Abstract
The relationship between PLIN2 expression and prognosis, and clinicopathological significance of various cancers has been extensively studied, but the results are not completely consistent. This review followed the guidelines for systematic reviews of prognostic factors studies and was reported under the Preferred Reporting Program for Systematic Reviews and Meta-Analysis (PRISMA). We searched PubMed, Embase, Cochrane Library, Web of Science, and Google Academia for relevant articles up to September 2, 2022, and calculated the pooled hazard ratios (HR) with 95% confidence intervals (CI) to determine the association between PLIN2 expression and the prognosis of various cancers. The meta-analysis ultimately included 17 studies. The quality of all included cohort studies was evaluated using the Quality in Prognosis Studies (QUIPS) tool, and an adaptation of Grading of Recommendations Assessment, Development and Evaluation (GRADE) method was used to assess the certainty of the results. High expression of PLIN2 was associated with poorer overall survival (HR = 1.65; 95% CI = 1.14, 2.38; P = 0.008), metastasis-free survival (HR = 1.48; 95% CI = 1.12, 1.94; P = 0.005), progression-free survival (HR = 2.11; 95% CI = 1.55, 2.87; P < 0.0005) and recurrence-free survival/relapse-free survival (HR = 2.21; 95% CI = 1.64, 2.98; P < 0.0005) in cancers. The clinicopathological parameters of digestive system malignancies suggested that high expression of PLIN2 was notably associated with distant metastasis ( + ) (odds ratio (OR) = 3.37; 95% CI = 1.31, 8.67; P = 0.012), lymph node metastasis ( + ) (OR = 1.61; 95% CI = 1.01, 2.54; P = 0.004), and tumor stage (III–IV) (OR = 1.96; 95% CI = 1.24, 3.09; P = 0.006). In summary, overexpression of PLIN2 is significantly associated with a poor prognosis in various human cancers, especially in respiratory and digestive malignancies. Thus, PLIN2 expression may be a potential prognostic biomarker in cancer patients.
Introduction
The latest global cancer statistics, GLOBOCAN-2020, show that there are about 19.3 million cancer cases and nearly 10 million cancer-related deaths each year. 1 Cancer could be the leading cause of premature death in most countries in this century,2,3 in part because approximately 50% of cancers are detected in an advanced stage. No country can eliminate cancer by treatment alone. Large-scale investments to ensure long-term cancer prevention and strategies for early detection, diagnosis, and treatment are fundamental to minimize the cancer burden. 4 The discovery and validation of a useful cancer biomarker can provide sufficient information for early diagnosis to make clinical decisions and achieve a better prognosis for patients. 5
PLIN2, 6 also known as Perilipin 2, Adipophilin, 7 and Adipose Differentiation-Related Protein (ADRP/ADFP),8,9 is a protein-coding gene with a molecular weight of 50 kDa, located on human chromosome 9p22.1, encoding 437 amino acids. 6 PLIN2 exists in lipid droplets (LD) in cells and relate to substances present on the surface of LD and plays a key role in the regulation of a variety of cellular functions. 10 The biophysical properties of cell biofilms determine the efficiency of signal transmission, leading to changes in lipid metabolism pathways, which are closely related to cancer and are the biochemical basis for the carcinogenicity and malignancy of tumor cells. 11 Cancer cells exhibit uncontrollable changes in cellular lipid metabolism, including endogenous lipid synthesis, storage, and degradation, and disruption of exogenous uptake of membrane-forming materials. PLIN2 regulates cell function and lipid metabolism, leading to the transformation of the membrane properties of cancer cells. 12
The in-depth study of PLIN2 found that it is highly expressed in Burkitt lymphoma, 13 colorectal cancer, 14 liver cancer, 15 skin malignant melanoma, 16 gastric carcinoma, 17 and more. Therefore, PLIN2 may affect the carcinogenic mechanism as an oncogene. Conversely, however, the higher expression of PLIN2 in the urine and kidney tissue of patients with clear-cell renal cell carcinoma (ccRCC) represents a better prognosis. 18 Similarly, high expression of PLIN2 in hepatoblastoma (HB) has also been shown to have a better prognosis. 19 Inconsistency in study outcomes has resulted in an indeterminate prognostic value of PLIN2 in various cancers. Therefore, we combined data from all relevant studies in a systematic review and meta-analysis to assess the association of PLIN2 expression with cancer prognosis and clinical pathological characteristics.
Materials and methods
Guidance protocol
This review followed the guidelines for systematic reviews of prognostic factors studies 20 and was reported under the Preferred Reporting Program for Systematic Reviews and Meta-Analysis (PRISMA) 21 (Supplemental material Table S1). We registered our protocol on PROSPERO (National Institute for Health Research, International Prospective Register of Systematic Reviews https://www.crd.york.ac.uk/PROSPERO) with number CRD42022375842.
All the work of this systematic review was carried out by two investigators (ML-L and HY-L) independently, and differences that occurred during the period were resolved by discussion and consultation. If no consensus was reached, a third reviewer (LT-H) was involved in resolving them.
Literature search strategies
We clearly defined the question of systematic review using the PICOTS 20 system (Population, Index prognostic factor, Comparator prognostic factors, Outcome, Timing, Setting) system (Table S2) and comprehensively searched PubMed, Embase, Cochrane Library, Web of Science, and Google Academia for relevant prognostic articles from cohort studies investigating the relationship between PLIN2 expression and clinical outcomes in cancer patients. The endpoint of the search was September 2, 2022. Using a combination of three keywords (and their synonyms) as a search topic, we entered (“plin2” or “PLIN2” or “Perilipin 2” or “ADRP” or “Adipose Differentiation-Related Protein” or “Adipophilin” or “ADFP” or “Perilipin-2”) and (“cancer” or “tumor” or “carcinoma” or “malignancy”) and (“prognos*” or “outcome” or “survival”) in the advanced search engines of various databases to retrieve literature that met the requirements of our study. To avoid omissions caused by the manual retrieval process, we carefully read the relevant reference lists for the included articles.
Inclusion and exclusion criteria
The following inclusion criteria were performed for this systematic review: (a) published studies in English; (b) original articles describing prognostic studies of PLIN2 (or other synonym) expression levels in cancer; (c) articles describing prognostic indicators and/or clinical characteristics of cancer; and (d) hazard ratio (HR) value and 95% confidence interval (CI) can be directly extracted from the text or indirectly calculated through the Kaplan–Meier (K–M) survival curve; odds ratio (OR) value, and 95% CI can be directly extracted from the text or calculated by extracting the corresponding sample size of the experimental group and the control group in the text.
This study complied with the following exclusion criteria: (a) non-English language publications; (b) non-prognostic studies; (c) lack of prognostic data; (d) animal studies; (e) reviews, case reports, letters, editorials, meta-analyses, and conference reports; and (f) repeated data articles with smaller sample sizes.
Data extraction and quality assessment
We performed extraction using CHARMS-PF 20 (a modification of CHARMS 22 (checklist for critical appraisal and data extraction for systematic reviews of prediction modelling studies)) checklist for prognostic factors. The main information extracted from eligible studies was: first author, year, country, data sources, genetic-testing methods, cutoff value for PLIN2 expression, tumor type, tumor stage, age, outcome measures, sample sizes (high expression/ low expression), HR (95% CI) with its P value, follow-up months, overall risk of bias of QUIPS (Quality in Prognosis Studies) tool, and clinicopathological parameters. If the HR (95% CI) value was not directly available in the included study text, the survival data were extracted from the K–M curve by Engauge Digitizer software (produced by Mark Mitchell et al., Webpage: http://markummitchell.github.io/engauge-digitizer/) and then the Tierney's method 23 was used to calculate the HR (95% CI) value.
The quality of all included cohort studies was evaluated using the QUIPS tool, 24 which includes six domains for bias and applicability: study participation, study attrition, prognostic factor measurement, outcome measurement, adjustment for other prognostic factors, and statistical analysis and reporting. The quality of individual study reporting was assessed using the Reporting Recommendations for Tumor Marker Prognostic Studies (REMARK) checklist 25 and informed the QUIPS assessments. We defined the overall risk of bias as: (a) low: low risk of bias in all domains or moderate risk in only one domain; (b) moderate: two to four domains were considered to have moderate risk of bias; and (c) high: at least five domains are considered moderate risk or at least one domain is high risk.
Statistical analysis
Stata 15.0 software (STATA Corporation, College Station, TX, USA) was chosen for this review. The relationship between PLIN2 expression and each prognostic indicator was represented by the respective HR and 95% CI, respectively. Additionally, OR and 95% CI were appropriately used to assess clinicopathological parameters. Subgroup analyses were cleverly performed to explore sources of heterogeneity. The Higgins I² test and the Cochran Q test were used to assess heterogeneity. 26 Publication bias was assessed using funnel plots, and if significant asymmetry was revealed, the Begg test and Egger test were used for further validation. Sensitivity analyses were performed to verify the robustness of the results by omitting each study in turn. Statistical significance was set at P < 0.05.
Confidence in summary results
An adaptation of Grading of Recommendations Assessment, Development and Evaluation (GRADE) instrument, which is based on study design, risk of bias, inconsistency, indirectness, imprecision, and other considerations, was used to examine confidence in the main summary results of systematic reviews of prognostic factor research. 27 The quality of evidence was assessed through a guideline development tool (http://gradepro.org) and ultimately graded as high, moderate, low, or very low.
Results
Study identification and characteristics
Details of the search strategy for all database (Table S3) and screening processes are shown in Figure 1. Twenty-four articles were evaluated in full text, and we eliminated seven after consultation. Five studies14,28–31 had no survival data, one 32 included patients with non-cancer, and one 33 included duplicate data. Ultimately, 17 articles were included. Four studies were conducted in China, 10 in Japan and 1 each in the United States, Germany, and South Korea. There were 4848 patients with sample sizes ranging from 45 to 1402. The included studies used quantitative real-time-polymerase chain reaction or immunohistochemistry to detect the expression level of PLIN2, and all patients were divided into high or low PLIN2 expression groups by cutoff value. In the included studies, there were 10 different malignancies: breast cancer,34–38 ccRCC,18,39,40 cutaneous malignant melanoma, 41 HB, 19 hepatocellular carcinoma, 42 high-grade serous ovarian carcinoma, 43 lung adenocarcinomas,44,45 oral squamous cell carcinoma, 46 pancreatic ductal adenocarcinoma, 47 and salivary duct carcinoma. 48 The result of analysis of survival data from the included studies were obtained following patients for 40–300 months. Analysis of survival data results in the form of prognostic indicators contained 10 overall survival (OS), 4 disease-free survival (DFS), 3 disease-specific survival (DSS), 3 metastasis-free survival (MFS), 3 progression-free survival (PFS), and 5 recurrence-free survival/relapse-free survival (RFS). The main characteristics of all included articles are summarized in Table 1.
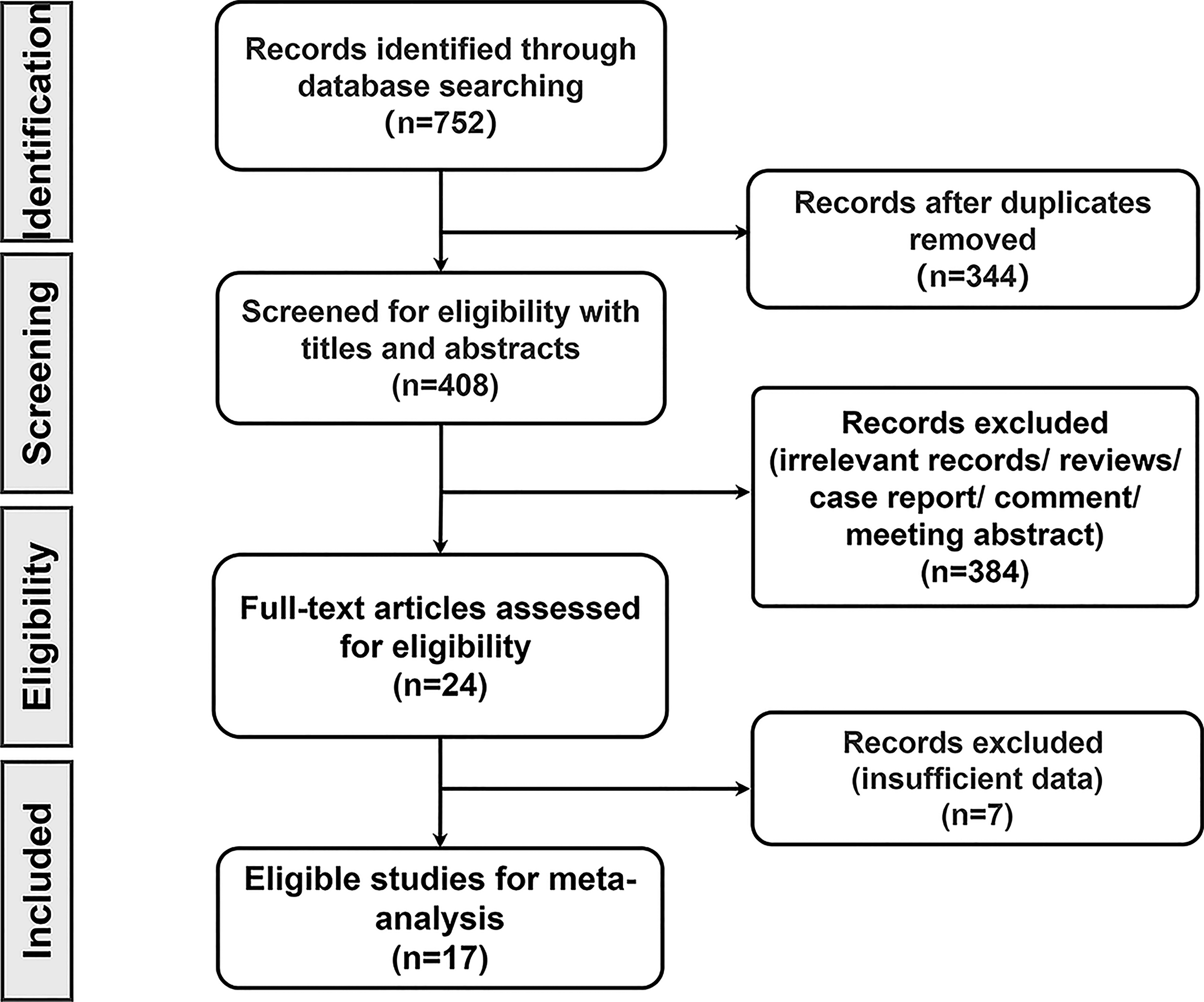
Flow diagram of literature search.
The main characteristics of eligible studies.
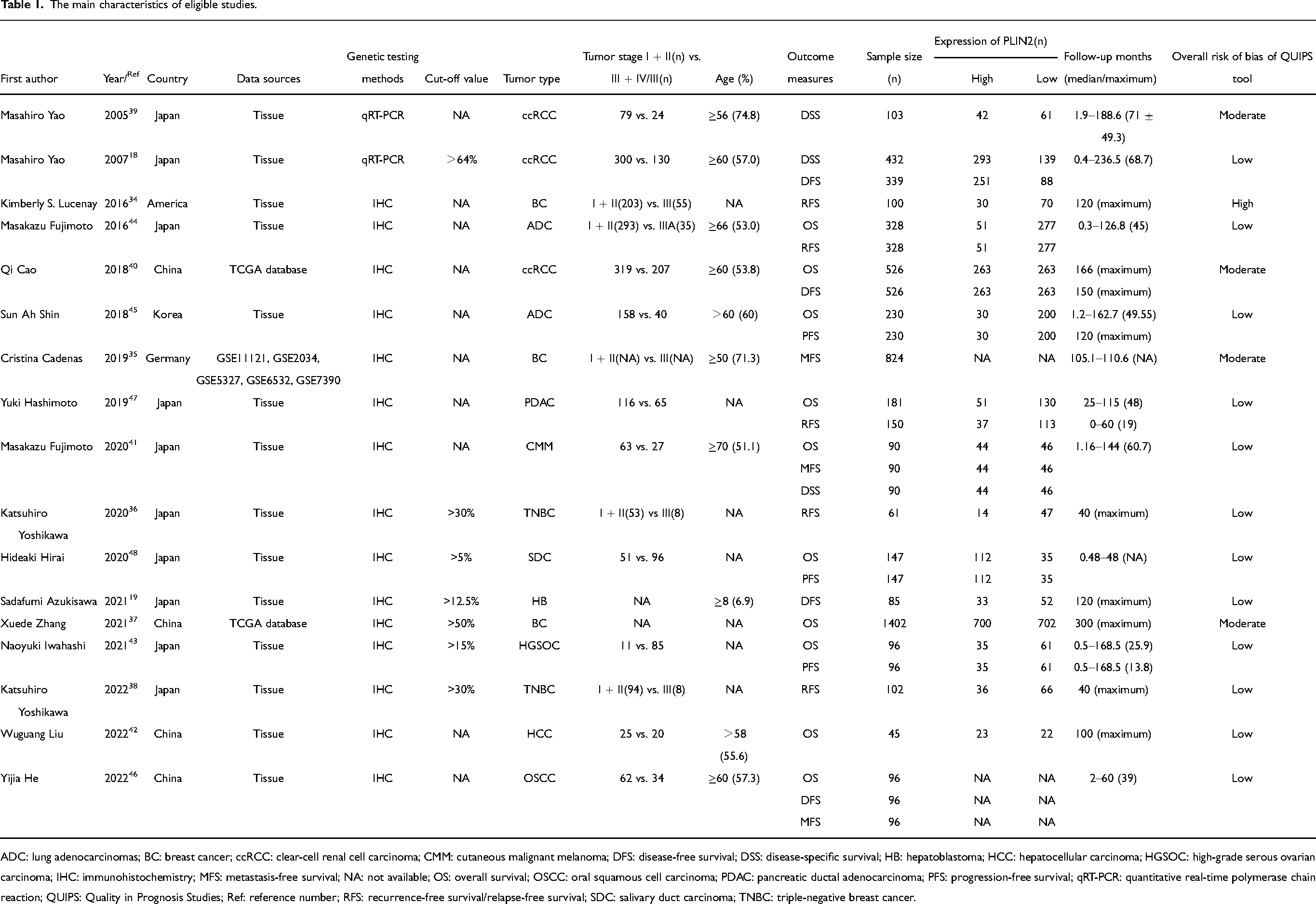
ADC: lung adenocarcinomas; BC: breast cancer; ccRCC: clear-cell renal cell carcinoma; CMM: cutaneous malignant melanoma; DFS: disease-free survival; DSS: disease-specific survival; HB: hepatoblastoma; HCC: hepatocellular carcinoma; HGSOC: high-grade serous ovarian carcinoma; IHC: immunohistochemistry; MFS: metastasis-free survival; NA: not available; OS: overall survival; OSCC: oral squamous cell carcinoma; PDAC: pancreatic ductal adenocarcinoma; PFS: progression-free survival; qRT-PCR: quantitative real-time polymerase chain reaction; QUIPS: Quality in Prognosis Studies; Ref: reference number; RFS: recurrence-free survival/relapse-free survival; SDC: salivary duct carcinoma; TNBC: triple-negative breast cancer.
Quality assessment
The overall risk of bias of each study assessed by the QUIPS tool is presented in Table 1. Twelve studies were at low risk of bias, four studies were at moderate risk of bias, and one study was considered at high risk. Details of the risk of bias in each domain are provided in Table S4.
Relationship between high expression of PLIN2 and OS
Data from 3141 patients in 10 studies were used to calculate HR and 95% CI of OS. Pooled HR indicated that patients with high expression of PLIN2 had a significantly worse prognosis (pooled HR = 1.65; 95% CI = 1.14, 2.38; P = 0.008, random effects model) (Figure 2(a)).
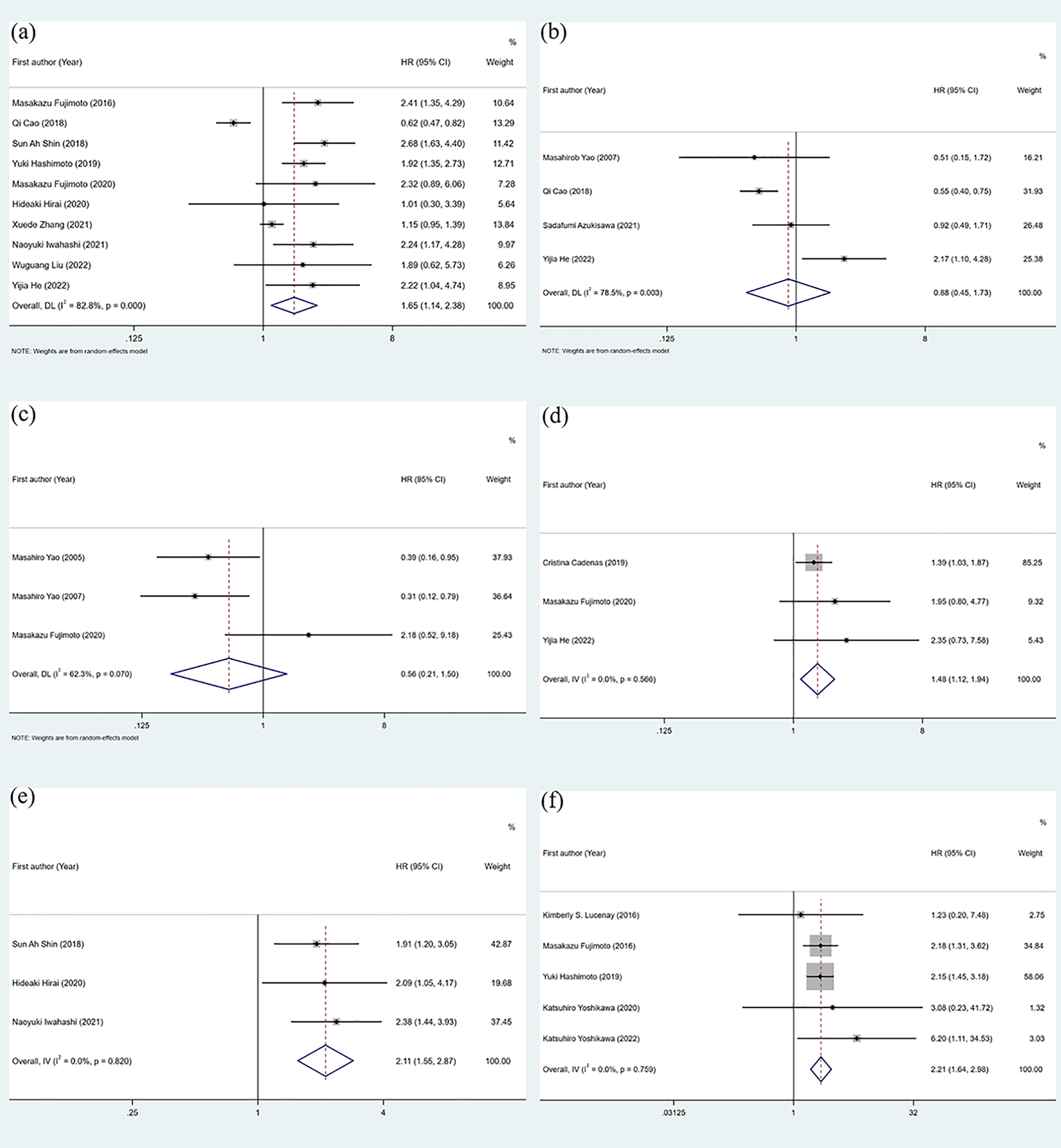
Forest plots of studies evaluating the association between PLIN2 expression with (a) overall survival (os); (b) disease-free survival (DFS); (c) disease-specific survival (DSS); (d) metastasis-free survival (MFS); (e) progression-free survival (PFS); and (f) recurrence-free survival/relapse-free survival (RFS).
Subgroup analysis for OS
To find out which studies caused heterogeneity, we performed subgroup analysis of cancer type (Figure 3(a)) and sample size (Figure 3(b)). Subgroup analysis showed that PLIN2 overexpression in the respiratory system cancer (pooled HR = 2.56; 95%CI = 1.76, 3.73; P < 0.0005, fixed effects model) and the digestive system cancer (pooled HR = 1.89; 95% CI = 1.40, 2.54; P = 0.000, fixed effects model) had a significantly worse prognosis, while in other systems cancer (pooled HR = 1.01; 95% CI = 0.87, 1.18; P = 0.877, random effects model) expression of PLIN2 was not associated with a significant difference in OS. After stratification by sample size, whether the sample size was <150 patients (pooled HR = 2.03; 95% CI = 1.38, 2.99; P < 0.0005, fixed effects model), or ≥150 patients (pooled HR = 1.19; 95% CI = 1.04, 1.36; P = 0.012, fixed effects model), high expression of PLIN2 and poor patient prognosis showed both a significant relationship.
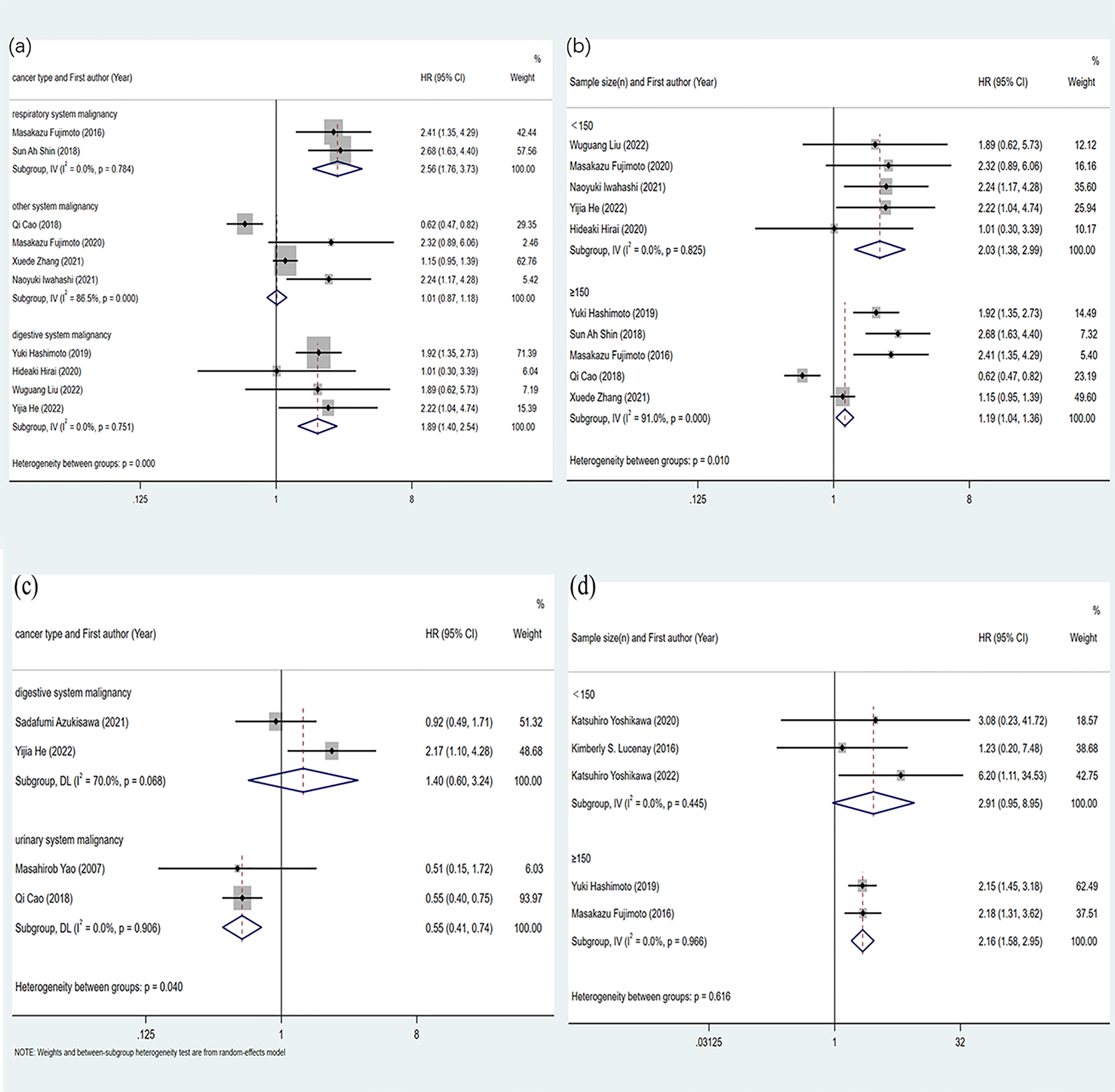
Forest plots of the subgroup analysis evaluating HRs of PLIN2 for overall survival (OS) by the factors of (a) cancer type; and (b)sample size. Forest plots of the subgroup analysis evaluating HRs of PLIN2 for (c) disease-free survival (DFS) by the factors of cancer type; and (d) recurrence-free survival/relapse-free survival (RFS) by the factors of sample size.
Relationship between high expression of PLIN2 and other outcome measures
Other outcome measures include DFS, DSS, MFS, PFS, and RFS. Four studies which included 1046 patients assessed the HR and 95% CI of DFS (Figure 2(b)), and the results (pooled HR = 0.88; 95% CI = 0.45, 1.73; P = 0.715, random effects model) showed that the expression of PLIN2 had no statistical significance on cancer patients. A total of 625 patients were included in three studies that evaluated DSS (Figure 2(c)), and the results were also not statistically significant (pooled HR = 0.56; 95% CI = 0.21, 1.50; P = 0.247, random effects model). In addition, three studies analyzed MFS (pooled HR = 1.48; 95% CI = 1.12, 1.94; P = 0.005, fixed effects model), including 1010 patients, the results of which suggested that high expression of PLIN2 was significantly associated with poor MFS (Figure 2(d)). Similarly, three studies were performed to assess PFS in 473 patients (Figure 2(e)), and the results (pooled HR = 2.11; 95% CI = 1.55, 2.87; P < 0.0005, fixed effects model) indicated a significant correlation between high expression of PLIN2 and poor PFS. Finally, five studies that included 741 patients assessed RFS (pooled HR = 2.21; 95% CI = 1.64, 2.98; P < 0.0005, fixed effects model), which also suggested a significant association between high expression of PLIN2 and poor RFS of patients (Figure 2(f)).
Subgroup analysis for DFS and RFS
Similarly, we further optimized the predictive power of PLIN2 expression on patient prognostic outcomes by subgroup analysis for DFS and RFS. First, the DFS group was divided into digestive system malignancy and urinary system malignancy according to tumor types. The results in the digestive system malignancy group (pooled HR = 1.40; 95% CI = 0.60, 3.24; P = 0.436, random effects model) showed that high expression of PLIN2 was not significantly associated with DFS. In the group of urologic malignancy, high expression of PLIN2 was associated with better DFS (pooled HR = 0.55; 95% CI = 0.41, 0.74; P < 0.0005, fixed effects model) (Figure 3(c)). Second, in the analysis of the RFS sample size subgroup, the results of the group of <150 patients showed that the expression level of PLIN2 was not significantly correlated with RFS (pooled HR = 2.91; 95% CI = 0.95, 8.95; P = 0.062, fixed effects model). The results for groups with sample sizes ≥150 indicate that high expression of PLIN2 was significantly associated with poor RFS (pooled HR = 2.16; 95% CI = 1.58, 2.95; P < 0.0005, fixed effects model) (Figure 3(d)).
Relationship between PLIN2 expression and clinicopathological characteristics
We extracted clinicopathological parameters from the included studies, including age, distant metastasis, gender, lymph node metastasis, and tumor differentiation (Table S5 and Figure S1). Due to the large heterogeneity of the data, random effects models were used in all cases. Our findings showed that high expression of PLIN2 was not significantly associated with age (older vs. younger) (pooled OR = 1.32; 95% CI = 0.95, 1.83; P = 0.095), distant metastasis ( + vs. −) (pooled OR = 1.84; 95% CI = 0.76, 4.44; P = 0.177), gender (male vs. female) (pooled OR = 0.90; 95% CI = 0.55, 1.48; P = 0.685), lymph node metastasis ( + vs. −) (pooled OR = 1.16; 95% CI = 0.73, 1.83; P = 0.528) and tumor stage (GIII–IV vs. GI–II) (pooled OR = 1.52; 95% CI = 0.89, 2.57; P = 0.123).
Considering the heterogeneity of the pan-cancer data sources, we separately analyzed the related data of digestive system malignancies to further explore the correlation of clinicopathological parameters with a high expression of PLIN2. Due to the low heterogeneity, we used fixed effects models. The results (Table S5 and Figure S2) showed that high expression of PLIN2 was not significantly associated with age (older vs. younger) (pooled OR = 1.09; 95% CI = 0.56, 2.12; P = 0.808), and gender (male vs. female) (pooled OR = 0.93; 95% CI = 0.59, 1.45; P = 0.735). The results suggested that high PLIN2 was notably associated with distant metastasis ( + ) (pooled OR = 3.37; 95% CI = 1.31, 8.67; P = 0.012), and lymph node metastasis ( + ) (pooled OR = 1.61; 95% CI = 1.01, 2.54; P = 0.004) and tumor stage (GIII–IV) (pooled OR = 1.96; 95% CI = 1.24, 3.09; P = 0.006).
Publication bias and sensitivity analysis
Publication bias in the included studies was detected by drawing a funnel plot or by further performing Begg and Egger tests. For OS, the shape of the funnel plot showed asymmetry (Figure 4(a)), but the P-values of Begg test (Figure 4(b)) and Egger test (Figure 4(c)) were 0.371 and 0.120, respectively, indicating that PLIN2 expression does not have a significant publication bias for patients’ OS. Likewise, for DFS (Figure S3(a)), DSS (Figure S3(b)), MFS (Figure S3(c)), PFS (Figure S3(d)), and RFS (Figure S3(e)), the points on both sides of the funnel plot were evenly distributed, suggesting that there is no apparent publication bias between PLIN2 expression and these metrics.
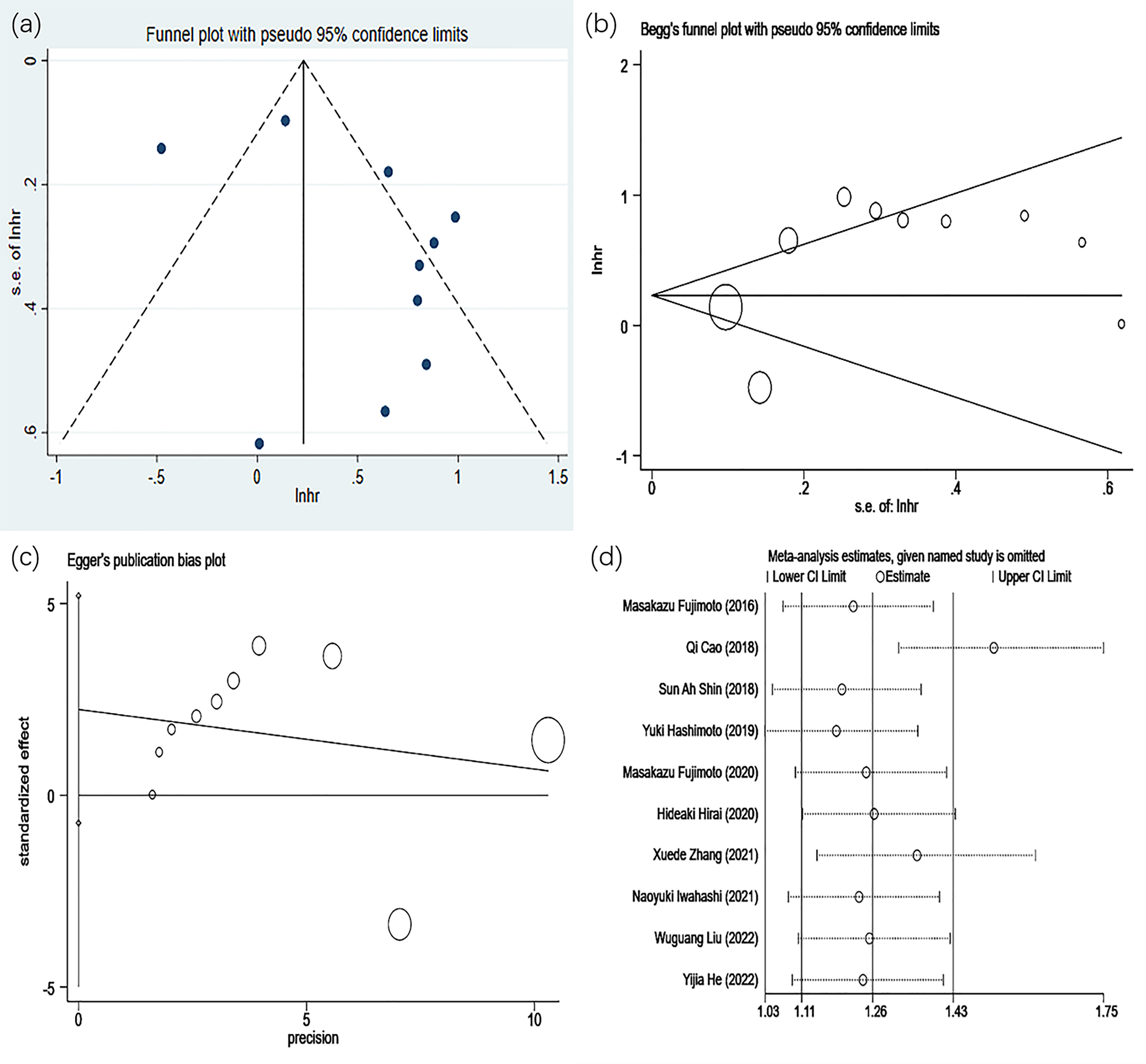
The funnel plot of publication bias of OS. (a) Funnel plot. (b) Begg's funnel plot. (c) Egger's publication bias plot. (d) Sensitivity analysis of the included studies concerning PLIN2 and OS.
To examine the effect of individual studies on the overall pooled results, a sensitivity analysis was performed to test the robustness of the results by sequentially excluding individual studies. In OS (Figure 4(d)) and DFS (Figure 4S(a)), when the Cao et al.40 study was removed, a more significant effect was found in the pooled HR, suggesting that this study may make the aggregated HR less robust However, in DSS (Figure S4(b)), MFS (Figure S4(c)), PFS (Figure S4(d)), and RFS (Figure S4(e)), when all studies were sequentially deleted, no significant effect was found, suggesting that pooled HR is reliable.
Quality of evidence
The results of the GRADE instrument showed that the confidence of the main results of this systematic review was moderate to high, indicating that the estimated effect was close to the true effect, but there is a possibility that it is substantially different (Table S6).
Discussion
Analysis of results
In this review, the combined results showed that the high expression of PLIN2 in cancer patients was negatively correlated with OS, MFS, PFS, and RFS. Further subgroup analysis showed that, for OS, high expression of PLIN2 was associated with a worse prognosis in both the subgroup of the respiratory and digestive system malignant tumor and the subgroup of the large or small sample size. For RFS, high expression of PLIN2 in the subgroup with large sample size showed worse RFS. However, there was no significant correlation between the expression of PLIN2 and DFS and DSS. Notably, however, a subgroup of urologic malignancies showed that high expression of PLIN2 was inversely associated with bad DFS. Besides, the expression of PLIN2 in pan-cancer was not significantly associated with clinicopathological features (age, distant metastasis, gender, lymph node metastases, and tumor grade). However, in gastrointestinal malignancies, high PLIN2 expression was associated with distant metastases ( + ), lymph node metastasis ( + ), and stage III–IV tumors.
A key feature of lipogenic metabolism in tumor biology is the maintenance of sustained cell proliferation, and the metabolic mechanisms may differ across tumor types. PLIN2 is found at the start of LD formation and is upregulated concurrently with lipid storage. 49 It exerts tumorigenic or tumor suppressive effects in a tumor-dependent manner. 46 As mentioned above, PLIN2 is associated with a poor prognosis in most cancers. However, other studies, as well as our systematic review and meta-analysis, suggest that inhibition of PLIN2 promotes tumor cell proliferation, tissue entry, and distant metastasis in ccRCC, uterine leiomyoma, 29 and HB. First, the LD content in cancer cells is much higher than that in normal cells 50 because the proliferative metabolism of cancer cells requires an ever-expanding lipid substrate and a continuous energy source to increase the LD content. LDs as energy reservoirs can trigger the development of metastatic clones in aggressive cancers. 51 PLIN2 is the only constitutive LD protein involved in LD synthesis. 52 This reveals the mechanism by which PLIN2 promotes the development of advanced cancer and promotes the metastasis of cancer cells. Second, autophagy triggers intracellular degradation, a process that inhibits tumor cell proliferation, 53 and decreased autophagy increases cancer cell proliferation. 54 It has been shown that the activity of the intracellular AMPK/ULK1 pathway can be decreased by PLIN2 to inhibit autophagy or prevent autophagy by PLIN2 through other mechanisms that have not yet been elucidated.42,55 Additionally, advanced solid tumors are more likely to suffer from hypoxia than normal tissues because of their low blood supply. Tumor cells can adapt to hypoxic conditions by adjusting metabolism and growth mechanisms, which are largely influenced by the hypoxia-inducible factor (HIF) 1 pathway. 56 The glycolysis and glucose uptake processes are activated by HIF1α protein, causing cancer cell proliferation. Some studies have observed that PLIN2 promotes the HIF1 signaling pathway by accelerating the production of the HIF1α protein. 42 It has also been shown that choline kinases (CHKs) phosphorylate PLIN2 to dissociate it from LDs and to promote lipolysis of LD, β-oxidation and tumor growth.31,57
Conversely, studies have shown that the expression of PLIN2 is higher in low-stage, low-grade, or von Hippel–Lindau (VHL) alteration-positive ccRCC. 18 A study found that the prognosis of patients with ccRCC was better when the VHL mutation was detected in tumor cells, and VHL mutation may induce the overexpression of PLIN2, which may explain the low invasiveness of tumors overexpressing PLIN2.18,33 Alterations in the VHL pathway in ccRCC are based on hypoxia, which further induces expression of PLIN2.58,59 Furthermore, overexpression of PLIN2 has been shown to lead to lipid overload. 49 It has been shown that the protective overexpression of PLIN2 is a cellular defense mechanism against lipid overload, which can cause lipid toxicity and even cell apoptosis. 60 Therefore, PLIN2 may attenuate renal cell carcinoma (RCC) cell migration and invasion, acting as an anti-cancer gene. 40 Similarly, in HB, in hyperproliferative or chemotherapy-induced stressful environments, increased PLIN2-induced LDs leads to dysregulation of autophagy, enhanced tumor cell apoptosis, and ultimately a better prognosis. 19 Other studies have shown that in uterine leiomyomas, PLIN2 gene expression is significantly reduced. When PLIN2 was knocked down, cell oxygen consumption and extracellular environment acidification were significantly increased. This results in a more vigorous metabolic basal activity and stress response of the cells, with a significant increase in proliferation. 29 In conclusion, the role of PLIN2 in tumor lipid biology should continue to be actively explored to explain the above paradoxical pathogenesis. Exploring the signaling pathways regulated by PLIN2 in various cancer models is an important verification direction in the future.
Important reasons for the heterogeneity
First, in order to explore the source of heterogeneity between studies, we performed subgroup analysis and sensitivity analysis. In the subgroup analysis, we found that the HR (95% CI) value of the Cao et al. 40 study in tumor subgroups of other systems showed better OS correlation with RCC. However, the HR values of the other three studies in this subgroup are all >1. Although the results are not statistically significant, it also indicates to some extent that the high expression of PLIN2 has a greater tendency to negatively correlate with OS. At the same time, we also performed a subgroup analysis in DFS, and the pooled results of two studies (both ccRCC) in the subgroup of urological malignancies suggested that the high expression of PILN2 was associated with better prognostic outcomes. In addition, in the sensitivity analysis for OS, our pooled results were statistically significant regardless of which study was deleted. However, we observed a significant increase in the pooled HR (95% CI) value when the study of Cao et al. 40 was deleted. In summary, we can reasonably suspect that studies of ccRCC are an important source of data heterogeneity. Second, subgroup analysis also found that studies with small sample sizes may also lead to heterogeneity in studies. Third, the analysis of PLIN2 and clinicopathological features indicated that in the pan-cancer analysis, various cancers may have different mechanisms for the occurrence and development of various clinicopathological features, and may also lead to heterogeneity. Our analysis of the association of digestive malignancies with clinicopathological features can counterevidence this point. Likewise, inconsistencies in study inclusion criteria and data collection processes across cancers, among other things, make it impossible to unify all confounding factors, which may be an unavoidable source of heterogeneity.
New discoveries and trends
Research on PLIN2 has never been interrupted in recent years, and current studies not only demonstrate that PLIN2 expression correlates with the prognosis of cancer patients, but can also be used as a diagnostic marker for early cancer detection. 14 Recent research by Rios Garcia et al. 61 demonstrated that PLIN2 levels can validate the responsiveness of tumor-targeting drugs. Additionally, studies have shown that PILN2 can serve as a biomarker for multiple age-related metabolic syndromes 62 such as fatty liver, diabetes, cardiovascular disease, and sarcopenia, and more. Therefore, whether PLIN2 can be regarded as a potential pharmacological target against one or more of the above diseases could be considered. In addition, Jung et al. 32 demonstrated that PLIN2 positivity was an independent factor for shorter OS and DFS in patients with a breast phyllodes tumor. The study by Tang et al. 28 demonstrated that PLIN2 is significantly associated with poor survival in patients with non-small cell lung cancer. Studies by Hoffman et al. 30 suggest that elevated PLIN2 can be detected in liposarcoma, etc. This gives us full confidence that PLIN2 can be used as an effective biomarker to indicate the prognosis of most cancer patients. However, our study also found that patients with high expression of PLIN2 in ccRCC had a better prognosis. In addition to the three articles on ccRCC included in this paper, the studies by Tolkach et al. 33 and Okeigwe et al. 29 came to the same conclusion. Because the current research data on the relationship between PLIN2 and RCC is insufficient, we were unable to conduct a further integrated analysis. More large-scale cohort studies are needed to explore the role of PLIN2 in different cancers. Other noteworthy studies have shown that urine PLIN2 rapid dipstick test, which is an economical, rapid, and promising test, can be used as a primary screening test for RCC, but the diagnostic efficacy and clinical practicability of the biomarker need to be further verified.63,64 Efforts to develop truly non-invasive biomarkers for the diagnosis of early-stage renal cancer may be a key future direction.
Strengths and weaknesses
We conclude this first systematic review and meta-analysis of the association between PLIN2 and pan-cancer by strictly adhering to the recommendations of the guidelines for the meta-analysis of prognostic studies and by using the GRADE tool to assess the credibility of the overall results. To ensure that relevant studies were not omitted, we conducted multiple literature searches. In order to ensure the authenticity and accuracy of the data, we carefully read the full texts and the data in a number of proof-reading. Through reasonable subgroup analysis, we have come to more comprehensive and detailed conclusions. We have also achieved a satisfactory result. However, the study has some limitations. First, the adoption of different cutoff values between studies resulted in different positive correlation rates for PLIN2, the presence of confounding factors between various variables that could not be controlled uniformly, and so on—all of which may have introduced heterogeneity. Second, we excluded studies that were not published in English, which may have contributed to selection bias. Third, the concentration of research in Asia limits the global applicability of our findings.
Conclusion
Overexpression of PLIN2 is significantly associated with a poor prognosis in various human cancers, especially in respiratory and digestive malignancies. In addition, in malignant tumors of the digestive system, attention should be paid to monitoring the expression of PLIN2 in patients with advanced tumors, lymph node metastases, and distant metastasis in order to pursue early intervention. Thus, PLIN2 expression may be a potential prognostic biomarker in cancer patients.
Supplemental Material
sj-docx-1-jbm-10.1177_03936155221147536 - Supplemental material for Prognostic and clinicopathologic significance of PLIN2 in cancers: A systematic review with meta-analysis
Supplemental material, sj-docx-1-jbm-10.1177_03936155221147536 for Prognostic and clinicopathologic significance of PLIN2 in cancers: A systematic review with meta-analysis by Ming-Lin Li, Han-Yong Luo, Zi-Wei Quan, Le-Tian Huang and Jia-He Wang in The International Journal of Biological Markers
Supplemental Material
sj-docx-2-jbm-10.1177_03936155221147536 - Supplemental material for Prognostic and clinicopathologic significance of PLIN2 in cancers: A systematic review with meta-analysis
Supplemental material, sj-docx-2-jbm-10.1177_03936155221147536 for Prognostic and clinicopathologic significance of PLIN2 in cancers: A systematic review with meta-analysis by Ming-Lin Li, Han-Yong Luo, Zi-Wei Quan, Le-Tian Huang and Jia-He Wang in The International Journal of Biological Markers
Supplemental Material
sj-docx-3-jbm-10.1177_03936155221147536 - Supplemental material for Prognostic and clinicopathologic significance of PLIN2 in cancers: A systematic review with meta-analysis
Supplemental material, sj-docx-3-jbm-10.1177_03936155221147536 for Prognostic and clinicopathologic significance of PLIN2 in cancers: A systematic review with meta-analysis by Ming-Lin Li, Han-Yong Luo, Zi-Wei Quan, Le-Tian Huang and Jia-He Wang in The International Journal of Biological Markers
Supplemental Material
sj-docx-4-jbm-10.1177_03936155221147536 - Supplemental material for Prognostic and clinicopathologic significance of PLIN2 in cancers: A systematic review with meta-analysis
Supplemental material, sj-docx-4-jbm-10.1177_03936155221147536 for Prognostic and clinicopathologic significance of PLIN2 in cancers: A systematic review with meta-analysis by Ming-Lin Li, Han-Yong Luo, Zi-Wei Quan, Le-Tian Huang and Jia-He Wang in The International Journal of Biological Markers
Supplemental Material
sj-docx-5-jbm-10.1177_03936155221147536 - Supplemental material for Prognostic and clinicopathologic significance of PLIN2 in cancers: A systematic review with meta-analysis
Supplemental material, sj-docx-5-jbm-10.1177_03936155221147536 for Prognostic and clinicopathologic significance of PLIN2 in cancers: A systematic review with meta-analysis by Ming-Lin Li, Han-Yong Luo, Zi-Wei Quan, Le-Tian Huang and Jia-He Wang in The International Journal of Biological Markers
Supplemental Material
sj-docx-6-jbm-10.1177_03936155221147536 - Supplemental material for Prognostic and clinicopathologic significance of PLIN2 in cancers: A systematic review with meta-analysis
Supplemental material, sj-docx-6-jbm-10.1177_03936155221147536 for Prognostic and clinicopathologic significance of PLIN2 in cancers: A systematic review with meta-analysis by Ming-Lin Li, Han-Yong Luo, Zi-Wei Quan, Le-Tian Huang and Jia-He Wang in The International Journal of Biological Markers
Supplemental Material
sj-docx-7-jbm-10.1177_03936155221147536 - Supplemental material for Prognostic and clinicopathologic significance of PLIN2 in cancers: A systematic review with meta-analysis
Supplemental material, sj-docx-7-jbm-10.1177_03936155221147536 for Prognostic and clinicopathologic significance of PLIN2 in cancers: A systematic review with meta-analysis by Ming-Lin Li, Han-Yong Luo, Zi-Wei Quan, Le-Tian Huang and Jia-He Wang in The International Journal of Biological Markers
Footnotes
Acknowledgments
Not applicable.
Author contributions
The final manuscript has been read and approved by all authors. JH-W, LT-H, and ML-L proposed this study after considerable deliberation. ML-L and HY-L gathered and integrated data for meta-analysis. ML-L, HY-L, ZW-Q, LT-H, and JH-W collaborated to compile the manuscript. ML-L and HY-L have contributed equally to this work and share first authorship
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was funded by grant number 2018YFC2002100, by National Key Research and Development Program of China, grant number 2018YFC2002104 and by Key Research and Development Program of Liaoning Province, grant number 2019JH8/10300021.
Ethical approval and informed consent
Not applicable.
Consent to publish
Not applicable.
Data availability
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
