mVAF change was determined from postoperative MRD1 to MRD2. (d) The number of SNVs change was determined from postoperative MRD1 to MRD2. (e) Analysis of changes in VAF in individual patients.
Twenty-three patients underwent ctDNA testing at least twice post-operatively, and we analyzed the dynamic of post-operative ctDNA changes in these patients. Detailed clinicopathologic characteristics of these patients are shown in Supplementary Table 6. Nine patients tested positive for ctDNA at post-operative MRD1, and 14 were negative. However, the second postoperative MRD (MRD2) testing revealed that 10 patients were positive and 13 were negative for ctDNA. Notably, the mVAF clearance was observed in 3 patients whose ctDNA at MRD1 was positive but became negative at MRD2 (Figure 2(c)). The numbers of SNV decreased in 5 patients, increased in 10 patients, and remained stable in 8 patients (Figure 2(d)). Six patients underwent ctDNA testing at baseline and at least twice post-surgery (Figure 2(e)). We found an upward trend in the VAF of one patient across two postoperative ctDNA tests. Over time, MRD might dynamically change in response to medications or autoimmune function. We suspected that MRD could provide further insights into disease progression after recurrence; however, this conjecture needs to be confirmed by further prospective and specialized studies. This result implied the necessity of dynamic monitoring.
Association of VAF levels, SNV numbers, and clinicopathologic characteristics
We explored the correlation between VAF levels, SNV numbers, and clinicopathologic characteristics (Figure 3 and Supplementary Figure 1). Our results revealed that mVAF levels were significantly higher in BTC compared to HCC (P = 0.0003, Figure 3(a). However, no significant correlation was observed between the number of SNVs and tumor subtypes. Patients with stages II–III disease exhibited a relatively higher mVAF level compared to those at stage I (P = 0.087); whereas patients at stage IV had significantly higher mVAF levels compared to both stage I (P = 0.0059) and stages II–III (P = 0.013). Although the mean number of SNVs was progressively increased with stage, there were no statistically significant differences (Figure 3(b)). Patients with hepatitis B (HBV) or liver cirrhosis had significantly lower mVAF levels compared to those without HBV or cirrhosis; however, the number of SNVs was opposite in both groups of patients, with dramatically higher SNVs in patients with HBV or cirrhosis than those without those conditions, suggesting that the number of SNVs might reflect the characteristics of patients with HBV or cirrhosis (Figure 3(c) and (d)). Patients with vascular invasion had dramatically elevated mVAF levels (P = 0.046) and SNV numbers (P = 0.015) (Figure 3(e)). Furthermore, we showed that mVAF levels were significantly associated with gender (P < 0.0001, Figure 4(a)) age (P < 0.0001, Figure 4(b)), tumor subtype (P < 0.0001, Figure 4(c)), stage (P < 0.0001, Figure 4(d)), metastasis (P < 0.0001, Figure 4(e)), vascular invasion (P < 0.0001, Figure 4(f)), HBV (P < 0.0001, Figure 4(g)), liver cirrhosis (P < 0.0001, Figure 4(h)), and differentiation (P < 0.0001, Figure 4(I)). However, our analysis did not reveal a significant relationship between SNV numbers and clinicopathologic characteristics (data not shown). The mVAF levels seemed to provide more information and refine layers, and the overall trend of SNV numbers was similar to that of mVAF and played a certain role in providing valuable indications. Nevertheless, we found that some results were different from those of mVAF. Further exploration is required to determine which indicator is accurate and which scenarios they applied to.
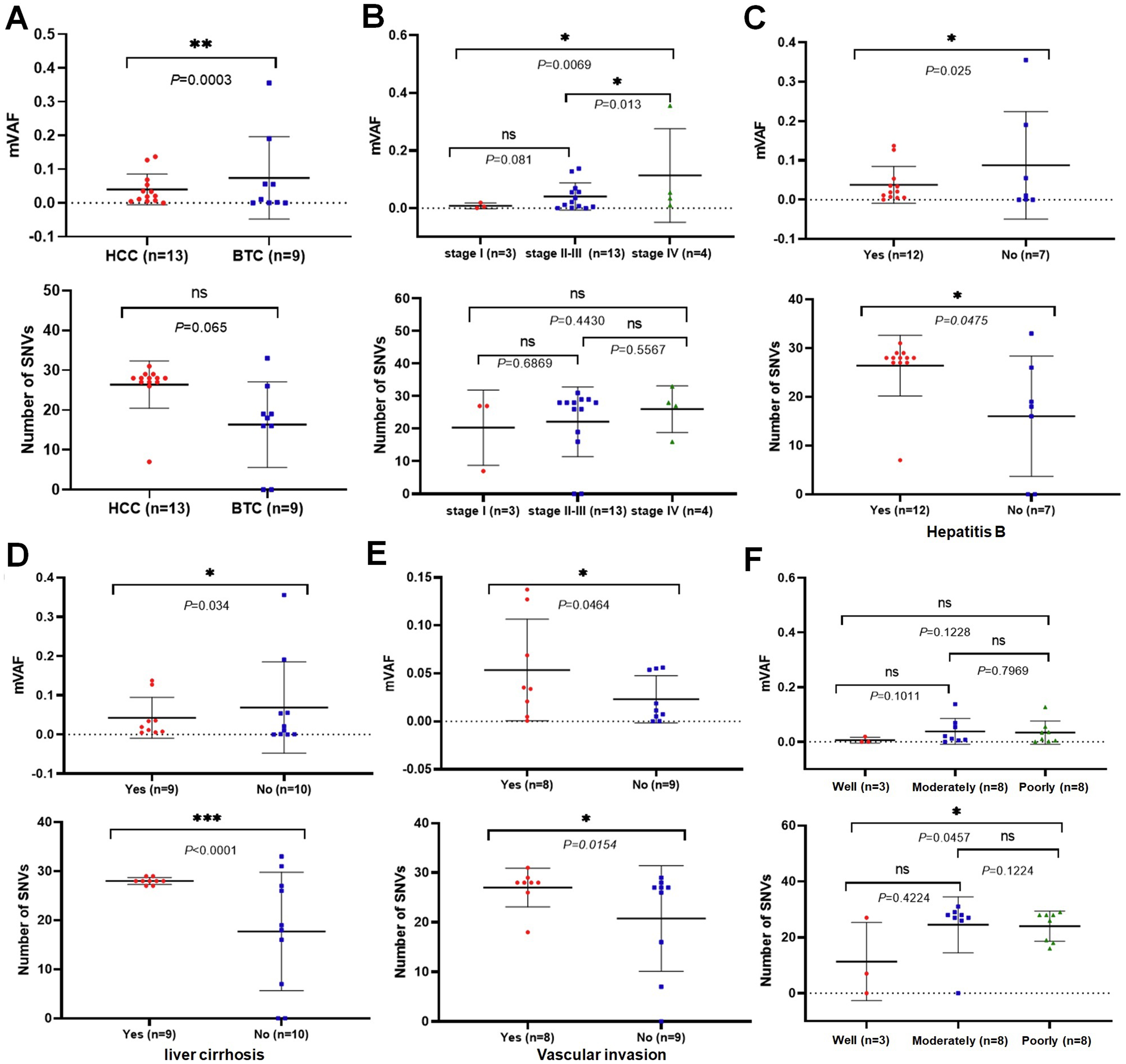
Correlation of baseline ctDNA and clinical parameters. The correlation between baseline mVAF (upper), SNV numbers (lower) and tumor subtype (a), stage (b), HBV (c), liver cirrhosis (d), vascular invasion (e), and tumor differentiation (f) were analyzed separately.
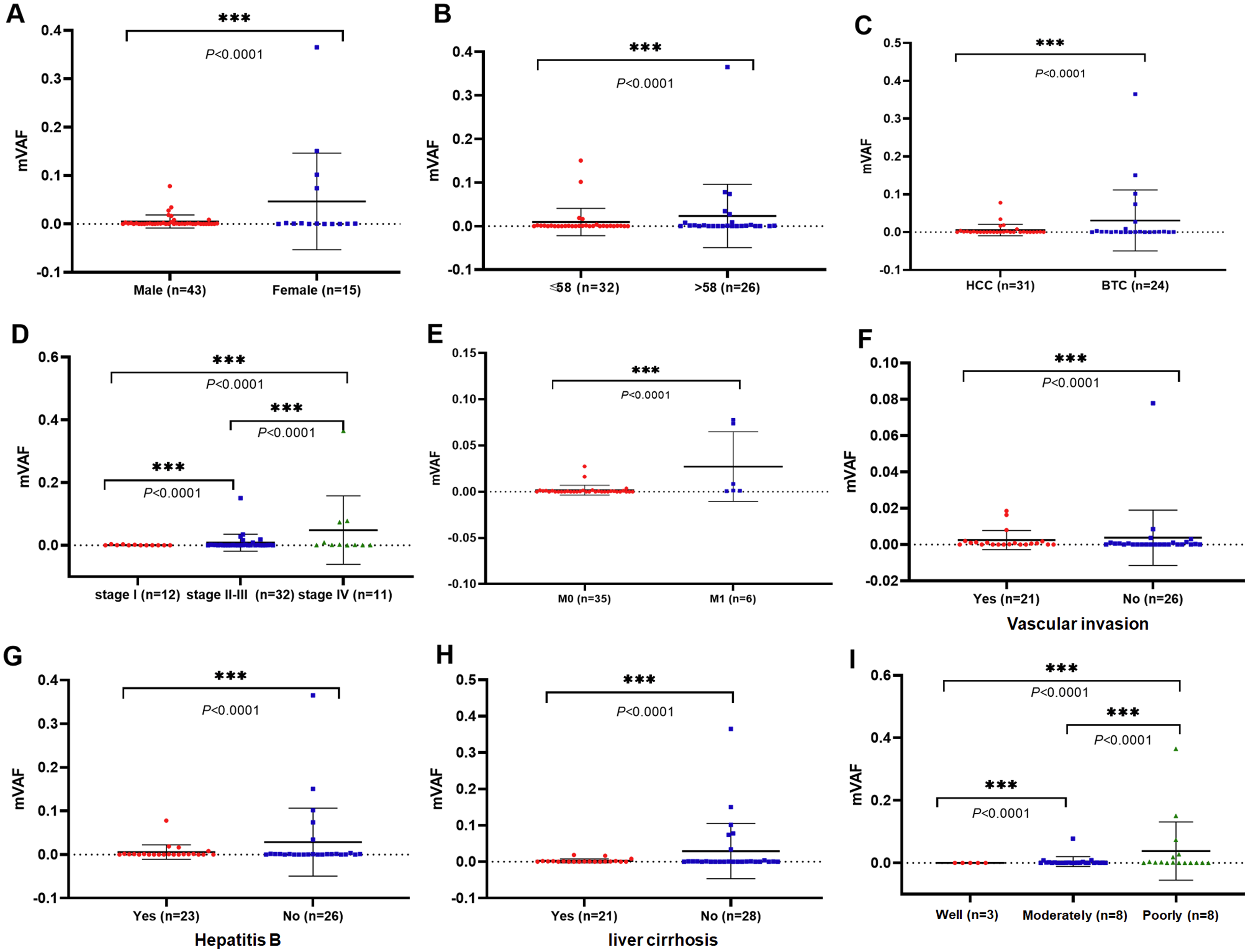
Correlation of postoperative ctDNA and clinical parameters. The correlation between baseline mVAF and gender (a), age (b), tumor subtype (c), stage (d), metastasis (e), vascular invasion (f), HBV (g), liver cirrhosis (h), and tumor differentiation (i) were analyzed separately.
MRD detection guided medication regimen modification to achieve clinical complete response (CR)
A 54-year-old male was admitted to our hospital in January 2021 due to right upper quadrant pain. Imaging showed a large tumor in the right lobe of the liver with significantly elevated AFP. Comprehensive pre-operative treatment with TACE + lenvatinib + treprinumab was administered, and resection of a massive cancerous growth in the liver and ablation of S5 nodules of the liver (local resection) were undertaken in August 2021. Post-operative pathology identified the tumor as HCC. From August to December 2021, the patient received trastuzumab/treprinumab treatment, but MRD dynamic monitoring showed that all three tests were positive (Figure 5(a)). Both AFP and mVAF levels continued to increase, with mVAF peaking earlier than AFP, suggesting that mVAF predicts poor post-operative prognosis and risk of tumor recurrence earlier than AFP (Figure 5(b)). During this period, enlarged pulmonary lymph nodes (lesions too small to be punctured) were detected in November 2021, raising the suspicion of pulmonary metastasis (Figure 5(c)). Based on the MRD results and clinical features, the physician switched the regimen to regorafenib combined with treprinumab in January 2022. After 1 month, the MRD turned negative and AFP levels decreased to 7.41 ng/mL. After 6 months of the new regimen, the MRD remained negative, AFP levels decreased to 5.57 ng/mL, and pulmonary lymphadenopathy disappeared. The physician adjusted the treatment based on the MRD results, and the patient has now achieved a quality life of more than 12 months.
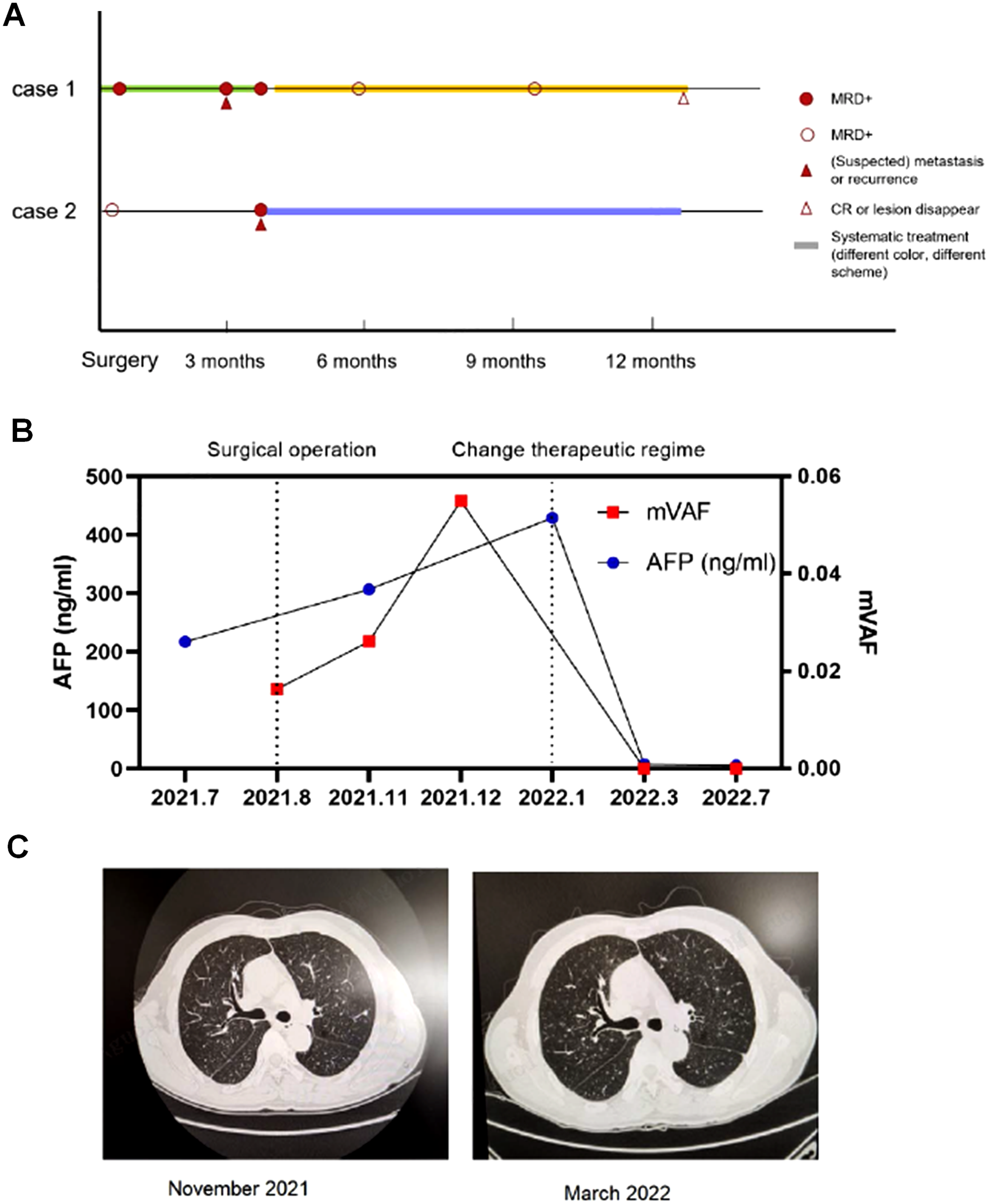
MRD detection guided medication regimen modification. (a) Dynamic changes in ctDNA in two patients from two cases. (b) Dynamic changes in mVAF and AFP. (c) CT scan (November 2021 and March 2022).
Another case involved a 58-year-old female who was admitted in August 2021 due to dull pain in the right upper abdomen for more than 10 days. A computed tomography (CT) scan revealed space-occupying lesions in the S7 segment of the right posterior lobe of the liver and multiple small cysts. The patient underwent right hemi hepatectomy and cholecystectomy Post-operative pathology identified early intrahepatic cholangiocarcinoma (stage I). Post-operative MRD1 was negative, and AFP (3.66 ng/mL) and CA199 were normal (< 2.06 ng/mL). However, in December 2021, post-operative MRD2 testing showed positivity, while AFP (3.32 ng/mL) and CA199 (< 2.06 ng/mL) values remained normal. Further examination showed re-expansion of the right lower lobe, local small nodules, and CT showed pulmonary metastasis (Figure 5(c)). Huaier granule was given for intervention, and the disease was well controlled with no progression for 6 months. This case suggests that MRD can better respond to disease progression, and for some patients with insensitive tumor markers, it can also better indicate distal metastasis.
Discussion
Tumor-informed ctDNA assays have exhibited superior sensitivity and specificity compared to tumor agnostic ctDNA assays, making them useful for detecting low disease volumes such as MRD and early recurrence, as well as monitoring advanced disease. 24 In this retrospective study, we present preliminary findings from the first tumor-informed ctDNA assay, customized according to WES test results, which demonstrated high sensitivity and specificity. We confirmed the applicability and feasibility of a tumor-informed ctDNA assay for MRD evaluation in hepatobiliary cancers. The MRD evaluation guided the disease progression and medication planning in two clinical cases.
NGS methodologies have been extensively utilized in tumor sequencing and have been applied to cfDNA for ctDNA detection. 25 A tumor-informed ctDNA assay first identified 16 tumor unique truncal mutations and subsequently employed a custom multiplex PCR assay to detect these mutations in the plasma of patients for the presence of residual tumor.6,17 The ultradeep multiplex PCR-based NGS ctDNA assay (SignateraTM, Natera, Austin, TX, USA) utilizes the WES of tumors to select 16 high-ranked patient-specific somatic mutations. 22 In our study, the performance of the ctDNA assay was evaluated based on NGS using the WES of tumors to select for 20–40 high-ranked patient-specific somatic mutations. Custom-designed NGS panel is a fast, precise, robust, and affordable way to screen for tumor-related mutations in a set of genes for precision medicine purposes. 26 Tumor tissue from 63 patients were sequenced using WES testing, and the most frequently altered genes were TP53, TERT, TTN, CDKN2A, OBSCN, ARID1A, TSC2, AGO2, MUC16, and SLC45A4. WES identified a total of 1952 SNVs, with only 6 loci (0.3%) shared among at least two patients, which indicates that over 95% of the 20–40 loci screened were unique to each patient. In addition, our analysis revealed high-frequency mutated genes in ctDNA post-surgery, with only 17 gene alterations common to at least two patients, which suggests a large variation in gene mutation between individuals. The MRD surveillance sites vary widely among individuals, which highlights the necessity of personalized customization.
Patients who exhibited a decrease in mVAF (dVAF < 0) after 6 weeks of durvalumab treatment persisted on therapy for a considerable duration compared to those with an increase (dVAF ≥ 0). 27 The reduction in mVAF was strongly associated with tumor response. In addition, patients with decreased mVAF demonstrated significant improvements in progression-free survival (PFS) and overall survival (OS) compared to those with dVAF ≥ 0. Zhang et al. 12 observed that higher pretreatment VAF were associated with poorer OS and other known prognostic factors, but not objective response, suggesting a prognostic role for patient outcomes. Raja et al. 27 previously reported the utility of mVAF, demonstrating that a reduction in mVAF was associated with prolonged PFS and OS in patients with non–small-cell lung cancer (NSCLC) and urothelial cancer treated with durvalumab. Reduced survival has been reported in NSCLC patients with somatic alterations exceeding 5% VAF . 28 In our study, we demonstrated that three patients exhibited a decrease in mVAF (dVAF < 0), transitioning from ctDNA positive to negative status, whereas patients had increased mVAF that continued to be positive. Therefore, changes in mVAF have the potential to aid in making treatment decisions in the early stages. Moreover, it is anticipated that ctDNA dVAF data and imaging data are expected to complement each other in future clinical practice by providing non-redundant characterizations. The assessment of ctDNA dVAF may assist in the detection of pseudo-progressions, particularly in indications where radiographic response assessment is problematic (e.g., pancreas, liver), and may serve as a predictive tool for relapse in an adjuvant setting. They assigned weightage somatic SNVs and indels from the Guardant360 report while calculating the mean of VAF. 12 In our studies, we observed that while the number of SNVs typically increases or decreases in line with the corresponding rise or fall in mVAF, there are exceptions, such as cancer types, HBV infection and cirrhosis where the trend is reversed (Figure 3). In practice, qualitative MRD assessment should be prioritized to judge disease progression and prognosis. When qualitative results are inconclusive, a comprehensive consideration of mVAF and the number of SNVs is necessary. If there is a discrepancy between the two parameters, comprehensive evaluation should be made in combination with clinical features (e.g., tumor markers, imaging, etc.). Serum AFP is the most widely used biomarker of HCC, which can be used for early diagnosis, efficacy, and prognosis evaluation of HCC. 29 Our case study revealed that ctDNA trends were consistent with AFP levels, prompting physicians to adjust therapeutic regimens, which resulted in improved response to disease progression, particularly in patients with insensitive tumor markers. It is noteworthy that in a small number of cases, the decrease in mVAF and objective response did not align in our study, which emphasizes the need for further research to assess whether ctDNA assessment can enhance the precision of clinical response evaluations in such situations. These findings suggest that the mean-change in VAF and SNV numbers may provide additional information for conventional tumor markers. The limitations of this study are primarily related to the fact that only a subset of patients underwent baseline ctDNA testing. Additionally, the frequency of ctDNA testing varied among patients, with some having only a single time point assessed, while others had up to five time points evaluated post-surgery. Previous studies have demonstrated that ctDNA clearance can serve as a surrogate marker for therapeutic efficacy, particularly in adjuvant therapy, and post-operative ctDNA negativity has better results than that in shown in ctDNA-positive patients. 30 We acknowledge that our small dataset limits the strength of our findings, but we did observe a discernible pattern in VAF values and the conversion of ctDNA-positive patients to -negative. In future studies, we aim to incorporate regular monitoring of ctDNA dynamics using serial assays to delve deeper into these findings.
Overall, our findings support the growing importance of ctDNA measurements in clinical practice. Our work provides direct evidence for the predictive and prognostic value of ctDNA, which can help clinicians and researchers design their clinical studies and support therapeutic decisions in the adjuvant setting. Furthermore, serial testing allows for treatment modification, treatment escalation in patients with disease progression, and the opportunity for early therapeutic intervention in patients with positive ctDNA but not yet progressive disease.
Conclusions
Personalized ctDNA-based MRD analysis is an increasingly valuable tool in the clinic. Ongoing prospective interventional studies will further assess the performance of this detection of MRD and help guide treatment decisions.
Supplemental Material
sj-docx-1-jbm-10.1177_03936155251315500 - Supplemental material for Molecular residual disease assessment based on tumor-informed assay predict disease progression and postoperative recurrence of hepatobiliary cancer: A preliminary study
Supplemental material, sj-docx-1-jbm-10.1177_03936155251315500 for Molecular residual disease assessment based on tumor-informed assay predict disease progression and postoperative recurrence of hepatobiliary cancer: A preliminary study by Xiaobing Zhang, Huiguo Shan, Hongyu Pan, Qian Zhong, Qiang Fang, Yun Xu, Yun Liu and Shuping Qu in The International Journal of Biological Markers
Footnotes
Abbreviations
Acknowledgments
We thank OrigiMed Co., Ltd for their technical support.
Author contributions
Methodology, Q.F., X.B.Z., H.G.S., and H.Y.P.; Project administration, S.P.Q. and Y.L.; Resources, X.B. Z., H.G.S., and H.Y.P.; Software: Q.Z.; Formal analysis: Q.F.; Data curation and validation: Y.X.; Writing-original draft, X.B.Z., H.G.S., and H.Y.P.; Writing review and editing, S.P.Q. and Y.L. All authors have read and approved the submitted version of the manuscript. X.B.Z., H.G.S., and H.Y.P. contributed equally to the study.
Data availability statement
The data that support the findings of this study are available upon request from the corresponding author.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Institutional review board statement
This study was approved by the Ethical Committee of Eastern Hepatobiliary Surgery Hospital, Second Military Medical University (Shanghai, China) with reference number of EHBHKY2023-K002-P001.
Informed consent statement
All patients provided written informed consent.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
