Abstract
Background:
The KRAS mutations are high-frequency somatic mutations found in colorectal cancer patients from Western and Asian countries however, with the exception of exon 2 of KRAS, other prevalence and prognostic values have not been adequately assessed in Asian patients. The aim of this study was to determine the mutation frequencies of whole exon mutations of KRAS in Chinese colorectal cancer patients and to investigate their impact on prognosis.
Methods:
A total of 7189 tumor tissue samples (iCohort) were subjected to next-generation sequencing for detection of KRAS mutations. All pathologic or likely pathologic mutations of KRAS were considered. In addition, clinical features and prognostic dates were collected from 145 patients at The Third Affiliated Hospital of Soochow University, China (sCohort) and used droplet digital™ polymerase chain reaction to detect KRAS mutations.
Results:
In the iCohort, 2706 patients (37.6%) were confirmed harboring KRAS mutations. The most frequent of these mutations were G12D (32.19%), G12V (17.96%), and G13D (17.59%). In the sCohort, 51 colorectal cancer patients (35.17%) had KRAS mutations, among which KRAS G12D (64.71%), G13D (29.41%), and G14D (3.92%) were high-frequency. The KRAS mutations were associated with shorter median overall survival than wild-type tumors (69 vs. 55 months; HR 1.80; 95% Cl 1.22, 2.64; P=0.0003). In the Cox multivariate analysis, age (HR 1.562; 95% Cl 1.10, 2.22; P=0.013), tumor differentiation (HR 0.417; 95% Cl 0.19, 0.90; P=0.026), and KRAS mutation (HR 1.897; 95% Cl 0.19, 0.90; P=0.001) remained independent predictors of shorter overall survival. Among the common KRAS mutations, G12D was significantly associated with shorter overall survival (HR 2.17; 95% Cl 1.31, 3.58; P < 0.0001) compared with KRAS wild-type patients.
Conclusions:
Our findings indicate that KRAS genes are frequently mutated, and over 30% harbored the KRAS G12D mutation subtype. We found that the KRAS G12D mutation is associated with inferior survival and is a biomarker of poor prognosis in Chinese patients. Our data emphasize the importance of molecular features in colorectal cancer patients, which could potentially be improved by G12D-specific related inhibitors.
Keywords
Introduction
Colorectal cancer (CRC) is one of the most common causes of cancer-related death worldwide. Also, it is the second most common cancer in females and the third in males. 1 Since China is the most populous country in the world, the environment and local lifestyles in various regions are quite different. In addition, the early detection rate of cancer is low and there is no national cancer screening program, which has resulted in a rapidly increasing burden of CRC in China. 2 With the emergence of targeted drugs, such as epidermal growth factor (EGFR) or vascular endothelial growth factor (VEGF) agents (cetuximab, panitumumab, and bevacizumab), the treatment of metastatic CRC (mCRC) has entered the era of personalized therapy; the overall survival (OS) of mCRC has increased from the traditional 8–12 months to 30 months.3-5 However, this personalized treatment is appropriate for patients with a specific genetic variant of CRC. Currently, targeted therapies for the treatment of mCRC approved by the US Food and Drug Administration include bevacizumab, cetuximab, ramucirumab, panitumumab, aflibercept, and regorafenib. In China, bevacizumab and cetuximab are widely used as first- and second-line targeted therapies for mCRC. 6 In China, a recent study that reviewed medical records from 2011 to 2017, revealed that use of anti-VEGF therapy in first- and second-line treatment of mCRC was 17.8% and 23.3%, respectively. 7 However, with the introduction of reimbursement for bevacizumab in 2017 in China, there is an increasing trend for bevacizumab to be added to chemotherapies as first- and/or second-line treatment.
CRC is a multistep carcinogenic process accompanied by the accumulation of epigenetic and genetic changes. The adenoma–carcinoma pathway and the serrated tumor pathway are the main pathways for the occurrence of CRC, among which the activation of RAS-RAF was considered to be one of the most common mechanisms involved in colorectal carcinogenesis.1,8 KRAS mutation is the most common typical functional mutation in CRC, with mutated frequencies of 30%–50%. 4 Although the predictive effect of KRAS mutations on EGFR resistance in advanced CRC patients has been widely accepted,9,10 the predictive effect of KRAS mutations on survival in CRCs remains controversial.11,12 The MRC FOCUS trial showed that a KRAS mutation was a poor prognostic factor for OS (hazard ratio (HR), 1.40; 95% confidence interval (CI) 1.20, 1.65; P < 0.0001), but had minimal impact on progression-free survival (PFS; HR 1.16; 95% CI 1.00, 1.36; P = 0.05). 11 In another study, univariate and multivariate analyses showed that KRAS mutations have no major prognostic value for relapse-free survival or OS. 12
Most studies on the frequency and prognostic value of KRAS mutations, especially the efficacy of targeted therapy, have been carried out among non-Asian populations. The mutation frequency and prognostic value of CRC patients in China are not consistent.13,14 In our study, we aimed to determine the frequency of KRAS mutations in 7189 Chinese CRC patients and to assess their association with clinicopathological characteristics. In addition, we collected follow-up data from 145 patients in another cohort to determine their potential prognostic role in survival.
Methods
Patients and tumor samples
A total of 7189 CRC patients (iCohort) were collected from January 2013 to December 2019.The following clinical characteristics were abstracted: age, sex, and tumor histology.
Included in the study were 145 patients diagnosed with stage II–IV CRC at The Third Affiliated Hospital of Soochow University (sCohort) from January 2010 to December 2019. The clinical data of these patients were pooled retrospectively, and the factors included in the analysis were age, sex, pathology, clinical stage, and survival. In addition, the KRAS state in the sCohort were detected by droplet digital™ polymerase chain reaction (ddPCR). These data were approved by the Institutional Ethics Committee of The Third Affiliated Hospital of Soochow University, and written informed consent was obtained from the participants. The study was conducted according to the institutional guidelines and the regulations set by Chinese law for the use of human material for research.
DNA extraction
Formalin-fixed paraffin-embedded (FFPE) tissue sections were evaluated for tumor cell content using hematoxylin and eosin (H&E) staining. Only samples with a tumor content of ⩾20% were eligible for subsequent analyses. The thickness of each section was 4–5 μm, and a total of 15 sections were taken from each sample. FFPE tissue sections were placed in a 1.5 microcentrifuge tube and deparaffinized with mineral oil. Samples were incubated with lysis buffer and proteinase K at 56°C overnight until the tissue was completely digested. The lysate was subsequently incubated at 80°C for 4 hours to reverse formaldehyde crosslinks. Genomic DNA was isolated from tissue samples using the ReliaPrep™ FFPE gDNA Miniprep System (Promega) and quantified using the Qubit™ dsDNA HS Assay Kit (Thermo Fisher Scientific) following the manufacturers’ instructions.
Library preparation and DNA sequencing
DNA extracts (30–200 ng) were sheared to 250 bp fragments using an S220 focused-ultrasonicator (Covaris). Libraries were prepared using the KAPA Hyper Prep Kit (KAPA Biosystems) following the manufacturer’s protocol. The concentration and size distribution of each library were determined using a Qubit 3.0 fluorometer (Thermo Fisher Scientific) and a LabChip GX Touch HT Analyzer (PerkinElmer), respectively.
KRAS mutations were detected by whole exome sequencing with 800× sequencing depth in a College of American Pathologists (CAP) and Clinical Laboratory Improvement Amendments (CLIA) certified laboratory of 3D Medicines Inc. All pathologic or likely pathologic mutations of KRAS were considered. KRAS mutations, including single nucleotide variation, insertions/deletions, copy number variations, gene rearrangement, and fusions were assessed. Germline alterations were excluded.
Statistical analysis
Progression-free survival was defined as the time from the date of first-line therapy administration to the progression of cancer, or death from any cause. OS was calculated from the date of first-line therapy administration to the date of death from any cause. Disease-free survival (DFS) was calculated from the date of first-line therapy administration to the time of first-confirmed recurrence. Survival description was illustrated by Kaplan–Meier curves, with the P-value determined by a log-rank test. HR was determined through univariate and multivariate Cox regression. The associations between response and variables were examined by a univariate logistic regression. Variables with significant P-values or interest were included in the multivariate logistic regression. Continuous variables were compared by Mann–Whitney U test. For all analyses, a P-value < 0.05 was considered to be statistically significant, and a CI of 95% was used. All statistical analyses were performed using SPSS22.0 software (SPSS, Inc., Chicago, IL, USA).
Results
Patient characteristics in the iCohort and sCohort
In the iCohort, 7189 CRC patient characteristics were collected, of which 2706 (37.6%) had KRAS mutations (Figure 1). Among CRC patients with a KRAS mutation, 1571 (58.1%) were male, and 1572 (32.7%) patients were more than 65 years (Supplemental Table S1). In this cohort, the proportion of male patients was higher in the KRAS mutation group than in the KRAS wild-type (KRASwt) group (62.6% vs. 58.1%; P=0.0002).
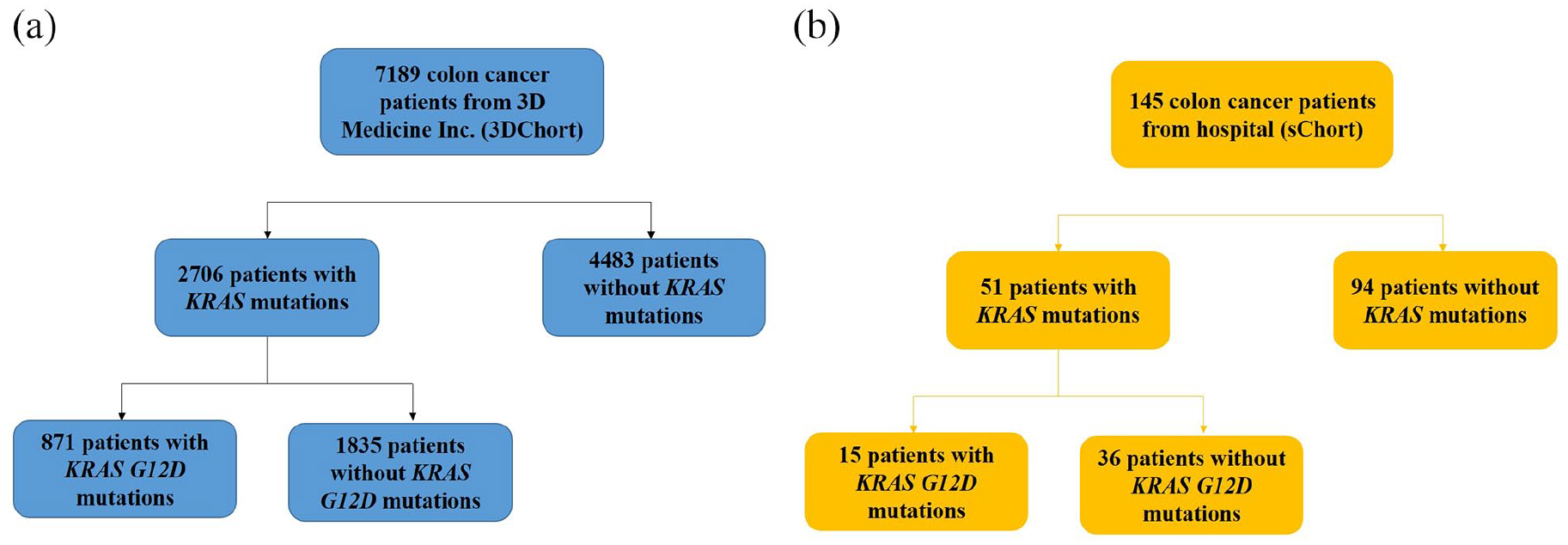
Flow charts of CRC patient enrollment in the study. Patients from iCohort (a) and sCohort (b).
In the sCohort, 145 patients diagnosed with CRC were included in this analysis. The baseline characteristics are summarized in Table 1. Of these 145 patients, 90 (62.1%) were male and 40.1% of the cohort were ⩾65 years old. There was no significant difference in tumor node metastasis (TNM) stage, tumor differentiation, and tumor location between KRAS mutation and KRASwt.
Clinicopathological characteristics of colorectal cancer patients in sCohort.
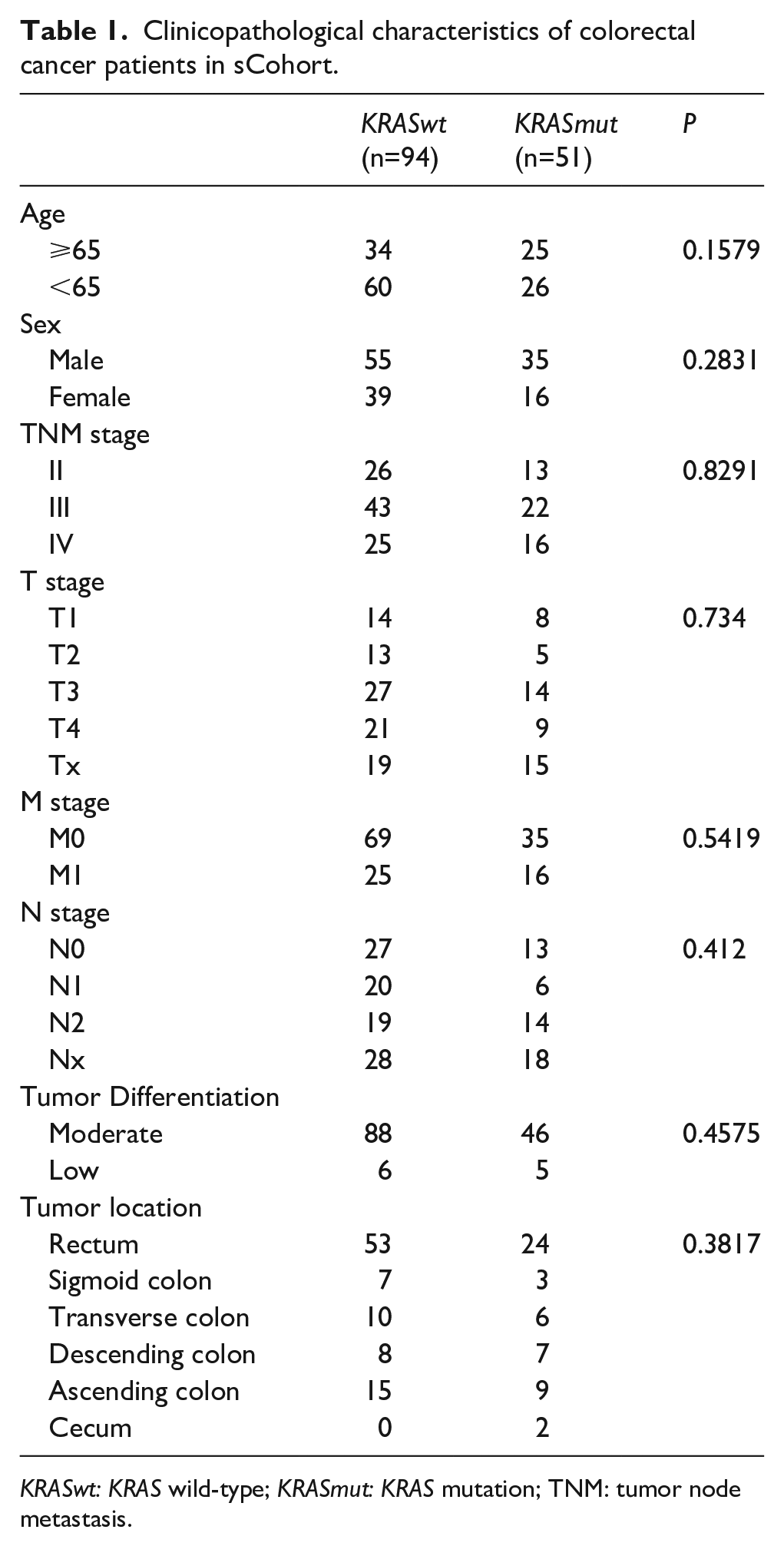
KRASwt: KRAS wild-type; KRASmut: KRAS mutation; TNM: tumor node metastasis.
KRAS mutations in the iCohort and sCohort
In the iCohort, the frequency of KRAS mutations was 37.6% (Figure 2(a)). Of the patients with KRAS mutations, approximately 80.9% of these mutations occurred in codons 12 or 13. The remaining mutations occurred mainly in codon 61 (3.7%) and exon 4 (7.1%). Also, 871 (32.2%) CRC patients were confirmed harboringG12D mutations (Figure 2(b)). The proportions of the other major KRAS mutations were as follows: G12V (18.0%), G13D (17.6%), and G12C (4.8%) (Figure 2(b)).
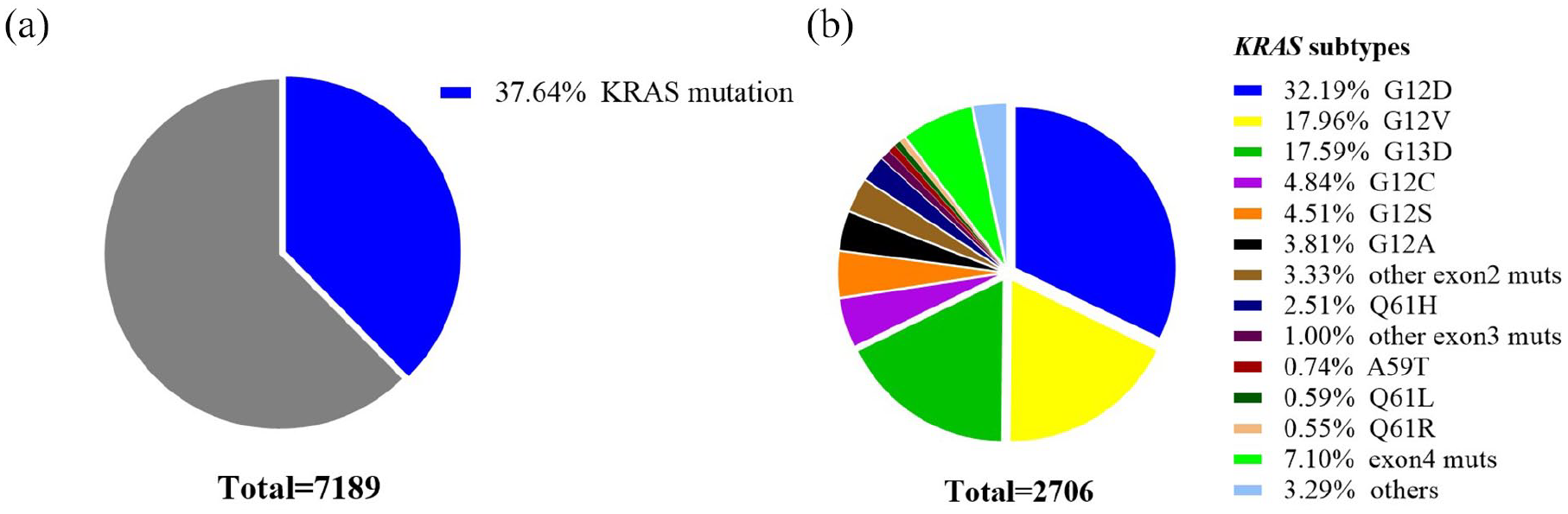
Pie charts of CRC patients with KRAS mutations. Pie charts showing the proportions of KRAS mutation (mut) and wild-type (wt) tumors in the iCohort (a), and the proportions of different KRAS mutation subtypes in the iCohort (b).
In the sCohort, 51 of 145 CRC patients were confirmed to have KRAS mutations (Figure 3(a)). G12D (64.7%) was the high-frequency mutation site in the KRAS mutation, followed by G13D (29.4%), G14D (3.9%), and Q16H (2.0%) (Figure 3(b)). The clinical and pathological characteristics of CRC with KRAS G12D and other KRAS mutations were compared in the sCohort (Supplemental Table S2). There were no significant differences in age, sex, TNM stage, and tumor differentiation between the KRAS G12D mutation and other KRAS mutations.
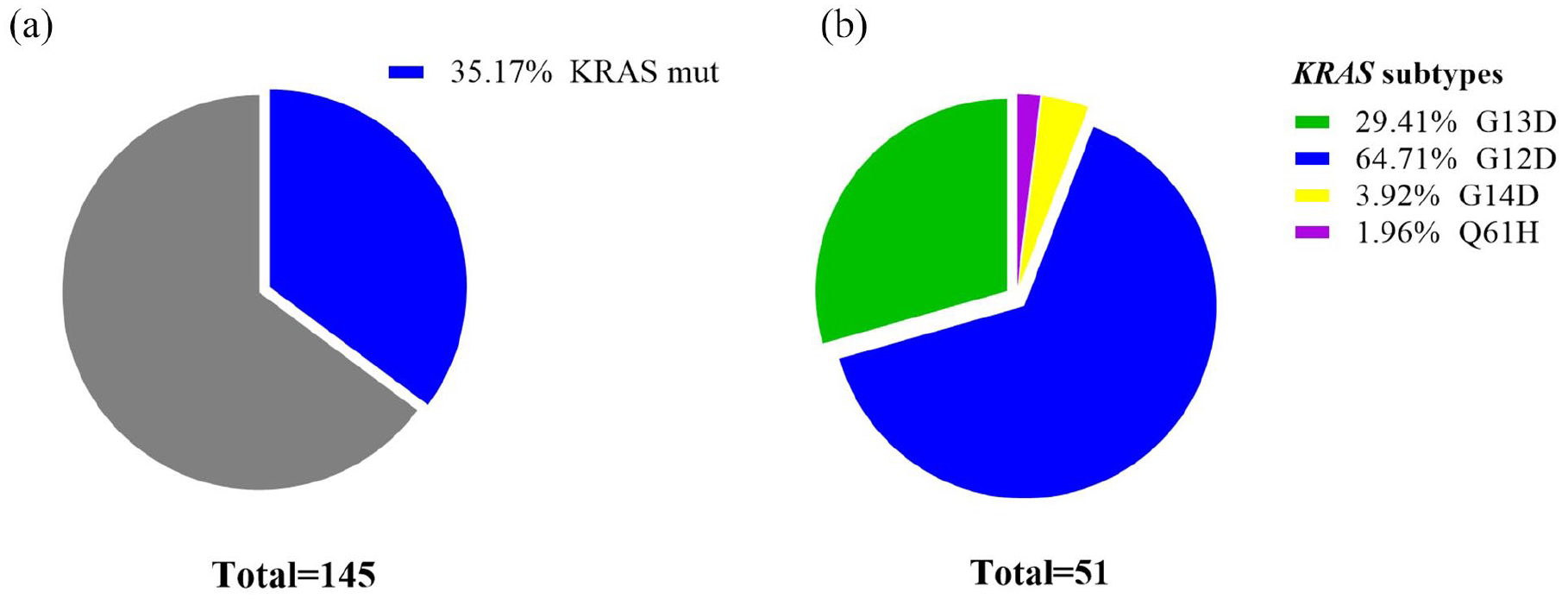
Pie charts of CRC patients with KRAS mutations. Pie charts showing the proportions of KRAS mutation (mut) and wild-type (wt) tumors in the sCohort (a), and the proportions of different KRAS mutation subtypes in the sCohort (b).
Association between KRAS G12D and survival outcomes
We first analyzed whether KRAS mutation was associated with the survival outcomes in the sCohort. The median OS was 69 months in patients with KRASwt versus 55 months in patients with KRAS mutation (HR 1.80; 95% Cl 1.22, 2.64; P=0.0003; Supplemental Figure S1(a)). KRAS G12D mutation was associated with poorer OS (69 vs. 55 months; HR 2.17; 95% Cl 1.31, 3.58; P<0.0001; Supplemental Figure S1(b)).
To identify the prognostic values of the KRAS mutations on OS, clinical and molecular variables were included in the Cox regression analysis. Univariable analysis revealed significant association between poorer OS and KRAS, age, moderate differentiation, and TNM stage; however, there was no association between OS and sex (Table 2). In a multivariate model, KRAS mutation, moderate differentiation, and TNM stage remained independent prognostic indicators for OS (Table 2; P<0.05).
Univariate and multivariate analyses of overall survival.
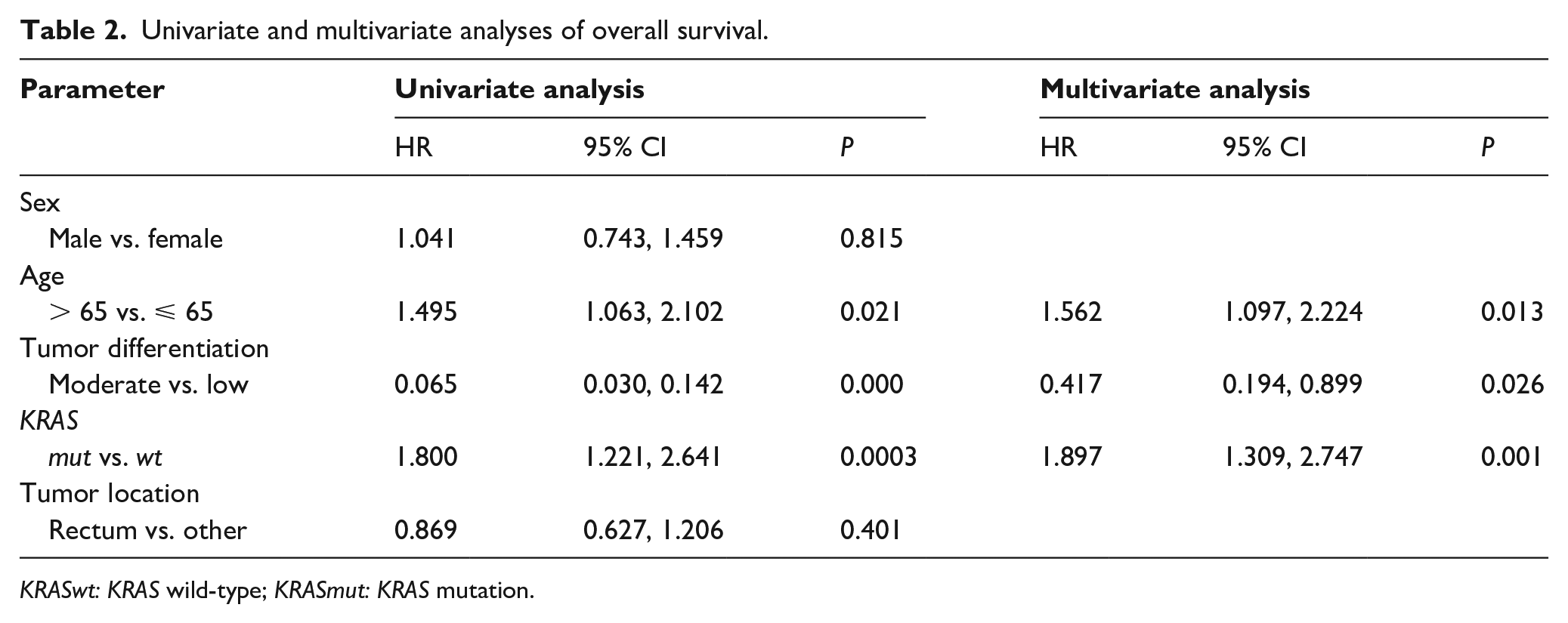
KRASwt: KRAS wild-type; KRASmut: KRAS mutation.
In addition, we analyzed the PFS in stage IV CRC patients and DFS in stage II–III CRC patients. As a data cutoff, the median PFS (mPFS) was 7.5 months in KRASwt patients versus 4.0 months in KRAS mutation patients (HR1.71; 95% Cl 0.84, 3.50; P=0.061, Supplemental Figure S2(a)). No differences in PFS were found between patients with KRAS G12D/G13D and without KRAS mutation (mPFS: 7.5 vs. 6.0 months; HR 1.64; 95% Cl 0.68, 3.95; P=0.1616; mPFS: 7.5 vs. 4.0 months; HR 2.03; 95% Cl 0.64, 6.37; P=0.0719; Supplemental Figure S2(b)). KRASwt CRC patients had longer median DFS (mDFS) than KRAS mutation patients (50.7 vs. 40.5 months; HR 2.15; 95% Cl 1.31, 3.53; P<0.0001; Supplemental Figure S3(a)). In the KRAS mutation, G12D were associated with shorter mDFS than KRASwt (57.0 vs. 38.0 months; HR 3.12; 95% Cl 1.56, 6.22; P<0.0001; Supplemental Figure S3(b)). In the multivariate analysis, KRAS mutation was an independent prognostic indicator for DFS (HR 2.16; 95% Cl 1.372, 3.395; P=0.001; Supplemental Table S3).
Discussion
There have been numerous studies on the clinicopathological characteristics and prognostic value of KRAS in patients with CRC both in China and other countries, which suggests that KRAS mutations may be associated with a poor prognosis, but this conclusion is inconsistent, especially with the data on local disease.14–17 However, some studies have demonstrated that KRAS mutations have no effect on survival, including stage II or III CRC.18,19 In addition, the RASCAL-II study has shown that among the 12 possible mutations on codon 12 or 13 of KRAS, only the G12V mutation (glycine to valine) was associated with inferior survival. 20 This contradiction may be due to limitations with other studies such as patient number and variation in regional environment and populations. In this study, we determined the frequency of KRAS mutation in 7189 Chinese CRC patients with resectable tumors. Our data show that KRAS G12D mutation is the highest frequency mutation. In addition, we elucidate the prognostic value of KRAS mutations in other cohorts. To the best of our knowledge, we are the first to assess the impact of KRAS mutations on the frequency and prognosis of CRC patients in China in a large sample cohort.
The frequency of KRAS is similar compared to Western data. 21 In our iCohort, the incidence of KRAS was 38% compared with 44% of the Chinese data reported in the previous small sample. 8 The distribution of KRAS mutations in Western populations shows that G12D is the most common mutation subtype, followed by G12V, G13D, G12C, G12S, and G12A. 22 In our data, the high-frequency mutation sites of KRAS were G12D, G12V, G13D, G12C, and G12S, which are consistent with the results of Shen et al. 23
Although the predictive role of KRAS mutation in anti-EGFR antibody therapy has been recognized, its prognostic value for survival remains controversial. This may be related to differences in study size, patient selection, surgical procedures, chemotherapy regimens, genetic variation detection methods, and data analysis. For example, cetuximab—a monoclonal antibody that binds the extracellular domain of epidermal growth factor receptor (EGFR)—is effective in a subset of KRASwt metastatic CRC. 24 In the CO.17 trial, patients with tumors harboring KRAS mutations did not benefit from cetuximab, while patients with KRASwt tumors benefited from cetuximab. 25 However, the relationship between different KRAS mutation sites and prognosis remains controversial. In a retrospective study, it was found that in patients with chemotherapy-refractory CRC with p.G13D mutations, the use of cetuximab was associated with longer OS and PFS, but not other tumors with KRAS mutations. 26 A meta-analysis compared the efficacy of anti-EGFR monoclonal antibody therapy in KRAS G13D and other KRAS-mutated metastatic CRC tumors and found that for other KRAS mutations, the HR of OS benefit with the addition of anti-EGFR treatment was 1.06 (95% CI 0.96, 1.17), compared to 1.08 (95% CI 0.73, 1.60) for KRAS G13D (interaction test P = 0.99). For PFS, the HR of other KRAS mutations was 1.07 (95% CI 0.92, 1.26), and 0.96 for KRAS G13D (95% CI 0.73, 1.27). This result showed that there was no significant difference between KRAS G13D and other KRAS mutated tumors in terms of the therapeutic benefit of anti-EGFR therapy for mCRC. 27 In addition, significant interactions for individual mutant KRAS alleles were observed with G13D negatively and G12V positively associated with OS in the panitumumab-containing arm. 28 Our study mainly found that a KRAS mutation had a worse prognosis than KRASwt for patients. In both univariate and multivariate analysis, we further confirmed KRAS mutation as an independent negative prognostic factor for OS. However, in our study, KRAS codon 12 and codon 13 mutations had no effect on PFS.
Our study included the largest sample size to date of CRC patients with KRAS mutations. However, it also has some limitations. First, the genetic variation data of CRC patients obtained from the iCohort have only partial clinical characteristics data and lack of survival data. Therefore, the prognostic effect of KRAS G12D mutation in poor OS was obtained from only one center. Second, KRAS mutations in the sCohort were detected by ddPCR, which only contained four loci. However, this does not affect the data of KRAS G12D, because G12D is the main type of variation. Thus, results regarding the prognostic roles of KRAS mutations and G12D mutations were reliable.
In conclusion, our study found KRAS mutations in approximately 37.6% of CRC patients in China. Among them, nearly 32.2% had the KRAS G12D mutation subtype, which usually occurred at in patients <65 years old. Our study suggests that the KRAS mutation is an independent prognostic factor in colon cancer patients.
Supplemental Material
sj-pdf-1-jbm-10.1177_17246008211017152 – Supplemental material for Clinical characteristics and prognostic value of the KRAS mutation in Chinese colorectal cancer patients
Supplemental material, sj-pdf-1-jbm-10.1177_17246008211017152 for Clinical characteristics and prognostic value of the KRAS mutation in Chinese colorectal cancer patients by Ye Yuan, Yingting Liu, Ye Wu, Junling Zhang, Chunti Shen, Feng Zhang, Changping Wu and Wenwei Hu in The International Journal of Biological Markers
Footnotes
Authors’ contributions
Conception and design of the study: Ye Yuan, Junling Zhang, Changping Wu, and Wenwei Hu; acquisition of clinical data: Ye Yuan, Yingting Liu, Ye Wu, Junling Zhang, Chunti Shen; analysis and interpretation of the data: Ye Yuan, Yingting Liu, and Feng Zhang; Manuscript drafting and revision: Chunti Shen, Ye Yuan, Junling Zhang, Changping Wu, and Wenwei Hu; final approval of the manuscript: all authors. Ye Yuan and Yingting Liu contributed equally to this work.
Declaration of conflicting interest
The author(s) declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: Junling Zhang is employee of 3D Med Medicine. No potential conflicts of interest were disclosed.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Applied Basic Research Program of Changzhou Science and Technology Project (CJ20180061 to Yingting Liu), the Funding from Young Talent Development Plan of Changzhou Health Commission (2020-233) (CZQM2020044to Yingting Liu) and the Science and Technology Project of Jiangsu Traditional Chinese Medicine (YB2017059 to Changping Wu).
Availability of data and materials
The datasets used and analyzed during the current study are available from the corresponding author upon reasonable request.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
