Abstract
Background
Fidgetin-like 1 (FIGNL1) participates in tumor resistance by playing the function of homologous recombination repair(HRR). However, the role of FIGNL1 in non-small cell lung cancer (NSCLC) is still unclear. This study aims to understand the expression of FIGNL1 in NSCLC and preliminarily explore its relationship with cisplatin resistance.
Methods
FIGNL1 messenger RNA (mRNA) was analyzed in 1018 NSCLC tissues and 111 adjacent tissues using The Cancer Genome Atlas program. FIGNL1mRNA in cisplatin-resistant and cisplatin-sensitive cell lines was analyzed by the Gene Expression Omnibus project. FIGNL1 protein was detected in 58 NSCLC tissues and 58 adjacent tissues by immunohistochemistry. The relationship between FIGNL1, clinical pathological characteristics and disease-free survival was retrospectively analyzed. Gene ontology was used to analyze the biological process mainly involving FIGNL1, and STRING online constructed its protein interaction network and screened the key genes (hub genes).
Results
The Cancer Genome Atlas showed that FIGNL1mRNA was higher in 1018 NSCLC tissues than in 111 adjacent tissues (P < 0.05). In the dataset “GSE157692,” FIGNL1mRNA was higher in cisplatin-resistant cell lines (P = 3.80e-05). The hub genes in FIGNL1 and HRR directions are RAD51 and CCDC36. Immunohistochemistry showed that the FIGNL1 protein in 58 NSCLC tissues was higher than that in 58 adjacent tissues (P < 0.01). FIGNL1 is associated with gender, histopathological type, and nerve invasion in NSCLC. The disease-free survival in NSCLC patients with high FIGNL1 expression was shorter (P = 0.032).
Conclusion
FIGNL1 is associated with poor prognosis in NSCLC, and cisplatin resistance may be involved. These observations provide a clinical basis for exploring FIGNL1 as a potential biomarker for cisplatin resistance in NSCLC.
Introduction
According to the 2021 global cancer burden data, lung cancer is the second-highest morbidity and mortality malignant tumor, which seriously threatens human health and life. 1 Histologically, lung cancer can be divided into small cell lung cancer and non-small cell lung cancer (NSCLC). NSCLC accounts for 80–85% of lung cancer histological types. NSCLC is comprised of approximately 40% adenocarcinoma and 25% squamous cell carcinoma, and the remaining histological types account for 35%. 2 Chemotherapy is one of the main treatments of lung cancer, and cisplatin is the cornerstone of NSCLC chemotherapy.3 However, the 2-year and 5-year survival rates of NSCLC after adjuvant chemotherapy were only 70.3% and 44.5%, respectively. 4 Cisplatin resistance was one of the reasons for treatment failure.
FIGNL1 is a member of the AAA-ATPase protein family involved in the depolymerization of multiple protein complexes. 5 The overexpression of FIGNL1 activates the HRR pathway, which can lead to chemotherapy resistance and contribute to the rapid progress and recurrence of SCLC and its poor prognosis. 6 FIGNL1 promotes cisplatin resistance through its homologous recombination repair (HRR) function of cisplatin-induced tumor cells. 6 However, it is unclear whether FIGNL1 participates in the development of NSCLC and its function affects the sensitivity of NSCLC to platinum-based chemotherapy. This study explores the correlation between FIGNL1 expression in NSCLC and clinical–pathological characteristics and cisplatin resistance.
Material and methods
Bioinformatics analysis
FIGNL1 expression in NSCLC was analyzed by UALCLN 7 (http://ualcan.path.uab.edu). We retrieved survival analysis in the Kaplan–Meier Plotters database 8 (http://kmplot.com/analysis). We analyzed the correlation between FIGNL1 expression and cisplatin resistance by Gene Expression Omnibus (GEO) 9 (http://www.ncbi.nlm.nih.gov/geo). Gene ontology analysis of FIGNL1 expression was performed by Metascape 10 (http://metascape.org/gp/). The FIGNL1 protein interaction network was constructed by STRING v11.0 11 (https://string-db.org/). The minimum interaction score was set at 0.4, and the maximum number was set at 20. We downloaded the protein interaction data and screened the Hub gene by Cystoscope 3.8.
Patients and clinical data
We retrospectively collected clinical data and tissue wax blocks from 58 patients who received surgery at the Department of Cardiothoracic Surgery, Second Affiliated Hospital of Anhui Medical University, from January 1, 2016, to December 31, 2019, and who were postoperatively pathologically diagnosed with NSCLC. All patients had postoperative tumor node metastasis staging by the American Joint Committee on Cancer 7th edition staging system. The inclusion criteria were: (a) patients aged ≥18 years; (b) patients whose postoperative pathological diagnosis was NSCLC; (c) patients received postoperative cisplatin-based adjuvant chemotherapy; and (d) patients did not receive radiotherapy (44 cases without radiotherapy indication or 14 refused radiotherapy). The exclusion criteria were: (a) patients who received a preoperative anti-tumor treatment; (b) patients who had serious perioperative complications; (c) patients who received postoperative radiotherapy; (d) patients who had a postoperative adjuvant chemotherapy; and (e) patients who accepted other platinum drugs besides cisplatin.
In this study, 74 patients met the inclusion criteria. Finally, 58 tissue specimens were collected, and the clinical pathological data, follow-up data, and tissue wax blocks of 58 patients were complete. Cisplatin adjuvant chemotherapy regimens included 18 cases with gemcitabine combination, 17 with pemetrexed, 9 with docetaxel, 12 with paclitaxel, and 2 with other drugs (1 case of vinorelbine, 1 case of etoposide).
Study endpoint
The primary endpoint was disease-free survival (DFS) which is defined as the time from the end of the last adjuvant chemotherapy to the first onset of disease progression, recurrence, or death. The secondary endpoint was overall survival (OS), which is defined as the time from the end of the last adjuvant chemotherapy to death.
Follow-up
Through medical records’ inquiries and telephone follow-up, the follow-up date was set at June 1, 2021, and the follow-up time was from July 15, 2016 to June 1, 2021. The median follow-up time was 23.9 months. During the period, 12 cases were lost, and 4 were retained without tissue waxing. Patients who had not progressed, relapsed, or been lost follow-up at the end of the follow-up period were regarded as “missing data.” After the follow-up, 28 patients were followed-up to the endpoint and 30 did not reach the endpoint.
Immunohistochemistry methods
Immunohistochemical reagents
The primary antibody was a rabbit anti-human FIGNL1 polyclonal antibody (Batch No.:bs-9433R, Bioss) at a concentration of 1:200. Antigen retrieval was performed by ethylenediaminetetraacetic acid (EDTA) antigen retrieval solution (pH 8.0) (Zhongshan Jinqiao Biological Technology Co. Ltd., Beijing, China). The endogenous peroxide was blocked by a 3% H2O2 solution. The secondary antibody was an enzyme-labeled goat anti-rabbit immunoglobulin-G polymer. A DAB chromogenic kit and hematoxylin dye were used for staining.
Immunohistochemistry
The immunohistochemical staining was performed by an Elivision two-step method. Paraffin-embedded tissues were sectioned and dried. The tissue sections were placed in the oven at 60°C for 2h, dewaxed with xylene, and hydrated with gradient ethanol solutions. The EDTA antigen repair solution (pH 8.0) was used for high-pressure antigen repair. After natural cooling, an endogenous peroxidase inhibitor was added dropwise and incubated at room temperature for 10 min. A solution containing the primary antibody FIGNL1 was added dropwise, and the sections were incubated at 37°C for 60 min. The sections were rinsed three times in phosphate-buffered saline and for 3 minutes each time. A secondary antibody was added, and the sections were incubated at room temperature for 20 min. DAB was used for color development for 2.5 min, and hematoxylin staining. The sections were rinsed with anti-blue, then dehydrated, transparentized, and sealed. The expression of FIGNL1 protein was observed under the microscope by selecting the appropriate visual field for photography.
Interpretation of the immunohistochemical results
The proportion of positive cell scores divided into five grades is 0 points: staining cells 75%. The percentage of positive tumor cells ≤5% were defined as negative tumor tissue samples. Image J software was used for semi-quantitative analysis of FIGNL1 protein expression. A score of 0 was negative and ≥1 was positive. All immunohistochemical sections were assessed by two senior pathologists by the double-blind method. The same result was the final interpretation result.
Statistical analysis
The SPSS 26.0 was used for statistical analysis. The measurement data were expressed as mean ± standard deviation, and the t-test was used for statistical analysis. The relationship between FIGNL1 expression and clinicopathological characteristics was analyzed by Chi-square test or Fisher exact probability method. Chi-square test was used to analyze the expression difference of FIGNL1 in NSCLC tissues and adjacent tissues. The Log-rank test and Kaplan–Meier survival curve were used for single factor survival analysis. By estimating the sample size of survival analysis, 73 subjects were obtained (1-β = 0.90). A P-value of <0.05 showed a statistically significant difference.
Results
Analysis of the relationship between FIGNL1 mRNA expression and clinical prognosis in NSCLC
The result showed that FIGNL1 expression in lung adenocarcinoma and lung squamous cell carcinoma is significantly higher than that in adjacent tissues (P<0.05) (Figures 1(a) and (b)). We found that the median OS time in the FIGNL1 high expression group was 48 months shorter than that in the low expression group (99.43 months). The prognosis in the FIGNL1 high expression group was worse than that in the low expression group (Figure 1(c)) (P = 1e-11, hazard ratio (HR) = 1.79, 95% confidence interval (CI) = 1.51–2.12).
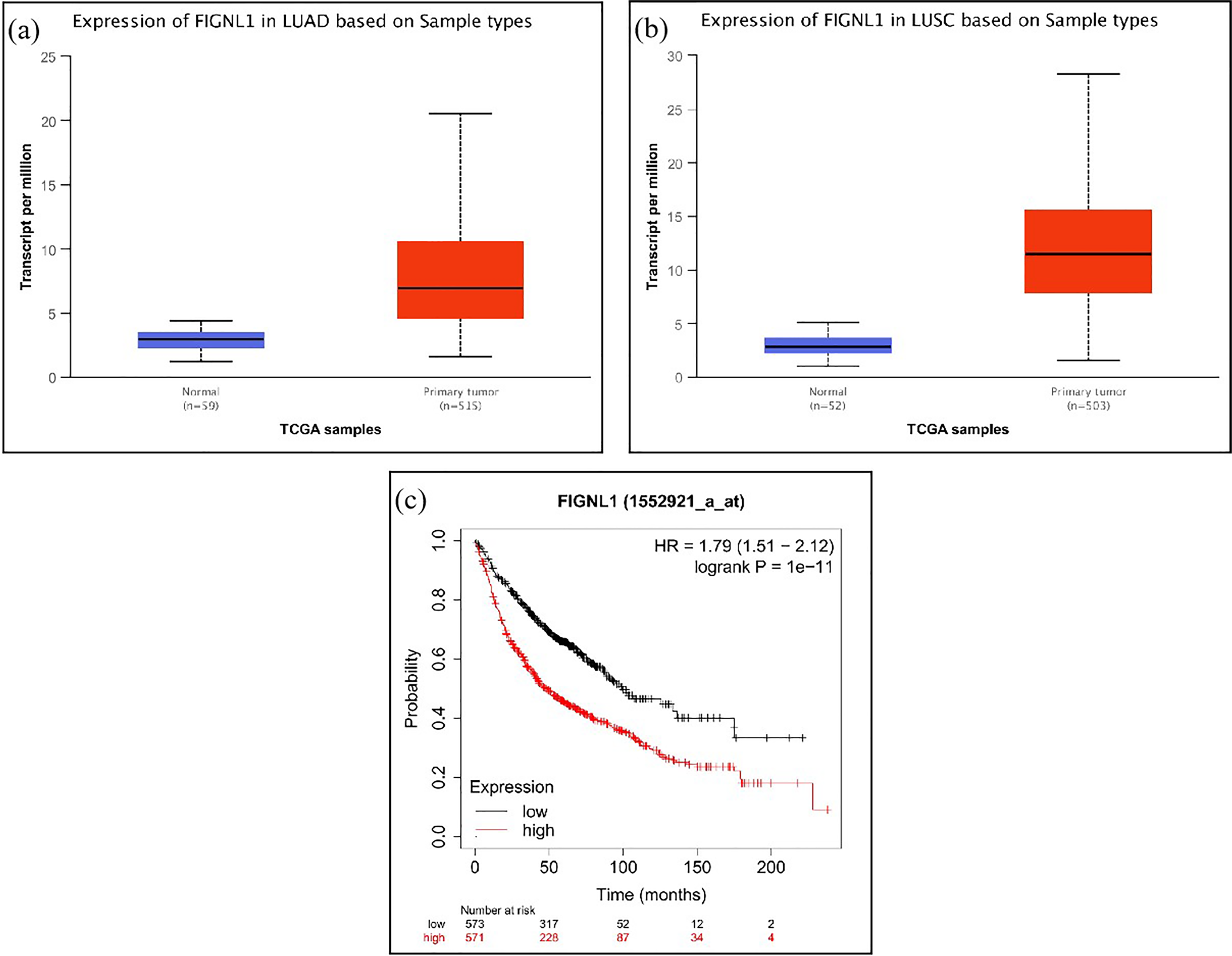
Correlation between
Searching cisplatin resistance dataset from the GEO database to explore the relationship between FIGNL1 expression and cisplatin resistance
We retrieved two NSCLC cisplatin-resistant microarray datasets, GSE21656 and GSE157692, and further assessed the expression of the FIGNL1 gene in the two datasets. The statistical analysis of the t-test showed that in the data set “GSE21656,” the relative expression level of FIGNL1 messenger RNA (mRNA) in the cisplatin-resistant group was higher than that in the cisplatin-sensitive group (P = 0.822); however, the difference was insignificant. In the dataset “GSE157692,” the relative expression in the cisplatin-resistant group was significantly higher than that in the cisplatin-sensitive group (Table 1; P = 3.80e-05).
The expression of FIGNL1 in cisplatin-resistant cell lines.
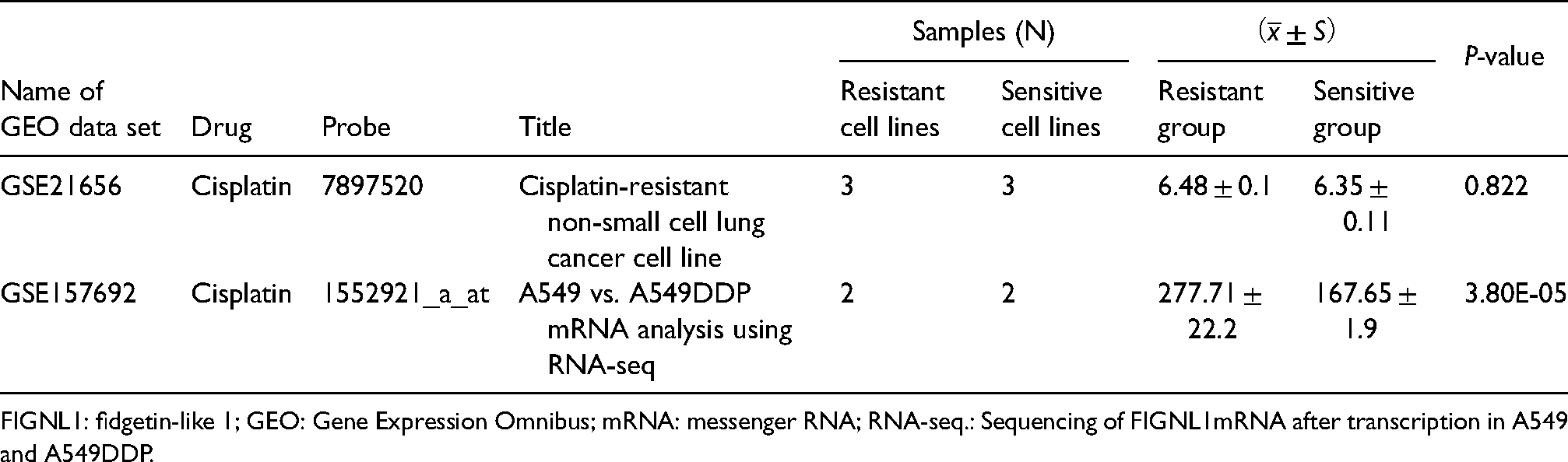

FIGNL1: fidgetin-like 1; GEO: Gene Expression Omnibus; mRNA: messenger RNA; RNA-seq.: Sequencing of FIGNL1mRNA after transcription in A549 and A549DDP.
Immunohistochemical detection of FIGNL1 protein expression in NSCLC tissues and adjacent tissues
The immunohistochemical analysis of tissues from 58 NSCLC patients showed that FIGNL1 was positive in 42 cases (42/58, 72.41%) and negative in 16 cases (16/58, 27.59%). FIGNL1 expression was mainly located in the nucleus as shown by the nuclear brown granules under the microscope (Figure 2). The expression of FIGNL1 in NSCLC was further semi-quantitatively analyzed, and the results showed that in the NSCLC tissues, 42 cases scored ≥1, and 16 scored 0; in the NSCLC adjacent tissues, FIGNL1 scores were 0 in 48 cases and ≥1 in 10 (Figure 3). The positive expression rate of FIGNL1 in NSCLC tissues was 72.41%, which was significantly higher than that in adjacent tissues (17.24%, 10/58; P < 0.01) (Table 2).
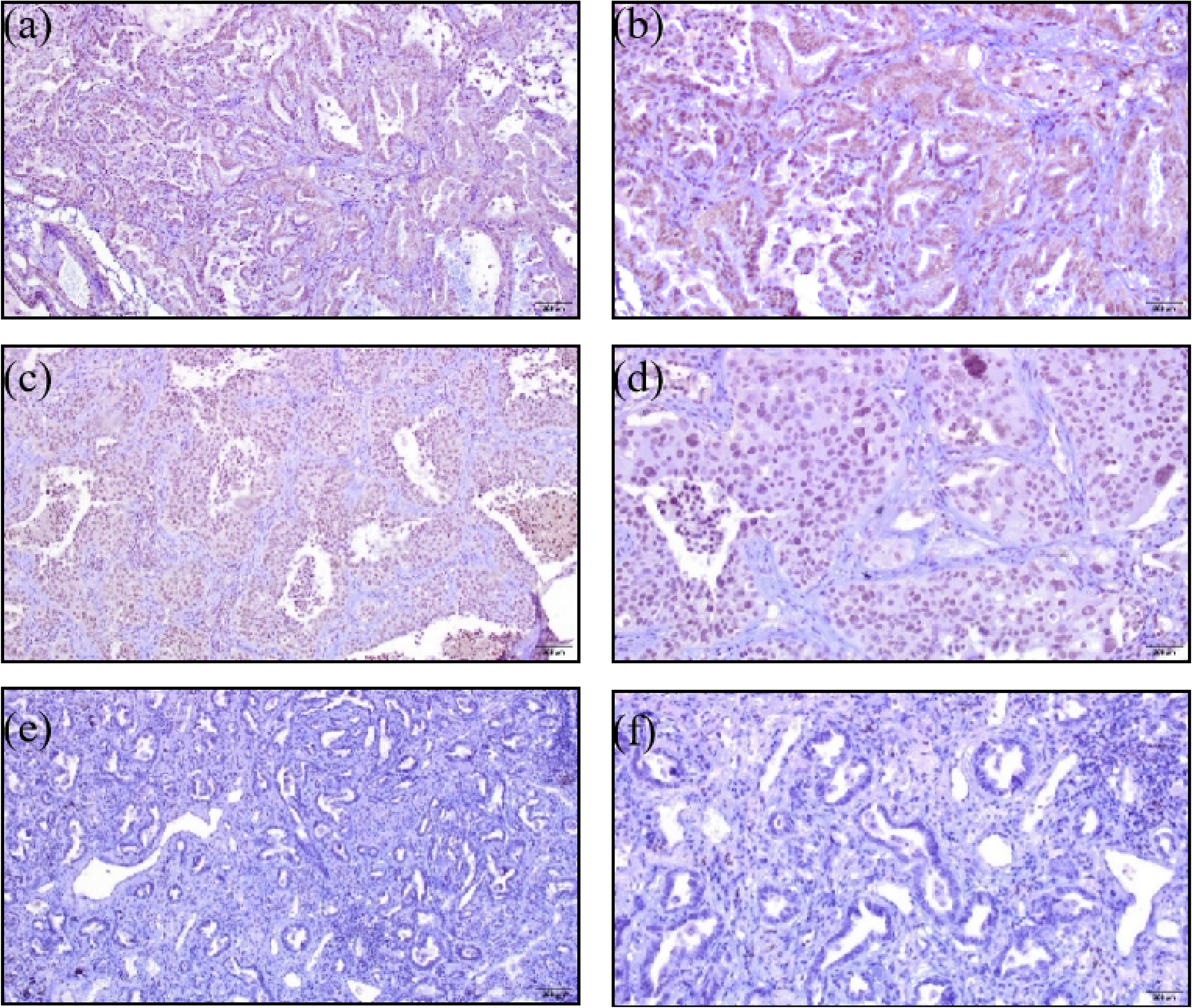
FIGNL1 expression in NSCLC. (a) FIGNL1 positive expression in lung adenocarcinoma ( × 100). (b) FIGNL1 positive expression in lung adenocarcinoma ( × 200). (c) FIGNL1 positive expression in lung squamous cell carcinoma ( × 100). (d) FIGNL1 positive expression in lung squamous cell carcinoma ( × 200). (e) FIGNL1 negative expression in cancer tissues ( × 100). (f) FIGNL1 negative expression in cancer tissues ( × 200).
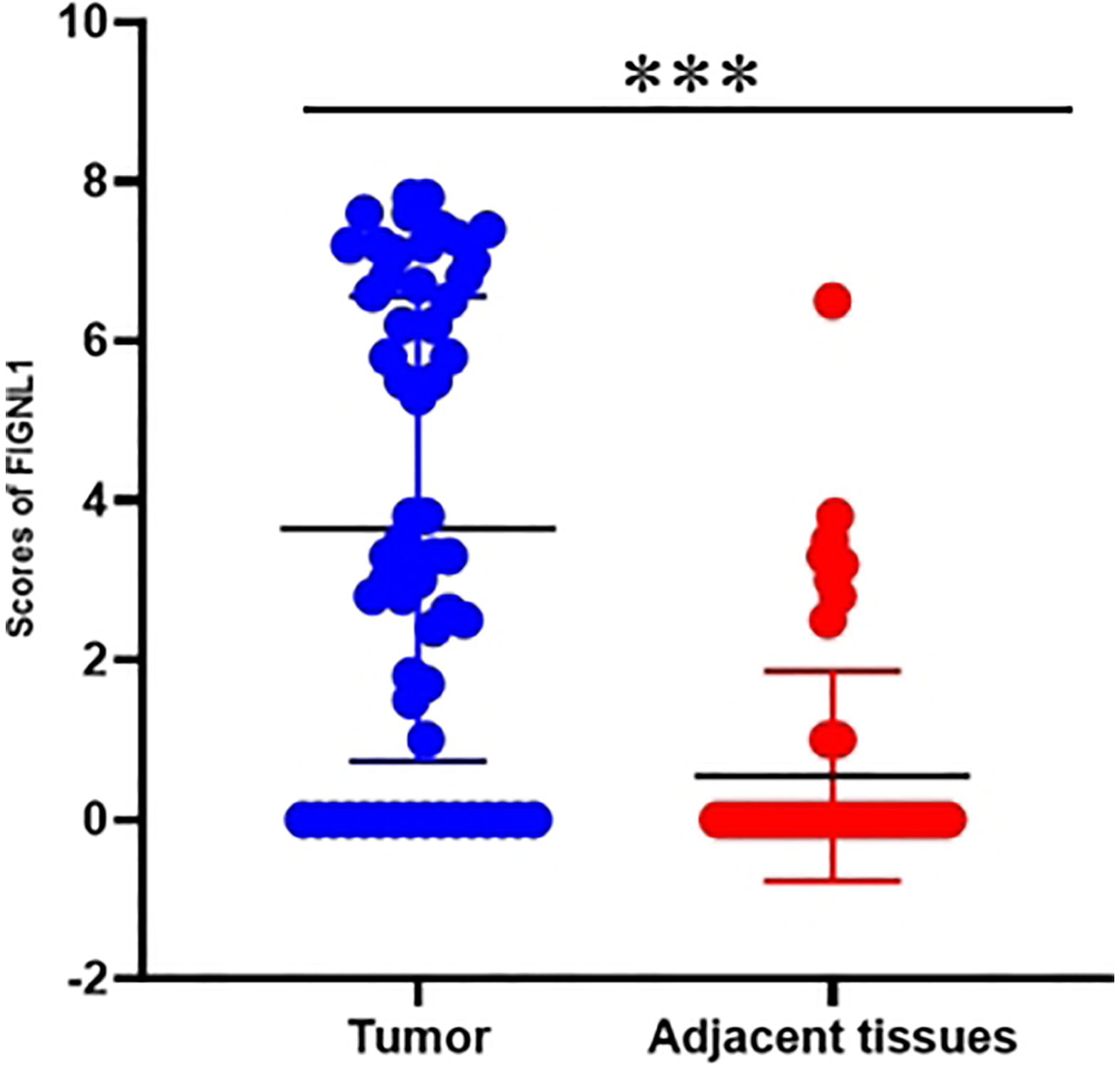
Differential expression of FIGNL1 protein in NSCLC tissues and adjacent tissues. FIGNL1 expression levels were significantly higher in NSCLC tissues compared to that in adjacent normal tissues (N = 58, ***P < 0.05).
Expression difference of FIGNL1 protein between NSCLC tissues and adjacent tissues.
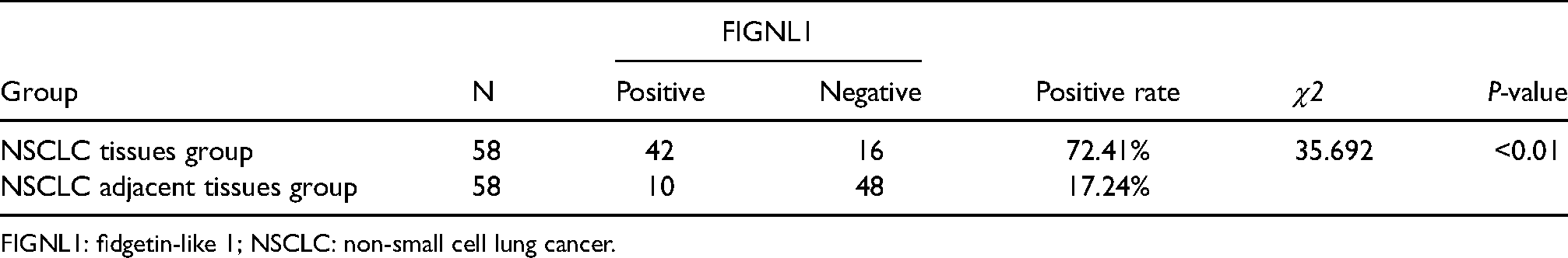

FIGNL1: fidgetin-like 1; NSCLC: non-small cell lung cancer.
FIGNL1 protein expression in NSCLC and its relationship with clinicopathological features
In this study, 74 patients met the inclusion criteria. However, during the follow-up, 12 were lost, and tissues from 4 patients were not waxed. Finally, 58 tissue specimens meeting the above inclusion criteria were collected. The clinico-pathological data, follow-up data, and waxed tissues from 58 patients were completed (Table 3). The relationship between FIGNL1 expression and clinicopathological features showed that FIGNL1 expression in NSCLC was closely related to gender, histological type, and nerve invasion (P < 0.05).
The correlation between expression of FIGNL1 and clinicopathological features in NSCLC.
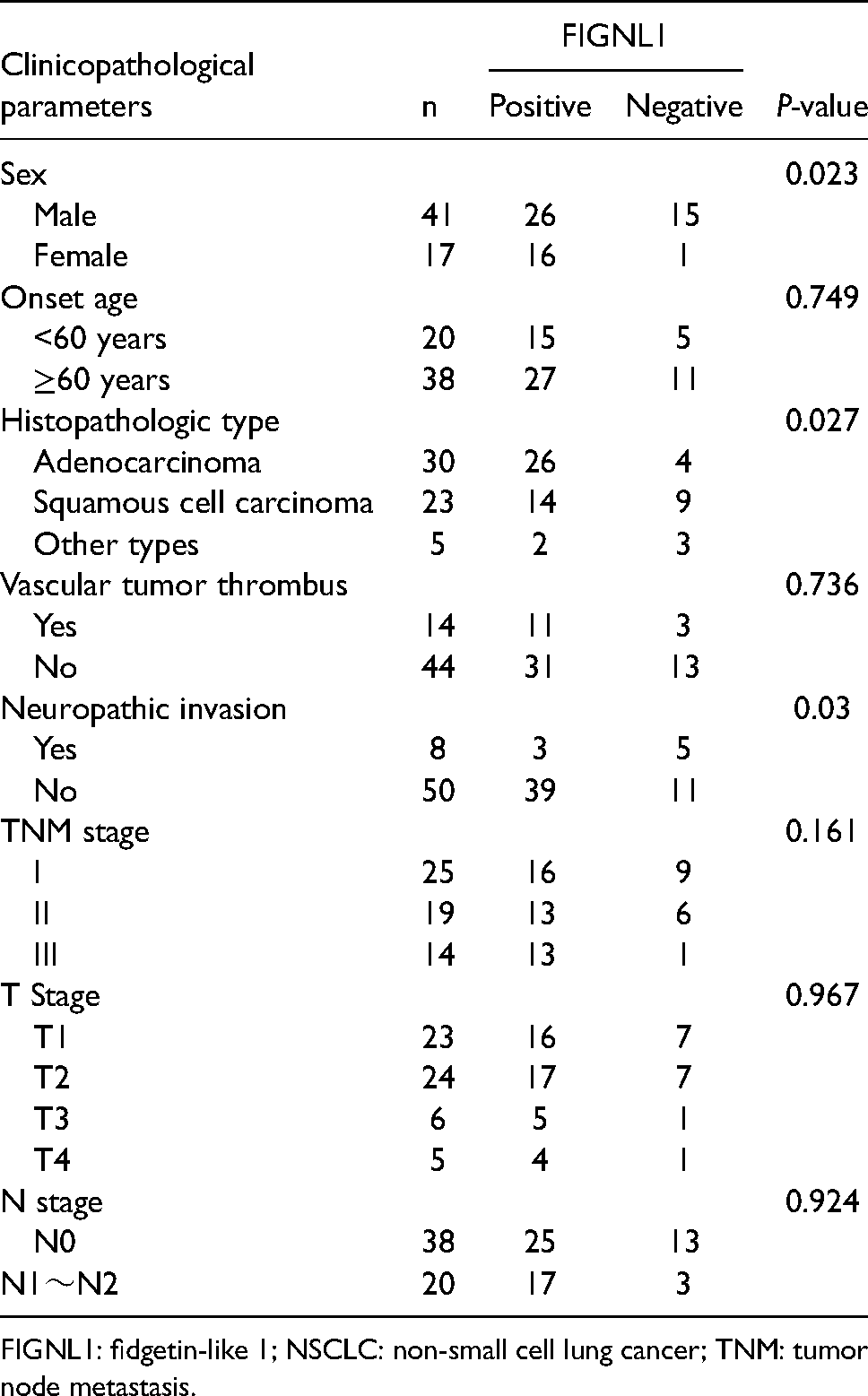
FIGNL1: fidgetin-like 1; NSCLC: non-small cell lung cancer; TNM: tumor node metastasis.
Correlation between FIGNL1 expression and DFS in NSCLC tissues
In the FIGNL1 positive and negative groups, each combination scheme was balanced and there was no correlation between different cisplatin combinations and DFS duration (Table 4) (χ2 = 3.402, P = 0.509).
The correlation between different combination regimes of cisplatin and DFS.
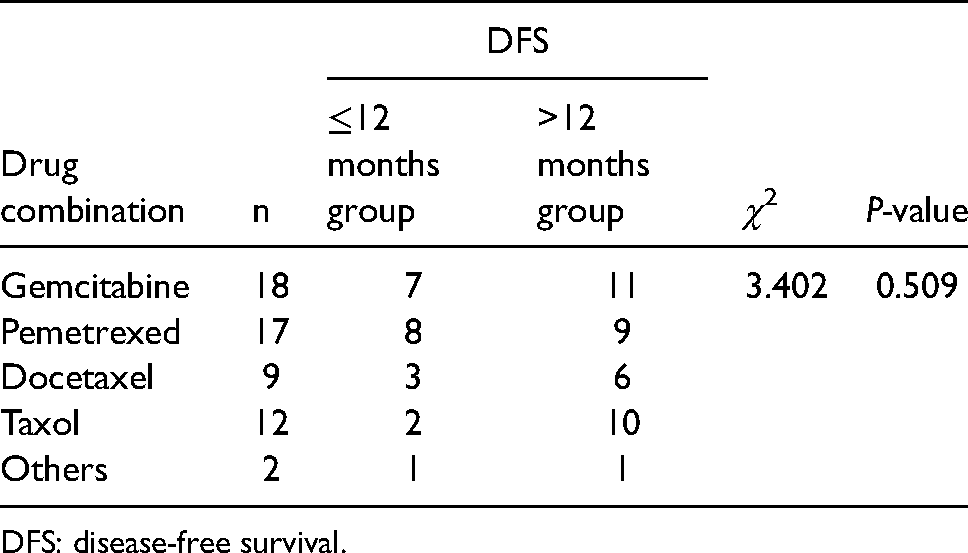

DFS: disease-free survival.
We analyzed the correlation between FIGNL1 expression and DFS using the Kaplan–Meier method (Figure 4). The median DFS of 58 NSCLC patients was 28.90 months (18.05∼39.75 months). The median DFS of NSCLC patients with FIGNL1 positive expression was 22.00 months (9.67∼34.34 months). The median DFS of NSCLC patients with FIGNL1 negative expression was 42.00 months (25.21∼58.79 months). The DFS of NSCLC patients with high expression of FIGNL1 was significantly shorter (P = 0.032).
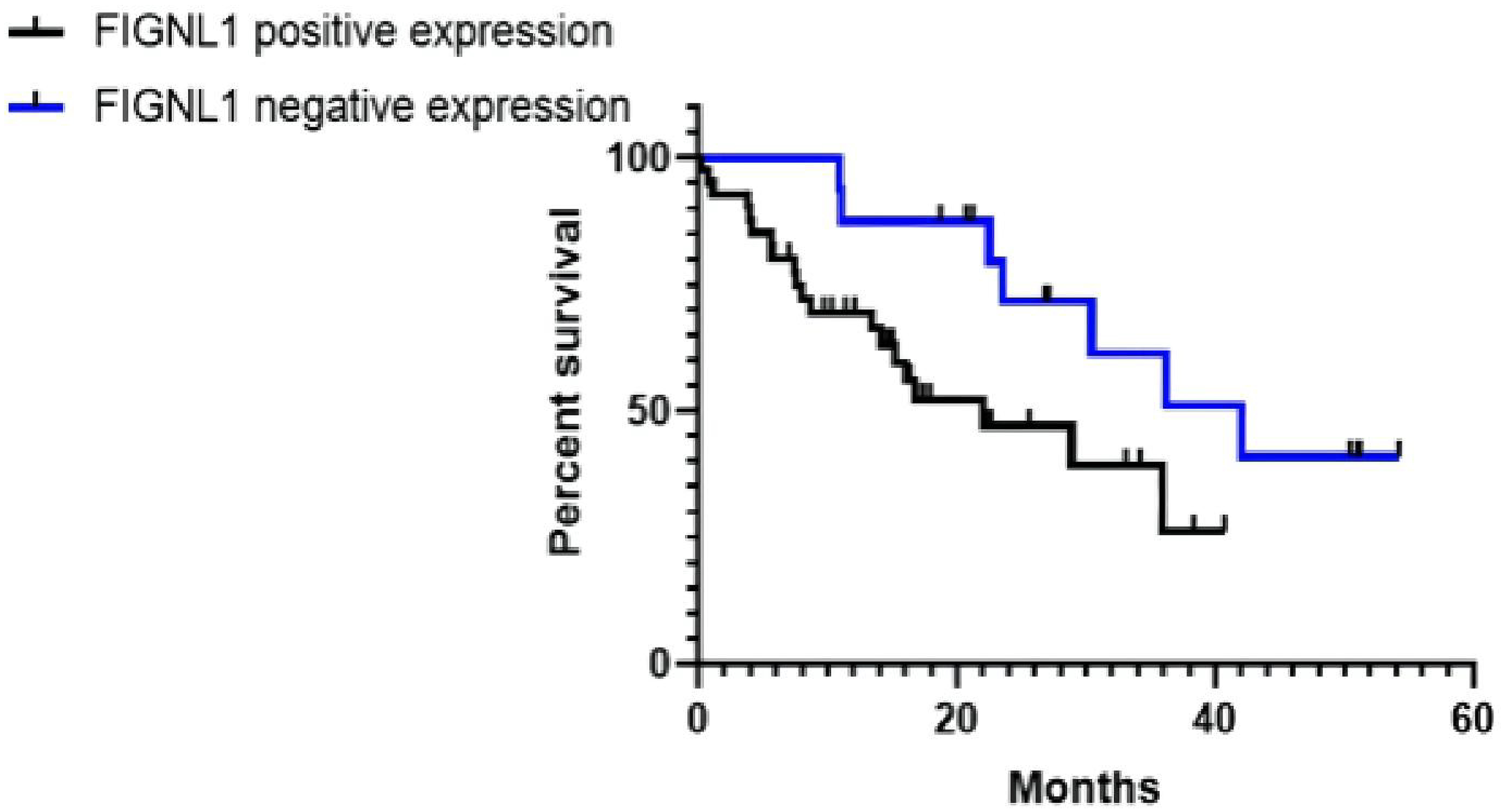
The correlation between expression of FIGNL1 and DFS in NSCLC.
GO analysis of FIGNL1 function in biological processes, building protein interaction networks, and screening hub gene
We first obtained FIGNL1 differential expression between NSCLC cisplatin-resistant and cisplatin-sensitive from the GSE157692 dataset (Table 1). The dataset was downloaded, and FIGNL1 differential expression was investigated by single gene GO analysis. We found that FIGNL1 is mainly involved in nuclear division, the cell response to radiation, DNA recombination, and the cytosol division biological process (Figure 5). The results showed that FIGNL1 can promote cisplatin resistance of NSCLC cells by participating in DNA recombination. We constructed the FIGNL1 protein interaction network, and obtained 105 FIGNL1 protein interaction networks by STRING (Figure 6).
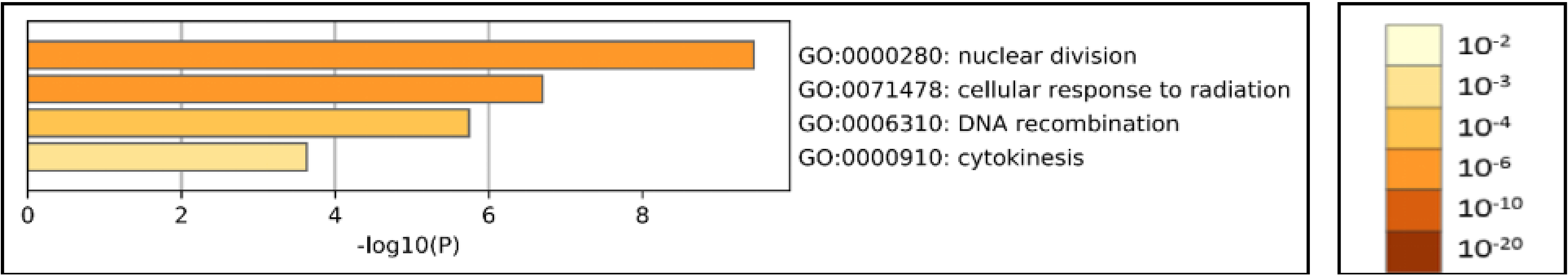
Go analysis of FIGNL1. FIGNL1 is mainly involved in nuclear division (GO:0000280), cell response to radiation (GO: 0071478), DNA recombination (GO: 0006310), and cytoplasmic division (GO: 0000910).
The interacting protein data were visualized by the Cystoscope software, and hub genes were screened. We identified the Top 20 hub genes, which are UBXN7, RAB11FIP3, TRAPPC6B, C7orf72, KBTBD12, VWC2, KCNG3, SIPA1L2, KLHDC8B, PBK, CDCA8, RAD51, KNTC1, TOP2A, RAB11FIP3, DECR2, AURKB, CCDC36, MEM204 and FIGNL1 There was a strong interaction between these genes. RAD51 and CCDC36 are key genes for FIGNL1 in HRR.
Discussion
According to China's Cancer Incidence and Mortality Statistics in 2015, lung cancer is a malignant tumor with the highest incidence and mortality among males in China.12,13 Although surgery is the main treatment for early NSCLC, patients have postoperative recurrence and metastasis. Therefore, adjuvant chemotherapy is used to reduce the risk of recurrence and metastasis and improve survival. 14 In an international multicenter randomized controlled trial, the median DFS for the chemotherapy group of patients (vinorelbine combined with cisplatin) was 36.3 months (range 28.0–52.1), a duration that was significantly higher than that of the observation group, which was 20.7 months (16.1–28.6) (HR = 0.76 (95% CI = 0.64–0.91); P = 0.017). 15 Yang et al. 16 analyzed 1128 NSCLC patients based on The Cancer Genome Atlas (TCGA) database and found that the 5-year OS rate of the high-risk group after adjuvant chemotherapy was 32.4%, and that of the low-risk group was 53.7%. Cisplatin resistance was one of the risk factors; therefore, improving the efficacy of adjuvant chemotherapy for NSCLC has become significant in clinical prognosis.
Cisplatin is the cornerstone drug for adjuvant chemotherapy after NSCLC surgery.3 Its primary anti-tumor mechanism consists in entering cancer cells, where it binds DNA to form Pt-DNA adducts, causing DNA damage and inducting cancer cell necrosis or apoptosis. Cisplatin resistance is attributed to three molecular mechanisms: (a) abnormal DNA damage repair function; (b) a decrease in intracellular cisplatin concentration; and (c) cisplatin cytoplasmic inactivation. 17 Abnormal DNA damage repair is a fundamental reason for cisplatin resistance.
In 2000, fidgetin was found by Cox et al. 18 in mammalian developmental mutants, and FIGNL1 and FIGNL2 genes were also identified after a homologous sequence comparison. FIGNL1 main biological functions are related to its three conserved domains: (a) the AAA-ATP enzyme domain; (b) the VPS4 domain; and (c) the FRBD domain. So far, the FRBD domain has only been found in the human FIGNL1 gene. This domain can combine with the recombinant enzyme RAD51 (the key enzyme in the HRR pathway), co-act on homologous recombination-mediated DNA repair, and enhance RAD51 HRR function. 19 A tissue microarray study by Schraenen et al. 20 reported that FIGNL1 is highly expressed in colorectal cancer, cervical cancer, lung adenocarcinoma, and other tumors. A study by Ma et al. 6 from Shanghai Jiao Tong University found that FIGNL1 is highly expressed in small-cell lung cancer (SCLC) patients. After FIGNL1 silencing, the sensitivity of H446 cells to cisplatin increased by 20.1 % and 15.6 %, respectively, and the inhibition rates by silencing were 48.6 % and 13.5 %, respectively. The lower the expression of FIGNL1 in SCLC, the more the HRR pathway was abnormally enhanced, and the higher the sensitivity of H446 cells to cisplatin. An in vitro study showed that FIGNL1 enhances the cisplatin resistance in NCI-H446 cells.
Our study investigated the clinical relationship between FIGNL1 expression in NSCLC tissues and cisplatin resistance. For this, bioinformatics methods were used to retrieve the TCGA and GEO data. TCGA database is an extensive cancer gene database with rich and standardized clinical data; the GEO database is a public repository of experimental data. We analyzed the clinical data of the TCGA and the experimental data of the GEO databases, which showed that FIGNL1mRNA was highly expressed in NSCLC tissues, and that this high expression may be related to the occurrence of cisplatin resistance. Immunohistochemistry analysis showed that FIGNL1 was highly expressed in NSCLC cancer tissues where it was mainly located in the nucleus. The nuclear localization of FIGNL1 is consistent with its HRR function. Therefore, we speculated that there may be a structure related to FIGNL1 HRR function in the nucleus, which opens up a future research direction in further investigating the role of FIGNL1 in DNA repair.
We preliminarily analyzed the relationship between FIGNL1 expression and NSCLC clinicopathological features. The results showed that FIGNL1 expression was closely related to gender, histopathological type, and nerve invasion. Female gender, lung adenocarcinoma, and the absence of nerve invasion were independent influencing factors in NSCLC. However, our sample size is small, and the relationship between FIGNL1 and clinicopathological characteristics of NSCLC would require a larger cohort.
We concluded that there was no correlation between the duration of DFS and the combination chemotherapy of the above six different chemotherapy drugs (Table4). This is consistent with the conclusion of previous large clinical studies which reported the absence of a significant difference in the efficacy of cisplatin adjuvant chemotherapy when combined with various chemotherapeutic drugs after NSCLC surgery.21,22 The correlation between FIGNL1 expression and DFS was further analyzed. The results showed that the DFS with FIGNL1 positive expression was shorter, suggesting that FIGNL1 is associated with NSCLC poor prognosis. They also showed that cisplatin resistance may be involved. We speculated that the potential mechanism may involve cisplatin forming Pt-DNA adducts in NSCLC cells causing DNA damage. When FIGNL1 is highly expressed in NSCLC, HRR is abnormally activated. FIGNL1 binds to RAD51, which promotes HRR of damaged DNA. Therefore, the abnormal DNA-repaired NSCLC cells can normally survive, causing cisplatin resistance, potential recurrence, and metastasis. 23 The abnormal DNA repair in tumors is often closely related to occurrence development and chemotherapy. Since cisplatin kills tumor cells through DNA damage, an abnormal HRR pathway will cause cisplatin resistance in tumor cells.
In exploring the mechanism of FIGNL1 in NSCLC cisplatin resistance, we considered a correlation between FIGNL1 and DNA damage repair. The differential expression of FIGNL1 in the cisplatin-resistance of NSCLC was analyzed by GO, which indicated that FIGNL1 is involved in DNA recombination. Through the FIGNL1 protein interaction network, we identified the hub genes that involved HRR are RAD51 and CCDC36. In addition to the reported relationship between FIGNL1 and RAD51, we also found that FIGNL1 and CCDC36 co-participate in HRR, which lays a foundation for exploring the mechanism of FIGNL1 and NSCLC cisplatin resistance at the cellular and molecular levels.
The study limitation is the small size of the sample which only had 58 cases and this may have impacted the scientific hypothesis results. In future, we will expand the sample size to provide more clinical evidence for further exploring tumor markers related to cisplatin resistance. Finally, intracellular DNA damage repair has attracted much attention in the study of cisplatin resistance in NSCLC, and targeting DNA repair molecules may be a new direction in solving cisplatin resistance.
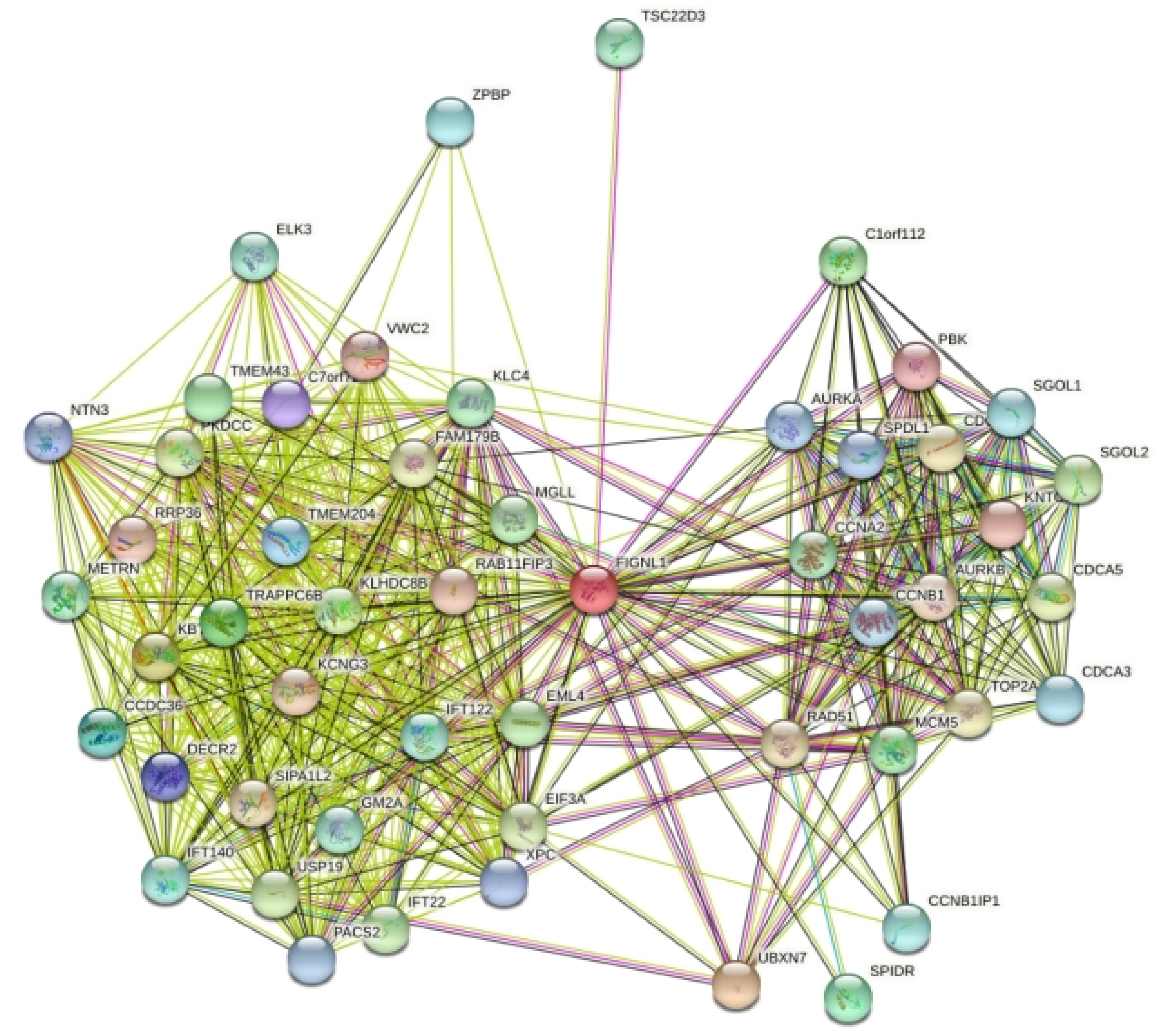
Diagrams FIGNL1 protein interaction network in STRING.
Supplemental Material
sj-jpg-1-jbm-10.1177_03936155221110249 - Supplemental material for FIGNL1 is a potential biomarker of cisplatin resistance in non-small cell lung cancer
Supplemental material, sj-jpg-1-jbm-10.1177_03936155221110249 for FIGNL1 is a potential biomarker of cisplatin resistance in non-small cell lung cancer by Chenxu Meng, Yang Yang, Pengfei Ren, Qian Ju, Xiangting Jin, Qihe Long, Xiangyu Chen, Xian Wang and Fanfan Li in The International Journal of Biological Markers
Supplemental Material
sj-docx-2-jbm-10.1177_03936155221110249 - Supplemental material for FIGNL1 is a potential biomarker of cisplatin resistance in non-small cell lung cancer
Supplemental material, sj-docx-2-jbm-10.1177_03936155221110249 for FIGNL1 is a potential biomarker of cisplatin resistance in non-small cell lung cancer by Chenxu Meng, Yang Yang, Pengfei Ren, Qian Ju, Xiangting Jin, Qihe Long, Xiangyu Chen, Xian Wang and Fanfan Li in The International Journal of Biological Markers
Supplemental Material
sj-docx-3-jbm-10.1177_03936155221110249 - Supplemental material for FIGNL1 is a potential biomarker of cisplatin resistance in non-small cell lung cancer
Supplemental material, sj-docx-3-jbm-10.1177_03936155221110249 for FIGNL1 is a potential biomarker of cisplatin resistance in non-small cell lung cancer by Chenxu Meng, Yang Yang, Pengfei Ren, Qian Ju, Xiangting Jin, Qihe Long, Xiangyu Chen, Xian Wang and Fanfan Li in The International Journal of Biological Markers
Supplemental Material
sj-docx-4-jbm-10.1177_03936155221110249 - Supplemental material for FIGNL1 is a potential biomarker of cisplatin resistance in non-small cell lung cancer
Supplemental material, sj-docx-4-jbm-10.1177_03936155221110249 for FIGNL1 is a potential biomarker of cisplatin resistance in non-small cell lung cancer by Chenxu Meng, Yang Yang, Pengfei Ren, Qian Ju, Xiangting Jin, Qihe Long, Xiangyu Chen, Xian Wang and Fanfan Li in The International Journal of Biological Markers
Supplemental Material
sj-docx-5-jbm-10.1177_03936155221110249 - Supplemental material for FIGNL1 is a potential biomarker of cisplatin resistance in non-small cell lung cancer
Supplemental material, sj-docx-5-jbm-10.1177_03936155221110249 for FIGNL1 is a potential biomarker of cisplatin resistance in non-small cell lung cancer by Chenxu Meng, Yang Yang, Pengfei Ren, Qian Ju, Xiangting Jin, Qihe Long, Xiangyu Chen, Xian Wang and Fanfan Li in The International Journal of Biological Markers
Supplemental Material
sj-docx-7-jbm-10.1177_03936155221110249 - Supplemental material for FIGNL1 is a potential biomarker of cisplatin resistance in non-small cell lung cancer
Supplemental material, sj-docx-7-jbm-10.1177_03936155221110249 for FIGNL1 is a potential biomarker of cisplatin resistance in non-small cell lung cancer by Chenxu Meng, Yang Yang, Pengfei Ren, Qian Ju, Xiangting Jin, Qihe Long, Xiangyu Chen, Xian Wang and Fanfan Li in The International Journal of Biological Markers
Footnotes
Acknowledgments
We gratefully acknowledge all the members of Department of the Second Hospital of Anhui Medical University. We also thank all patients for their participation in this study. We gratefully acknowledge Fanfan Li, Yang Yang for their support.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Anhui Medical University, (grant number 2019xkj044).
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
