Abstract
Introduction
In this paper, we studied several serum clinical chemistry tests of cardiovascular disease (CVD), iron deficiency anemia, liver and kidney disorders in migraine.
Methods
We first explored the association of 22 clinical chemistry tests with migraine risk in 697 migraine patients and 2722 controls. To validate and interpret association findings, cross-trait genetic analyses were conducted utilising genome-wide association study (GWAS) data comprising 23,986 to 452,264 individuals.
Results
Significant associations with migraine risk were identified for biomarkers of CVD risk, iron deficiency and liver dysfunction (odds ratios = 0.86–1.21; 1 × 10−4 < p < 3 × 10−2). Results from cross-trait genetic analyses corroborate the significant biomarker associations and indicate their relationship with migraine is more consistent with biological pleiotropy than causality. For example, association and genetic overlap between a lower level of HDL-C and increased migraine risk are due to shared biology rather than a causal relationship. Furthermore, additional genetic analyses revealed shared genetics among migraine, the clinical chemistry tests, and heart problems and iron deficiency anemia, but not liver disease.
Conclusions
These findings highlight common biological mechanisms underlying migraine, heart problems and iron deficiency anemia and provide support for their investigation in the development of novel therapeutic and dietary interventions.
Keywords
Glossary
ALB: serum albumin; ALT: alanine aminotransferase; ALP: alkaline phosphatase; AST: aspartate aminotransferase; Ca: calcium; CI: confidence interval; CRP: C-reactive protein; CVD: cardiovascular disease; GGT: gamma- glutamyltransferase; GWAS: genome-wide association study; HDL-C: high-density lipoprotein cholesterol; ICHD-2: International Classification of Headache Disorders 2nd edition; IHGC: International Headache Genetics Consortium; IQR: interquartile range; LCV: latent causal variable; LD: linkage disequilibrium; LDL-C: low-density lipoprotein cholesterol; LDSC: LD score regression; MA: migraine with aura; matSpD: matrix spectral decomposition; MO: migraine without aura; MR: Mendelian randomisation; OR: odds ratio; RICDF: rank-based inverse cumulative distribution function; CREAT: serum creatinine; FERRI: serum ferritin; SECA: SNP effect concordance analysis; SNP: single nucleotide polymorphism; UA: serum uric acid; TB: total bilirubin; TC: total cholesterol; TP: total protein; TS: transferrin saturation; TF: transferrin; TG: triglyceride
Introduction
Despite massive efforts in migraine research, many uncertainties surround our understanding of migraine. In this regard, identification of new biomarkers, including serum clinical chemistry tests, will provide new insights into migraine biology.
The level of serum clinical chemistry (biochemistry) tests has been clinically utilised for diagnosing unfavorable health conditions and monitoring body organs’ function (1). Since migraine is associated with increased risk of cardiovascular disease (CVD), ischemic stroke, and other cardiovascular events (2), there is a particular interest in testing associations between migraine and CVD morbidity tests such as lipid levels and circulating inflammatory biomarkers such as C-reactive protein (CRP). While associations between migraine and increased serum levels of lipids and CRP have been shown (3), conflicting evidence still exist (4).
To extend biological understanding of migraine and its possible connection with other conditions, we tested routine CVD risk, liver function, kidney function, and iron status chemistry tests for association with migraine status. Furthermore, we used genome-wide association study (GWAS) summary statistics data to interpret the findings by addressing three questions. First, do the epidemiological findings reflect a shared genetic basis underlying migraine and the associated chemistry tests? Second, do the findings link migraine to the relevant medical disorders? Third, are the identified associations and genetic correlations due to cause-and-effect relationships or shared biological mechanisms (biological pleiotropy)?
Materials and methods
In the current study, we first aimed to test the association between migraine and 22 clinical chemistry tests traditionally used to screen CVD risk, iron homeostasis status, liver and kidney functions using epidemiological data. Second, we carried out cross-trait genetic analyses utilising GWAS summary statistics of migraine, the observationally associated biomarkers and their medically relevant disorders.
Epidemiological data
We used up to 22 clinical chemistry results of 3419 unrelated individuals randomly drawn from three QIMRB (QIMR Berghofer Medical Research Institute) Australian twin cohorts ascertained from and representative of the Australian general population, between January 1991 and December 2005. Only individuals with self-reported European ancestry were included (5). Clinical chemistry tests were measured using Hitachi or Roche 917 or Modular P analysers and Roche reagents (Roche Diagnostics Australia, North Ryde NSW 2113) at Royal Prince Alfred Hospital, Sydney. The study was approved by the QIMRB Human Research Ethics Committee.
Migraine was diagnosed according to diagnostic criteria of the 2004 International Classification of Headache Disorders (ICHD-2) (6). To collect detailed ICHD-2 diagnostic criteria, participants who answered “yes” to “ever having migraine or recurrent attacks of headache” (screening positive), were asked several questions regarding their symptoms. Subjects also answered questions regarding demographic characteristics (e.g. sex, date of birth, etc.) via a semi-structured telephone interview and/or online questionnaire. This study includes a total of 3419 unrelated individuals, of 697 migraine cases (age 25–78 years; sex, 550 females and 147 males) and 2722 migraine-free individuals (age 25–91 years; sex, 1512 females and 1210 males). Migraine cases satisfied ICHD-2 criteria for migraine without aura (MO) and/or migraine with aura (MA), using visual prodromal symptoms as an index of MA.
The following serum clinical chemistry panels were investigated: Traditional markers of CVD, including lipid panel (total cholesterol, TC; high-density lipoprotein cholesterol, HDL-C; low-density lipoprotein cholesterol, LDL-C; triglyceride, TG) and an inflammation marker (C-reactive protein, CRP), iron status panel (serum iron, IRON; serum ferritin, FERRI; transferrin, TF; transferrin saturation, TS), liver panel (total bilirubin, TB; alkaline phosphatase, ALP; alanine aminotransferase, ALT; aspartate aminotransferase, AST; gamma- glutamyltransferase, GGT), renal panel (serum creatinine, CREAT; blood urea, UREA; serum uric acid, UA), and also biomarkers that their abnormal values can be caused by liver or kidney problems (7), including proteins (total protein, TP; serum albumin, ALB) and calcium (Ca). In addition to these tests, the ratios of TC/HDL-C, known as cholesterol ratio, and TG/HDL-C were calculated and added into the clinical chemistry tests (Table 1).
Characteristics of individuals and clinical chemistries laboratory (raw) values presented as median (IQR) among studied groups.

*The concentrations of TG and HDL-C were converted to mg/dL prior to taking the ratio, and multiplied by 88.57 and 38.67, respectively.
n: number of observations; IQR: interquartile range; CVD: cardiovascular disease; TC: total cholesterol; HDL-C: high-density lipoprotein cholesterol; LDL-C: low-density lipoprotein cholesterol; TG: triglyceride; CRP: c-reactive protein; IRON: serum iron; FERRI: serum ferritin; TF: transferrin; TS: transferrin saturation; TB: total bilirubin; ALP: alkaline phosphatase; ALT: alanine aminotransferase; AST: aspartate aminotransferase; GGT: gamma-glutamyltransferase; CREAT: serum creatinine; UREA: blood urea; UA: serum uric acid; TP: total protein; ALB: serum albumin; Ca: calcium.
Genetic data
Genetic analyses were only performed for migraine, migraine-associated chemistry tests (HDL-C, LDL-C, TG, IRON, TS, TB and GGT) and their medically relevant disorders (heart/cardiac problems, iron deficiency anemia and diseases of the liver). We used GWAS summary statistics, all from European ancestry, like epidemiological data.
Migraine GWAS
In 2016, the International Headache Genetics Consortium (IHGC) performed meta-analysis of several European GWAS datasets. Briefly, migraine was diagnosed according to the ICHD criteria after either self-reported questionnaires or clinical interviews and association tests between each SNP and migraine status (8). Herein, we analysed summary statistic meta-analysis data after excluding results from the 23andMe GWAS. This data includes association results of 8,094,884 SNPs from analysis of 29,209 migraine cases and 172,931 migraine-free controls.
Clinical chemistry tests GWAS
GWAS summary statistics of five blood chemistry tests, including HDL-C, LDL-C, TG, TB and GGT (phenotype codes are “30760_irnt”, “30780_irnt”, “30870_irnt”, “30840_irnt” and “30730_irnt”, respectively) were obtained from round 2 of analysing UK Biobank data, released on 1 August 2018 by Ben Neale’s lab (9). These data include allelic association results for 13,791,467 SNPs from analysis of quantile normalised chemistry test values in 315,133 to 344,104 individuals.
Allelic association data of iron status biomarkers, IRON and TS, were gained from the published meta-analysis of 11 GWAS results comprising 23,986 participants of European ancestry in total and included association results for 2,096,456 to 2,102,225 SNPs (10).
Notably, there are no GWAS results publicly available for TC/HDL-C and TG/HDL-C ratios.
Medically relevant disorders GWAS
GWAS summary statistics for three potentially relevant disorders, “heart/cardiac problem” (UK Biobank fields: 20002; cases = 32,474; controls = 419,790), “D50 Iron deficiency anaemia” (UK Biobank fields: 41202, 41204; cases = 8405; controls = 443,859) and “K70-K77 Diseases of liver” (UK Biobank fields: 41202, 41204; cases = 5472; controls = 446,792), were taken from UK Biobank Gene Atlas (11) comprising 452,264 participants and 34,649,508 SNPs.
Epidemiological association analysis
All pre-processing and tests were performed with the statistic software R version 3.6.2 (2019-12-12).
Initially, the raw measurements were transformed using the rank-based inverse cumulative distribution function (RICDF) given as Φ−1 (rank (xi)/(length (x) + 1)), where Φ−1 donates the quantile of the normal distribution and x is a vector of the measurements.
A logistic regression model was fitted using the migraine status as the binary response and each chemistry test transformed measurement as a potential predictor. To adjust for biological and technical confounders, age, age2, sex, the interaction of sex and age, the interaction of sex and age2, and the study batches (i.e. a five-level factor variable was used to represent the five separate batches) were added into the model as potential covariates.
Moreover, because of sex-based differences in migraine biology and clinical chemistry measurements, sex-stratified analyses were performed. Notably, the dataset was stratified by sex before the normalisation in order to avoid any leak of information between sexes. In sex-stratified analyses, we used the same regression model but without sex, the interaction of sex and age, the interaction of sex and age2 covariates.
To deal with the multiple testing problem, Bonferroni correction with an effective number of independent tests estimated using the spectral decomposition of a correlation matrix method (matSpD) to adjust p values (12). Briefly, the 22 clinical chemistry measurements were estimated to be equivalent to 17.09 effective independent measurements in the full dataset. We also used a permutation approach with 50,000 iterations to empirically correct for multiple tests. In each iteration, labels of migraine status were randomly shuffled and then the mentioned multivariable logistic regression models were fitted to calculate corresponding p values for testing association of the 22 clinical chemistry tests with migraine. Afterwards, we determined that how often there would be the same number of associated findings with p < 0.05 to estimate permuted (empirical) p value.
Genetic analyses
Where it was needed to include only the independent SNPs, linkage disequilibrium (LD) reference data of the European panel of 1000 Genomes Project Phase 1 version 3 with minor allele frequency (MAF) of > 0.01 was used to perform p value informed clumping using PLINK 1.9 (13).
Estimation of cross-trait genetic overlap
To further investigate epidemiological findings, cross-trait LD score regression (LDSC) (14) was applied to estimate genetic correlation (rg) between migraine and those biochemistry tests that reached significance level of p < 0.05 in the epidemiological analyses and their corresponding medical traits. Briefly, the cross-product of two z-scores from two GWAS is regressed on the LD score pre-calculated using the European panel of 1000 Genomes Project. The slope from this regression is used to estimate the genetic covariance between two traits.
Moreover, to further examine directions of the genetic overlaps among subsets of SNPs, SNP effect concordance analysis (SECA) was performed (15). SECA tests 144 associations between effect directions of 12 independent SNP subsets in GWAS summary statistics of trait 1 and trait 2 (a matrix of 12 × 12 combinations) using Fisher’s test and calculates an empirical p value for the number of significant associations (adjusted for testing 144 subsets). Notably, the independent SNP subsets are a product of two rounds of p-value informed LD clumping (first clumping: –clump-r2 0.1; –clump-kb 1000; second clumping: –clump-r2 0.1; –clump-kb 10000) and then categorising them into 12 groups based on p-value thresholds of ≤ (0.01, 0.05, 0.1, 0.2, 0.3, 0.4, 0.5, 0.6, 0.7, 0.8, 0.9, 1.0) (15).
Test of causality and biological pleiotropy
Comorbidity and genetic correlation between traits point to the pleiotropy that may be a part of common genetic factors underlying traits (known as biological or horizontal pleiotropy) or may indicate a causal relationship between traits (known as vertical pleiotropy). To investigate whether the epidemiological and genetic associations are more consistent with causality than biological pleiotropy, the Mendelian Randomization (MR) methods (16) and latent causal variable (LCV) model (17) were implemented.
To assess the effect of the associated biochemistry tests (as exposures) on migraine (as an outcome), we implemented two-sample MR analysis. We implemented three different MR methods: The inverse variance weighted (IVW), MR-Egger regression and weighted median (WM) using a two-sample MR package in R (16). In addition, as sensitivity analyses, heterogeneity and horizontal pleiotropy (e.g. biological pleiotropy) were tested by Q statistic and the intercept term of MR-Egger regression. We used the effect estimates of the SNPs suggestively associated with biochemistry levels (p < 1 × 10−5) as genetic instruments. To include only the independent instrumental variables, SNPs in LD of r2 > 0.01 with a more significantly associated SNP not farther than 1000 kb were considered as dependent instruments (–clump-r2 0.01; –clump-kb 1000) and were therefore excluded from MR analyses.
Moreover, the LCV model was carried out using the available tutorial and R codes (17). This approach calculates a genetic causality proportion (GCP) of phenotype 1 on phenotype 2 that GCP of zero means no genetic causality and GCP of one means full genetic causality (17). Briefly, the LCV model produces latent variables that mediate genetic correlation between two phenotypes of interest. Where phenotype 1 has a genetic correlation with the latent variables stronger than phenotype 2, phenotype 2 is genetically caused by phenotype 1.
To study the presence of pleiotropy at the gene level, we next applied multi-marker analysis of genomic annotation (MAGMA) v1.07b (18) to assign SNPs to protein-coding genes by adding an annotation window of 2 kb around genes and then performing gene-based analysis using the matched Ensembl build version of the SNP-gene annotations and the genome reference panel (1000 Genomes Project Phase 3). This analysis is limited to autosomal genes presenting in all GWAS summary statistics (16,879 genes). We tested whether genes associated with migraine risk are statistically over-represented in the genes associated with clinical chemistry test levels and medically relevant traits by estimating the Fisher’s exact test one-sided p-value.
Data availability
To express your interest in epidemiological data, please contact the corresponding author. To express your interest in migraine GWAS data, please contact the data access committee. All other GWAS summary statistics except migraine are publicly available. All codes used to perform statistical analyses are available from the corresponding author upon request.
Results
Epidemiological associations
The baseline characteristics of 3419 samples and their recorded raw laboratory values of 22 clinical chemistry tests and two ratios are summarised in Table 1.
The multivariable-adjusted association between migraine status and normalised values of chemistry tests are shown in Table 2. Nine out of 22 chemistry tests, including levels of five CVD risk biomarkers, levels of two iron status biomarkers and two liver function biomarkers, reached the nominal significance threshold (p < 0.05). The strongest association was between a higher TC/HDL-C ratio and increased migraine risk across all analyses (both sexes, OR = 1.16, p = 0.0001; female, OR = 1.17, p = 0.0038; male, OR = 1.29, p = 0.0044). We note they did not markedly differ from the non-sex-stratified results (results are not shown).
Epidemiological association between migraine status and clinical chemistries level.
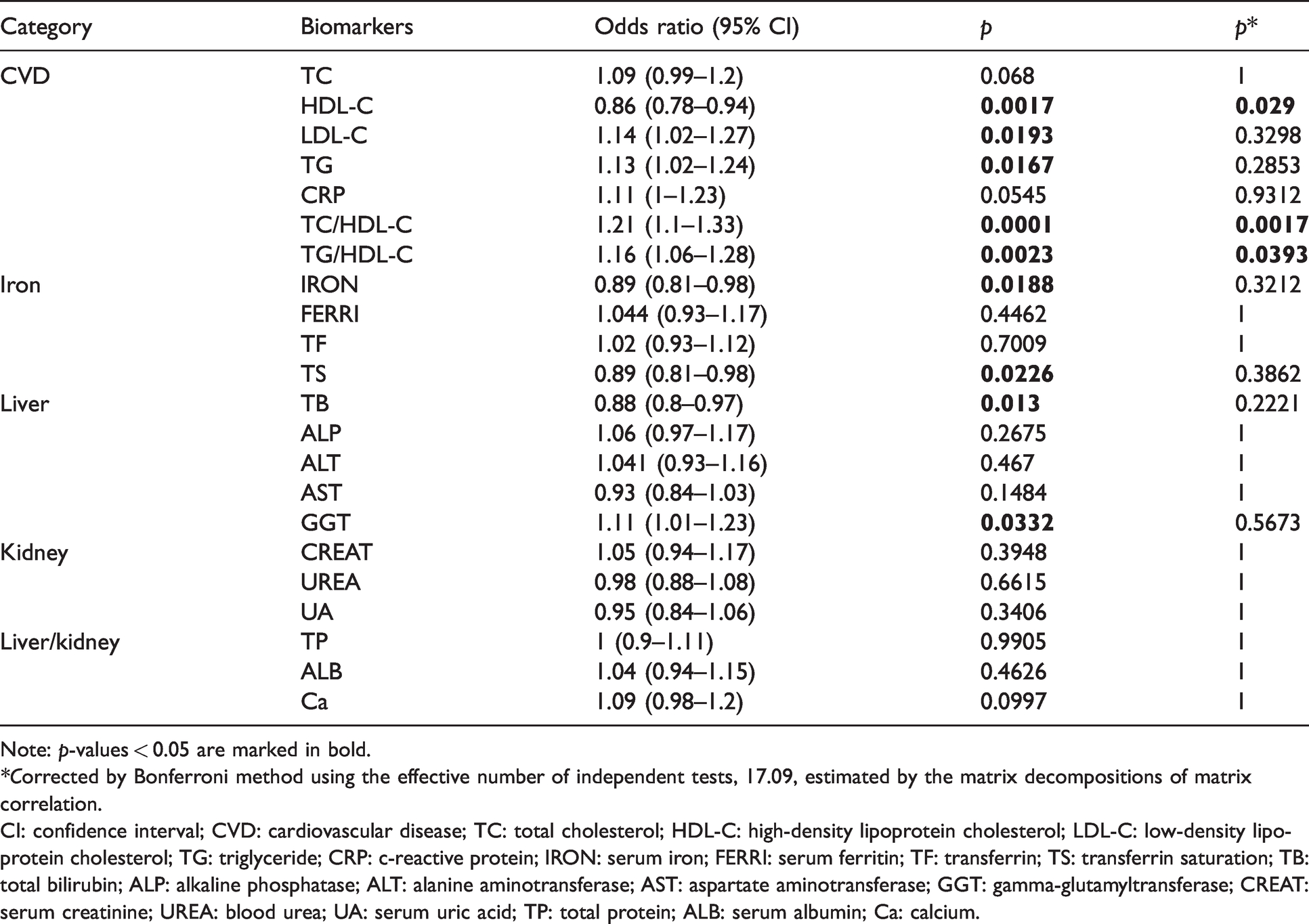
Note: p-values < 0.05 are marked in bold.
*Corrected by Bonferroni method using the effective number of independent tests, 17.09, estimated by the matrix decompositions of matrix correlation.
CI: confidence interval; CVD: cardiovascular disease; TC: total cholesterol; HDL-C: high-density lipoprotein cholesterol; LDL-C: low-density lipoprotein cholesterol; TG: triglyceride; CRP: c-reactive protein; IRON: serum iron; FERRI: serum ferritin; TF: transferrin; TS: transferrin saturation; TB: total bilirubin; ALP: alkaline phosphatase; ALT: alanine aminotransferase; AST: aspartate aminotransferase; GGT: gamma-glutamyltransferase; CREAT: serum creatinine; UREA: blood urea; UA: serum uric acid; TP: total protein; ALB: serum albumin; Ca: calcium.
Although only three chemistry tests (HDL-C, TC/HDL-C and TG/HDL-C) passed the modified Bonferroni correction threshold (p < 0.05/17.09), a 50,000-iterations permutation test showed that the chance of observing nine associated biomarkers (p < 0.05) is rare and in fact is 10/50,000. Therefore, all migraine-associated biomarkers with nominal p < 0.05 were subjected to further genetic analyses.
Among CVD risk biomarkers, HDL-C was significantly lower, and LDL-C, TG, TC/HDL-C and TG/HDL-C ratios were significantly higher in migraine cases compared to controls. Notably, we observed marginal association between a higher level of CRP and migraine risk (OR = 1.11, p = 0.0545). Moreover, two iron status biomarkers, IRON and TS, were significantly decreased in migraine compared to controls. Among liver function biomarkers, lower levels of TB and higher levels of GGT were associated with migraine status.
Genetic overlap
Where GWAS summary statistics were available, we tested cross-trait genetic overlap between migraine and its seven nominally associated chemistry tests and their related medical traits to verify our epidemiological associations.
LDSC estimated genetic correlations between migraine and four observationally associated chemistry tests, including HDL-C (rg = −0.13, p = 0.0001), TG (rg = 0.14, p = 5.8 × 10−6), TB (rg = −0.07, p = 0.0483) and GGT (rg = 0.13, p = 1.1 × 10−5) with the direction of effects (rg) agreeing with our observational findings (Table 3). Notably, LDSC was unable to estimate genetic correlation between serum levels of IRON and migraine risk, likely due to the inadequate power of the IRON GWAS. Consistent with epidemiological findings and LDSC results, SECA identified that the genetic effects on migraine are concordant with lower levels of three tests, including HDL-C, IRON, TS, and higher levels of two tests, TG and GGT (Table 3). Together, genetic correlations and concordances resembled phenotypic associations.
Genetic correlation and concordance between migraine and clinical chemistry tests.

Note: p-values < 0.05 are marked in bold.
aThe ratio of the number of SNP subsets with significant concordant effects between migraine and the other trait.
bPermuted P for observing ≥ significant SNP subsets as the observation.
LDSC: linkage disequilibrium score regression; SECA: SNP effect concordance analysis; rg: genetic correlation; s.e.: standard error; CI: confidence interval; CVD: cardiovascular disease; HDL-C: high-density lipoprotein cholesterol; LDL-C: low-density lipoprotein cholesterol; TG: triglyceride; IRON: serum iron; TS: transferrin saturation; TB: total bilirubin; GGT: gamma-glutamyltransferase.
To extend these results, we estimated genetic correlations for 24 pairs of the four binary traits, including migraine, heart/cardiac problems, iron deficiency anemia and diseases of the liver, and five significantly associated chemistry tests (Figure 1). To minimise the number of tests, genetic correlations of LDL-C with other traits were not tested because neither LDSC nor SECA found significant genetic overlap between migraine and LDL-C. Likewise, IRON GWAS was not tested because it has a low SNP-based heritability, likely due to its small sample size.

Genetic correlations among migraine, the medically relevant disorders, and five biochemistry tests. Different statistical significance thresholds are shown as asterisks.
We identified positive genetic correlations between migraine and two medical traits of heart problems (rg = 0.16, P = 0.0057) and iron deficiency anemia (rg = 0.21, P = 0.0444) (Figure 1). Interestingly, we found that migraine, heart/cardiac problems and iron deficiency anemia traits have a strong similar pattern of significant genetic correlations with clinical chemistry tests (Figure 1). Although two liver function tests, TB and GGT, were phenotypically and genetically correlated with migraine risk, no genetic correlation was found between migraine and liver diseases by LDSC (rg = −0.01, p = 0.9114). Moreover, we found no significant genetic correlation between iron deficiency anemia and level of TS, which is likely due to the limited power of the TS GWAS (e.g. low heritability and/or small sample size). Despite TB level being a well-known biomarker for assessing liver function, we found no genetic correlation between TB level and liver disease, whereas we found that TB and GGT liver function biomarkers have strong genetic correlations with heart problems and iron deficiency anemia traits (p < 0.05/24) suggesting evidence against involvement of liver function and in favor of CVD and iron homeostasis in migraine (Figure 1).
Potential causal relationships
We performed two-sample MR and LCV model analyses to identify potential credible causal influences of the clinical chemistry tests and the medical traits on migraine risk. While the LCV modelled no cross-traits genetic causality (0.9798 < p < 0.2108), two-sample MR methods indicate weak causal effects of lower HDL-C level (IVW method, β = −0.06, p = 0.0219) and higher levels of LDL-C (IVW method, β = 0.08, p = 0.0128) and GGT (IVW method, β = 0.08, p = 0.0031) on migraine risk. However, strong evidence for SNP heterogeneity was indicated by the Cochran’s Q value of the MR-Egger regression test, suggesting pleiotropic SNPs simultaneously influencing levels of the chemistry tests and migraine risk (8.8 × 10−12 < p < 1.3 × 10−5). In other words, our analyses revealed that the relationships between migraine and the tested traits are most likely due to biological pleiotropy (horizontal pleiotropy) rather than causality (vertical pleiotropy). Causal inference analyses are presented in Table 4 and 5.
Two-sample Mendelian randomization results between migraine (as the outcome) and the other traits (as the exposures).

Note: p-values < 0.05 are marked in bold.
β: beta coefficient; s.e.: standard error; INT: intercept; HDL-C: high-density lipoprotein cholesterol; LDL-C: low-density lipoprotein cholesterol; TG: triglyceride; IRON: serum iron; TS: transferrin saturation; TB: total bilirubin; GGT: gamma-glutamyltransferase.
Testing causality between migraine and the other traits by Latent Causal Variable (LCV) model.

GCP: genetic causality proportion; s.e.: standard error; ; HDL-C: high-density lipoprotein cholesterol; LDL-C: low-density lipoprotein cholesterol; TG: triglyceride; IRON: serum iron; TS: transferrin saturation; TB: total bilirubin; GGT: gamma-glutamyltransferase.
Genes influencing both migraine and other traits
To investigate biological pleiotropy at the level of genes (as genes are more connected to the biology), we implemented gene-based analyses using MAGMA (18). We performed Fisher’s exact test to examine whether the significant genes of migraine are enriched into the significant genes of the other traits (Table 6). For obtaining pleiotropic genes, we considered two levels of p < 0.05 and p < 0.05/16,879 (0.05/total number of genes). Out of 16,879 genes, 1746 and 27 genes are associated with migraine risk at p < 0.05 and p < 0.05/16,879, respectively. This analysis revealed that nominally associated genes (p < 0.05) of HDL-C, TG, TB and GGT are enriched in the migraine-associated genes (p < 0.05), replicating identified epidemiological and genetic overlaps (Table 6). Interestingly, while we found no SNP-based genetic overlap between the level of LDL-C and migraine (rg = 0.07, p = 0.1375), a gene-based pleiotropy between LDL-C and migraine was identified (OR = 1.28, p = 8.5 × 10−6), replicating our epidemiological association.
Genes associated with migraine risk overlapping with genes associated with other traits.

Note: p-values < 0.05 are marked in bold. Associated genes were selected at two different thresholds, p < 0.05 and p < 0.05/16,879. Gene a denotes the ratio of the number of genes associated with both migraine and the trait at p < 0.05 to the total number of genes are associated only with the trait at p < 0.05. Gene b denotes the ratio of the number of genes are associated with both migraine and the trait at p < 0.05/16,879 to the total number of genes are associated only with the trait at p < 0.05/16,879. One-sided p-values were estimated using Fisher’s Exact test.CVD: cardiovascular disease; HDL-C: high-density lipoprotein cholesterol; LDL-C: low-density lipoprotein cholesterol; TG: triglyceride; IRON: serum iron; TS: transferrin saturation; TB: total bilirubin; GGT: gamma-glutamyltransferase.
Moreover, although we found no enrichment for IRON and TS, associated genes of “iron deficiency anaemia” are enriched into the migraine-associated genes (OR = 1.20, p = 0.0274). Consistent with genetic correlation results, there are no pleiotropic genes between migraine and liver disease, suggesting that liver function is unlikely to be linked to migraine risk (p = 0.7795).
Interestingly, genes controlling the level of HDL-C are enriched into the genes influencing migraine risk in the stringent threshold analysis (OR = 3.67, p = 0.0072), suggesting strong shared biology between migraine and HDL-C level (Table 6). Six genes on chromosome 12, including MAP3K12, TARBP2, NAB2, STAT6 and LRP1, and one gene on chromosome 5, PRR13, are involved in controlling both the level of HDL-C and migraine risk with p < 0.05/16,879.
Discussion
The present study aimed to identify and characterise the association between migraine and 22 serum clinical chemistry tests traditionally used to monitor CVD, iron deficiency anaemia, liver, and kidney functions. Significant associations with migraine risk were identified for biomarkers of CVD risk, iron deficiency and liver dysfunction. Furthermore, cross-trait genetic analyses identified a shared genetic basis underlying migraine and the associated chemistry tests; linked migraine to heart problems and iron deficiency anemia, but not liver diseases; and indicated that the relationships between the identified biomarkers and migraine are more consistent with biological pleiotropy than causality. A summary of analyses workflow and findings is presented in Figure 2.

A summary of study workflow and findings.
The information in the literature on the relationship between migraine and CVD risk chemistry tests are inconsistent and limited to migraine subtypes. For example, the TC/HDL-C ratio was significantly higher in migraine without aura (MO) (4) patients in one study and in another study was limited to migraine with aura (MA) patients (19). Similarly, a level of TG was significantly higher in only patients with MA (20) and elsewhere shows no significant differences (21). This study strongly highlights higher risk of CVD among migraineurs, as we identified phenotypic and genetic overlap between migraine and a lower level of HDL-C, known as good cholesterol, and increased levels of CVD risk chemistry tests (22), including TC/HDL-C (cholesterol ratio), TG/HDL-C ratio, LDL-C and TG.
Consistent with our finding, a recent metabolome study found decreased levels of apolipoprotein A1 and free cholesterol to total lipid ratio in small HDL-C in migraine patients (23), reflecting a reduced level of HDL-C metabolism.
Since our study yielded the strongest significant result for TC/HDL-C ratio and it is a core risk factor in CVD risk-assessment models (24), we believe that this chemistry test is an ideal marker for investigating comorbidity and genetic overlap between migraine and CVD.
Due to lack of availability of TC/HDL-C ratio GWAS, we were unable to examine the shared genetic basis between migraine and this biomarker; hence, its role remains to be determined.
It is worthy of note that we observed a marginally significant association between a higher level of CRP, a risk factor incorporated in some CDV risk-prediction models (24), and migraine risk (OR = 1.11, p = 0.0545). Several studies found no significant epidemiological association between CRP and migraine risk (25). However, a significant higher level of CRP in self-reported female migraineurs was found (26). Similarly, significantly higher values of CRP in migraine patients have been observed by pooling 12 studies through a parametric meta-analysis (6980 migraine cases and 38,975 controls) (27). Together, CRP level may have a very small or negligible effect on migraine risk and therefore, its association with migraine is highly varied across different studies due to technical and biological confounders.
In the most common types of anemia, anemia of chronic disease and iron deficiency anemia, levels of serum IRON and TS are reduced (28). It has been determined that iron deficiency anemia is significantly more common in female patients with menstrual migraine attack than non-menstrual migraine attack (29). Likewise, a study in a small sample size suggested that comorbidity between migraine and iron deficiency anemia is limited to women (30). Here, we showed an association between lower levels of IRON and TS and migraine risk, revealing a role of anemia phenotype in migraine that is not limited to female migraineurs alone. This is supported by our epidemiological association findings, which were adjusted by sex-related covariates (sex, the interaction of sex and age, the interaction of sex and age2). Importantly, our male-stratified analysis showed that the two identified iron biomarkers are significantly lower among male migraineurs (IRON, OR = 0.79, p = 0.014; TS, OR = 0.77, p = 0.007). Consistently, genetic analyses support the epidemiological findings and suggest shared genetic factors underlying migraine risk and IRON, TS, and iron deficiency anemia in both males and females. Interestingly, migraine patients showed significantly higher amounts of accumulated iron in red nucleus and periaqueductal gray matter (25), which can be hypothetically connected to the loss of serum iron hemostasis. For instance, it has been reported that Parkinson’s disease patients have a significant increased level of iron in a region of the brain and a lower level of serum iron compared to controls (31).
Moreover, lower and higher levels of TB and GGT, two liver function biomarkers, were associated with migraine risk, respectively. A low level of TB is not a concern for liver disorders, but it is associated with CVD risk factors (32) and stroke (33), which again supports higher risk of CVD among migraineurs. Interestingly, low TB levels are associated with higher risk of deep white matter lesions (34), which are more common among migraine patients (35). Clinically, increased serum level of GGT reflects liver dysfunction but epidemiologically it is associated with higher risk of CVD (36). We conclude that the involvement of liver function in migraine biology less likely fits our genetic analyses’ findings.
Our genetic correlation findings of migraine, biomarkers and medically relevant disorders show that not only heart/cardiac problems and iron deficiency anemia traits have positive significant genetic correlations with migraine risk, but also, they mirror the pattern of genetic correlations of migraine with clinical chemistry tests. We also showed that epidemiological associations and genetic correlations between migraine and liver function biomarkers, TB and GGT, are unlikely to be due to the involvement of liver function but likely intermediated by heart or iron homeostasis-related traits.
Genetic correlation between two traits implies pleiotropy accruing when a gene or variant influences both traits mostly in two scenarios, horizontal and vertical pleiotropy. In horizontal pleiotropy (biological pleiotropy), a genetic locus affecting both traits is common, indicating a shared biological mechanism underlying both traits. In vertical pleiotropy, a genetic factor contributes to the risk of trait 1 and then trait 1 alters the risk of trait 2, reflecting a causal cascade (37). Here, we show that all epidemiological and genetic overlaps are more likely due to biological pleiotropy rather than causality.
Three possible limitations of this study need to be mentioned. First, the sample size for male migraineurs is relatively small in epidemiological data; however, we utilized well-powered genetic data to validate our epidemiological findings. Second, there are no GWAS results publicly available for TC/HDL-C and TG/HDL-C ratios; hence, they are absent from our genetic analyses. Third, the information needed for robustly classifying migraine patients into MO and MA was available in only a small number of patients; therefore, the involvement of the identified chemistry tests in MO and MA remains to be investigated in future studies.
In conclusion, we performed a large epidemiological study and identified several clinical chemistries associated with migraine risk. Furthermore, using current statistical genetic analysis approaches, we verified our epidemiological findings and showed that genetic factors influencing these common clinical chemistry tests are shared across migraine, heart problems and iron deficiency anemia, providing novel insight into the biological mechanisms underlying these often co-occurring disorders. The approach of utilizing genetic data to validate observational epidemiological findings make our study robust and powerful, as genetic data is considered to be immune to environmental confounding. Our results are in line with the observed increased risk of CVD among migraine patients. Also, our finding of reduced markers of iron status, IRON and TS, among migraine cases suggests a role for anemia phenotypes in migraine and provide support for their investigation in the development of novel therapeutic and dietary interventions.
Article highlights
Chemistry tests of cardiovascular disease risk, iron deficiency, and liver dysfunction were significantly associated with migraine risk. Migraine has shared genetics with the clinical chemistry tests, heart problems, iron deficiency anemia, but not liver diseases. Migraine patients have an increased risk for heart problems and iron deficiency anemia compared with non-migraine controls.
Footnotes
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
