Abstract
Background
Colorectal cancer (CRC) is often diagnosed late and has a poor prognosis. Circular RNAs (circRNAs) have been identified as prognostic biomarkers in various cancers, including CRC.
Objective
The objective was to elucidate the role of circLPHN3 (hsa_circ_0069865) in CRC progression and to provide a promising prognostic marker for CRC.
Methods
CircLPHN3 was identified through bioinformatics analysis of the GSE121842 dataset. The levels of circLPHN3 in CRC samples were analyzed by real time-quantitative polymerase chain reaction. Its clinical significance was assessed using the Kaplan–Meier curve and multivariate Cox regression. Downstream microRNAs of circLPHN3 were predicted with the RNAhybrid, Circular RNA Interactome, and starBase online databases. The target of miR-142-5p was predicted using miRDB, TargetScanHuman, starBase, and miRWalk databases. The relationship between circLPHN3, miR-142-5p, and LDB2 was verified by dual luciferase reporter assay. The function of circLPHN3 on CRC cell growth and metastasis was measured using Transwell and the cell counting kit-8 assay.
Results
Significant downregulation of circLPHN3 was found in CRC. CircLPHN3 was closely related to higher tumor node metastasis stage, lymph node metastasis, and predicted unfavorable prognosis. miR-142-5p was highly expressed in CRC and its expression was negatively regulated by circLPHN3. Overexpression of circLPHN3 curbed CRC cell growth, migration, and invasion, mediated by miR-142-5p. Moreover, LDB2 was identified as a target of miR-142-5p.
Conclusion
CircLPHN3 acted as a prognostic biomarker and tumor suppressor for CRC via modulating miR-142-5p.
Introduction
Colorectal cancer (CRC) is a common tumor of the digestive tract, ranking third in incidence and second in mortality worldwide, after lung cancer. 1 In the latest survey, there were over 1.93 million new CRC cases in 2020, with a death toll of 935,000. 2 In China, due to an aging population and increasing risk factors, the incidence and mortality of CRC have risen, making it the second most frequent cancer. 3 Despite advances in surgical and chemoradiotherapy techniques, the high rate of CRC metastasis, drug resistance, and side effects from chemoradiotherapy continue to threaten patients’ survival and quality of life.4–6 Therefore, monitoring CRC progression and finding effective therapeutic targets are crucial for the treatment and prognosis of CRC patients.
Accumulating evidence indicates that non-coding RNAs, including circular RNAs (circRNAs), microRNAs (miRNAs), and long non-coding RNAs (lncRNAs), are critical regulators of biological processes and are closely associated with disease progression in malignant tumors.7,8 CircRNAs are covalent, closed-loop, single-stranded RNAs that are ubiquitous in mammals and bind directly to specific miRNAs to regulate their activity. 9 CircRNAs have been shown to act as biomarkers of CRC in several reports and exert an inhibitory effect on the development of CRC.10,11 For example, circ_0000826 was reported to be markedly downregulated in CRC tissues and to inhibit CRC cell growth, suggesting its potential for CRC treatment. 12 Additionally, several circRNAs have been implicated in CRC development in other studies.13–15 Furthermore, these CRC-related circRNAs are often closely associated with the prognostic status of patients. 16 Previously, researchers constructed a circRNA expression profile (GSE121842) in CRC based on RNA deep sequencing and identified circLPHN3 (hsa_circ_0069865) as a down-regulated circRNA. 17 Circ_0069865 is one of the circRNAs derived from the LPHN3 gene, and its function in CRC has not been reported in other studies. CircLPHN3 (mmu_circ_0001358) has been explored in a mouse model of traumatic brain injury, where it bound to miR-185-5p and protected the blood–brain barrier. 18 Whether circLPHN3 affects CRC development and the downstream miRNAs it regulates in CRC is currently unknown.
CircLPHN3 was hypothesized to act as a biomarker of CRC progression. Through a series of in vitro experiments and clinical analyses, the biological role of circLPHN3 and its clinical significance in CRC was elucidated. This study may provide a promising prognostic marker and therapeutic target for CRC.
Materials and methods
Study subjects
Cancer and paracancer tissue samples were collected from 113 CRC patients (69 males and 44 females) who underwent surgery at Shanghai Baoshan Luodian Hospital from April 2016 to March 2018 and were preserved at −80°C. Inclusion criteria: (a) Patients had been diagnosed with CRC before surgery and confirmed with CRC by postoperative histopathologic examination; (b) patients’ clinical data (gender, age, tumor node metastasis (TNM) stage, etc.) were complete; (c) patients did not undergo chemotherapy, radiotherapy, or other therapeutic measures before diagnosis. Exclusion criteria: (a) patients suffered from blood diseases or serious heart, lung, and liver diseases; (b) patients died during the perioperative period; (c) patients had undergone neoadjuvant therapy before surgery. A 5-year follow-up was conducted after surgery to investigate the patients’ survival status. Consent for this study was obtained from all patients or informants, and approval was received from the Ethics Committee of Shanghai Baoshan Luodian Hospital.
Cell lines
FHC (human colorectal mucosa cell) and CRC cell lines (LoVo, Caco2, HCT8, SW480) were obtained from Oricellbio (Guangzhou, China). DMEM/F12 was prepared for culturing FHC cells, and DMEM containing 10% fetal bovine serum (FBS) and 1% penicillin/streptomycin (Gibco/ThermoFisher Scientific, Lenexa, KS, USA) was used for the other several cells. The culture conditions were set at 37°C with 5% CO2.
Cell transfection
For transient transfection, miR-142-5p mimic or inhibitor, and their negative control (miR-NC) were bought from RiboBio (Guangzhou, China). The overexpression plasmid pCD5-circLPHN3 (oe-circ) and pCD5-ciR vector (oe-NC) were acquired from Warbio (Nanjing, China). These plasmids and/or oligonucleotides were introduced into LoVo and SW480 cells using Lipofectamine 3000 reagent (Invitrogen/ThermoFisher Scientific, Carlsbad, CA, USA) once the cells reached 70% confluence. After 48 h of transfection, cell growth continued, with cells harvested at the times required for different experiments.
RNA extraction
Cells were lysed by adding TRIzol reagent (Invitrogen) directly. CRC tissues were sheared and fully ground with a small amount of liquid nitrogen, then TRIzol reagent was added, mixed well, and left on ice for 10 min. The mixtures were treated with chloroform, left on ice for 5 min, and then centrifuged. The supernatant was mixed with isopropanol, left on ice for 10 min, and then centrifuged to obtain RNA. The RNA was washed with 70% ethanol, and total RNA was obtained by removing the ethanol and adding RNase-free water. The RNA concentration was measured using a Nanodrop2000 and recorded. The RNA was stored at −80°C.
Real time-quantitative polymerase chain reaction
Extracted RNA was reverse-transcribed to obtain cDNA using the reaction system and conditions provided in the RevertAid RT kit (ThermoFisher Scientific, Waltham, MA, USA). Relative quantitative analysis of circLPHN3, miR-142-5p, and LDB2 mRNA levels was performed using GoTaq qPCR Master Mix (Promega, Madison, WI, USA) on the LightCycler 480 real time-quantitative polymerase chain reaction (RT-qPCR) instrument (Roche, Basel, Switzerland). The data were calculated using the 2−ΔΔCt method. The levels of circLPHN3 and LDB2 mRNA were normalized by GAPDH, and the levels of miR-142-5p were normalized by U6.
Bioinformatics analysis
RNA-seq data from tumor tissues and adjacent normal colorectal tissues of three CRC patients were downloaded from the GEO database (GSE121842 dataset, https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE121842). Volcano plot and cluster heatmap were then created using the Bioinformatics online platform (https://www.bioinformatics.com.cn/). Downstream miRNAs of circLPHN3 were predicted using the online databases RNAhybrid (https://bibiserv.cebitec.uni-bielefeld.de/rnahybrid/submission.html/), Circular RNA Interactome (https://circinteractome.nia.nih.gov), and starBase (https://rnasysu.com/encori/agoClipRNA.php?source=circRNA). Target genes for miR-142-5p were predicted using the databases miRDB (https://mirdb.org/cgi-bin/search.cgi), TargetScanHuman (https://www.targetscan.org/vert_72/), starBase (https://rnasysu.com/encori/agoClipRNA.php?source=mRNA), and miRWalk (http://mirwalk.umm.uni-heidelberg.de/). The predicted results were visualized with Venn diagrams using the Bioinformatics online platform. The binding sites of circLPHN3 and miR-142-5p were predicted by the starBase database. The binding sites of miR-142-5p and LDB2 were predicted by the TargetScanHuman database. The expression of LDB2 in CRC and normal samples was analyzed using the GEPIA database (http://gepia.cancer-pku.cn/).
Cell counting kit-8 assay
The transfected cells were digested and resuspended at 2 × 103/well, then inoculated in 96-well plates and cultured. After 0, 24, 48, and 72 h of incubation, 10 μL of cell counting kit-8 (CCK-8) solution (Beyotime, Shanghai, China) was added, and the cells were incubated for 30 min. The OD450 value was measured using the Synergy H1 Full-Featured Enzyme Labeler (BioTek, Winooski, VT, USA) to assess cell viability.
Transwell assay
Cells were collected after transfection and resuspended in FBS-free DMEM. The cell suspension was seeded into the upper chamber of the Transwell coated with matrix gel (not used in the migration assay), and the complete culture medium was added to the lower chamber. After 24 h of incubation, the migrated and invaded cells were fixed with 4% paraformaldehyde and stained with 0.1% crystal violet. The number of migrating/invasive cells was counted under a BX53 microscope (Olympus, Tokyo, Japan) in several random fields, and the mean cell number was recorded as the number of migrating/invasive cells.
Dual-luciferase reporter assay
The luciferase reporter plasmids circLPHN3-wild type (WT), circLPHN3-mutant (MUT), LDB2-WT, and LDB2-MUT were constructed. LoVo and SW480 cells were inoculated in 24-well plates. When the cell confluence was 60%, the luciferase reporter vectors were co-transfected with miR-142-5p mimic, inhibitor, or negative control (miR-NC). Cells were lysed after 48 h of incubation and assayed using the Dual-Luciferase Reporter Gene Assay Kit (Promega).
Statistical analysis
The significance of the difference between the data from the control and experimental groups was analyzed using Student's t-test and ANOVA with GraphPad Prism 9.3. The correlation between circLPHN3 and miR-142-5p was assessed by Spearman correlation analysis. Statistical analysis of clinical data was performed using SPSS 23.0. The relationship between circLPHN3 expression and the clinical characteristics of patients was evaluated using the Chi-square test. Kaplan–Meier and Cox regression analyses were used to predict CRC prognosis based on circLPHN3. All data were expressed as mean ± SD. A statistically significant difference was noted at P < 0.05.
Results
Downregulated circLPHN3 was associated with CRC prognosis
Volcano plot (Figure 1(a)) and cluster heatmap (Figure 1(b)) were plotted based on RNA-seq results of CRC tissues versus adjacent normal colorectal tissues from the GEO database (GSE121842). CircLPHN3 was downregulated in CRC compared to normal colorectal tissues. The RT-qPCR results of this study showed that the expression of circLPHN3 was markedly lower in CRC tissues (Figure 1(c)). The same difference was also observed in CRC cell lines (LoVo, HCT8, Caco2, SW480) (Figure 1(d)).
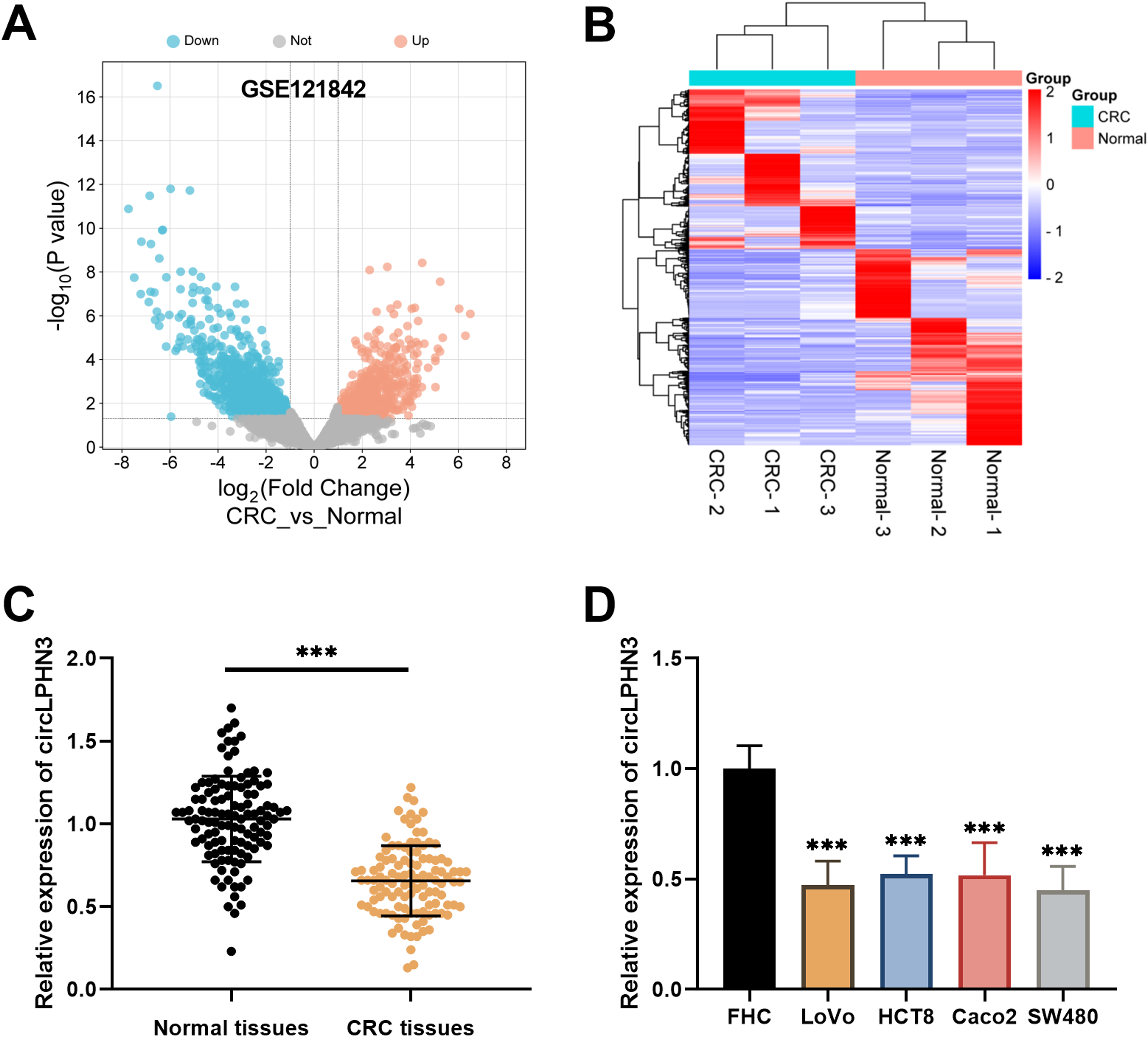
CircLPHN3 was downregulated in CRC tissues. The volcano plot (a) and cluster heatmap (b) were generated based on the GSE121842 dataset. (c) In CRC tissues, circLPHN3 was expressed at decreased levels compared to normal tissues. (d) Compared to FHC cells, circLPHN3 expression in CRC cell lines (LoVo, Caco2, HCT8, SW480) was also decreased. ***P < 0.001.
Using the average expression of circLPHN3 in CRC tissues as the cut-off value, CRC patients were divided into high-circLPHN3 (n = 53) and low-circLPHN3 (n = 60) groups. Clinical data collected from patients showed that circLPHN3 was related to TNM stage (P = 0.048) and lymph node metastasis (P = 0.021) in CRC patients (Table 1). Kaplan–Meier results showed that overall survival was lower in the low-circLPHN3 group than in the high-circLPHN3 group (P = 0.030, Figure 2(a)). Further analysis confirmed that circLPHN3 (hazard ratio (HR) = 4.026, 95% confidence interval (CI) = 1.312–12.349), as well as TNM stage (HR = 3.253, 95% CI = 1.130–9.366) and lymph node metastasis (HR = 3.457, 95% CI = 1.175–10.173, Figure 2(b)), were independent predictors for CRC prognosis.
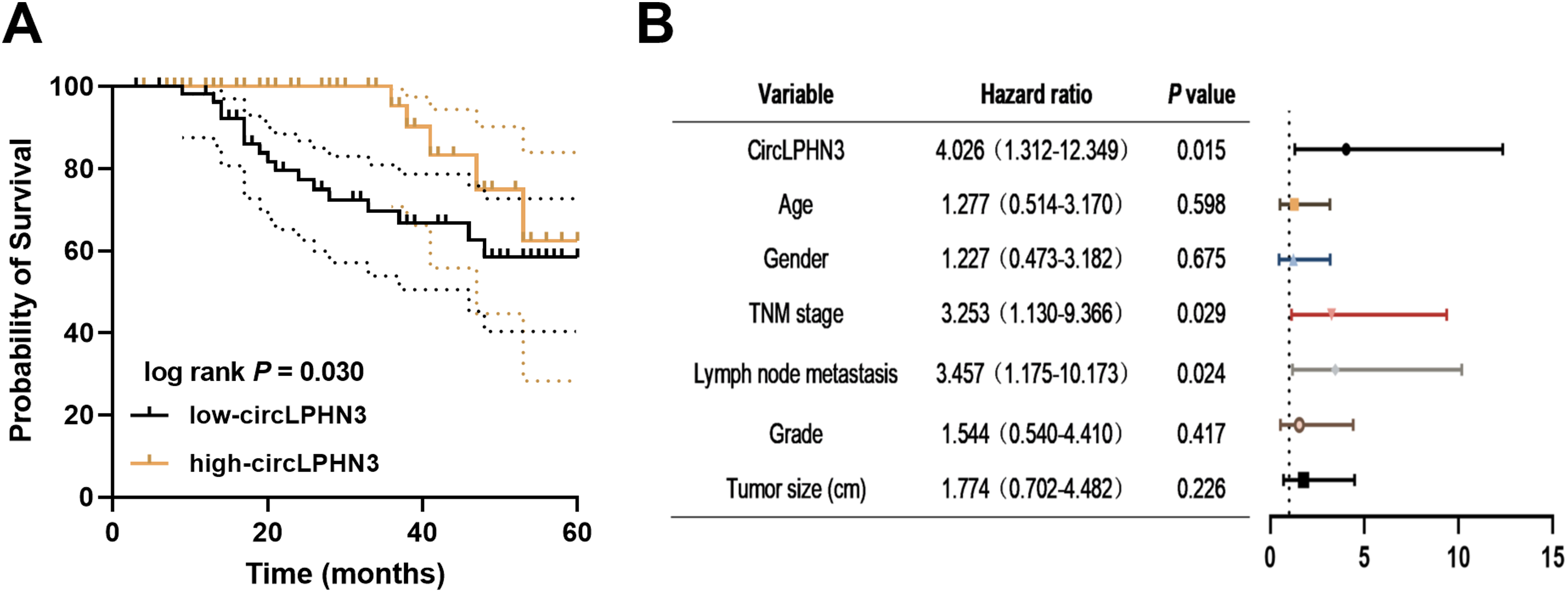
Clinical significance of downregulated circLPHN3 in CRC. A. Comparatively low expression of circLPHN3 was more strongly related to lower survival in patients with CRC. B. CircLPHN3 served as an independent predictor of CRC prognosis.
Association between circLPHN3 expression in CRC tissues and patients’ clinical features.
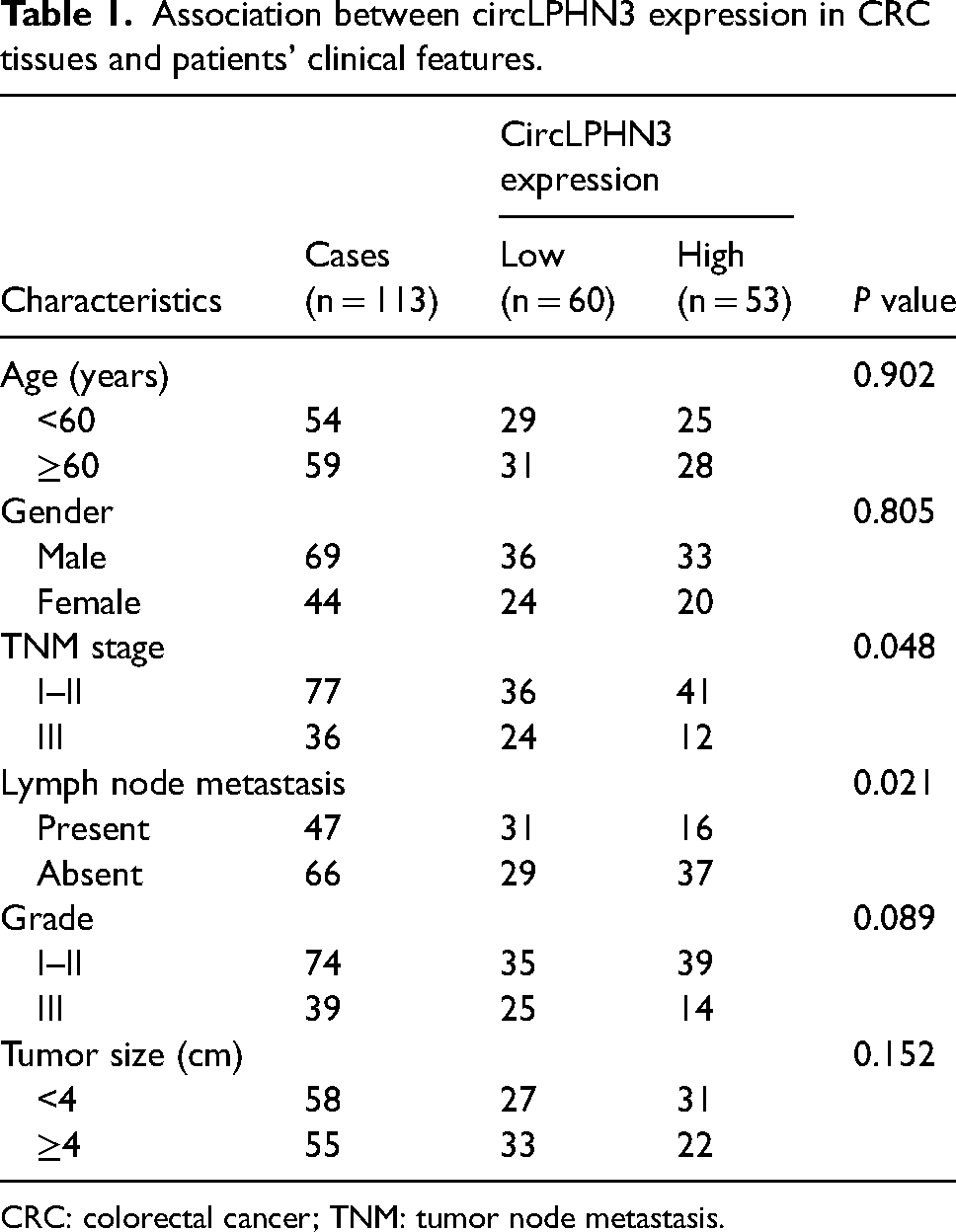
CRC: colorectal cancer; TNM: tumor node metastasis.
CircLPHN3 targeted and negatively regulated mir-142-5p
The miRNAs that bind to circLPHN3 were predicted by the RNAhybrid, Circular RNA Interactome, and starBase databases. Seven overlapping miRNAs were identified (Figure 3(a)). Among these, miR-142-5p was significantly upregulated in LoVo and SW480 cells compared to FHC cells (Figure 3(b)). In CRC tissues, miR-142-5p expression was elevated (Figure 3(c)). A strong negative correlation between circLPHN3 and miR-142-5p levels was found in CRC (r = −0.830, P < 0.001, Figure 3(d)). The binding sites of circLPHN3 to miR-142-5p were predicted by starBase database (Figure 3(e)). Then, the targeting relationship between circLPHN3 and miR-142-5p was validated through the dual-luciferase reporter assay. The results indicated that miR-142-5p mimic dramatically decreased the luciferase activity of circLPHN3-WT in LoVo and SW480 cells, but not circLPHN3-MUT. Conversely, miR-142-5p inhibitor elevated this activity (Figure 3(f) and (g)).
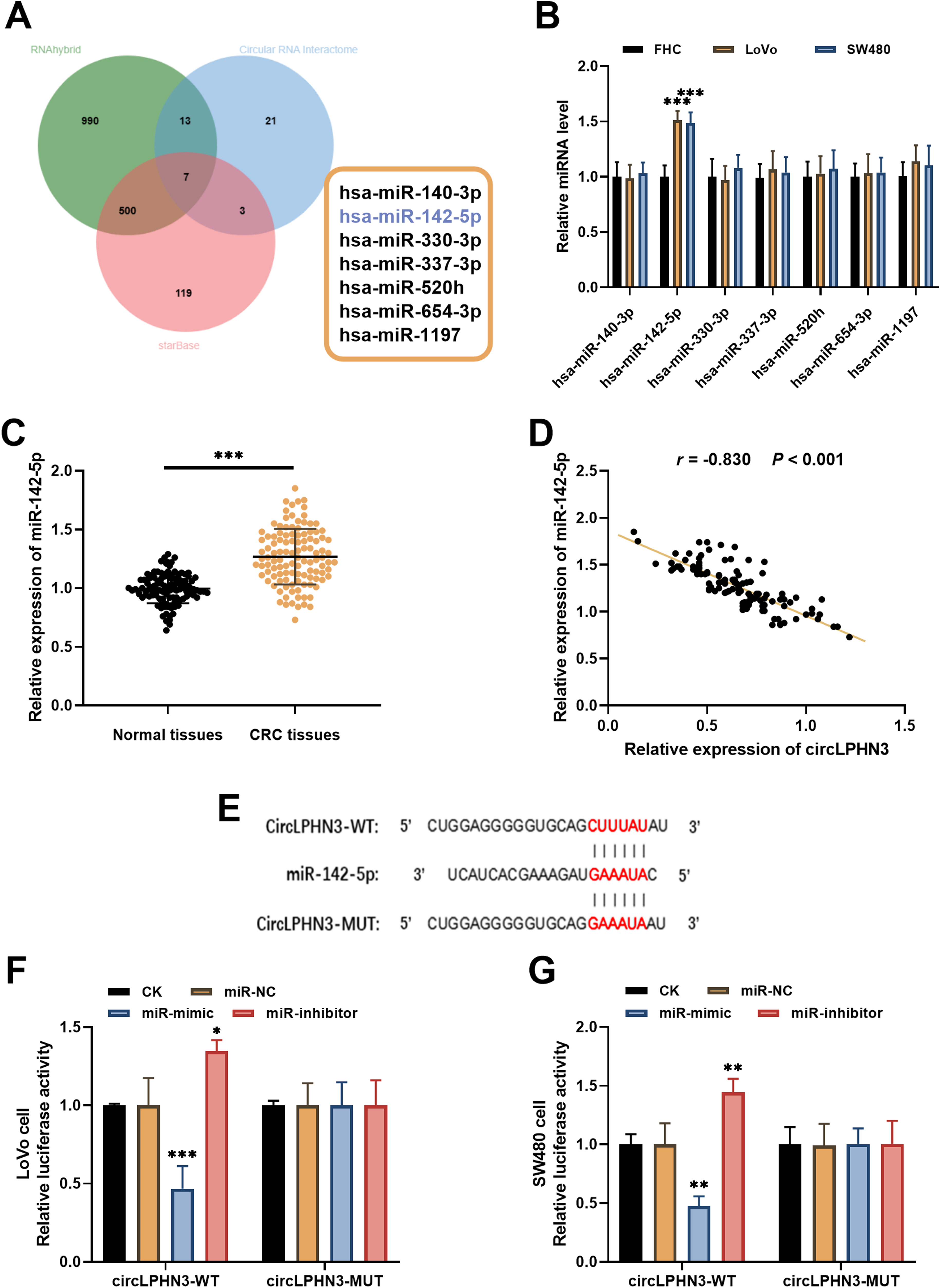
CircLPHN3 targeted miR-142-5p in CRC. (a) Target miRNAs of circLPHN3 were predicted by RNAhybrid, Circular RNA Interactome, and starBase databases. (b) miR-142-5p was upregulated in LoVo and SW480 cells. (c) miR-142-5p was present at elevated levels in CRC tissues. (d) In CRC, miR-142-5p and circLPHN3 showed a negative correlation. (e) Predicted binding sites of circLPHN3 and miR-142-5p. (f) and (g) Overexpression of miR-142-5p declined the luciferase activity of circLPHN3-WT in LoVo (f) and SW480 (g) cells, while downregulation of miR-142-5p had the opposite effect. *P < 0.05, **P < 0.01, ***P < 0.001.
CircLPHN3 regulated CRC cell progression via mir-142-5p
Transfection of LoVo and SW480 cells with pCD5-circLPHN3 (oe-circ) significantly increased circLPHN3 expression, while miR-142-5p mimic exerted no effect on this trend (Figure 4(a)). miR-142-5p levels were suppressed as the result of circLPHN3 overexpression, but miR-142-5p mimic partially restored it (Figure 4(b)). Overexpression of circLPHN3 notably curbed the proliferation of LoVo and SW480 cells, while increasing miR-142-5p expression reversed this suppression (Figure 4(c) and (d)). CircLPHN3 upregulation also markedly repressed CRC cell migration and invasion, which was notably attenuated by miR-142-5p mimic (Figure 4(e) and (f).
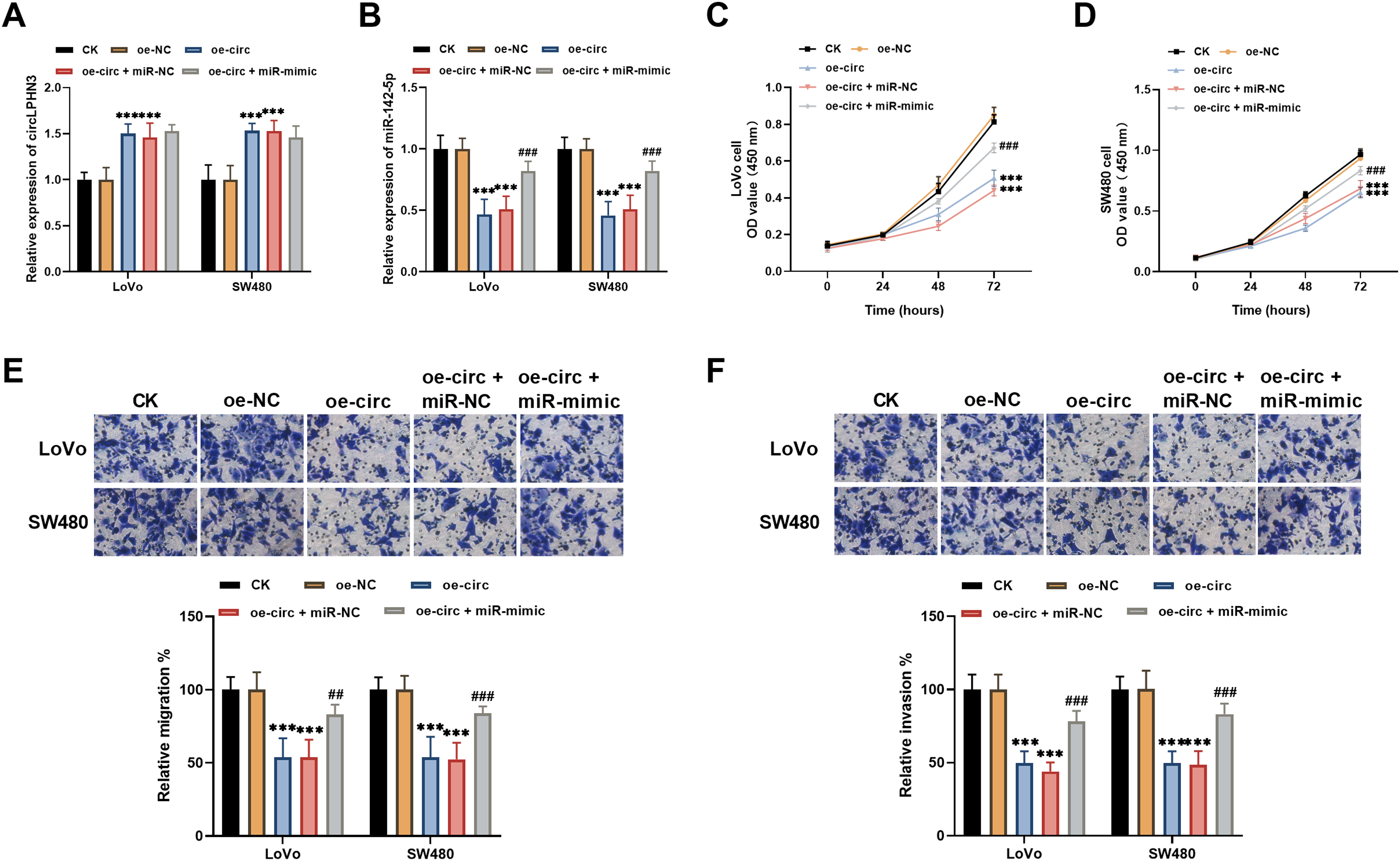
CircLPHN3 regulated CRC cell progression via miR-142-5p. (a) Transfection of pCD5-circLPHN3 (oe-circ) in LoVo and SW480 cells enhanced circLPHN3 expression. (b) In LoVo and SW480 cells, overexpression of circLPHN3 resulted in a reduction in miR-142-5p levels, which was reversed through the introduction of miR-142-5p mimic. (c) and (d) Suppression of proliferation of the LoVo (c) and SW480 (d) cells induced by circLPHN3 overexpression was lessened by miR-142-5p mimic. (e) and (f) Upregulated miR-142-5p attenuated the suppression effect of circLPHN3 overexpression on the migration (e) and invasion (f) of both cells. ***P < 0.001 (vs. oe-NC group), ##P < 0.01, ###P < 0.001 (vs. oe-circ group).
LDB2 was a target gene of mir-142-5p
To find the downstream genes of miR-142-5p, the prediction was performed using miRDB, TargetScanHuman, starBase, and miRWalk databases. A total of five potential targets of miR-142-5p were identified (Supplementary Figure 1(a)). Among them, LDB2 was remarkably reduced in LoVo and SW480 cells compared to FHC cells (Supplementary Figure 1(b)). Furthermore, an examination of the GEPIA database revealed that LDB2 expression was diminished in CRC tissues compared to normal colorectal tissues (Supplementary Figure 1(c)). The TargetScanHuman database was used to predict the binding sites of miR-142-5p and LDB2 (Supplementary Figure 1(d)). The overexpression of miR-142-5p resulted in a notable reduction in the luciferase activity of LDB2-WT in LoVo and SW480 cells. Conversely, the inhibition of miR-142-5p expression led to an increase in the luciferase activity of LDB2-WT (Supplementary Figure 1(e) and (f)). Nevertheless, the overexpression or inhibition of miR-142-5p demonstrated no significant impact on the luciferase activity of LDB2-MUT (Supplementary Figure 1(e) and (f)).
Discussion
CRC progression is a complex process involving oncogene inactivation, oncogene activation, and epigenetic dysregulation. circRNAs are a major research focus in epigenetic regulation, particularly for their function in disease progression. Numerous findings have uncovered that aberrant expression of circRNAs engaged in the development of multiple cancers such as CRC. For example, hsa_circ_0001955 is considered a promoter of tumorigenesis and a possible therapeutic target. 19 CircCAMSAP1 appeared to drive tumorigenesis in CRC and had clinical value in the diagnosis and prognosis of CRC. 20 According to RNA deep sequencing results, circLPHN3 was aberrantly expressed in CRC. 17 This study was the inaugural investigation into the expression and clinical significance of circLPHN3 in CRC, and it elucidated the biological function of this molecule.
Previously, circLPHN3 was appraised to be down-regulated in hyperplastic scars and involved in signaling pathways modulating HS. 21 More recently, a study identified circLPHN3 as a potential target for treating neuropathic pain by systematically characterizing circRNAs in SNT mice. 22 In this study, circLPHN3 was examined in CRC tissues and was observed to be dramatically lower than in normal tissues. Decreased circLPHN3 was associated with lymph node metastasis and advanced TNM stage, which may indicate the malignant progression of CRC. Additionally, the analysis of patients’ 5-year survival rates showed that lower circLPHN3 expression predicted shorter survival time. Moreover, the study found circLPHN3 to be an independent prognostic factor in CRC, with lower circLPHN3 linked to a higher risk of poor prognosis. Collectively, these results disclose that circLPHN3 is involved in CRC development and has potential as a therapeutic target.
Since researchers demonstrated that ciRS-7 could regulate miRNA activity, 23 numerous circRNAs interacting with miRNAs have been observed in human diseases. Previous research reported that circLPHN3 modulated miR-185-5p to enhance ZO-1 expression for the protection of the blood–brain barrier. 18 However, few studies have investigated the regulation of miRNAs by circLPHN3 in cancers. In this study, we used bioinformatics tools to identify miRNAs that may bind to circLPHN3 and identified miR-142-5p as being upregulated following validation in CRC cells. Furthermore, this study corroborates previous findings that miR-142-5p expression was increased in CRC. 24 It has been reported that miR-142-5p can be sponged by various circRNAs (e.g., circ_0000020, circ_0062491, and circ_0006873), thereby exerting different functions in diseases such as glioma, ulcerative colitis, and osteoporosis.25–27 Here, circLPHN3 exerted a negative modulation effect on the levels of miR-142-5p in CRC cells.
Evidence continues to confirm that circRNAs regulate proliferation, apoptosis, metastasis, and other cellular processes in CRC. circDDX17 was also downregulated in CRC, and circDDX17 overexpression inhibited CRC cell apoptosis and promoted proliferation. 28 Herein, circLPHN3 overexpression curbed CRC cell proliferation, migration, and invasion, suggesting that circLPHN3 might suppress CRC malignant progression. miR-142-5p was reported to regulate the progression of various cancers. For example, miR-142-5p was recognized as an oncogene that contributed to the development of cervical cancer. 29 Additionally, miR-142-5p, as a potential therapeutic target, could also modulate energy metabolism for CRC progression. 30 In this study, miR-142-5p attenuated the inhibition of CRC cell growth and metastasis by circLPHN3 overexpression, proving that miR-142-5p mediated the regulation of CRC cellular processes by circLPHN3. Further bioinformatics analysis and in vitro experiments confirmed that LDB2 was a downstream target of miR-142-5p. Previously, methylation of LDB2 has been shown to be useful for early detection of CRC. 31 In addition, LDB2 has been reported to play an inhibitory role in suppressing the progression of lung cancer and hepatocellular carcinoma.32,33 Therefore, LDB2 is hypothesized to be a suppressor of CRC progression. Further investigation is required to elucidate the role of LDB2 in CRC progression and associated mechanisms.
However, the remaining limitations of this study call for future modifications. Firstly, the clinical samples were sourced from a single center and the sample size was relatively small, reducing the reliability of the results. Tumor size, a known risk factor for CRC in previous studies, 34 did not show significant prognostic value in this study. Future research should include more samples from multiple centers. Additionally, in vivo experiments were performed with Balb/c nude mice (6 weeks old) to validate the inhibitory effect of circLPHN3 on CRC progression. However, subcutaneous transplantation tumor modeling in nude mice was not successful, which was possibly due to animal or cell line selection, experimental conditions, and so on. In animal experiments, it is necessary to improve the experimental protocol and strictly control the experimental conditions to determine whether circLPHN3 can inhibit tumor growth in vivo.
Taken together, circLPHN3 was significantly down-regulated in CRC and predicted poor prognosis. Moreover, this study concluded that circLPHN3 targeting to negatively modulate miR-142-5p might be critical for regulating CRC progression and had the potential to aid in CRC treatment.
Supplemental Material
sj-docx-1-jbm-10.1177_03936155241287219 - Supplemental material for CircLPHN3 correlates with prognosis in colorectal cancer and regulates cellular processes by targeting miR-142-5p
Supplemental material, sj-docx-1-jbm-10.1177_03936155241287219 for CircLPHN3 correlates with prognosis in colorectal cancer and regulates cellular processes by targeting miR-142-5p by JiWen Zhang and Yan Cai in The International Journal of Biological Markers
Footnotes
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Ethics approval and consent to participate
The study protocol was approved by The Ethics Committee of Shanghai Baoshan Luodian Hospital and followed the principles outlined in the Declaration of Helsinki. In addition, informed consent has been obtained from the participants involved.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
