Abstract
BACKGROUND:
Long intergenic non-coding RNA (lincRNA) belongs to a special type of RNA that is unable to encode proteins but has been proved to play a role in gene regulation and differentially expressed in various malignant tumors.
OBJECTIVE:
In this study, we aimed to identify whether lincRNA LINC00173 was differentially expressed in non-small-cell lung cancer (NSCLC) and whether it could serve as a potential diagnostic biomarker.
METHODS:
The quantification real-time quantitative polymerase chain reaction (qRT-PCR) was used to detect the expression of LINC00173 in serum and cultured cells. For large sample analysis, the lncRNA expression matrix in TCGA database were generated via R software. To evaluate the diagnostic performance of serum LINC00173, the receiver operating characteristic (ROC) curve was used.
RESULTS:
The qRT-PCR analysis showed that the serum LINC00173 expression level in 108 NSCLC patients was higher than that in 91 healthy donors and 55 patients with benign pulmonary disease (BPD). And the area under the curve (AUC) of serum LINC00173 was 0.809 for the diagnosis of NSCLC (95% CI: 0.750–0.868,
CONCLUSION:
Serum LINC00173 may prove to be a potential non-invasive auxiliary diagnostic biomarker for NSCLC patients.
Introduction
Lung cancer is the leading cause of cancer-related death in the world [1], especially in men, and smoking is thought to induce high risk of lung cancer. Both late diagnosis and low 5-year survival rate bring challenge for the diagnosis of lung cancer [2]. Adequate tissue material used for histological diagnosis and molecular testing is the “gold standard” for the diagnosis of lung cancer [3]. Other Standard tests, including routine hematology, renal and hepatic function, and bone biochemistry tests, are also recommended [4]. Besides, serum biomarker detection, such as carcinoembryonic antigen (CEA), squamous cell carcinoma antigen (SCC) and cytokeratin 19 fragment (Cyfra21-1), is increasingly used in the diagnosis of lung cancer due to its non-invasiveness, and concomitant detection of multiple biomarkers is more conducive for early diagnosis. However, several tumor markers currently used in clinical practice still have certain defects, such as insufficient sensitivity and low specificity. Therefore, it is of great value to identify new potential biomarkers for early diagnosis of lung cancer.
Long non-coding RNA (lncRNA) is a kind of special RNA transcripts with more than 200 nucleotides, and long intergenic non-coding RNA (lincRNA) is a specific type of lncRNA that does not intersect with any protein-coding locus [5]. Although unable to encode proteins, lincRNAs have proved to have a great power in regulating protein coding genes [6]. In addition, some specific lincRNAs in the peripheral blood circulation were reported to be aberrantly expressed in certain diseases, and underwent dynamic changes at different stages, endowing them with great potential to become useful biomarkers for diagnostic and prognostic predictions. For example, serum lncRNA AFAP1-AS1 was found to act as a molecular marker for distinguishing non-small cell lung cancer (NSCLC) patients from healthy people with an area under the curve (AUC) of 0.759, and combination of FAP1-AS1 and Cyfra21-1 exhibited a better effect [7].
LINC00173 is an uncharacterized lincRNA belonging to a cluster at 12q24.22. Some current studies reported that LINC00173 was associated with several cellular pathological processes. Postler et al. [8] found both transcript variants of LINC00173 (transcript variant 1 and 2) were reproducibly up-regulated during infection, suggesting that LINC00173 may play a role in regulating cytokines in T cells. Besides, the expression of LINC00173 in granulocytes was 10-fold that in any other human tissue, suggesting that this lincRNA may participate in the regulation of granulocyte proliferation and maturation [9]. Mao et al. [10] observed that knockdown of LINC00173 boosted the colony number and decreased the G1/G0 population in esophageal squamous cells, indicating its potential role in the development of malignant tumors. In small-cell lung cancer (SCLC), LINC00173 was found to be upregulated in chemoresistant cell lines, and promoted SCLC cells chemoresistance, proliferation, and migration-invasion by sponging miR-218 [11]. Besides, our previous study discovered that miR-182-5p was negatively regulated by LINC00173 and facilitated cell growth and migration in NSCLC cells via AGER/NF-
In this study, we investigated the serum expression level of LINC00173 in a cohort of 108 NSCLC patients, 91 healthy donors, 55 patients with benign pulmonary diseases (BPDs), and 37 small-cell lung cancer (SCLC) patients. A diagnostic model consisting of serum LINC00173 and the existing diagnostic indicators (CEA and Cyfra21-1) was conducted with a view of improving the diagnostic efficiency. Besides, we also examined the utility of LINC00173 in monitoring tumor dynamics in NSCLC patients.
Methods and materials
Patient selection and sample collection
Serum samples were obtained from 108 NSCLC patients, 37 SCLC patients, 91 healthy donors and 55 BPD patients at the Department of Laboratory Medicine (Affiliated Hospital of Nantong University, Nantong, China) between October 2017 and November 2019. Besides, additional 60 surgery- or chemotherapy-treated NSCLC patients were also obtained, including 20 with tumor recurrence and metastasis as confirmed by oncology. The study protocol was approved by the Clinical Research Ethics Committee of the local hospital and written informed consent was obtained from all subjects participating in the study. The collected serum samples were stored in RNase-free centrifuge tubes at
Serum RNA extraction and cDNA synthesis
Serum RNA was extracted from 300
Cell culture
Human bronchial epithelial cell (16HBE) and NSCLC cell lines (A549, NCI-H1299 and SPC-A1) were purchased from the Cell Bank of the Chinese Academy of Sciences (Shanghai, China). The cells were cultured in DMEM/RPMI-1640 medium (Corning, USA) containing 10% fetal bovine serum (FBS) (Gibco, USA) in a humid incubator with 5% CO
Bioinformatics analysis
Microarray profiles related to Lung Adenocarcinoma (LUAD) and Lung Squamous Cell Carcinoma (LUSC) were obtained from the Cancer Genome Atlas (TCGA) database. The lncRNA expression matrix were generated via R version 3.5.1 (R: A Language and Environment for Statistical Computing, R Core Team, R Foundation for Statistical Computing, Vienna, Austria, 2018,
Quantitative real-time polymerase chain reaction (qRT-PCR) assays
All qRT-PCR assays were performed on the ABI 7500 PCR Detection System (ABI, USA), and SYBR Green I (Roche, Germany) was used as the fluorescent molecule. The procedure of PCR cycling program consisted of preamplification at 95
Construction of LINC00173 standard plasmid
The PCR products of serum LINC00173 were purified by a 3% agarose gel and connected with pMD18-T Vector (Takara, Japan). Then 10
Detection of CEA, SCC and Cyfra21-1 levels
Serum CEA, SCC and Cyfra21-1 were measured with ARCHITECT I2000 SR (Abbott, Chicago, USA). The cut-off value was developed for a distinction between positive and negative results under the protocols of our department (CEA: 5 ng/ml; SCC: 1.5 ng/ml; Cyfra21-1: 2.05 ng/ml).
Statistical analysis
All data are presented as the mean
Methodology evaluation of LINC00173 detection in serum samples. (A) Standard curves of serum LINC00173 in a ten-fold serial dilution of standard plasmid. (B) and (C) Stability of circulating LINC00173 under prolonged room temperature incubation time or multiple freeze-thaw cycles. Data are presented as raw Ct value (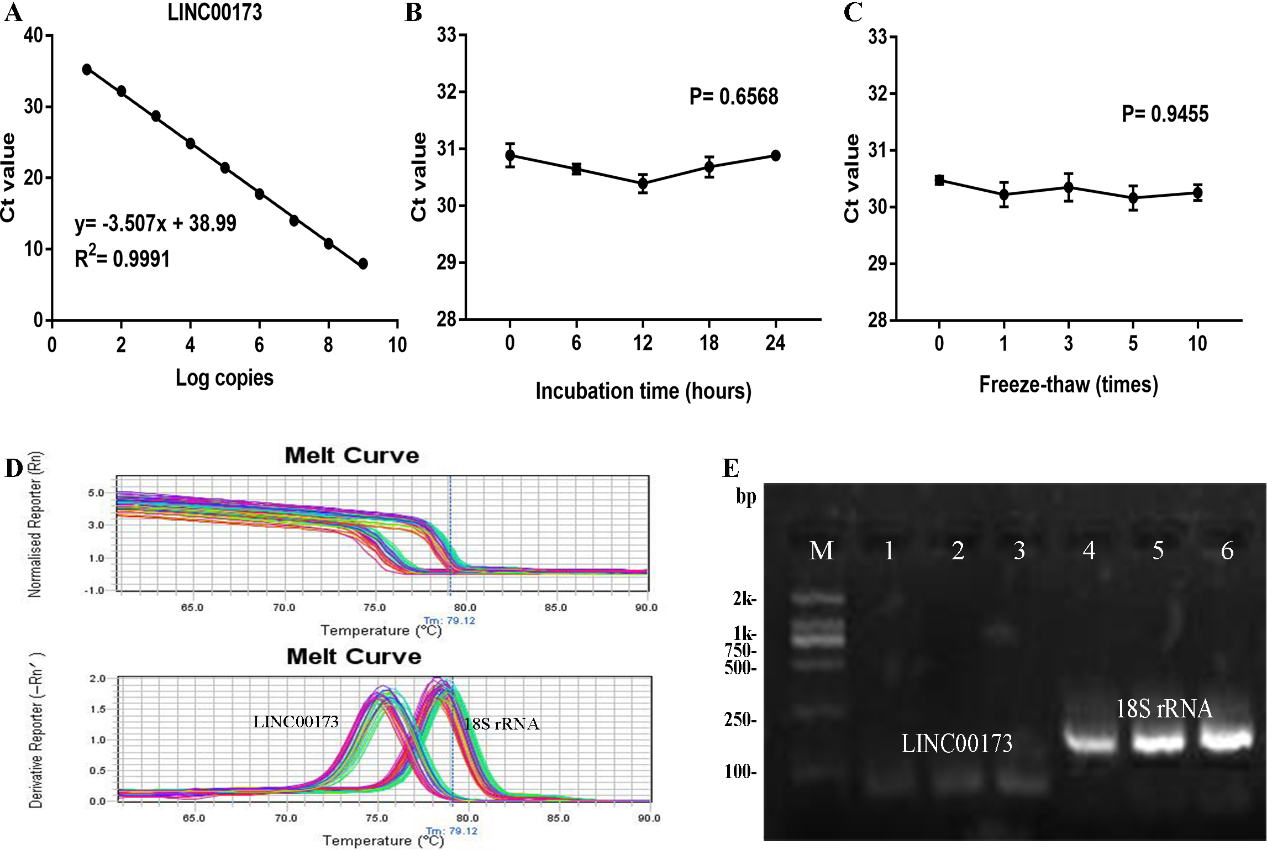
Methodology evaluation of circulating LINC00173 detection in serum samples
Due to no standard internal reference for serum lincRNA in lung cancer, we compared three most commonly used internal references (
To verify whether LINC00173 was suitable for clinical laboratory analysis, we performed methodology evaluation of LINC00173 detected by qRT-PCR assay. Firstly, to evaluate its linearity, we constructed the standard recombinant plasmid for LINC00173 and diluted the standard 10 times with DNA dilution to obtain 10 concentration gradients of 10
Intra- and inter-assay reproducibility of LINC00173 and 18S rRNA
Intra- and inter-assay reproducibility of LINC00173 and 18S rRNA
Abbreviations: SD, standard deviation; CV, coefficient of variation.
Concentrations of serum LINC00173, CEA, and Cyfra21-1 in NSCLC cases. (A) Detection of serum LINC00173 in NSCLC patients (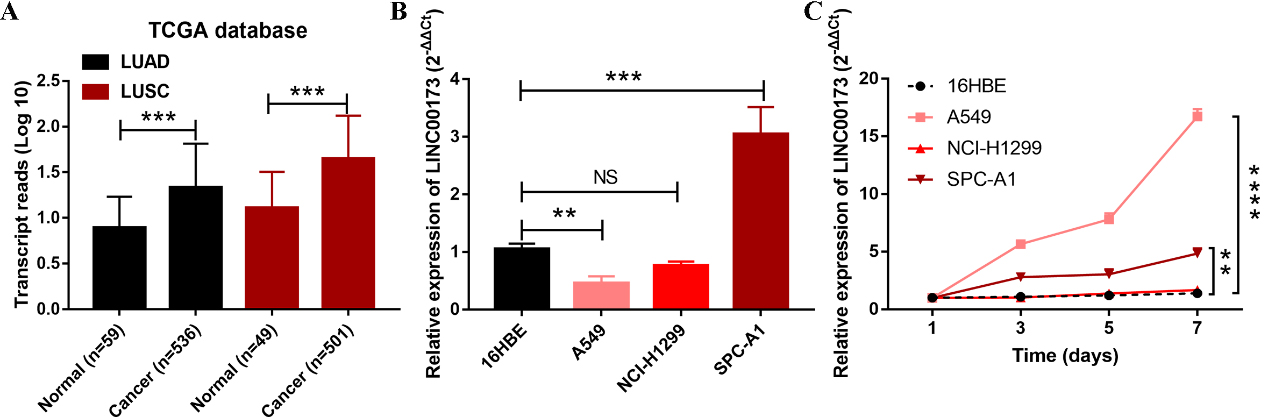
The origin of serum LINC00173 in NSCLC cases. (A) Expression levels of LINC00173 from TCGA database. (B) Expression levels of LINC00173 in 16HBE cells and three NSCLC cell lines. (C) LINC00173 was secreted into the culture medium by A549 and SPC-A1 cells in a time-dependent manner. Abbreviations: LUAD, lung adenocarcinoma; LUSC, lung squamous cell carcinoma.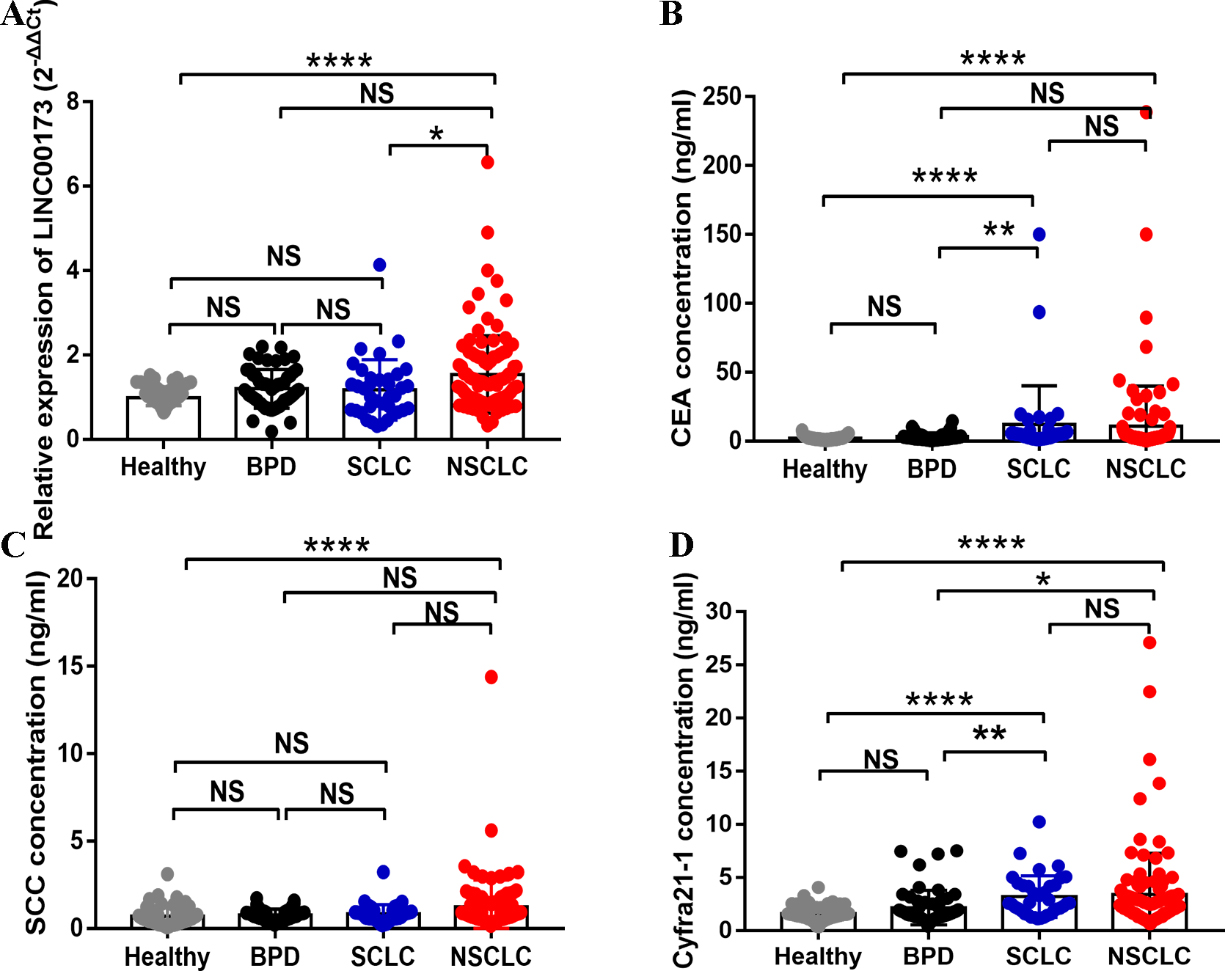
To explore the association between NSCLC and serum LINC00173, qRT-PCR was performed to detect the expression of LINC00173 in the 108 NSCLC patients, 55 BPD patients, 91 healthy donors and 37 SCLC patients. It was found that serum LINC0173 expression in the NSCLC patients was higher than that in the healthy donors and SCLC patients (Fig. 2A). However, there was no statistically significant difference between the healthy group, BPD group and SCLC group. Meanwhile, serum CEA, SCC and Cyfra21-1 levels were also elevated in the NSCLC patients, and CEA and Cyfra21-1 were also elevated in the SCLC group as compared with those in the healthy group (Fig. 2B–D).
Combination of serum LINC00173, CEA and Cyfra21-1 levels significantly improves the diagnostic sensitivity between NSCLC patients and healthy donors
Combination of serum LINC00173, CEA and Cyfra21-1 levels significantly improves the diagnostic sensitivity between NSCLC patients and healthy donors
Abbreviations: SEN, sensitivity; SPE, specificity; ACCU, overall accuracy; PPV, positive predictive value; NPV, negative predictive value.
Diagnostic utility of serum LINC00173 in NSCLC patients. (A) Evaluation of diagnostic efficiency of LINC00173 in differentiating NSCLC patients from healthy donors. (B) Evaluation of diagnostic efficiency of LINC00173 in differentiating NSCLC patients from patients with BPD. (C) Evaluation of diagnostic efficiency of LINC00173 in differentiating NSCLC patients from SCLC patients. (D) ROC analysis of single or combined use of serum LINC00173, CEA, SCC and Cyfra21-1 in differentiating NSCLC patients from healthy donors. (E) ROC analysis of single or combined use of above indicators in differentiating NSCLC patients from SCLC patients. Abbreviations: NSCLC, non-small-cell lung cancer; BPD, benign pulmonary disease; SCLC, small-cell lung cancer.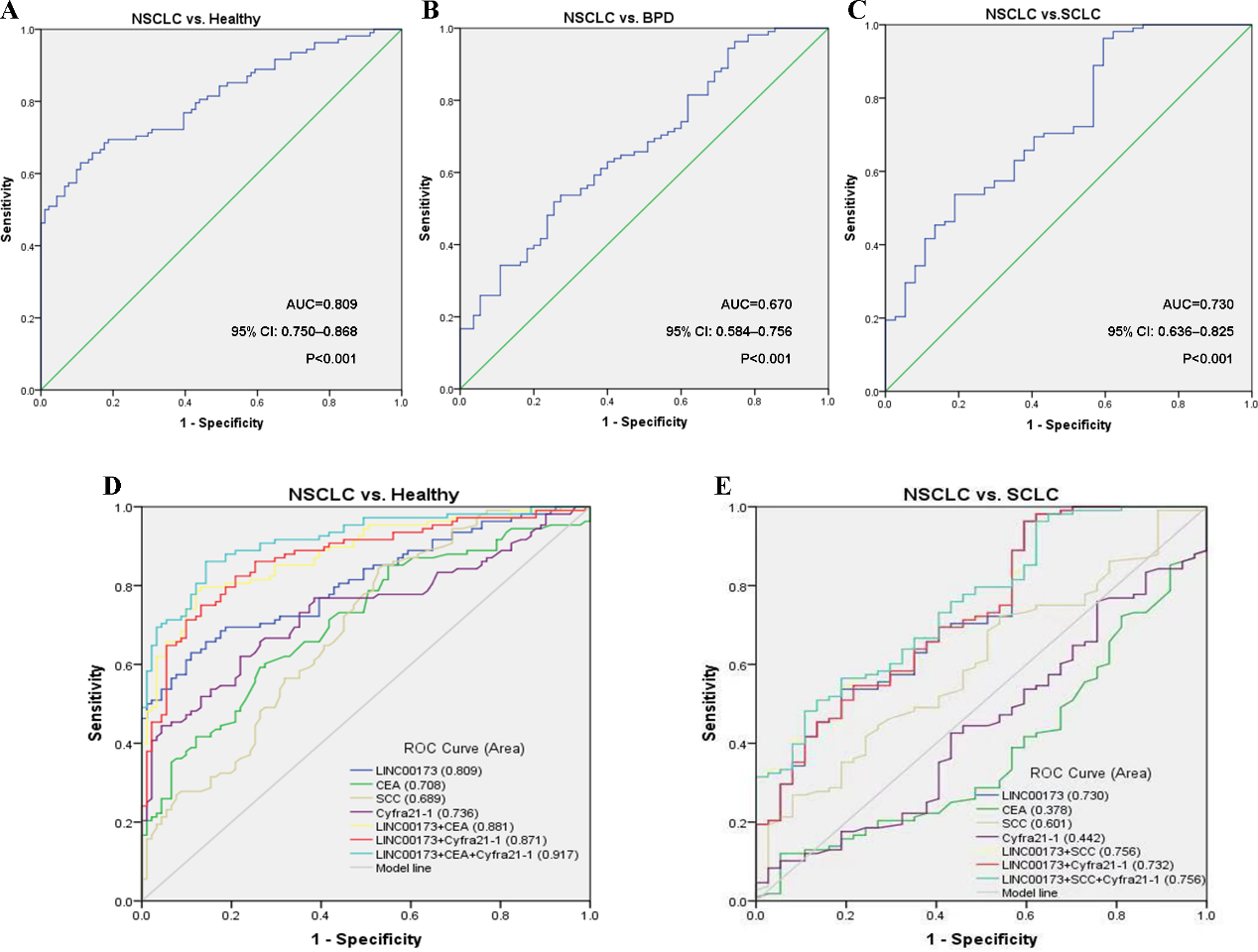
To figure out the origin of serum LINC00173 in NSCLC patients, we firstly search the TCGA database, including LUAD database and LUSC database. Results showed that LINC00173 was highly expressed in both LUAD and LUSC patients (Fig. 3A). We also detected the expression of LINC00173 in three NSCLC cell lines, and found LINC00173 was only upregulated in SPC-A1 cells (Fig. 3B). The discrepancy in LINC00173 expression trends in serum, cultured cells and tissue samples from the TCGA database caused us great curiosity. We then explored whether LINC00173 was secreted by NSCLC cells. We detected the expression of LINC00173 in the culture medium of three NSCLC cell lines (A549, NCI-H1299 and SPC-A1) and human bronchial epithelial cell 16HBE. Results showed that compared with 16HBE cells, the level of LINC00173 secretion in SPC-A1 and A549 cells increased with increasing incubation time (Fig. 3C). Therefore, we speculate that high levels of LINC00173 in peripheral blood circulation may originate from some certain tumor cells.
Diagnostic utility of serum LINC00173 in differentiating the NSCLC patients from the healthy donors, BPD and SCLC patients
To investigate the potential of serum LINC00173 as a diagnostic biomarker for NSCLC, ROC curves were constructed. It was found that serum LINC00173 effectively differentiated the primary NSCLC patients from the healthy donors, with an AUC of 0.809 (95% CI: 0.750–0.868,
Small-cell lung cancer (SCLC) was another type of lung cancer, accounting for 13% of all newly diagnosed lung cancers. Most SCLC patients were diagnosed at advanced stages when early and distant metastasis has occurred, resulting in a low cure rate [13, 14]. On the basis of the above findings, we wondered whether serum LINC00173 was differentially expressed in SCLC patients vs. NSCLC patients. As shown in Fig. 4C, serum LINC00173 showed a good diagnostic performance in discriminating between NSCLC and SCLC (AUC
Combination of LINC00173, CEA and SCC to construct a novel diagnostic model for NSCLC
Compared with the three existing clinical indicators, serum LINC00173 offered the highest AUC in differentiation between the NSCLC and the healthy donors (0.809, 95% CI: 0.750–0.868) (Fig. 4D), followed by Cyfra21-1 (0.736, 95% CI: 0.667–0.806) and CEA (0.708, 95% CI: 0.637–0.779). In this study, SCC displayed a relative low diagnostic value for NSCLC (AUC
Correlation between serum LINC00173 expression and clinicopathological parameters of NSCLC
The clinicopathological parameters of the 108 NSCLC patients are summarized in Table 3. According to the cut-off value of serum LINC00173, the 108 NSCLC patients were classified into two groups: a high-expression group (
The correlation between LINC00173 expression and clinicopathologic parameters of NSCLC patients
The correlation between LINC00173 expression and clinicopathologic parameters of NSCLC patients
Statistical analyses were carried out using Pearson
To verify the association between serum LINC00173 expression and tumor dynamics, we compared LINC00173 expression in the 108 primary NSCLC patients, 40 patients who received chemotherapy or surgical treatment, and 20 patients with tumor recurrences. It was found that the serum LINC00173 expression in the post-treatment patients (including surgical resection and chemotherapy) was significantly lower than that in the pre-treatment NSCLC patients and recurrent patients (
Role of LINC00173 in monitoring tumor dynamics in NSCLC patients. (A) Detection of serum LINC00173 expression in NSCLC patients (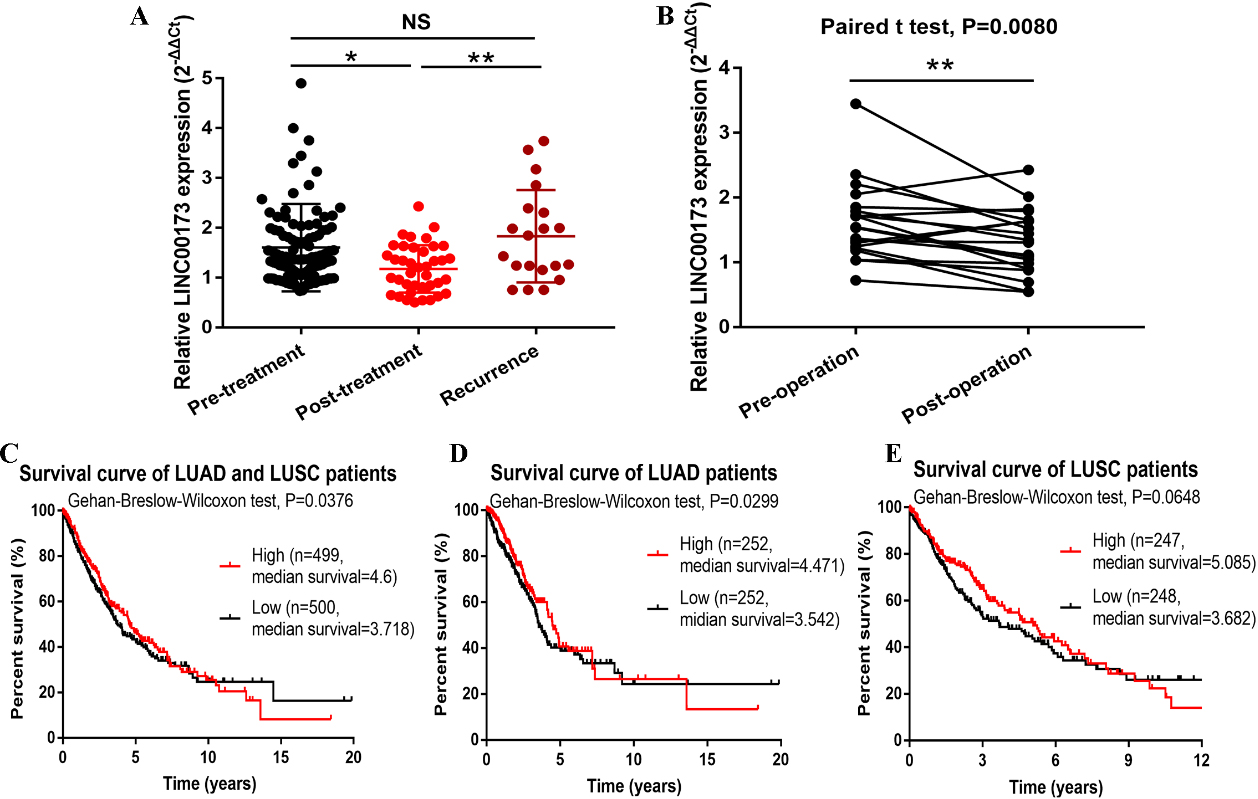
Lung cancer remains one of the most frequent malignancies in the world, contributing to almost one-quarter of all cancer-related deaths [15]. According to the histological classification, it contains two main groups: SCLC (15%) and NSCLC (85%), and the latter can be subcategorized into LUAD, LUSC and large cell carcinoma [16, 17]. Based on newly identified molecular profiles and targetable genetic alterations, the above categories can be further subdivided into several different types, symbolizing high heterogeneity and difficult treatment of the troublesome disease [16, 17]. Early diagnosis of lung cancer can not only make early treatment possible but also reduces the probability of tumor metastasis, thus achieving a better prognosis. Under this premise, exploring circulating tumor cells and circulating tumor DNA or RNA as a diagnostic marker, also known as “liquid biopsies”, is gaining widespread attention owing to their non-invasiveness and low cost [18, 19].
The whole genome and transcriptome sequencing has opened a new prelude to the exploration of lncRNA, which used to be considered as transcriptional “noise” or cloning artifacts [20]. According to the location relative to the nearest protein-coding transcripts, lncRNAs can be categorized as exonic, intronic, overlapping or intergenic [21]. The wide diversity of human lncRNAs is consistent with their diverse roles in gene expression, and therefore aberrant lncRNA expression can cause various human diseases and disorders [22]. Monitoring these differentially expressed lncRNAs is beneficial to perceive the abnormal state of the human body, so as to achieve the purpose of early diagnosis. Wang et al. [23] reported that pleural effusion lncRNA was effective in differentiating lung cancer-associated malignant pleural effusion from benign pleural effusion. Lin et al. [24] reported that 7 of the 26 lncRNAs could be reliably detected in the plasma and combined use of plasma lncRNA as a biomarker signature offered a 84.13% sensitivity and a 87.88% specificity for the diagnosis of lung cancer. Besides, some specific lncRNAs are also evaluated for diagnostic efficacy. LncRNA XLOC_009167 was found to be highly expressed in whole blood of lung cancer patients as compared with that in healthy controls or pneumonia patients, with an AUC of 0.7398 and 0.7005 respectively [25]. Li et al. [26] reported that linc00152 had high diagnostic accuracy for NSCLC, and combination of Linc00152 and CEA could provide more powerful diagnosis efficiency than Linc00152 or CEA alone.
LINC00173 is an uncharacterized lincRNA mapped to chromosome 12q24.22. Several studies have reported the involvement of LINC00173 in infections [8], granulocytes [9] and esophageal squamous cell carcinoma [10]. In the present study, we detected serum LINC00173 expression in the lung cancer patients by qRT-PCR with good linearity, specificity and reproducibility. We discovered that serum LINC00173 expression in patients with primary NSCLC was significantly higher than that in healthy donors and SCLC patients. However, there was no statistically significant difference between the healthy donor group, BPD group and SCLC group. We also found that serum LINC00173 could clearly distinguish NSCLC patients form SCLC patients, which increased its clinical utility. Additionally, clinicopathological parameters showed that high serum LINC00173 was associated with lymph node metastasis and distant metastasis. Therefore, we subsequently constructed a novel diagnostic model that combined the detection of LINC00173 with CEA and Cyfra21-1. The result showed that it significantly improved the diagnostic efficiency in distinguishing NSCLC patients from healthy donors. Besides, combined detection of LINC00173 and Cyfra21-1 to differentiate between NSCLC and SCLC achieved even higher diagnostic efficiency than either of them alone. Moreover, serum LINC00173 was then significantly decreased in patients after surgery or chemotherapy to a level similar to that in the healthy donors, which to some extent forebodes improvement or no progression of the disease. In addition, the expression of serum LINC00173 rebounded in patients with tumor recurrences, implying the ability of LINC00173 in monitoring tumor dynamics.
However, for comparison between pre- and post-operative patients, one patient showed an increase in LINC00173 after surgery. Whether this increase was due to an early relapse or just an accidental individual difference or other reasons needs to be further clarified in future larger-sample studies. Besides, we observed that NSCLC patients (especially LUAD patients) with high LINC00173 expression had a lower overall survival rate, though this difference is not statistically significant for patients with a survival time of less than 10 years. Moreover, it is doubtful that the median survival time of the high-expression group was longer than that of the low-expression group. This contradictory phenomenon may due to that the data downloaded from the TCGA database was mostly derived from tissue samples, and the dynamic change of LINC00173 expression in tumor tissues may not be sensitive as that in serum samples. In addition, the discrepancy in LINC00173 expression trends in serum samples and cultured cells suggested that high levels of LINC00173 in peripheral blood circulation might be derived from some certain tumor cells. But we cannot exclude the possibility that LINC00173 may also be secreted by other cells in the tumor microenvironment, such as tumor stromal cells.
In conclusion, increased serum LINC00173 could be a potential diagnostic biomarker for NSCLC and monitoring tumor dynamics. However, the clinical value of serum LINC00173 suggested in this preliminary study should be viewed prudently due to the limited sample size. Future larger-sample studies and long-term observation are required to gain new insights into the mechanism underlying the diagnostic and prognostic value of LINC00173.
Footnotes
Acknowledgments
This project was supported by grants from the National Natural Science Foundation of China [grant number: 81871720], Jiangsu Program for Young Medical Talents [grant number: QNRC2016695] and Jiangsu Innovation and Entrepreneurship Training Program for Undergraduates [grant number: 201810304076Y].
Conflict of interest
The authors declare no conflicts of interest in the publication of this paper.
Supplementary data
Selection of a suitable internal reference for LINC00173 Abbreviations: SD, standard deviation; CV, coefficient of variation.
Patient
Healthy donor
-actin
GAPDH
18S rRNA
-actin
GAPDH
18S rRNA
Mean
26.74
28.93
22.18
27.70
30.67
22.61
CV (%)
5.30%
7.22%
5.31%
1.32%
1.84%
1.29%
