Abstract
Background
Cervical cancer is the most prevalent malignant tumor in women. This study aims to detect collagen type V α1 chain (COL5A1) expression and its clinical relevance in the prognosis of patients with cervical cancer.
Methods
Cervical cancer tissues and their paired adjacent normal tissues were prepared for tissue microarray. The expression of COL5A1 protein and the scores of the expression were evaluated by immunohistochemistry (IHC) staining. The prognostic value of COL5A1 was analyzed by R software version 4.2.1 with “survival, survminer, ggplot2” packages and Gene Expression Profiling Interactive Analysis (GEPIA). The cBioPortal database was utilized for the analysis of COL5A1 gene mutations.
Results
COL5A1 protein was overexpressed in human cervical cancer tissues compared to their paired adjacent normal tissues detected by IHC (P < 0.001). High expression of COL5A1 tends to be in elderly patients with cervical cancer. Survival analyses of clinical data of patients with cervical cancer showed that a high level of COL5A1 expression was significantly correlated with shorter overall survival (P = 0.031) and disease-free survival (P = 0.042) of patients. Further analyses of The Cancer Genome Atlas-Cervical Squamous Cell Carcinoma and the GEPIA survival datasets confirmed the association of high COL5A1 expression with poor overall survival of patients (P = 0.040 and P = 0.018, respectively). The analysis of genomic alterations of COL5A1 using the cBioPortal tool revealed that the COL5A1 gene was altered in 4% of cervical cancer patients and COL5A1 corresponding protein alterations with post-translational modifications were hydroxylation.
Conclusion
COL5A1 is a tissue biomarker correlated with the poor prognosis of patients with cervical cancer, which may lead to a new clinical application.
Introduction
Cervical cancer is the most prevalent malignant tumor of the female reproductive tract worldwide. 1 According to the International Agency for Research on Cancer (IARC) from the World Health Organization (WHO), new cancer cases and cancer deaths of patients with cervical cancer in 2022 are 662,301 and 348,874, respectively (https://gco.iarc.fr/today/en/dataviz/tables?mode=cancer&group_populations=1&multiple_populations=1&types=0). In China, cervical cancer is the fifth most common cancer in females with 150,700 estimated new cases and 55,700 cancer deaths in 2022. 2 In the US, the estimated number of new cancer cases and cancer deaths of the uterine cervix are 13,820 and 4,360, respectively, in 2024. 3 It has been shown that early detection and timely and effective treatment are important to make improvements in the prognosis of patients with cervical cancer in the last two decades.4,5 However, the number of cervical cancer cases in China has not significantly declined. 6 The most common reasons are the slow growth of tumors and lack of symptoms at the early stage of cervical cancer. Some patients are often ignored, disregarded, and careless although cervical cytology and human papillomavirus (HPV) tests are performed. The sensitivity of cervical cytology in existing screening programs is low, and the HPV test has a high sensitivity but with low specificity.7,8 False-negative results can occur from cervical sampling limitations, whereas false-positive results can occur from laboratory contamination. On one hand, HPV infection is one of the risk factors; on the other hand, cervical cancer is not only caused by HPV infections. It has been reported that approximately 99.7% of cervical cancer cases are HPV-related, 9 whereas tongue squamous cell carcinoma can also be associated with HPV infection, 10 indicating that the positive of HPV infection is not a unique indicator of cervical cancer. The occurrence and progression of cervical cancer are complex and some gene and protein alterations may contribute to the tumorigenesis of the cervix uteri.11,12 Therefore, defining relevant biomarkers as useful molecular indicators may have the potential to develop clinical applications for the diagnosis, prevention, and prognosis of cervical cancer.
Type V collagen has three alpha peptide chains (α1, α2, and α3). The collagen type V α1 chain encoded by the COL5A1 gene and one α1 chain forms a heterotrimer with two α2 chains that usually control the initiation of collagen fibril assembly.13,14 It has been shown that COL5A1 protein can regulate extracellular glial remodeling and plays an important role in the development and progression of tumors involved in many cancers, including breast, ovarian, lung, and gastric cancers.15–18 However, the role of COL5A1 in cervical cancers is still unidentified and the prognostic and predictive value of COL5A1 in cervical cancer has not been adequately studied. In cancer, remodeling of the interstitial extracellular matrix (ECM), in which COL5A1 is involved, 19 induces a broad range of biophysical and biochemical changes affecting cell migration and metastatic progression at the late stage of the disease. 20 The prognosis of advanced cervical cancer is often poor. 21 Therefore, improving the early diagnosis of cervical cancer is crucial for prolonging the survival of patients.
In this study, the diagnostic value of COL5A1 as a tissue biomarker in cervical cancer was evaluated. The association of the expression of COL5A1 with cervical cancer at different stages was also analyzed. The clinical relevance of COL5A1 involved in tumorigenesis may provide theoretical and experimental bases for the diagnosis and treatment of cervical cancer.
Materials and methods
Human sample collection and tissue microarray
The ethics approval of using human subjects was approved by the Ethics Committee of Jinshan Hospital. The inclusion criteria for cervical cancer patients were (a) female; (b) age 18–75 years; (c) pathological confirmation of cervical cancer after operation; (d) neither received chemotherapy nor radiotherapy; and (e) archived tumor tissue samples within 5 years. The exclusive criteria were (a) incomplete specimens of patients; (b) paired samples without adjacent non-tumorous tissues; (c) acute genital tract inflammation; (d) combined with other genital tract tumors; and (e) incomplete clinical data. Freshly isolated tissues were collected and stored at −80°C until use. The histological and clinicopathological features with tumor, node, metastasis (TNM) classification and different stages were determined according to the International Federation of Gynecological and Obstetrics (FIGO) and the American Joint Committee on Cancer (AJCC) criteria. A tissue microarray was prepared by Shanghai Outdo Biotech Co. Ltd (Cat. number: CEC1601; Shanghai, China). A total of 80 formalin-fixed paraffin-embedded cervical cancer samples (FIGO stage I–IV, each 20 samples/group) were subjected to histopathological examination and immunohistochemistry. The adjacent non-tumorous tissues were used as controls.
Immunohistochemistry staining
Four micrometer-thick sections of tissue specimens were stained with hematoxylin and eosin (H&E) to confirm the histological characteristics. The Mouse/Rabbit Specific HRP/DAB Detection IHC Kit was purchased from Abcam (Cambridge, UK) and was used for immunohistochemistry (IHC). De-paraffin sections were incubated with rabbit polyclonal anti-COL5A1 antibodies (1:100 dilution, Abcam) at 4°C overnight. Non-immune rabbit serum (1:200 dilution, Abcam) in phosphate-buffered saline (PBS) instead of primary antibody served as the negative control. The cells that showed positive staining were counted in at least 12 sample fields under a microscope. Two independent pathologists who were blind to the clinical features and results evaluated the immunostaining. The IHC staining of COL5A1 was scored as described previously. 22 The score of the percentage of positive cells was 0 (0% positive cells), 1 (≤25% positive cells), 2 (26–50% positive cells), 3 (51–75% positive cells), and 4 (>75% positive cells). The score of intensity of immunoreactive staining was 0 (no staining), 1 (weak staining), 2 (moderate staining), and 3 (strong staining). The final IHC score of COL5A1 (0–7) was determined by the sum of the positive score and intensity score. Finally, the 0–1, 2–4, and 5–7 sum points were defined as negative, low expression, and high expression of COL5A1, respectively.
Detection of serum cancer antigen 125 and squamous cell carcinoma antigen
Whole blood samples from patients were collected into tubes under sterile conditions. After centrifugation at 3000 g for 10 min, the upper layer of serum was transferred to a new tube and stored at 4°C until use. Serum cancer antigen (CA125) and squamous cell carcinoma antigen (SCC-Ag) were measured by chemiluminescence immunoassay using a Roche E6000 analyzer (Roche Diagnostics, Basel, Switzerland) according to the manufacturer's instructions.
Prognostic value analysis
The prognostic value of COL5A1 was analyzed by R software version 4.2.1 with “survival, survminer, ggplot2” packages, using clinical data from cervical cancer patients of tissue microarray and The Cancer Genome Atlas-Cervical Squamous Cell Carcinoma (TCGA-CESC) dataset (https://portal.gdc.cancer.gov/). Gene Expression Profiling Interactive Analysis (GEPIA) (http://gepia.cancer-pku.cn/), an online database, was also used to conduct the prognostic value of COL5A1.
Cbioportal database analysis
The cBioPortal (http://cbioportal.org) was an open-access web resource that visualized and analyzed multidimensional cancer genomic data from TCGA. 23 “Cervical Squamous Cell Carcinoma (TCGA, PanCancer Atlas)” was chosen for the analysis of COL5A1 gene mutations, and copy number alterations (CNA) (278 samples/patients) were selected. The OncoPrint, cancer types summary, mutations, and comparison tabs were applied according to the online instructions.
Statistical methods
SPSS19.0 software was used for data processing and statistical analysis. To compare two groups, a Student's t-test was used. For multiple group comparisons, an analysis of variance (ANOVA) followed by Dunnett's multiple comparisons test was applied. For the association of two categorical variables, Yates’ correction test, Chi-squared test, or Wilcoxon test was used. Data are presented as the mean ± SD. A P-value < 0.05 was considered statistically significant.
Results
COL5A1 is overexpressed in cervical cancer
The tissues of the cervix from 80 patients with cervical cancer were collected and subjected to tissue microarray. The pathological types of cervical cancer were all squamous cell carcinoma. Among these, two pairs of tissues were lost during the preparation of microarray and IHC. The IHC staining showed that the COL5A1 protein was overexpressed in human cervical cancer tissues compared to their paired adjacent non-tumorous tissues (Figure 1(a)). Next, the staining scores of IHC were measured and compared between cervical cancer and paired adjacent non-tumorous tissues. In 78 cervical cancer tissues, COL5A1 staining was negative in 2 cases, low expression in 33 cases, and high expression in 43 cases. The IHC score of COL5A1 protein was significantly higher in cervical cancer tissues than in their paired adjacent non-tumorous tissues (Figure 1(b)). Further analysis of the staining scores of IHC at different tumor stages revealed that COL5A1 was elevated in tumor tissues at different stages (T) compared to adjacent non-tumorous tissues (N) but no statistical difference between tumor stages (T1–T4) (Figure 1(c)). These data suggest that COL5A1 is a tissue biomarker of human cervical cancer.
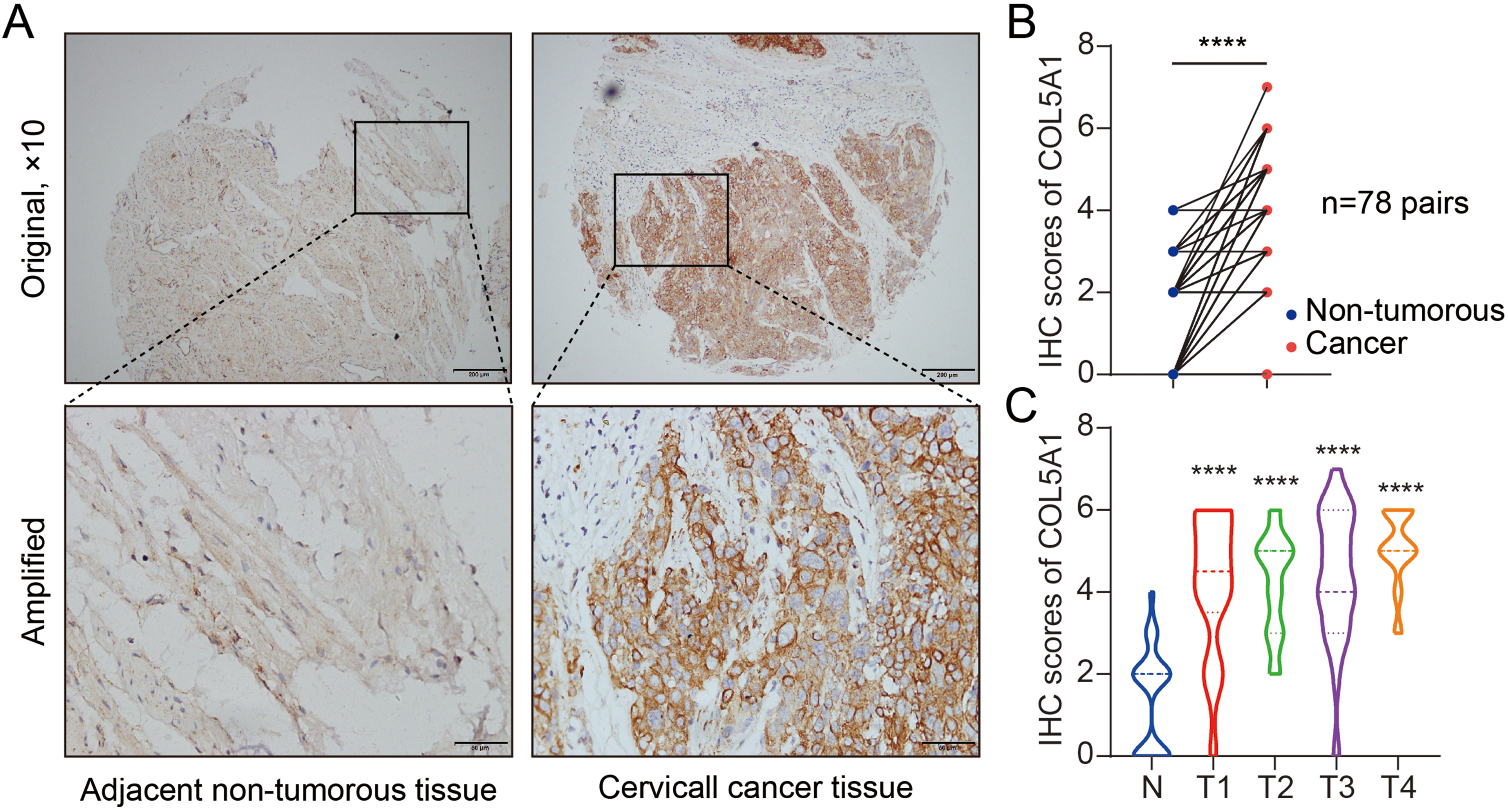
Detection of COL5A1 protein expression in tissue microarray by immunohistochemistry (IHC) staining. (a) Detection of COL5A1 protein expression in paired adjacent non-tumorous tissues and primary cervical cancer tissues. Representative images are shown. Original magnification ×200; Amplified ×400. (b) Comparison of the IHC score of COL5A1 protein expression between adjacent non-tumorous tissues and cervical cancer tissues among 78 pairs after IHC staining. Student's t-test was used. ****, P < 0.001 cancer vs. normal. (c) Analysis of the IHC score of COL5A1 at different stages. ANOVA followed by Dunnett's multiple comparisons test was applied. ****, P < 0.001 tumors at different stages (T) vs. non-tumorous tissues (N). T1-4, tumor stage 1–4. Number of cases: N = 78, T1 = 18, T2 = 26, T3 = 23, and T4 = 11.
High expression of COL5A1 tends to be in elderly patients with cervical cancer
Next, we analyzed the correlation between COL5A1 expression and clinicopathological features in patients with cervical cancer. Based on the expression levels of COL5A1, patients with cervical cancer were divided into two groups in which the high-expression group had an IHC score ≥5 (n = 43) and the low-expression group had an IHC score between 2 and 4 (n = 33). We found that although COL5A1 was overexpressed in cervical cancer, the high expression of COL5A1 was not correlated with tumor stages except for the factor of age. Higher levels of COL5A1 expression tended to be in elderly patients with cervical cancer (Table 1).
Correlation between COL5A1 expression and clinicopathological features in cervical cancer.
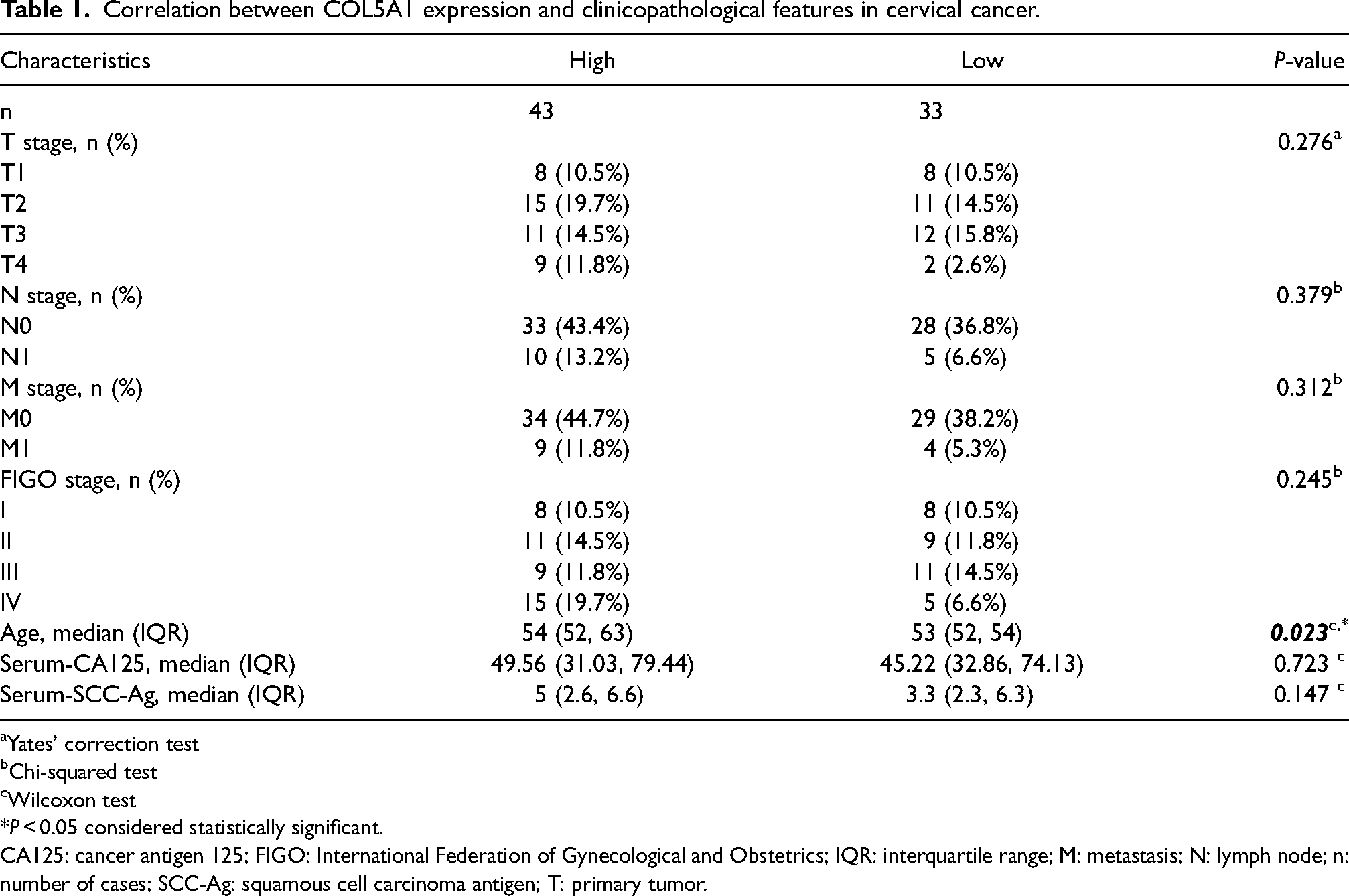
Yates’ correction test
Chi-squared test
Wilcoxon test
P < 0.05 considered statistically significant.
CA125: cancer antigen 125; FIGO: International Federation of Gynecological and Obstetrics; IQR: interquartile range; M: metastasis; N: lymph node; n: number of cases; SCC-Ag: squamous cell carcinoma antigen; T: primary tumor.
COL5A1 overexpression is correlated with poor prognosis in cervical cancer
Subsequently, survival analyses of the clinical data of patients with cervical cancer were performed. Again, patients with cervical cancer were divided into two groups: high-expression and low-expression based on the tissue microarray outcome. A high level of COL5A1 expression was significantly correlated with low overall survival (OS) (P = 0.031) and disease-free survival (DFS) (P = 0.042) rates of patients (Figure 2(a) and (b)). Similarly, high levels of COL5A1 expression had unfavorable OS time compared to those with low levels of COL5A1 expression in patients with cervical cancer observed after analyzing the TCGA-CESC dataset by R software (P = 0.040) (Figure 2(c)). In addition, analysis of OS data from the GEPIA survival panel revealed that high levels of COL5A1 expression were also correlated with poor OS of patients with cervical cancer (P = 0.018) (Figure 2(d)).
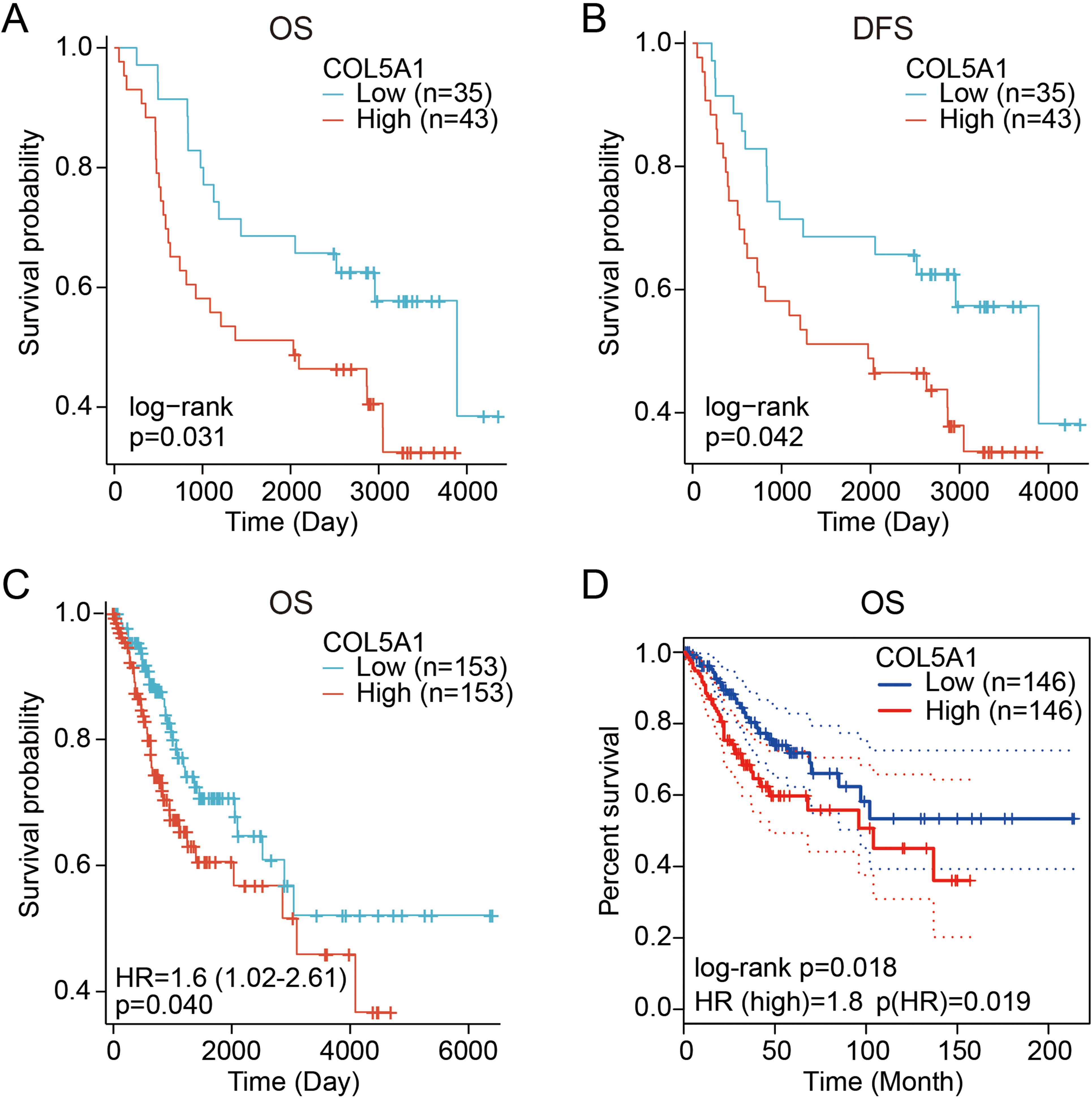
Plots of Kaplan–Meier curves showing the association between COL5A1 expression and survival status of patients with cervical cancer. (a) Overall survival (OS) analysis comparing the high-expression to low-expression of COL5A1 in patients with cervical cancer. Clinical and expression data were obtained from patients with tissue microarray. (b) Disease-free survival (DFS) analysis comparing the high-expression to low-expression of COL5A1 in patients with cervical cancer. Clinical and expression data were obtained from patients with tissue microarray. (c) OS analysis comparing the high-expression to low-expression of COL5A1 in patients with cervical cancer. Clinical and expression data were obtained from the TCGA-CESC dataset and analyzed by R software. (d
Genomic alterations of COL5A1 are associated with cervical cancer progression
Since high levels of COL5A1 protein expression were detected in cervical cancer, subsequently, we measured the frequency of COL5A1 gene alterations using the cBioPortal tool based on DNA sequencing data. The outcome of the analysis showed that the COL5A1 gene was altered in 11 of 278 (4%) cervical cancer patients. The alterations included mutation (3.6%) and deep deletion (0.36%) (Figure 3(a) and 3(b)). In addition, we found that the top 10 genes (TTN, HUWE1, LPR1B, PEAR1, FOXP1, SRP68, CASKIN2, TDRD6, MACF1, and MDN1) with the highest event frequency were differentially expressed between the altered group and the unaltered group (Figure 3(c)). These data indicated that these 10 genes were most likely to be associated with COL5A1 gene alteration. Further analysis of COL5A1 gene alteration on the corresponding protein positions with post translational modifications (PTMs), including phosphorylation, acetylation, ubiquitination, and hydroxylation, in 11 patients with cervical cancer revealed that hydroxylation might have more possibility to expose alterations (Figure 3(d)).
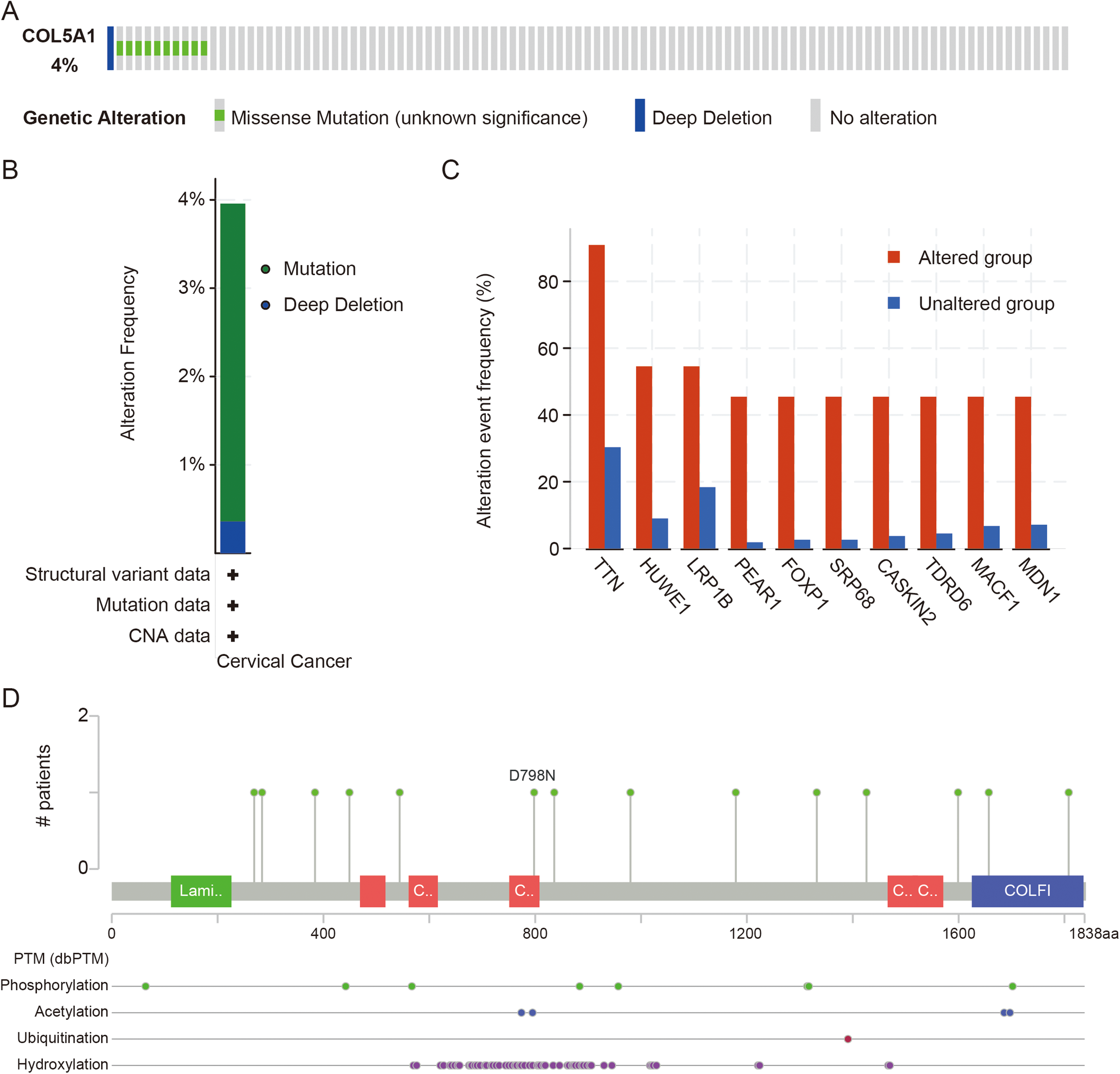
The genetic alterations of COL5A1 in TCGA-CESC patients. (a) OncoPrint summary of alterations on a query of COL5A1. (b) Summary of the alteration frequency derived from mutation and deep deletion in cervical squamous cell carcinoma. (c) Top 10 genes (TTN, HUWE1, LPR1B, PEAR1, FOXP1, SRP68, CASKIN2, TDRD6, MACF1, and MDN1) with the highest event frequency between the altered group and the unaltered group. (d) Eleven patients with COL5A1 gene alterations on the corresponding protein positions; for example, AA change at collagen (collagen triple helix repeat (20 copies) (751–808)): D798N. Green, Laminin_G_2: Laminin G domain (112–226); Red, collagen: Collagen triple helix repeat (20 copies) (469–517), (561–616), (751–808), (1466–1523), (1513–1571); Blue, COLFI: Fibrillar collagen C-terminal domain (1625–1836).
Discussion
The current study evaluates the clinical relevance of COL5A1 in cervical cancer. The value of COL5A1 as a tissue biomarker in cervical cancer and the association of the expression of COL5A1 with clinicopathological features in cervical cancer are also analyzed. COL5A1 tends to be useful as a molecular indicator to predict the prognosis of patients with cervical cancer.
Cervical cancer is an HPV-related disease, 9 but the HPV infection is also associated with other cancers, such as head and neck squamous cell carcinoma.24,25 Therefore, defining alternative biomarkers is important. Our experimental data demonstrated that the expression of the COL5A1 protein was elevated in cervical cancer. The positive expression rate of COL5A1 in cervical cancer tissues was significantly higher than that in their paired adjacent normal tissues. It has been shown that COL5A1 is overexpressed in glioblastoma and oral squamous cell carcinoma,26,27 which correlates with chemoresistance in ovarian and lung cancers16,28 and promotes metastasis in kidney and gastric cancers.29,30 Our previous studies have also shown the overexpression of COL5A1 in breast and ovarian cancers.15,16 However, little is known about COL5A1 expression in cervical cancer. The current study confirmed the overexpression of the COL5A1 protein in cervical cancer detected by IHC. The IHC technique has the advantages of cheap and rapid direct visualization of protein expression and, thus, has a clinical implication for personalized treatment. 31
A previous study on the relationship between clinicopathological characteristics and COL5A1 expression revealed that a high level of COL5A1 expression is correlated with advanced pathological stage and lymph node metastasis in patients with tongue squamous cell carcinoma. 32 However, our study showed that high levels of COL5A1 expression did not relate to the tumor stages. Furthermore, there was no association between high levels of COL5A1 expression and levels of two serum biomarkers such as CA125 and SCC-Ag in cervical cancer. The important clinical significance shown in the current study is that the high levels of COL5A1 expression tend to be in elderly patients.
The present study demonstrated that patients with cervical cancer had short OS and DFS, indicating that COL5A1 overexpression is correlated with poor prognosis in cervical cancer. It has been shown that high expression of COL5A1 is associated with a worse prognosis in patients with kidney cancer and accelerates the growth and metastasis of tumors. 29 The tumor progression and worse prognosis are also observed in gliomas and laryngeal squamous cell cancer.33,34 These data imply that COL5A1 may have the potential to be a prognostic indicator for cervical cancer patients.
Further analyses showed that genomic alterations of COL5A1 were associated with cervical cancer progression. In the present study, we found that the COL5A1 gene was altered in 11 of 278 (4%) cervical cancer patients. Interestingly, the COL5A1 gene was also altered in 32 of 817 (4%) breast cancer patients. 35 By analysis of COL5A1 gene alteration frequency, TTN, HUWE1, LPR1B, PEAR1, FOXP1, SRP68, CASKIN2, TDRD6, MACF1, MDN1 genes were most likely to be associated with COL5A1 gene alteration as they were different with alteration event frequency between the altered group and the unaltered group. The biological function of alterations may be involved in the process of post-translational modifications such as phosphorylation, acetylation, ubiquitination, and hydroxylation. A mechanism study showed that COL5A1 was involved in cancer cell migration and invasion is negatively regulated by miR-29b-3p and miR-582-5p.36,37 Furthermore, COL5A1 expression is mediated by the TGF-β signaling pathway 15 and is associated with metastasis and poor survival in ovarian cancer. 38 COL5A1 is also targeted by non-coding RNA such as long non-coding RNA (LINC00173) in oral squamous cell carcinoma 27 and circular RNA (circACAP2) in breast cancer. 39
There are some limitations in this study. The regulatory mechanism of COL5A1 in cervical cancer remains unclear. The serum levels of COL5A1 have not been measured in cervical cancer patients. The correlation of the serum-COL5A1 with clinicopathological features of cervical cancer has not been evaluated. Therefore, more rigorous studies with larger sample sizes should be performed in the future.
Conclusion
COL5A1 as a tissue biomarker is overexpressed in cervical cancer tissues compared to their paired adjacent normal tissues. High levels of COL5A1 expression are associated with poor prognosis in patients with cervical cancer. This clinical relevance of COL5A1 may provide theoretical and experimental bases to develop clinical applications for the diagnosis and prognosis of cervical cancer.
Footnotes
Acknowledgments
We thank the aforementioned databases for data access.
Author contributions
Xiaoling Duan: Conceptualization, data curation, software, formal analysis, validation, investigation, visualization, writing-original draft. Danjuan Ye, Jia Yuan, Bin Guan, Wencia Guan, Songping Liu, Jingyi Fang, and Qinmei Li: Formal analysis, validation, investigation, methodology, visualization. Jimin Shi and Yan Zhu: Validation, investigation, methodology. Qi Lu and Guoxiong Xu: Conceptualization, resources, data curation, supervision, funding acquisition, project administration, writing-review and editing. All authors read and approved the final manuscript.
Data availability statement
The data presented in this study are contained within the article.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Natural Science Foundation of Shanghai Municipality, Start-up Fund of Research from Jinshan Hospital, Project of the Science and Technology Commission of Shanghai Municipality, (grant number 23ZR1408900, JYQN-LC-202101, 20Y11914100).
Informed consent statement
Informed consent for the human participant was waived because the study was conducted on retrospective pathological tissue samples according to institutional requirements.
Institutional review board statement
The study was reviewed and approved by the Ethics Committee of Jinshan Hospital, Fudan University (No. JYLLKY-2019-01-01).
