Abstract
Background
This study aims to investigate the mutation status and protein expression of low-density lipoprotein receptor-related protein 1B (LRP1B) in endometrial cancer, and analyze its association with lymph node metastasis (LNM) in endometrial cancer.
Methods
Targeted next-generation sequencing (NGS) was conducted on both tumor tissues and paired blood DNA obtained from 94 endometrial cancer patients, followed by comprehensive analysis. Additionally, immunohistochemistry (IHC) was used to explore the correlation between LRP1B protein expression levels, its gene mutation status, and LNM.
Results
LRP1B mutation was observed in 19 patients (20.2%). Our results revealed that LRP1B mutation frequencies were significantly different between endometrial cancer with or without LNM (P = 0.038). Multivariate analysis indicated that LRP1B mutation was a favorable predictor (odds ratio 0.09; 95% confidence interval 0.01–0.95; P = 0.045) for LNM in endometrial cancer. Further analysis revealed that combination of LRP1B mutation with clinical variants (LVSI and histological subtype) yielded a higher area under the curve value of 0.871) and patients harboring LRP1B mutated-type were less likely to develop LNM. On integrated analysis, the concordance between LRP1B NGS and LRP1B IHC was 73.3%.
Conclusions
This study utilizes targeted NGS to uncover the relationship between LRP1B mutation and LNM status, contributing to the development of primary prevention and proactive treatment strategies.
Keywords
Introduction
Endometrial cancer (EC) is the most common gynecologic carcinoma. 1 Most patients are diagnosed with early-stage localized disease and typically have favorable long-term survival rates, with a 5-year survival of 95%, whereas those with recurrent disease tend to exhibit reduced response to therapy and have a worse prognosis. 2 The occurrence of lymph node metastasis (LNM) without proper treatment will result in increasing risk of recurrence and poor prognosis. However, the value of lymph node resection in the treatment of EC has been a subject of controversy, especially considering the low incidence of LNM in low-risk or stage IA patients.3,4
Obturator nodes and para-aortic lymph nodes are the primary sites of LNM in EC, followed by internal iliac nodes, common iliac nodes, and external lymph node metastasis. 5 LNM plays a crucial role in staging the disease and determining treatment options. 6 Early detection and identification of LNM help in formulating more effective treatment strategies to improve survival and prognosis of EC patients. Recent studies have revealed the molecular subtypes and potential therapeutic targets of EC through genomic, transcriptomic, and proteomic multi-omics analysis. Currently, The Cancer Genome Atlas (TCGA) has identified four novel molecular subtypes, including POLE ultramutated, microsatellite instability hypermutated (MSI), copy number low, and copy number high. 7 Despite these subtypes having been shown to effectively predict the prognosis of EC patients, few studies have explored LNM-associated genes. Given the significant role of LNM in the prognosis of EC patients, we collected clinical information and genomic data from 94 EC patients and found that low-density lipoprotein receptor-related protein 1B (LRP1B) mutations are common in EC and closely associated with LNM in EC.
Low-density lipoprotein receptor (LDLR)-related protein 1B (LRP1B) belongs to the LDLR protein family,8–10 consisting of transmembrane and intracellular domains, 9 and shares a highly similar structure with LRP1. 11 LRP1B is widely expressed in normal tissues such as the gastrointestinal tract,12,13 skeletal muscle, 14 ovaries, 15 and cervix, etc. 12 It participates in various biologic processes, including endocytosis, intracellular transport, immune response, cell proliferation, and adhesion, by specifically recognizing extracellular ligands.9,16 A cancer genomic analysis of cBioPortal indicated that LRP1B mutation frequency was higher across cancer cases.17,18 In addition, studies have shown that LRP1B is frequently altered or deleted in various solid tumors, suggesting that LRP1B may act as a putative tumor suppressor.19,20 A multicenter pan-cancer study conducted collaboratively by Duke University, Johns Hopkins University, and the University of Michigan indicated that patients with LRP1B mutations showed higher overall response rates and favorable outcomes (longer progression-free survival (PFS) and overall survival (OS)) after receiving immune checkpoint inhibitors (ICI) therapy, and the conclusion remained consistent after adjusting for microsatellite instability (MSI) and tumor mutational burden. 18 However, the mutation status of LRP1B in EC, its true biological relevance, and whether LRP1B can independently predict LNM in EC remain to be determined.
Herein, we conducted a comprehensive analysis of next-generation sequencing (NGS) in EC to investigate the LRP1B mutation frequency and status, as well as its prognostic significance on LNM in EC. The concordance of genetic mutation patterns and protein expression of LRP1B was analyzed.
Material and methods
Patient selection and clinical specimens
A total of 94 EC patients were enrolled and submitted to the Molecular Detection Center Laboratory of the First Affiliated Hospital of Chongqing Medical University for NGS between July 2019 and June 2022. A retrospective collection of clinical and pathological data from outpatient or inpatient electronic health records, including age at diagnosis, lymphovascular space invasion (LVSI), serum cancer antigen 125 (CA-125) level, depth of myometrial invasion, cervical stromal invasion, histological subtype, TCGA molecular classifications, and LNM.
NGS detection
All formalin-fixed and paraffin-embedded (FFPE) tumor tissues and matched blood samples were validated by two qualified pathologists. DNA was extracted using the QIAamp Genomic DNA Kit (Qiagen, Germany), and sequencing libraries were constructed. Subsequently, genomic DNA was sheared into 150–250 bp fragments, end-repaired, A-tailed, and ligated to adapters. The genomic libraries were hybridized with a specific gene panel (Roche NimbleGen, Madison, WI, USA), which included gene target regions (i.e., selected exons, hotspot regions, and complete coding regions). Then, the captured libraries were pooled and sequenced on the HiSeq2500 NGS platform (Illumina Inc., San Diego, CA, USA). After deleting the low-quality sequencing data, the reads were aligned to the human reference genome and the base quality scores in the sequencing data were corrected.
To further compare and analyze the somatic mutation rate, variant classifications, and the types and quantities of associated mutated genes of LRP1B gene in EC patients, we downloaded the STAR-counts data (data in TPM format), 21 mutation MAF data, and corresponding clinical information of EC patients from TCGA database (https://portal.gdc.cancer.gov). 22 Subsequently, we conducted somatic mutation analysis and visualization using the maftools package based on R software, and normalized using log2(TPM + 1). 23
Tissue microarray preparation and immunohistochemical staining analysis
Fresh tissues were fixed in formalin and embedded in paraffin. Paraffin specimens were marked based on the tumor location identified by hematoxylin and eosin staining of the slices, and tumor tissue regions were selected for sampling and preparation of tissue microarrays. After baking, deparaffinization, hydration, and high-pressure heat retrieval, the samples were blocked at room temperature for 30 min. The blocking solution was removed, and the primary antibody was incubated at room temperature for 2 h. Subsequently, the corresponding secondary antibody was added and incubated at room temperature for 15 min. After gradient alcohol dehydration, xylene transparency, and neutral resin sealing, the samples were observed and photographed under a microscope. The antibodies used in this study included: anti-LRP1B (1:200, HPA069094, Sigma). Five high-power field views (×400) were randomly observed for each sample, and scores were assigned based on the tumor cell positivity rate and staining intensity: (a) Scored according to the cell positivity rate: 0 for no staining; 1 for 6–25%, 2 for 26–50%, 3 for >51%; and (b) scored according to the staining intensity: 0 for no staining, 1 for light yellow, 2 for yellow, 3 for brown. The total score was calculated as the product of the cell positivity rate and staining intensity, with a total score ≥ 4 indicating moderately and significantly positive and a score < 4 indicating negative and weakly positive.
Statistical analyses
All statistical analyses were performed using SPSS software (version 24.0) and R (version 4.3.2). Continuous variables were expressed as mean ± SD, while categorical variables were presented as frequency (%). Differences between groups were evaluated using the Chi-square test or Fisher's exact test. Univariate and multivariate logistic regression analyses were conducted to determine the effects of gene mutation status and clinical variables on LNM, and the corresponding odds ratios (ORs) were calculated. Using two-tailed test, and the P < 0.05 was considered statistically significant.
Results
The frequency, types, and correlation of LRP1B mutation with TCGA molecular subtypes
Based on the TCGA database, we found that LRP1B mutations are frequently detected in various cancers, accounting for 45.1% (211/468) in skin cutaneous melanoma, 35.1% (170/485) in lung squamous cell carcinoma, 32.9% (170/517) in lung adenocarcinoma, 26.7% (117/439) in stomach adenocarcinoma, and 18.6% (99/531) in uterine corpus endometrial carcinoma (Figure 1(a)). Additionally, detailed genomic features of LRP1B mutation in EC can be found in Supplementary Figure 1.
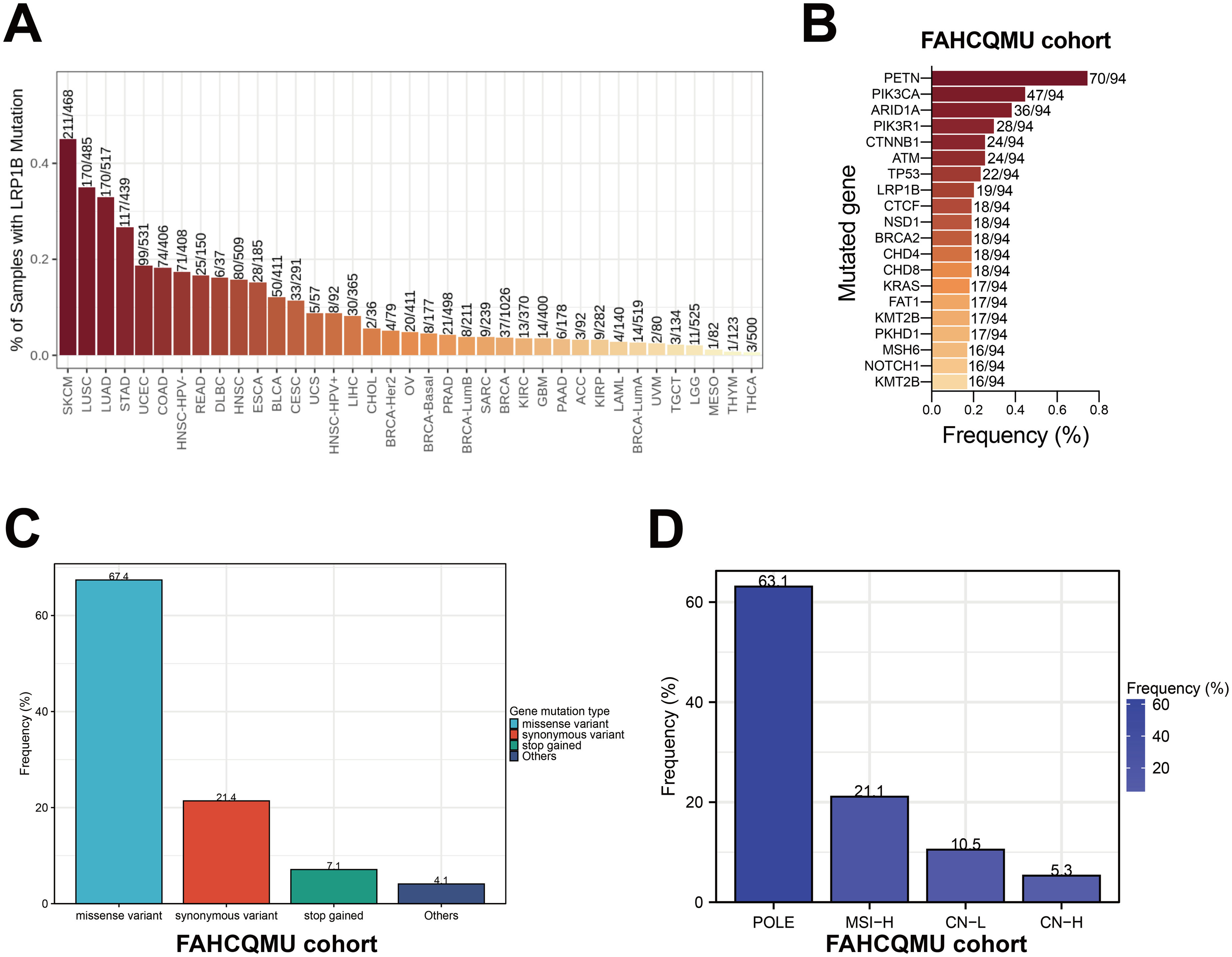
Frequencies and distributions of LRP1B aberrations. (a) Mutation frequency of LRP1B in pan-cancer based on TCGA database; (b) the gene mutation frequency in the FAHCQMU patient cohort; (c) LRP1B gene mutation types and frequency; and (d) proportion of TCGA molecular classifications in EC with LRP1B mutations (%).
Subsequently, through comprehensive analysis of NGS data from 94 Chinese EC cases in our cohort, we found that LRP1B is one of the most mutated genes in EC, accounting for 20.2% of all patients who underwent genomic testing (19 out of 94 cases) (Figure 1(b)). Additionally, the genetic profile revealed that missense variants were the primary type among all LRP1B mutations, representing 67.4% of all mutation types; followed by synonymous variants at 21.4%. Less common mutation types included frameshift variants, splice donor variants, stop gained mutations, and combined mutations (missense variant and splice region variant) (Figure 1(c)). According to TCGA molecular classification, we observed that among EC patients with LRP1B mutations, the highest proportion was in the POLEmut subtype (63.1%), followed by the microsatellite instability-high subtype (21.1%) (Figure 1(d)).
Demographic and clinicopathological characteristics of EC patients with LRP1B mutations
To further investigate the impact of LRP1B gene mutations on clinicopathological characteristics in EC patients, we collected and analyzed detailed clinical and pathological data from these patients. Chi-square tests revealed that the LRP1B mutation was significantly correlated with LNM (P = 0.038). However, there were no statistically significant differences in other clinical variables between the two groups (Supplementary Table 1).
LRP1B mutation is associated with LNM in EC patients
Previous studies have shown that LRP1B mutations are common in cancer tissues and are significant for clinical diagnosis, prognosis assessment, and treatment selection for the disease. However, the relationship between the mutation status of LRP1B and LNM remains unclear. Therefore, we further analyzed the relationship between LRP1B mutations and LNM in EC. The results showed that the rate of LNM was significantly lower in the LRP1B mutated-type group compared to the LRP1B wild-type group (P = 0.038, Figure 2(b)). Additionally, among EC patients with LNM, missense mutations were predominantly observed in LRP1B mutation carriers, whereas both missense and synonymous mutations were more prevalent in non-lymph node metastatic patients (Supplementary Table 2).
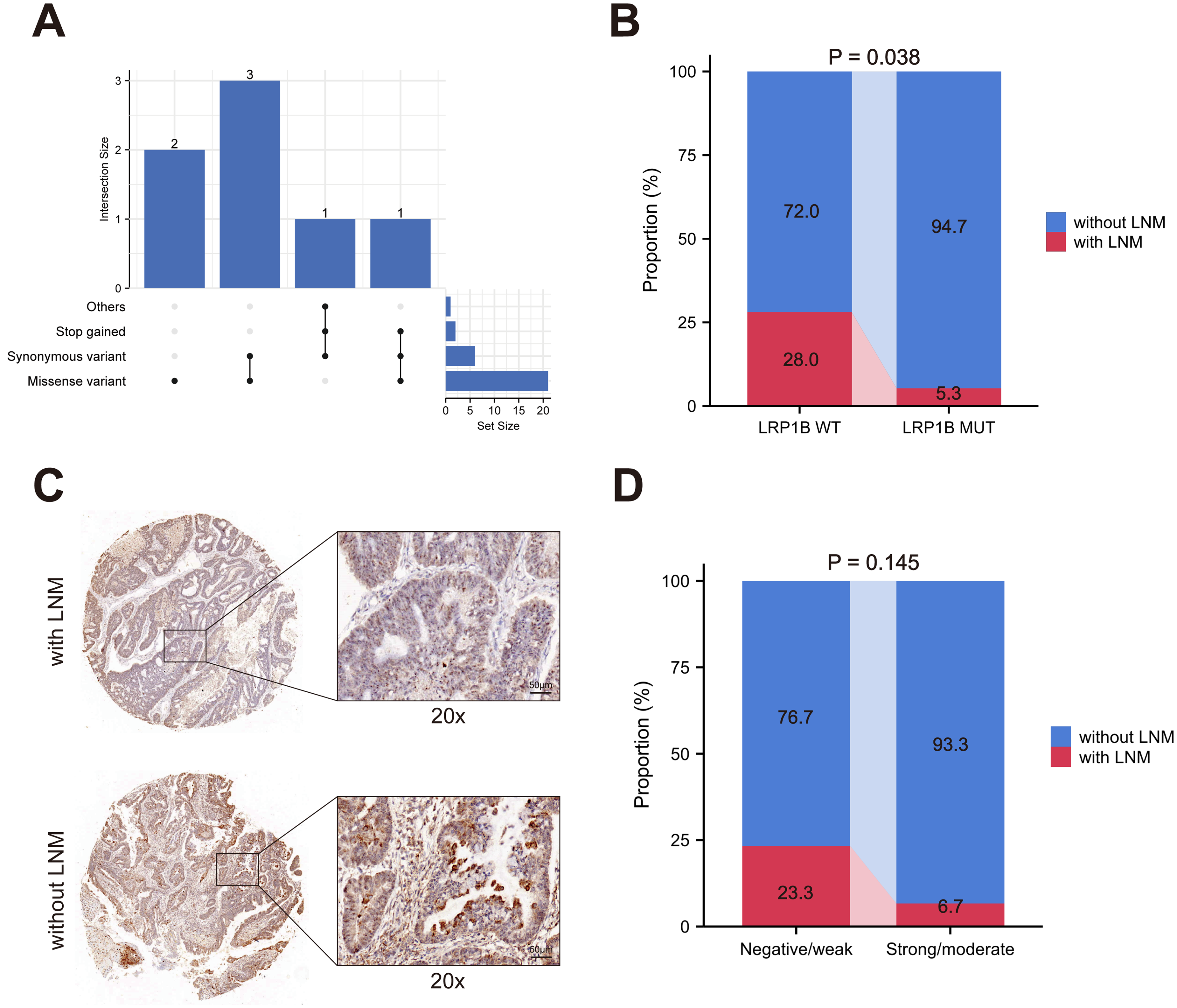
Relationship between LRP1B and LNM in EC. (a) Consistency of IHC LRP1B expression with LRP1B mutations; (b) distribution of LRP1B mutations in lymph node metastatic and non-lymph node metastatic EC patients in NGS; (c) representative images of tissue microarray; and (d) distribution of LRP1B protein expression in lymph node metastatic and non-lymph node metastatic EC patients in IHC.
Consistency between LRP1B expression and LRP1B mutations
We further conducted immunohistochemical (IHC) staining on 15 mutated LRP1B EC cases and 15 cases with LRP1B wild-type to analyze the relationship between LRP1B mutation status and LRP1B protein expression (Figure 2(a) to (c)). The results revealed a high consistency rate of 73.3% between LRP1B protein expression and LRP1B mutation status. In LRP1B mutated-type EC tissues, 73.3% of tumor samples showed positive LRP1B protein expression, while among the tumors with LRP1B positive by IHC, 69.2% of tumor samples harbored LRP1B missense mutations (Supplementary Figure 2). As expected, patients with LRP1B protein overexpression exhibited a similar predictive effect on the rate of LNM, although the difference was not statistically significant (Figure 2(d)). These findings indicated a significant level of consistency between LRP1B protein expression and LRP1B mutation status.
Clinical value of LRP1B mutation status in predicting LNM and prognosis in EC patients
As demonstrated above, LRP1B mutation is strongly associated with LNM in EC. However, it remains unclear whether LRP1B mutation is a concomitant effect of known clinical high-risk factors for LNM rather than an independent effect contributing to LNM. To address this issue, we performed univariate and multivariate logistic analysis to identify the independent risk factors for LNM, the results indicated that histological type II (OR = 5.24; 95% confidence interval (CI) 1.38–20.05) and positive LVSI (OR = 5.55; 95% CI 1.23–24.66) were independent risk factors for LNM in EC. Conversely, LRP1B mutated-type (OR = 0.09, 95% CI = 0.01–0.95) considered as an independent protective factor for LNM in EC (all P < 0.05) (Table 1). Further analysis revealed that combination of LRP1B mutation with clinical variants (LVSI and histological subtype) yielded a higher area under the curve (AUC) value of 0.871 (Figure 3).
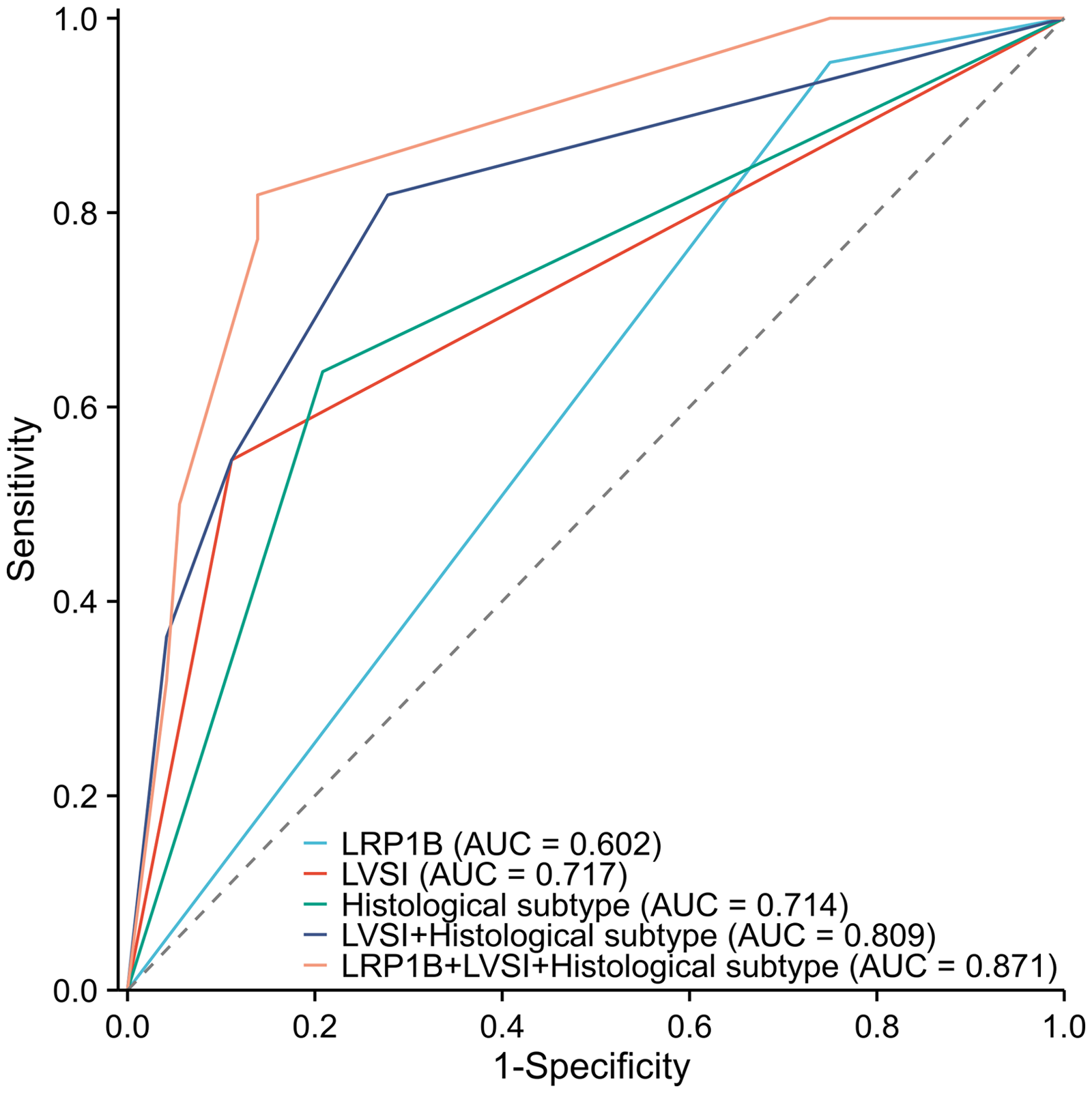
ROC analysis of LNM in different groups.
Univariate and multivariate logistic analysis of risk factors associated with LNM in EC.
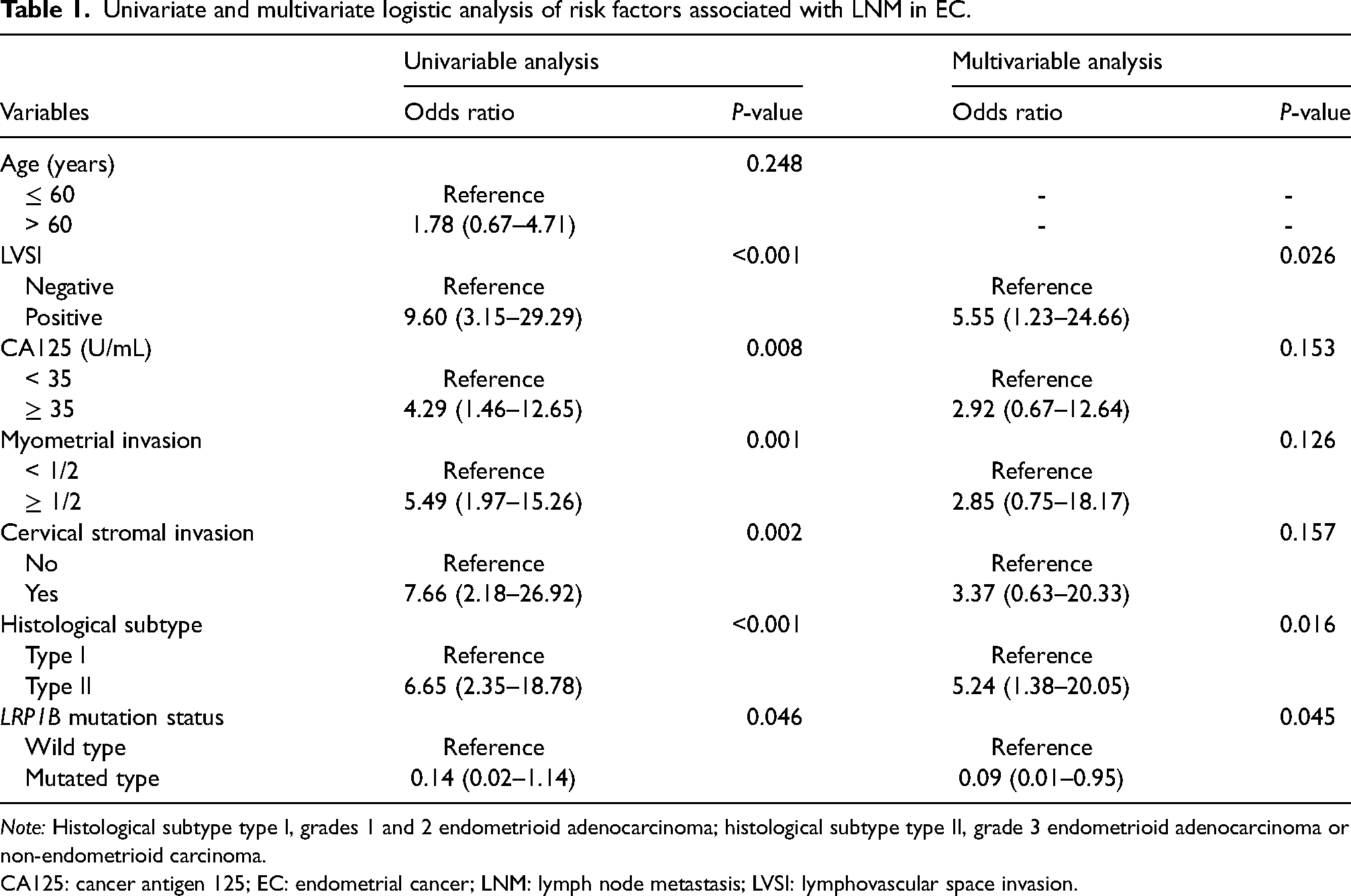
Note: Histological subtype type I, grades 1 and 2 endometrioid adenocarcinoma; histological subtype type II, grade 3 endometrioid adenocarcinoma or non-endometrioid carcinoma.
CA125: cancer antigen 125; EC: endometrial cancer; LNM: lymph node metastasis; LVSI: lymphovascular space invasion.
Furthermore, survival analysis based on TCGA database demonstrated that EC patients harboring LRP1B mutated-type had favorable OS and recurrence-free survival (RFS) than those with LRP1B wild-type (hazard ratio (HR) = 0.38, 95% CI 0.19–0.75, P = 0.0038; and HR = 0.4, 95% CI 0.17–0.93, P = 0.027) (Figure 4).
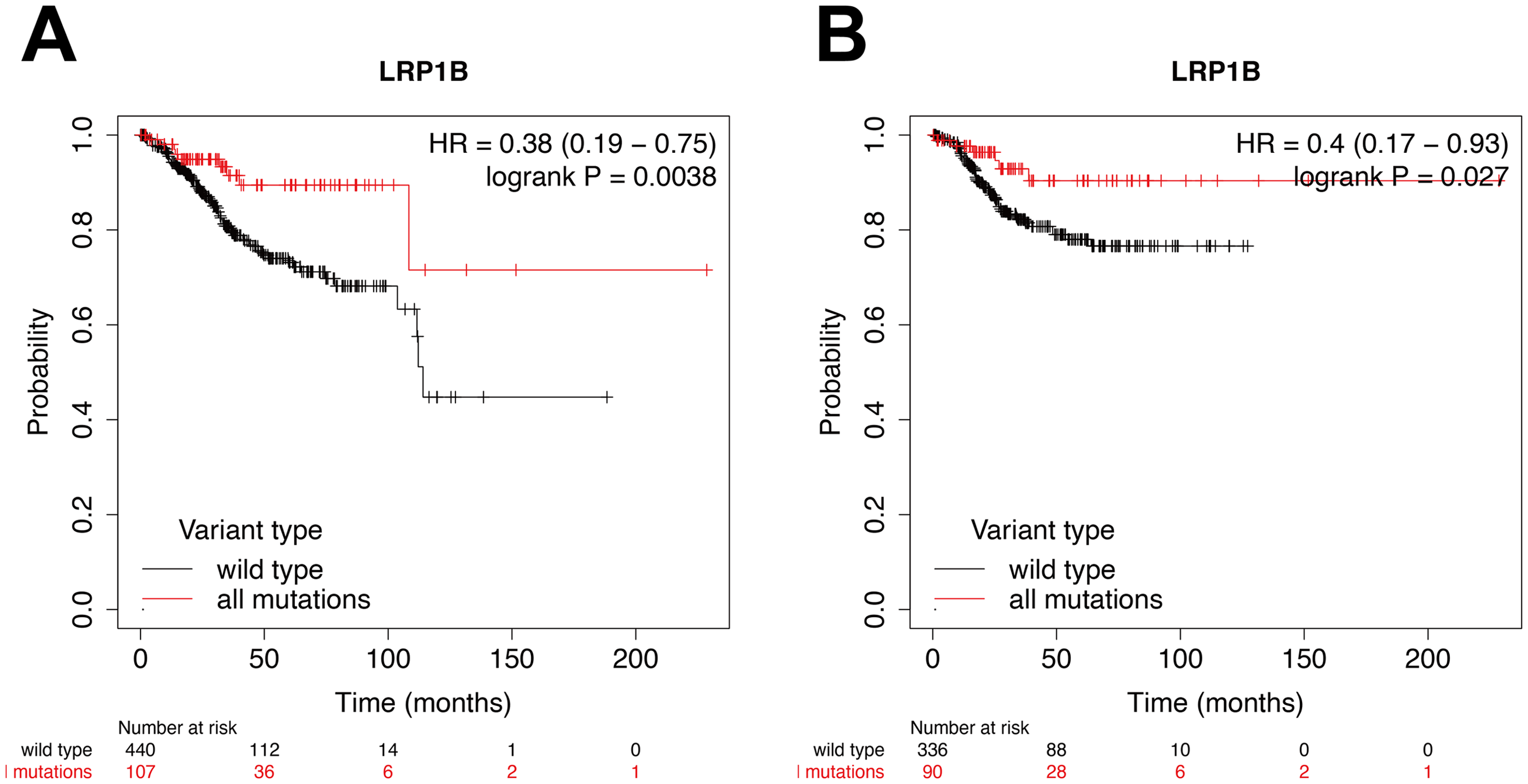
Impact of LRP1B mutation status on OS (a) and RFS (b) in EC patients from K–M plotter. OS: mutation group, n = 107; wild-type group, n = 440; Log rank P = 0.0038. RFS: mutation group, n = 90; wild-type group, n = 336. Log rank P = 0.027.
Collectively, these findings indicated that LRP1B mutation played an essential role in driving tumorigenesis and serves as an independent protective factor for LNM in EC.
Patient harboring LRP1B mutation treated with anti-PD-1 camrelizumab plus bevacizumab (targets vascular endothelial growth factor)
A 52-year-old female with no remarkable family history of cancer presented at an external hospital in March 2021 with postmenopausal vaginal bleeding. Upon hysteroscopic examination, she was diagnosed with EC. In April 2021, she came to our institute and underwent complete staging surgery, including extended total hysterectomy, bilateral salpingo-oophorectomy, pelvic and para-aortic lymphadenectomy, omentectomy, pelvic adhesiolysis, and intestinal adhesiolysis as primary treatment. The postoperative pathology revealed low-differentiated endometrioid adenocarcinoma (FIGO stage IIIC2) with invasion of the outer 1/2 layer of the uterine wall and tumor emboli in blood vessels (substantial LVSI). No cancerous tissue was found in the parametrium or greater omentum, and no obvious abnormalities were observed in the bilateral adnexa. LNM was detected in 1 of 30 pelvic lymph nodes and 1 of 12 para-aortic lymph nodes. The tumor exhibited deficient mismatch repair with complete loss of PMS2 protein expression and programmed death-ligand 1 (PD-L1) staining is > 1% (according to the tumor proportion score). Since April 2021, the patient has received three cycles of adjuvant chemotherapy with carboplatin (AUC6 at 3-week intervals) and paclitaxel (175 mg/m2), along with adjuvant radiotherapy to the pelvic and para-aortic lymph node regions (total radiation dose of 50.8 Gy, administered in 25 fractions of 2 Gy). In September, 2021, magnetic resonance imaging revealed multiple lymph nodes around the para-aorta with high signal intensity on diffusion-weighted imaging sequences, indicating tumor progression (LNM). The Patient was identified as MSI-High and harboring a missense variant in LRP1B gene (c.517C > T, p.H173Y). Subsequently, she commenced treatment with the anti-PD-1 camrelizumab (200 mg/dose, intravenously once every 2 weeks) in combination with bevacizumab (400 mg/dose, 5 mg/kg/every 2 weeks) in September 2021. After two cycles of immunotherapy, the para-aortic lymph nodes metastases decreased in size with no obvious enhancement (Supplementary Figure 3). Throughout the entire treatment process, no adverse events of grade 3 or higher were observed (according to the Common Terminology Criteria for Adverse Events version 5, by the National Cancer Institute). 24
Discussion
In this study, we analyzed the genomic signatures of 94 Chinese EC patients through NGS technologies and investigated their correlation with clinical pathology, uncovering potential etiologies and providing novel insights into the diagnosis and therapy strategies for EC. Intriguingly, systematic molecular profiling revealed that frequent mutations in LRP1B accounted for 20.2% of all gene alterations, slightly higher than the mutation rate of LRP1B (18.6%) in the TCGA database. LRP1B, acting as a tumor suppressor, is frequently altered in various types of cancer. 18 Previous studies have reported frequent inactivation mutations of LRP1B in multiple cancers, with a mutation frequency of 21.34% in lung adenocarcinoma 25 and 8% in hepatocellular carcinoma. 26 Additionally, LRP1B has been described as a common target gene for viral integration and has been observed in hepatitis B virus. Furthermore, LRP1B has been detected as a human papillomavirus integration hotspot in cervical carcinogenesis. 27 Therefore, exploring the role of LRP1B in the development and prognosis of EC is of significant clinical importance.
Furthermore, our study found that missense mutation was the most frequent mutation in LRP1B gene. Missense mutation was a specific type of point mutation caused by a single base change, resulting in alterations to the corresponding codon and encoded amino acid, ultimately affecting the structure and function of the protein. LRP1B, as an LDL receptor, plays a wide variety of roles in normal cellular function and cancer development. 11 On the basis of TCGA dataset, a recent study has found co-mutation of LRP1B and FAT3 happened in 11.16% EC patients. Importantly, the co-mutations of LRP1B and FAT3 not only activate the immune status of EC, but indicate an improvement in prognosis (prolonged PFS and OS). 28 However, there is currently no evidence to prove whether EC patients with mutations in either LRP1B or FAT3 alone can benefit from immunotherapy, and the exact relationship between LRP1B mutation status and LNM in EC remains unclear.
Due to the limited understanding of association between molecular characteristics and LNM in EC, few studies have explored effective biomarkers related to LNM in EC. To our knowledge, this is the first study describing the association between mutation status of LRP1B and LNM in EC patients. Compared to LRP1B wild-type, patients with LRP1B mutations have a lower LNM rate (LRP1B-MT vs. LRP1B-WT: 5.3% vs. 28.0%), and this association remained significant after adjusting for multiple variants such as age, LVSI, cervical invasion, and histological subtype. Further analysis using receiver operating characteristic curves revealed that the combination of LRP1B mutation status and clinicopathological features showed a strong predictive ability for LNM in EC patients with an AUC value of 0.871, surpassing multiple predictive indicators. Together, these findings suggest that LRP1B mutation serves as an independent protective factor for LNM. When integrated with traditional clinicopathological parameters, it enhances the effectiveness of predicting LNM in EC. In the past, numerous studies have confirmed the significant role of LRP1B in the occurrence and progression of cancer. The study by Ikari et al. 29 found that LRP1B is one of the most prominent somatic genes in liver cancer metastasis. In a report by Han et al., 30 genomic profiling was performed on tissue biopsies collected from 199 patients with esophagogastric junction adenocarcinoma. Notably, CDK6, LRP1B, MET, and NOTCH1 mutations were shown to significantly associate with LNM, in particular with LRP1B mutation frequency being significantly higher in patients without LNM compared to those with LNM (16% vs. 6%), which were consistent with our conclusions. Likewise, in a similar research on the association of LRP1B mutations with immunotherapy outcomes in lung adenocarcinoma, it was found that patients with LRP1B-mutated lung adenocarcinoma exhibited a significant prolongation of PFS after receiving ICIs treatment. 31 Additionally, our findings demonstrated a good concordance of 77.3% between the LRP1B mutation and IHC, and LRP1B protein expression was mainly localized to the cell membrane of the epithelial cells and endometrial glands; however, distinct positive staining was also observed in the cytoplasm. This was detected to be lower expression in EC tissues compared with adjacent normal tissues from individuals (Supplementary Figure 4). As expected, patients with LRP1B protein overexpression had a similar predicted effect on LNM rate. This suggests that LRP1B IHC staining score could serve as a potential biomarker for predicting LNM in EC.
Clinical management of EC is transitioning from histological subtype to molecular stratification, and the finding of novel biomarkers is essential for understanding and improving outcomes of EC. Survival analysis based on the TCGA database reveals that patients with LRP1B mutations have longer OS and RFS. Notably, molecular classification analysis indicates a significant enrichment of the POLEmut and MSI-H subtypes in patients with LRP1B mutations, which are frequently associated with favorable survival outcomes. This could be one of the reasons contributing to the prolonged survival in patients harboring LRP1B mutations. Nonetheless, the reasons behind how the wild-type LRP1B gene underpins the occurrence of LNM in these malignancies remain nebulous. Given the significantly increased risk of LNM and poor prognosis in patients with LRP1B wild-type compared to those with LPR1B mutations, close follow-up and active surveillance are recommended. In recent years, ICIs have shown promising prospects in cancer treatment, with some studies confirming the effectiveness of anti-PD-1 therapies in cancer patients with LRP1B mutations.25,31,32 Due to the relatively recent application of immunotherapy in gynecological cancer, the specific population for the application of ICIs in EC remains unclear. The limited evidence in the present study indicated LRP1B mutations might be a promising candidate biomarker for response to anti-PD-1 therapy among EC patients. Additionally, the development of more effective therapies for EC patients harboring LRP1B wild-type holds significant clinical importance.
While the present study conducted in-depth sequencing analysis, there are still inherent limitations. Firstly, since long-term follow-up data are not available for the cohort at this time, a survival analysis of EC patients could not be performed. Secondly, despite validating the genomic analysis of FFPE samples in this study to ensure the accuracy of our findings, the method of formalin fixation and paraffin embedding may still lead to DNA degradation and modification compared to flash-frozen tissue. Finally, the sequencing data obtained in the present study are derived from a single center, so the observed results will need further validation in independent, multicenter clinical studies. Nevertheless, we still hope that this study can provide valuable insights into the genetic mutations of EC and essential information for understanding, predicting, and making treatment decisions for LNM in EC.
Conclusion
This study utilizes targeted NGS to determine the predictive role of LRP1B mutation status in LNM of EC, and to clarify the consistency between LRP1B mutation and its protein expression. Through the evaluation of LRP1B mutation status and expression levels, high-risk individuals for LNM can be identified, leading to improved monitoring and tailored cancer-directed therapies. Additionally, larger datasets are required to validate these findings.
Supplemental Material
sj-docx-1-jbm-10.1177_03936155241304433 - Supplemental material for LRP1B mutation is associated with lymph node metastasis in endometrial carcinoma: A clinical next-generation sequencing study
Supplemental material, sj-docx-1-jbm-10.1177_03936155241304433 for LRP1B mutation is associated with lymph node metastasis in endometrial carcinoma: A clinical next-generation sequencing study by Yunfeng Zheng, Fan Yang and Jie Wu in The International Journal of Biological Markers
Footnotes
Acknowledgments
The authors acknowledge the support of the Molecular Medicine Testing Center (Department of Clinical Pathology and Medical Laboratory) at the First Affiliated Hospital of Chongqing Medical University.
Author contributions
Yunfeng Zheng: conceptualization, formal analysis, investigation, methodology, software, supervision, validation, writing—original draft, and writing—review and editing.
Fan Yang: conceptualization, formal analysis, investigation, methodology, validation, writing—revision, and editing.
Jie Wu: project administration, conceptualization, methodology, writing—revision and editing.
Consent for publication
All patients consented to the publication of the results of this study.
Data availability statement
The data can be obtained on request from the corresponding author, and not publicly available due to privacy or ethical restrictions.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethics statement
This study complied with the principles of the Declaration of Helsinki and was approved by the Ethics Committee of the First Affiliated Hospital of Chongqing Medical University (K2024-012-233). Meanwhile, the participants provided their written informed consent to participate in this study.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
