Abstract
Canine appendicular osteosarcoma (OSA) is a highly metastatic tumor in dogs. Mortality due to metastatic disease is common and frequently occurs within 1 year of diagnosis despite standard-of-care treatment. Treatment includes amputation for palliation and chemotherapy for metastatic disease. Current histologic grading schemes and biomarkers are poor at predicting clinical outcome. Novel prognostic and therapeutic markers are required to improve patient care. MicroRNAs (miRNAs) are small molecules expressed by all cells and released into bodily fluids. Studies in human and canine OSA cell lines, tissues from the primary site, and blood have demonstrated the role of miRNAs in metastatic progression of OSA and its prognostication. We sought to investigate the miRNA profile of primary OSA tissue and compare it to pulmonary metastases and normal lung tissue by real-time quantitative polymerase chain reaction (PCR). Multiple miRNA and multiple variable models were investigated in primary OSA tissue to predict clinical outcome. Thirteen miRNAs had similar expression between primary and metastatic OSA but were different from normal lung tissue. MiR-9-5p, miR-196a-5p, and miR-196b were expressed in metastatic OSA but lacked expression in almost all normal lung samples. In multiple variable models for overall survival and disease-free interval, only miRNAs were selected as significant variables. This study found miRNAs that are nearly exclusively expressed in metastatic pulmonary OSA and could serve as novel therapeutic targets. MiRNAs were also found to be important prognostic biomarkers in tissue and improved prognostic ability as miRNA signatures.
Osteosarcoma (OSA) is the most common tumor of bone in dogs and primarily affects the appendicular skeleton.6,12 Primary tumors are locally invasive and osteolytic and present as lameness with a localized swelling. 12 Although pathologic fracture is uncommon at the time of diagnosis, if only palliative treatment is elected, 38% of cases eventually fracture. 31 While most do not have radiographically detectable metastatic disease at diagnosis, up to 95% of patients already have micrometastases, since metastases still develop after removal of the primary tumor. 15 Standard-of-care therapy is amputation and adjuvant systemic chemotherapy. The addition of chemotherapy extends survival to about 1 year in most cases.28,37 Mortality is due to metastatic disease, most commonly of the lungs. 15 Despite extensive research into novel treatments, standard-of-care therapy has not changed over several decades.21,38 A greater understanding of metastatic disease is required to improve survival time and develop novel therapeutics.
Despite the overall short survival time, there is a wide variation in the disease-free interval (DFI); some dogs develop metastases within a few months, while others live for several years. 39 Better tools to help predict metastatic disease and assist owners and oncologists in clinical decision-making are required. Definitive diagnosis requires histologic evaluation of the affected limb, and as this is already a component of standard care, histologic grading schemes were investigated. Unfortunately, a recent study determined that the current grading schemes are unable to predict clinical outcome in patients receiving standard-of-care therapy. 36 Clinical characteristics suggested to predict outcome are a humeral location and serum alkaline phosphatase (ALP) activity. The humerus was a significant negative prognostic indicator in multiple meta-analyses.4,34 Two meta-analyses found serum ALP activity was predictive for overall survival (OS) and DFI, while other studies found no correlation.4,9,12,32,34,36 One study demonstrated that serum ALP activity was poor at predicting long-term survivors but performed well in combination with miR-214 to predict OS and DFI. 17
MicroRNAs (miRNAs) are non-coding RNA species of approximately 22 nucleotides in length that are widely expressed in tissues and bodily fluids in all species, including dogs.16,20 There is extensive evidence in numerous human and canine cancers, including OSA, that miRNAs are dysregulated and contribute to tumorigenesis and metastatic progression.23,33,41 Primary tumor and cell line expression studies demonstrate several key miRNAs in canine OSA, including miR-9 and miR-34a, which were shown to be dysregulated and contribute to invasion and migration of OSA cell lines.14,22,26,29 Dailey et al 11 found a 3 miRNA model (miR-223-3p, miR-103a-3p, and let-7b-5p) in primary OSA tissue that could separate high (median DFI = 123.5 days) and low (median DFI = 392 days) risk groups to predict DFI. This study also investigated serum miRNAs but did not discuss the expression differences between serum and tissue or whether the combined information could predict clinical outcome parameters.
There is a lack of literature investigating a multiple miRNA profile in pulmonary metastases in canine OSA patients. It is uncertain if the miRNAs are similar or different to those expressed in the primary tumor. A better understanding of the miRNAs expressed in metastatic lesions will further our knowledge of their development and progression and serve as potential therapeutic targets directed at the metastases. As such, this study sought to investigate miRNA expression profiles in both primary OSA and matched and non-matched pulmonary metastases. Some cases had matched plasma previously analyzed; 27 models including both tissue and plasma miRNA profiles were also investigated.
Materials and Methods
Samples
All animal experiments were performed according to Canadian Council of Animal Care guidelines. This study was approved by the Animal Care Committee of the University of Guelph (AUP 4409), and owners’ written consent was obtained to collect tissue. All cases had a histologic diagnosis of central appendicular OSA and primary cases had no clinically detectable (eg, thoracic radiographs) evidence of metastatic disease at the time of diagnosis. Subtype was not considered in this population, as it has been shown there is only fair agreement amongst pathologists in this determination. 36 Similarly, grade was not recorded due to its documented lack of prognostic significance. 36 For primary OSA cases, only cases that received standard-of-care treatment (amputation and at least 1 dose of cytotoxic chemotherapy) were included in this study. All patients received care at the Animal Cancer Centre (Ontario Veterinary College, Guelph, Ontario). All samples were collected within 24 hours from fresh specimens either at the time of limb amputation surgery (primary tissues) or at the time of postmortem examination (metastatic and normal lung tissues). Representative tissue was collected (approximately 2.0 cm3) and rinsed with phosphate-buffered saline and immediately submerged in 1.5 to 2 ml of RNAlater Stabilization Solution (RNAlater, Thermo Fisher Scientific, Waltham, Massachusetts) for 3 to 5 days at 4°C. The RNAlater was removed, and the tissue was stored in a cryovial at −80°C. An adjacent portion of tissue was examined by histology to confirm a diagnosis of OSA and/or normal lung tissue. Forty-two primary OSA tissue samples and 12 lung metastases, of which 7 also had primary OSA tissue available (matched), were included in this study. In addition, 5 lung samples were obtained from postmortem cases, which grossly and histologically did not have evidence of neoplasia or other significant disease in the lungs. These patients were euthanized for a reason unrelated to OSA and pulmonary disease. Fifteen cases had matched plasma previously investigated. 27 Complete follow-up information from diagnosis until death/euthanasia with cause was discernable from the medical records in 40/42 primary OSA cases. For the remaining 2 cases, only a last known date alive was available, each with no history of clinical metastatic disease at that time.
MicroRNA Isolation
MiRNAs were isolated from 14.1 to 15.9 mg of RNAlater preserved tissues using the QIAGEN miRNeasy Tissue/Cell Advanced Mini Kit (Hilden, Germany). A 1:50 working buffer was made by adding 2 M dithiothreitol (0.154 g dithiothreitol in 0.5 ml RNase-free water) to Buffer RLT. The working buffer (260 µl per sample) and a stainless-steel bead were added to the sample tube followed by tissue homogenization at 30 Hz for 4 minutes using the TissueLyser II (QIAGEN). The bead was removed, and the lysate was centrifuged for 3 minutes at 14 500 rpm (Eppendorf MiniSpin centrifuge). The supernatant was placed into a new tube without disturbing the pellet. The remaining steps followed the manufacturer’s protocol. The column was transferred to the final 1.5 ml storage tube and 50 µl of RNase-free water was added to the center of the column. It was then incubated for 1 minute at room temperature and centrifuged for 1 minute at 14 500 rpm. Two microliters of the sample was used to quantify the RNA concentration and assess purity (260 nm/280 nm and 260 nm/230 nm ratios) using the NanoDrop 2000 to detect UV-absorbance (Thermo Scientific, Supplemental Table S1). All samples had a 260 nm/280 nm ratio > 1.85 and variable 260 nm/230 nm ratios (0.66–2.15). Sixteen samples had a 260 nm/230 nm ratio < 1.5, and samples with lower concentrations tended to have a lower 260 nm/230 nm ratio. As there is no established cut-off in the literature, all samples were considered to have passed quality assessment for downstream experiments. Sample concentrations by NanoDrop 2000 ranged from 12.1 ng/µl to 1344.1 ng/µl with 7 samples having a concentration < 50 ng/µl. In addition, the RNA concentration was measured using the Qubit miRNA Assay Kit (Qubit) with the Qubit 2.0 Fluorometer (Invitrogen, Waltham, Massachusetts) (Supplemental Table S1); however, the upper limit of detection of the Qubit was insufficient for 27/59 (45.8%) samples despite maximum dilution per the manufacturer’s recommended protocol. NanoDrop 2000 concentration values were used for the reverse-transcription polymerase chain reaction (PCR) input.
Reverse-Transcription Polymerase Chain Reaction
Each sample isolate was diluted to 5 ng/µl in nuclease-free water using 2 µl of miRNA isolate stock based on the NanoDrop 2000 concentration. The QIAGEN miRCURY LNA RT kit was used for all samples following the manufacturer’s protocol to obtain 10 µl of complementary DNA (cDNA) with a standard input of 10 ng of template RNA. The samples were incubated in a C1000 Bio-Rad Thermo Cycler at 42°C for 60 minutes, 95°C for 5 minutes, and 4°C for at least 5 minutes with the lid temperature set to 95°C before storage at −20°C overnight. A no-template control was included for each run, replacing the sample volume with water.
Target MicroRNA Selection
As similar to Craig et al, 10 a pilot study of 5 pooled representative tissue samples per group was performed with the commercially available canine miScript miRNome miRNA arrays (QIAGEN). These arrays assess 277 miRNAs known to be expressed in dogs. MiRNAs that were differentially expressed between groups and that also had robust expression were identified. The primary OSA samples were evaluated in 2 groups, which included samples from patients with either greater or less than 1-year survival. The metastatic tissues were also evaluated in 2 groups as either those with matched primary samples or those without. All normal lung samples were run as a single pooled group. The miScript II RT Kit (QIAGEN) was used to convert isolated miRNAs into cDNA by reverse-transcription PCR using the manufacturer’s protocol. Each sample’s miRNA isolate was diluted from its original miRNA stock to 20 ng/µl, and 1 µl (for a total of 5 µl as the pooled sample) was added to the reverse-transcription mix. This 20 µl cDNA solution was diluted with 90 µl of nuclease-free water and stored at −20°C until used for real-time quantitative PCR (RT-qPCR). The corresponding miScript SYBR green kit was used per the manufacturer’s protocol to create a master mix. The Roche LC480 LightCycler was used for RT-qPCR and set to QIAGEN’s recommended settings. The LC480 software (release 1.5.1.62 SP3) calculated cycle threshold (Ct) values using the second derivative maximum method. A total of 59 miRNAs with the largest fold-changes between groups alongside miRNAs previously reported in the literature and controls were added to a custom miRCURY PCR array design (QIAGEN) (Supplemental Table S2) to be used for the individual samples described above.
Real-time Quantitative Polymerase Chain Reaction
The miRCURY SYBR green kit (QIAGEN) was used to create a master mix for each sample and consisted of 360 µl of SYBR green, 288 µl of the diluted cDNA sample (1:80 dilution of 1 ng/µl cDNA concentrate), and 72 µl of DNase-free water. Each plate well (miRCURY LNA miRNA custom PCR array 384-well plate [QIAGEN]) received 10 µl of the master mix for its corresponding sample. A Roche LC480 LightCycler was used for the RT-qPCR. The reaction volume was set to 10 µl with the SYBR green I/HRM dye detection format. Each array was incubated at 95°C for 2 minutes (ramp rate of 4.8°C/s) with 45 cycles of 10 seconds at 95°C (ramp rate of 4.8°C/s) followed by 60 seconds at 56°C (ramp rate of 2.5°C/s), with a single acquisition at the end of each cycle. At the end of the 45 cycles, a melting curve analysis was performed from 55 to 95°C (ramp rate of 0.11°C/s) with 5 acquisitions/s and a cooling step to 37°C. The LC480 software (release 1.5.1.62 SP3) calculated Ct values using the second derivative maximum method.
Quality Assessment of Real-time Quantitative Polymerase Chain Reaction Results
As per QIAGEN’s recommendations, the UniSp3 inter-plate calibrator was used to correct for potential differences between PCR arrays. The difference between the average UniSp3 of the no-template control (up to 6 replicates per plate) for each PCR array and the overall average was determined. Each Ct value was subtracted by this difference to get an inter-plate calibrator-corrected Ct value. All further analyses utilized the inter-plate calibrator-corrected Ct values, from now on simply referred to as Ct values.
A Ct value cut-off of 35.00 was employed for all miRNAs and controls. The amplification and melting curves were manually assessed to ensure appropriate Ct determination by the LC480 software. All undetectable Ct values (Ct = 0), those >35.00, and those with visually inappropriate amplification curves were changed to equal 35.00 to provide a conservative Ct determination and were considered to have no expression. Each sample and miRNA were evaluated for overall expression levels. All samples included in the study had an overall expression of miRNAs within the average of all samples isolated minus 2 standard deviations. MiRNAs expressed in a low number of samples compared to the average for all miRNAs minus 2 standard deviations were removed from further analyses. This included 4 miRNAs originally included on the custom panels (miR-211-5, miR-326, miR-592-5p, and miR-1844).
Statistical Analysis
Analyses used the R Studio environment, software version 4.1.0 (R Core Team, 2021). 30 The NormFinder R script was used to identify stably expressed miRNAs as endogenous controls. 1 MiRNAs included in NormFinder analysis were expressed in all samples (Ct < 35.00). As recommended by Vandesompele et al, 40 a minimum of 3 controls were selected. MiRNAs miR-93-5p, let-7b-5p, and miR-151a-5p were selected as endogenous controls based on their stability, which had standard deviations <1. Normalization was performed using the comparative Ct method, where the Ct for a miRNA of interest was subtracted from the average Ct value of the endogenous controls. 35 In place of the arithmetic average, the geometric average was employed as it less susceptible to outliers. 40 Normalization was completed on all data simultaneously.
Normality of the data could not be assumed due to the small sample size for each group. A Shapiro-Wilk test for normality was used to determine the distributions of each group for each miRNA. If 1 or more groups failed the normality test (P < .05), the miRNA was considered non-normally distributed. Twenty-nine percent of miRNAs were non-normally distributed; therefore, non-parametric tests were used. With normalized Ct values (∆Ct), the Wilcoxon-rank sum test was performed, and the Benjamini-Hochberg procedure was used to correct for multiple comparisons. 2 A P-value of < .05 was considered statistically significant. This test was repeated for each miRNA comparing each population independently to one another. Interquartile range was also determined and reported. Fold-changes were calculated using 2−∆∆Ct, where ∆∆Ct is the median ∆Ct for a miRNA of interest in one group subtracted from the ∆Ct of the second group. The median was utilized for each miRNA. Fold-differences were equivalent to fold-changes if >1, while negative fold-differences equaled the negative inverse of the reported fold-change (values between 0 and 1) for each miRNA.
The surv_cutpoint() and surv_categorize() functions, available in the survminer package in R, were used for each miRNA to establish the optimal cut-off ∆Ct point and categorize high or low expression groups based on survival data. 18 The default minprop value of 0.2 was used in surv_cutpoint() indicating a minimum proportion of 0.2, or 20%, of samples must be allocated to either expression group. Kaplan-Meier curves were generated from each miRNA and the log-rank test was used to calculate P-values, where < .05 was considered statistically significant. This was repeated for OS, 1-year survival, and DFI.
To investigate multiple miRNA models in predicting prognosis, survival trees were grown and pre-pruned using rpart() by specifying minsplit and maxdepth parameters. For all survival metrics, a minsplit of 15 was used, indicating a minimum of 15 samples must be present in a node to consider splitting again. For OS and DFI, a maxdepth of 3 was used and for 1-year survival a maxdepth of 2 was used. This indicates the maximum number of levels the tree may grow to. Both parameters were chosen to limit the number of variables selected to 1 per 10 events, as typically recommended. 5 Trees were made for the same survival metrics as listed above. For each survival metric, samples were assigned a group based on the corresponding tree and a hazard ratio and mean deviance were determined for each group. Kaplan-Meier curves were used to assess differences in survival between the groups. Log-rank tests were performed using survdiff() to calculate P-values for each pairwise group comparison. To account for multiple comparisons, the Bonferroni correction was applied. A P-value < .05 was considered statistically significant. Fifteen samples had tissue analyzed, as well as plasma previously analyzed, so multiple miRNA models using only the plasma and tissue Ct values were also investigated by these methods. 27
Finally, multivariate (multiple variable) models were created using both miRNA expression and patient demographic information. First, univariate Cox regression was used to calculate hazard ratios (HRs) and assess the significance of the patient variables independently. Categorical variables were assessed in a manner that allowed each category to serve as the reference population. That is, for a variable with n categories, there were n HRs calculated, except for categories with only a single sample where a meaningful P-value and HR could not be calculated. Using the same methods as described for the multiple miRNA models, survival trees were grown and pruned. Kaplan-Meier curves were made for significant models, and P-values were calculated using the log-rank test for every pairwise group comparison. The Bonferroni multiple comparisons correction was applied, and a P-value < .05 was considered statistically significant.
Fifteen samples had matched primary OSA tissue and plasma, and 22 miRNAs analyzed in this experiment and in a previous study. 27 Correlation between tissue and plasma expression of each miRNA was investigated. Using GraphPad Prism (version 10 for Mac OS X, GraphPad Software, Boston, Massachusetts, www.graphpad.com), the normalized Ct values were compared using a non-parametric 2-tailed Spearman correlation test and were considered statistically significant if P < .05.
To further investigate miRNAs similarly expressed between primary and metastatic OSA but differentially expressed from normal lung tissue, gene targets were searched for based on statistically significantly different miRNAs using TargetScan (https://www.targetscan.org/vert_80/) and miRDB (https://mirdb.org/).
Results
Population Statistics
The breeds represented in the primary OSA, pulmonary metastases, and normal lung groups are detailed in Supplemental Table S3. Mixed breed was the most common breed in the primary and pulmonary metastases groups. In the primary OSA group, Rottweiler, greyhound, golden retriever, and Doberman were represented by more than 1 case. Two cases were greyhounds in the pulmonary metastases group. For the primary OSA group, 26/42 affected the right limb. The proximal humerus (n = 10), followed by the distal radius (n = 9), was the most common location. The primary OSA group was represented by 21 neutered males, 18 spayed females, 2 intact males, and 1 intact female with a median age of 7.4 years (range = 1.4–12.5 years) at the time of diagnosis and a median weight of 36.1 kg (range = 13.9–69.0 kg) prior to amputation. MiRNA expression in the primary OSA group was compared between males (neutered and intact, n = 23) and females (neutered and intact, n = 19). No miRNAs had statistically significant differential expression between these 2 groups (Supplemental Table S4). The pulmonary metastases group was represented by 8 neutered males and 4 spayed females with a median age of 7.9 years (range = 5.4–13.8 years) at postmortem examination and a median weight of 37.2 kg (range = 25.4–66.0 kg) post-amputation at postmortem examination. Finally, the normal lung group was represented by 4 neutered males and 1 intact female with a median age of 6.2 years (range = 5.0–12.8 years) and a median weight of 15.2 kg (range = 5.7–25.6 kg) at postmortem examination.
All primary OSA cases were treated with amputation followed by at least 1 dose of cytotoxic chemotherapy (standard-of-care therapy). Most dogs received carboplatin (41/42, 98%), either alone (20/41, 49%) or in combination with another chemotherapeutic (21/41, 51%). Cyclophosphamide (12/21, 57%), toceranib phosphate (6/21, 29%), doxorubicin (2/21, 9%), or rapamycin (1/21, 5%) were used in combination with carboplatin. One case received doxorubicin with cyclophosphamide. In the pulmonary metastases group, treatment prior to death was variable. Eight of 12 cases (67%) received standard-of-care therapy. Seven of those cases received carboplatin, of which 6 were in combination with either doxorubicin (3/6), cyclophosphamide (2/6), or toceranib phosphate (1/6). One case received doxorubicin with cyclophosphamide. Four cases did not undergo amputation (4/12, 33%) with 2 of those cases receiving radiation therapy with carboplatin.
Of all causes of death, 12/40 dogs (30%) had a postmortem examination. Cause of death for the remaining cases was determined by confirmed or presumptive imaging (17/40, 43%) or clinical signs (11/40, 27%). Death due to OSA was recorded in 29/40 cases. Eleven of these dogs (38%) underwent postmortem examination to confirm progressive disease and metastases. Progressive disease and metastases were determined as the cause of death via confirmed or presumptive imaging (14/29, 48%) or clinical signs (4/29, 14%). The median OS time of this population was 335 days. The median DFI was 261 days, and 52% (22/42 cases) of patients were alive at 1 year.
Comparison of MicroRNAs in Primary Osteosarcoma and Pulmonary Metastases
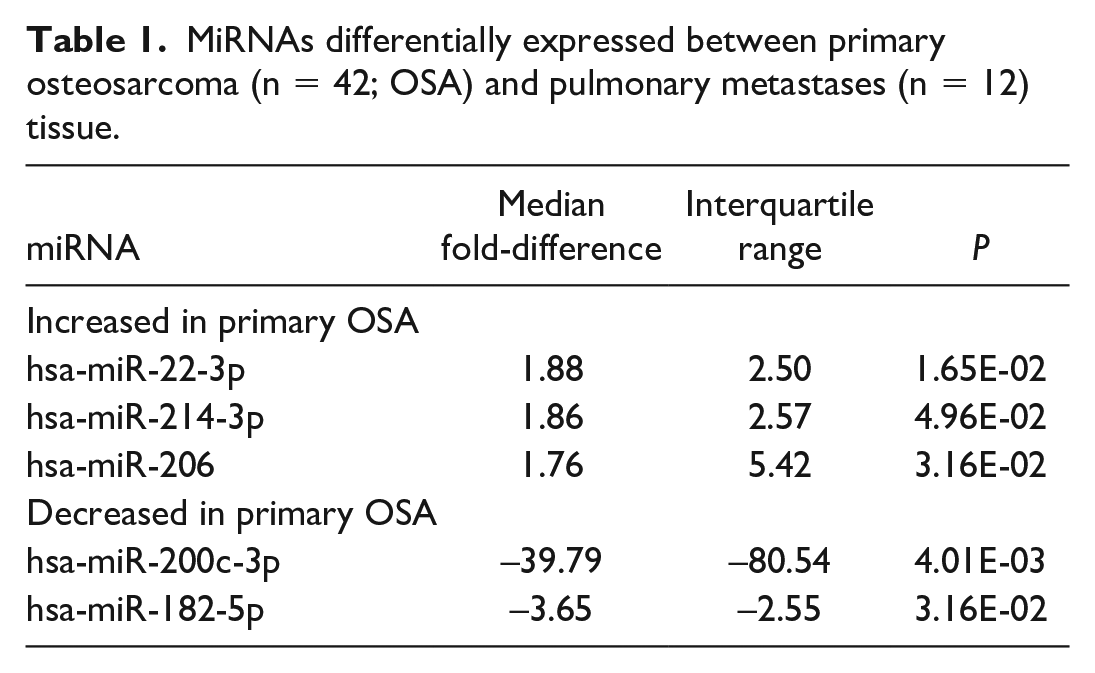
Of the 52 miRNAs included in final analyses, 5 had a significant (P < .05) median fold-difference between primary and pulmonary metastatic OSA samples (5/52, 10%; Table 1). Three miRNAs had increased expression and 2 had decreased expression in primary OSA tissue compared to the pulmonary metastases. MiR-200c-3p and miR-182-5p had a significant fold-difference >2 or <−2. MiR-200c-3p had the greatest fold-difference (39.79; interquartile range [IQR] = 80.54) in pulmonary metastases compared to primary OSA cases and was only expressed in 2 primary OSA samples at low levels (> 33.00 Ct) (Fig. 1). Therefore, 90% (47/52) of miRNAs investigated were not differentially expressed between the primary and pulmonary metastatic OSA samples (Supplemental Table S5).
MiRNAs differentially expressed between primary osteosarcoma (n = 42; OSA) and pulmonary metastases (n = 12) tissue.
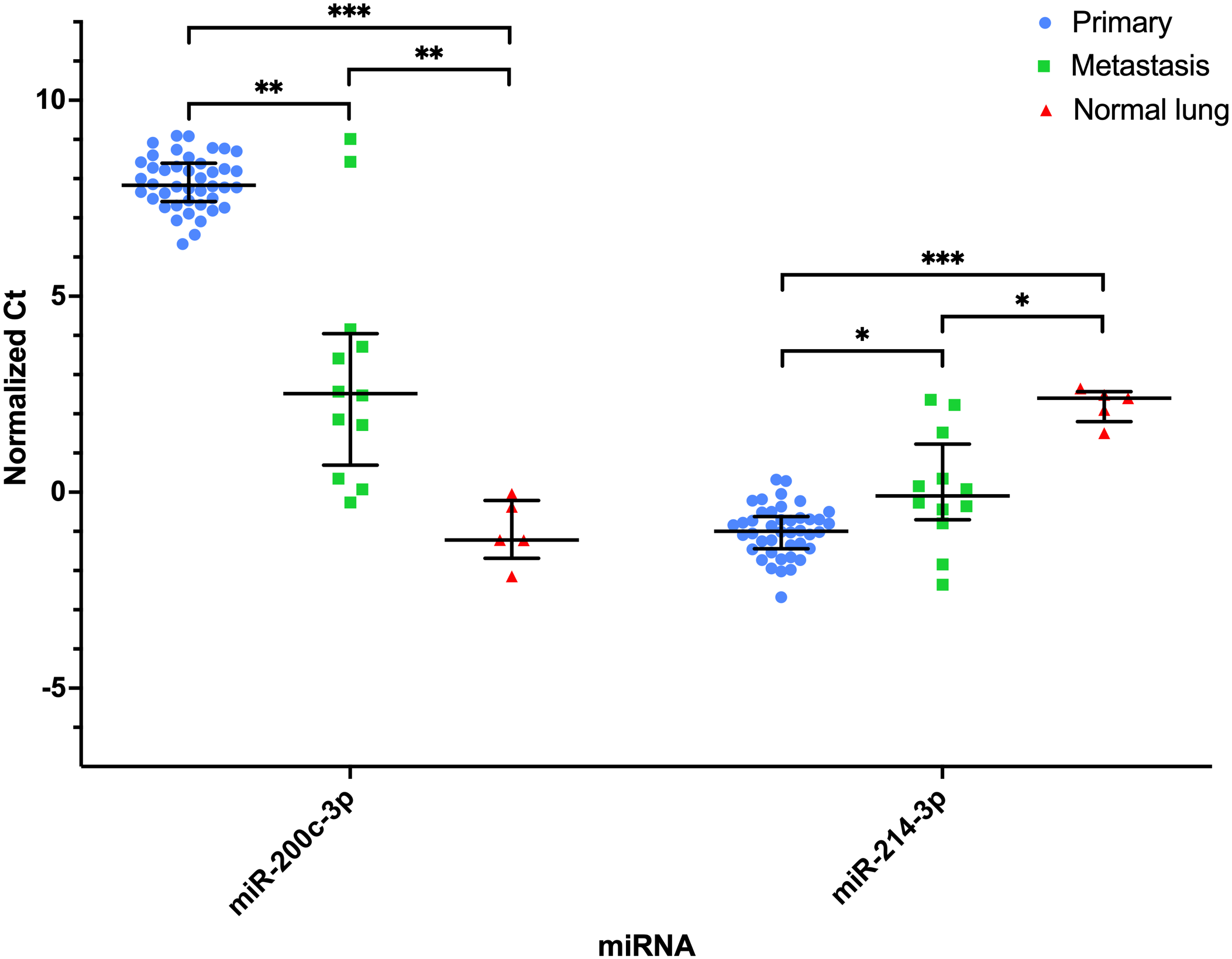
MiRNAs significantly differentially expressed between primary osteosarcoma (OSA) (n = 42), pulmonary metastases (n = 12), and normal lung (n = 5) tissues. Normalized Ct values plotted for each sample within their respective group (primary OSA, pulmonary metastatic OSA, and normal lung tissue). Bar and lines represent the median and interquartile range. Statistical significance determined based on median fold-change. *P < .05; **P < .01; ***P < .001.
Comparison of MicroRNAs in Pulmonary Osteosarcoma Metastases and Normal Lung
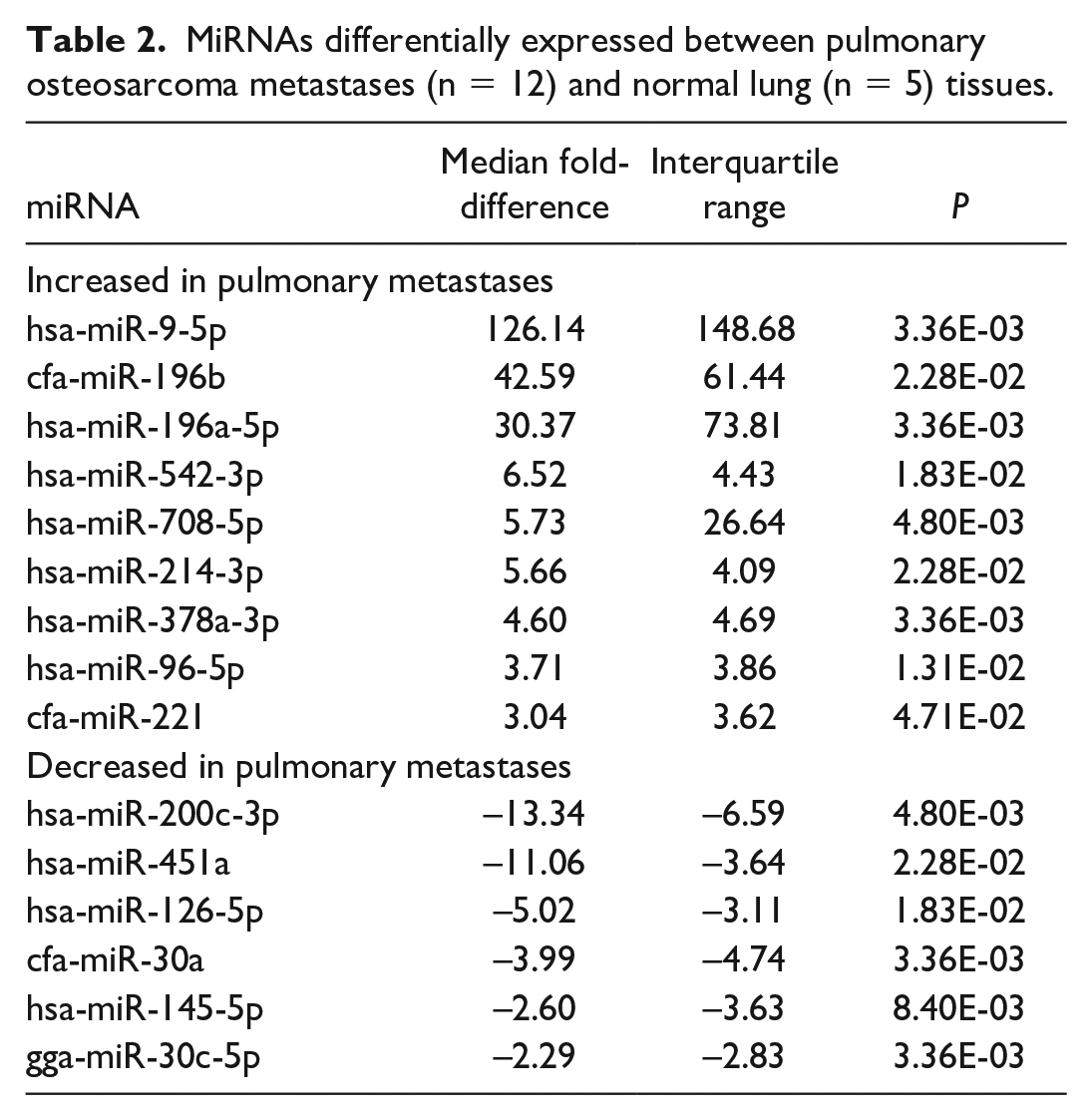
Fifteen miRNAs had a significantly different median fold-difference (15/52, 29%) between pulmonary OSA metastases and normal lung tissue (control) groups (Table 2). Nine of these miRNAs had increased expression in pulmonary metastases compared to controls. Six miRNAs had decreased expression. All had a significant fold-difference > 2 or < −2. MiR-9-5p had the greatest fold-difference (126.14; IQR = 148.67) and no expression (Ct = 35.00) in 4 of the 5 normal lung samples (Fig. 2 and Table 2). MiR-196a-5p and miR-196b had the next greatest fold-differences in pulmonary OSA metastases compared to normal lung tissue with median fold-differences of 30.37 (IQR = 73.81) and 42.59 (IQR = 61.44), respectively, and had no expression in any of the normal lung samples (Fig. 2). MiR-200c-3p had the greatest increase in expression in normal lung tissue (fold-difference of 13.34; IQR = 6.59) compared to the pulmonary OSA metastases (Fig. 1). Thirty-seven miRNAs (37/52, 71%) were not significantly differentially expressed between the pulmonary OSA metastases and normal lung tissue (Supplemental Table S5).
MiRNAs differentially expressed between pulmonary osteosarcoma metastases (n = 12) and normal lung (n = 5) tissues.
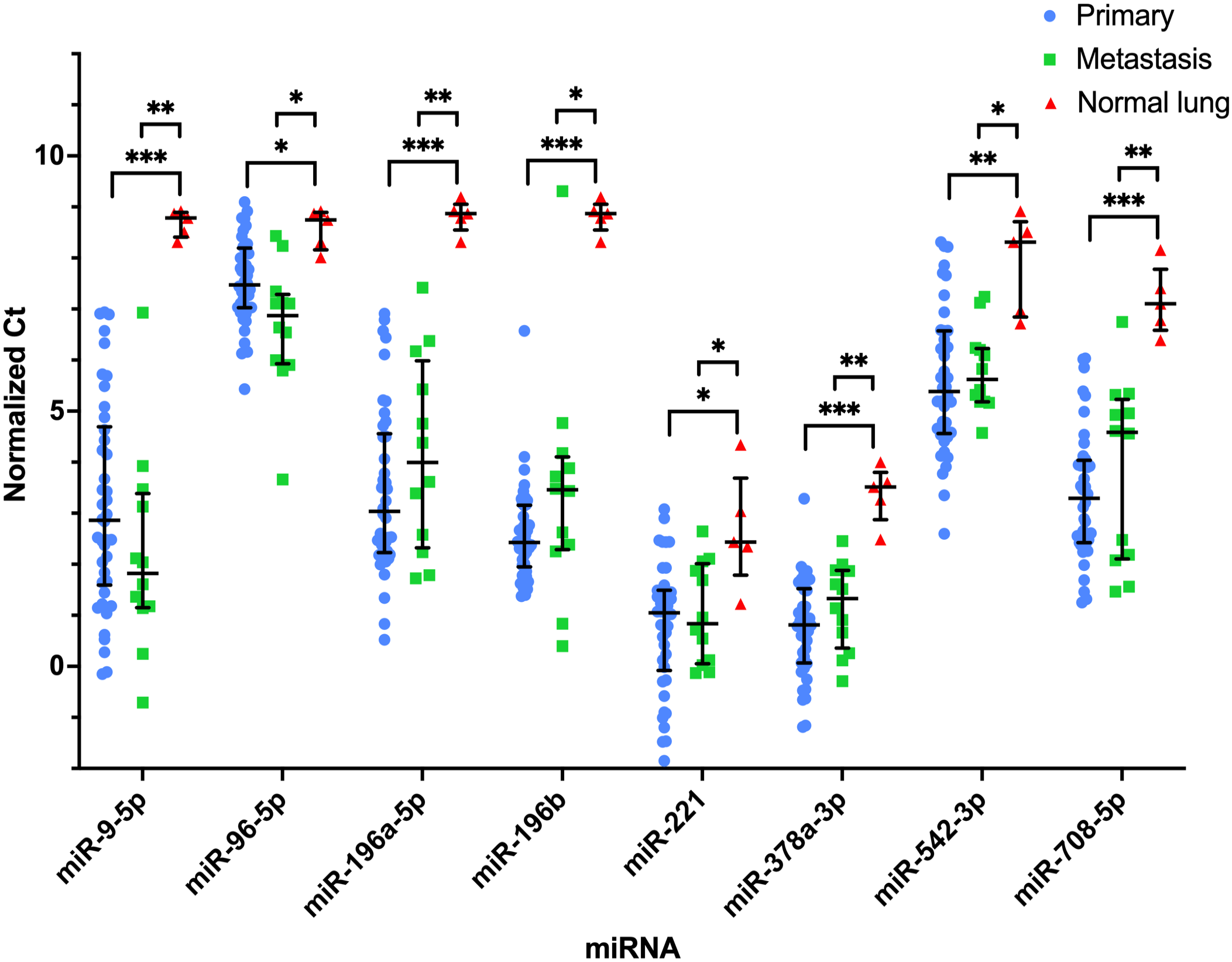
MiRNAs with significantly increased expression in primary (n = 42) and pulmonary metastatic (n = 12) osteosarcoma (OSA) compared to normal lung (n = 5) tissue. Normalized Ct values plotted for each sample within their respective group (primary OSA, pulmonary metastatic OSA, and normal lung tissue). Bar and lines represent the median and interquartile range. Statistical significance is determined based on median fold-change. *P < .05; **P < .01; ***P < .001.
Comparison of MicroRNAs in Primary and Pulmonary Metastatic Osteosarcoma and Normal Lung
Thirteen miRNAs were not significantly different between the primary OSA and pulmonary metastases, but both groups had significantly different expression from normal lung tissue: miR-9-5p, miR-30a, miR-30c-5p, miR-96-5p, miR-126-5p, miR-145-5p, miR-196a-5p, miR-196b, miR-221, miR-378a-3p, miR-451a, miR-542-3p, and miR-708-5p. Of these, 8 miRNAs had increased expression in the primary and metastatic OSA tissues compared to normal lung tissue (Fig. 2). Five of these miRNAs had decreased expression in the primary and metastatic OSA tissues compared to normal lung tissue (Fig. 3). Expression of miR-9-5p and miR-196a-5p in normal lung samples had no overlap with the primary or metastatic OSA tissues, and only 1 lung metastasis had expression of miR-196b in the range of the normal lung tissues (Fig. 2). The expression of miR-206 was not significantly different in the lung metastases and normal lung tissue but was statistically significantly different in both groups compared to the OSA primary tissues. MiR-200c-3p and miR-214-3p were significantly different across all 3 groups (Fig. 1), while 15 miRNAs had similar levels of expression across all 3 groups.
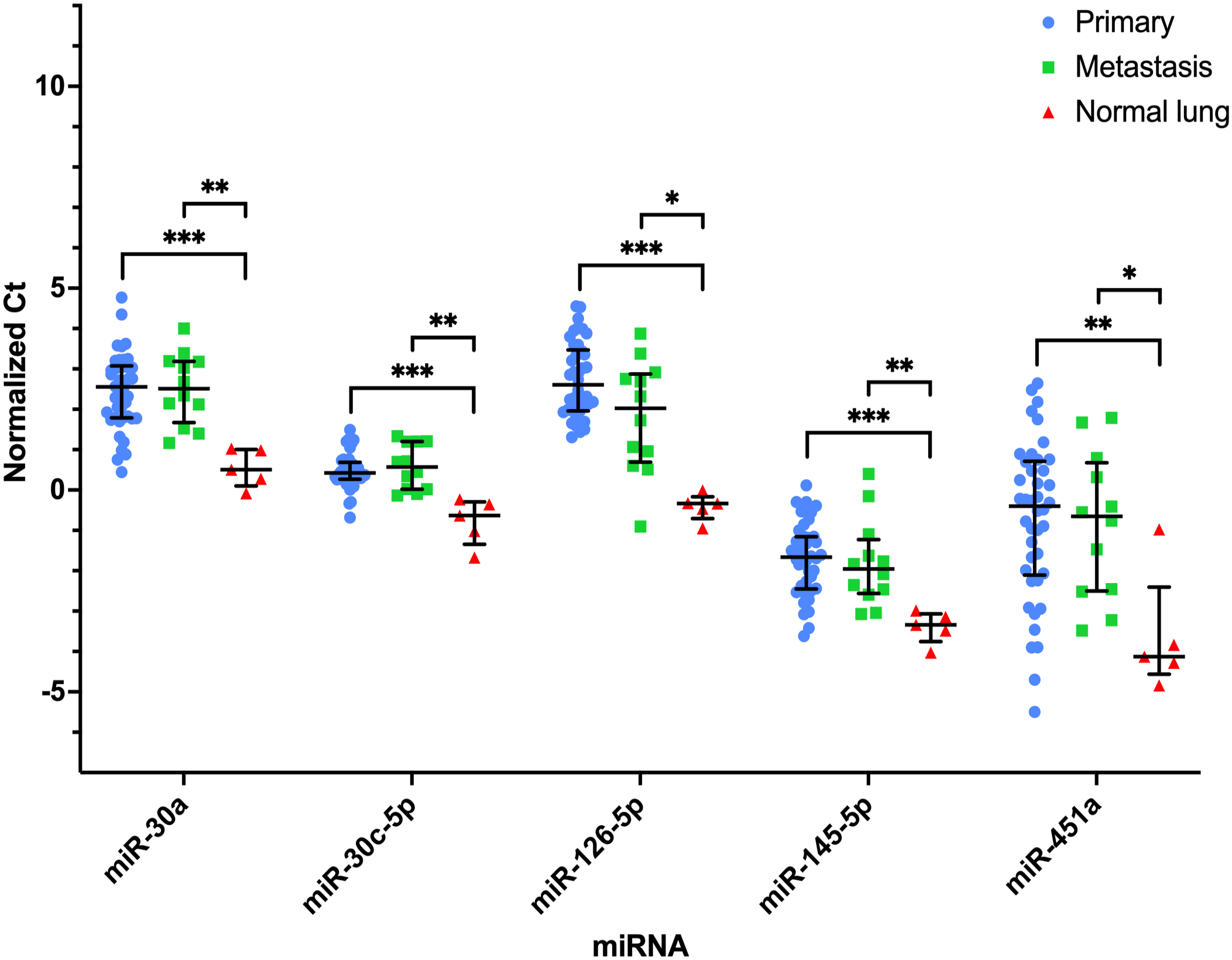
MiRNAs with significantly decreased expression in primary (n = 42) and pulmonary metastatic (n = 12) osteosarcoma (OSA) compared to normal lung (n = 5) tissue. Normalized Ct values plotted for each sample within their respective group (primary OSA, pulmonary metastatic OSA, and normal lung tissue). Bar and lines represent the median and interquartile range. Statistical significance is determined based on median fold-change. *P < .05; **P < .01; ***P < .001.
MicroRNAs Predictive of Clinical Outcome in Primary Osteosarcoma Tissues
Within primary OSA tissues, 14 individual miRNAs stratified patients with short and long DFI (Supplemental Table S6), and 15 individual miRNAs stratified patients with short and long OS (Supplemental Table S7). Twenty-two individual miRNAs stratified patients based on 1-year survival (Supplemental Table S8). Eleven individual miRNAs in primary OSA tissues similarly stratified patients for OS, DFI, and 1-year survival. For DFI, the 5 most statistically significant miRNAs were miR-101, miR-27b-3p, miR-128-3p, miR-451a, and miR-30c-5p. For OS, the 5 most statistically significant miRNAs were miR-101, miR-451a, miR-27b-3p, miR-128-3p, and miR-30c-5p. Finally, for 1-year survival, the 5 most statistically significant miRNAs were miR-7-5p, miR-542-3p, miR-27b-3p, miR-101, and miR-30a.
Univariate Analyses of Clinical Parameters
In univariate analyses of clinical parameters, including age, sex, weight, serum ALP activity status [low (below reference range), normal (within reference range), and high (above reference range)], and tumor location, only the scapular location of a primary tumor had an increased HR of 6.9 (95% confidence interval (CI) = 1.4-34) for DFI, but there were only 2 cases (Supplemental Table S9).
Multiple MicroRNA and Multiple Variable Models in Primary Osteosarcoma Tissue for Prediction of Clinical Outcome Parameters
Multiple miRNA models using primary OSA tissue miRNAs were investigated to predict DFI, OS, and 1-year survival. MiR-16-5p, miR-451a, and miR-885-5p were utilized for survival prediction for DFI (Fig. 4). MiR-16-5plow had the greatest HR (2.81; mean deviance (MD) = 0.6) and shortest DFI (96 days). MiR-16-5phigh with either miR-451ahigh (HR = 2.03; MD = 0.63; DFI = 112 days) or with miR-451alow and miR-885-5plow (HR = 1.7, MD = 1.46; DFI = 121 days) were not significantly different from the group miR-16-5plow alone and all indicated a poor DFI. MiR-16-5phigh/miR-451alow/miR-885-5phigh predicted a longer DFI (711 days) and had a protective HR (0.526; MD = 0.76). This model remained the same when the clinical variables were also considered.
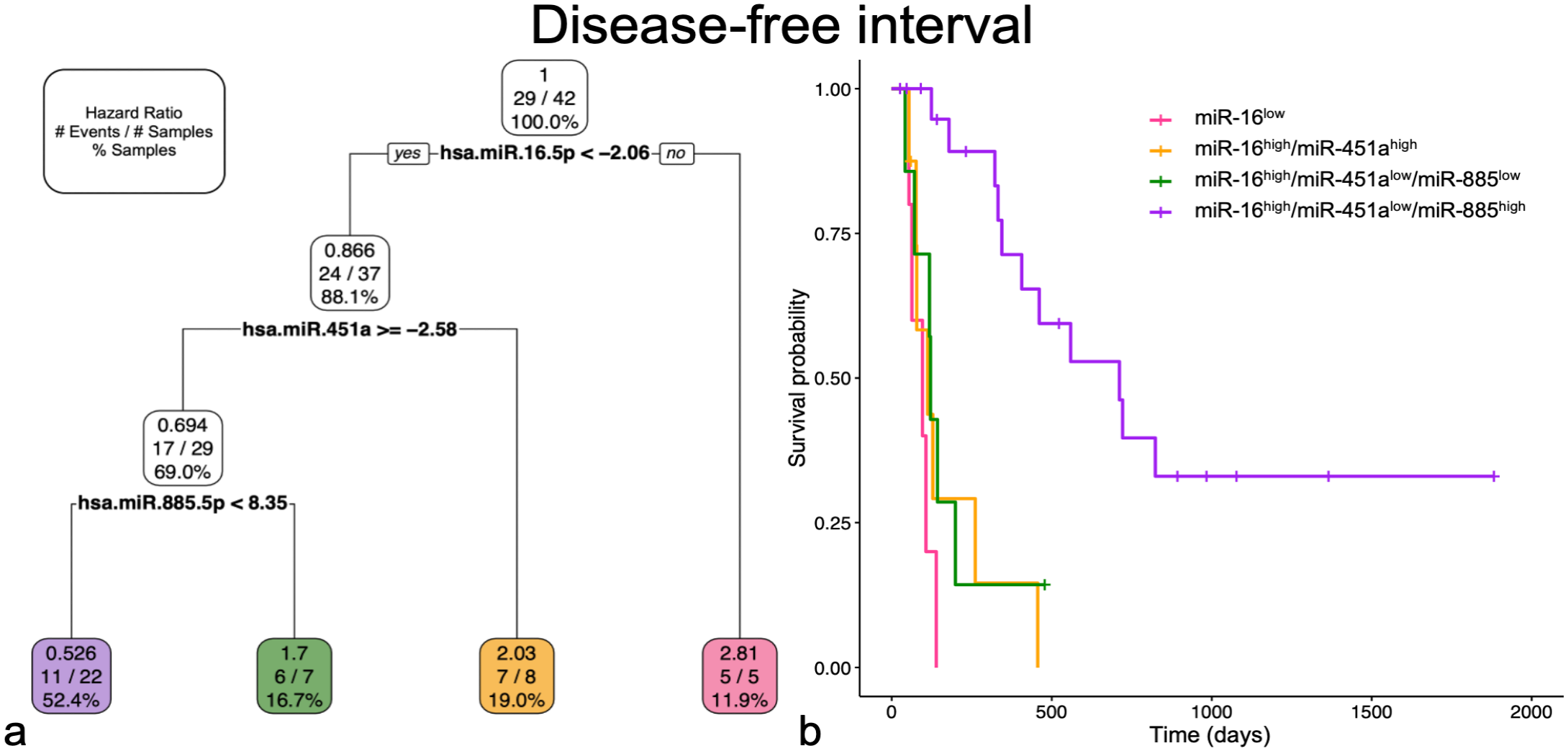
Multiple miRNA and multiple variable models for disease-free interval selected miR-16-5p, miR-451a, and miR-885-5p. Groups in both graphs are represented by corresponding miRNA profile and color. (a) Pruned decision tree for disease-free interval classified samples into 4 groups with different hazard ratios (HRs). Each step includes the HR, number of events defined by metastatic disease out of the total number of samples, and percentage samples in each group. Note that decision values are normalized Ct. A “yes” determination is always to the left. (b) Kaplan-Meier survival curve for each of the groups. The probability of survival for miR-16high/miR-451alow/miR-885high is significantly better (P < .001) compared to all other groups. The remaining 3 groups are not significantly different from each other.
For OS, miR-27b-3p, miR-128-3p, and miR-34a-5p were selected in the multiple miRNA and multiple variable models (Fig. 5). MiR-27b-3plow/miR-34a-5plow had an increased HR of 3.4 (MD = 0.59; OS = 118 days) with the worst probability of survival. This group was significantly different from all others. MiR-27b-3plow/miR-34a-5phigh (HR = 1.03; MD = 0.50; OS = 461 days) and miR-27b-3phigh/miR-128-3phigh (HR = 1.16; MD = 0.94; OS = 333 days) were not significantly different from each other and had an intermediate probability of survival. MiR-27b-3phigh/miR-128-3plow had a decreased HR (0.36; MD = 0.94; OS = <50% of cases experienced an event) and the greatest probability of survival, which was significantly different from all other groups.
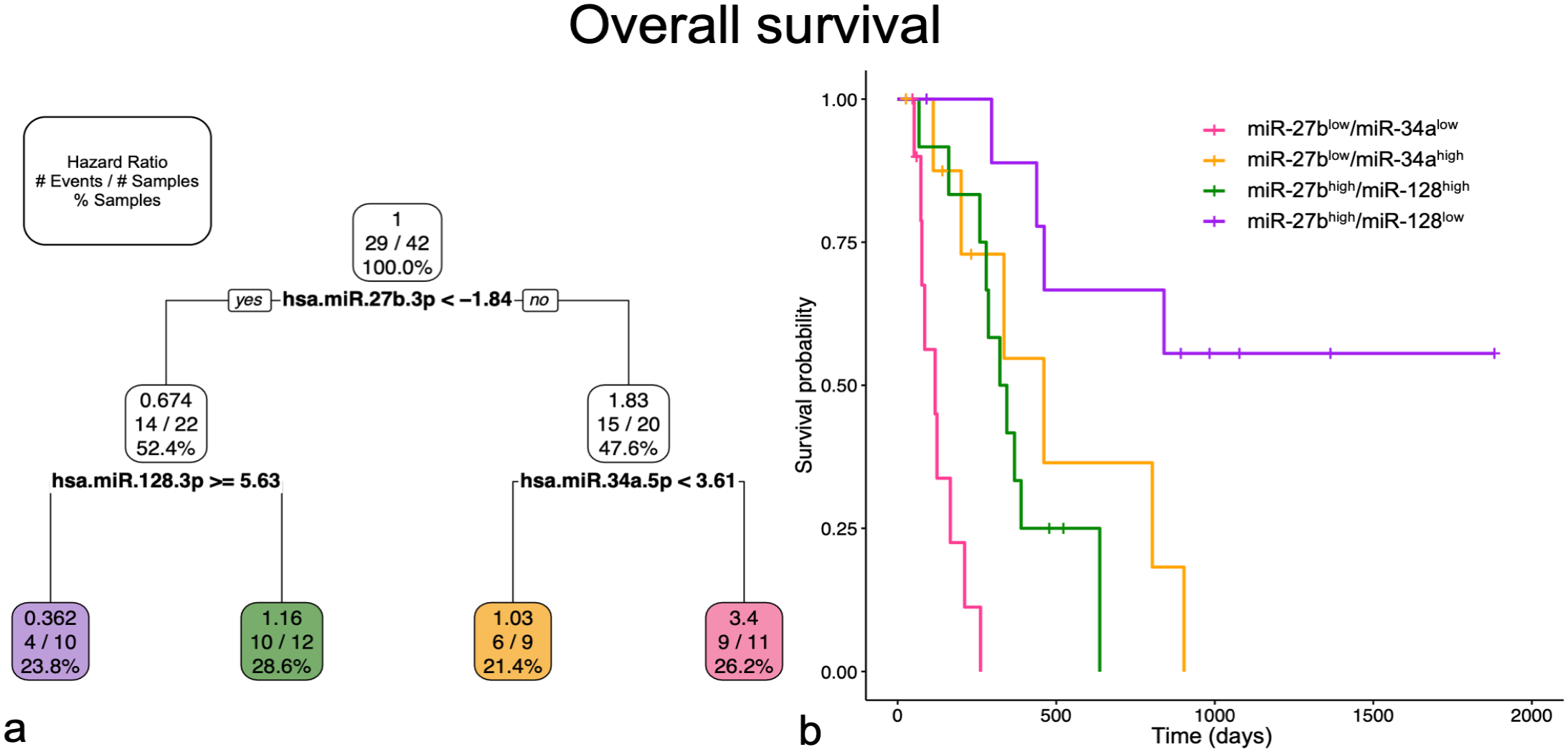
Multiple miRNA and multiple variable models for overall survival selected miR-27b-3p, miR-128-3p, and miR-34a-5p. Groups in both graphs are represented by corresponding miRNA profile and color. (a) Pruned decision tree for overall survival classified samples into 4 groups with different hazard ratios (HRs). Each step includes the HR, number of events defined by death due to osteosarcoma out of the total number of samples, and percentage samples in each group. Note that decision values are normalized Ct. A “yes” determination is always to the left. (b) Kaplan-Meier survival curve for each of the groups. The probability of survival for miR-27bhigh/miR-128low is significantly better compared to miR-27bhigh/miR-128high (P < .05) and miR-27blow/miR-34alow (P < .001). The probability of survival for miR-27blow/miR-34alow is also significantly worse compared to miR-27bhigh/miR-128high (P < .01) and miR-27blow/miR-34ahigh (P < .05). The other group comparisons are not statistically significantly different.
In the 1-year survival multiple miRNA and multiple variable models, both miR-7-5p and miR-30a were selected. In the multiple miRNA model, miR-101 was also included (Fig. 6a, b), but in the multiple variable model, weight categorization (greater or less than 32 kg) was selected instead (Fig. 6c, d). In the multiple miRNA model, miR-7-5phigh/miR-101low had an increased HR (3.16; MD = 0.87) but was only significantly different from miR-7-5plow/miR-30ahigh expression, which had a decreased HR (0.24; MD = 0.54). This decreased risk group was significantly different from miR-7-5plow/miR-30alow, which had an HR of 1.72 (MD = 0.63). MiR-7-5phigh/miR-101high (HR = 0.98; MD = 1.05) was not significantly different from any of the other groups. Eighteen samples were included in miR-7-5plow/high miR-30ahigh expression group, and only 2 cases died prior to 1-year; this was similar in the multiple variable model as well. In the multiple variable model, miR-7-5phigh/weightlow had an increased HR (3.87; MD = 1.04) and the worst probability of 1-year survival, which was significantly different from all other groups. MiR-7-5plow/miR-30alow (HR = 1.72; MD = 063) or high miR-7-5phigh/weighthigh (HR = 1.40; MD = 0.86) were not significantly different from each other and represented intermediate 1-year survival groups, which were significantly different from the low and high probability of 1-year survival groups. MiR-7-5plow/miR-30ahigh had the best probability of 1-year survival (HR = 0.24; MD = 0.54).
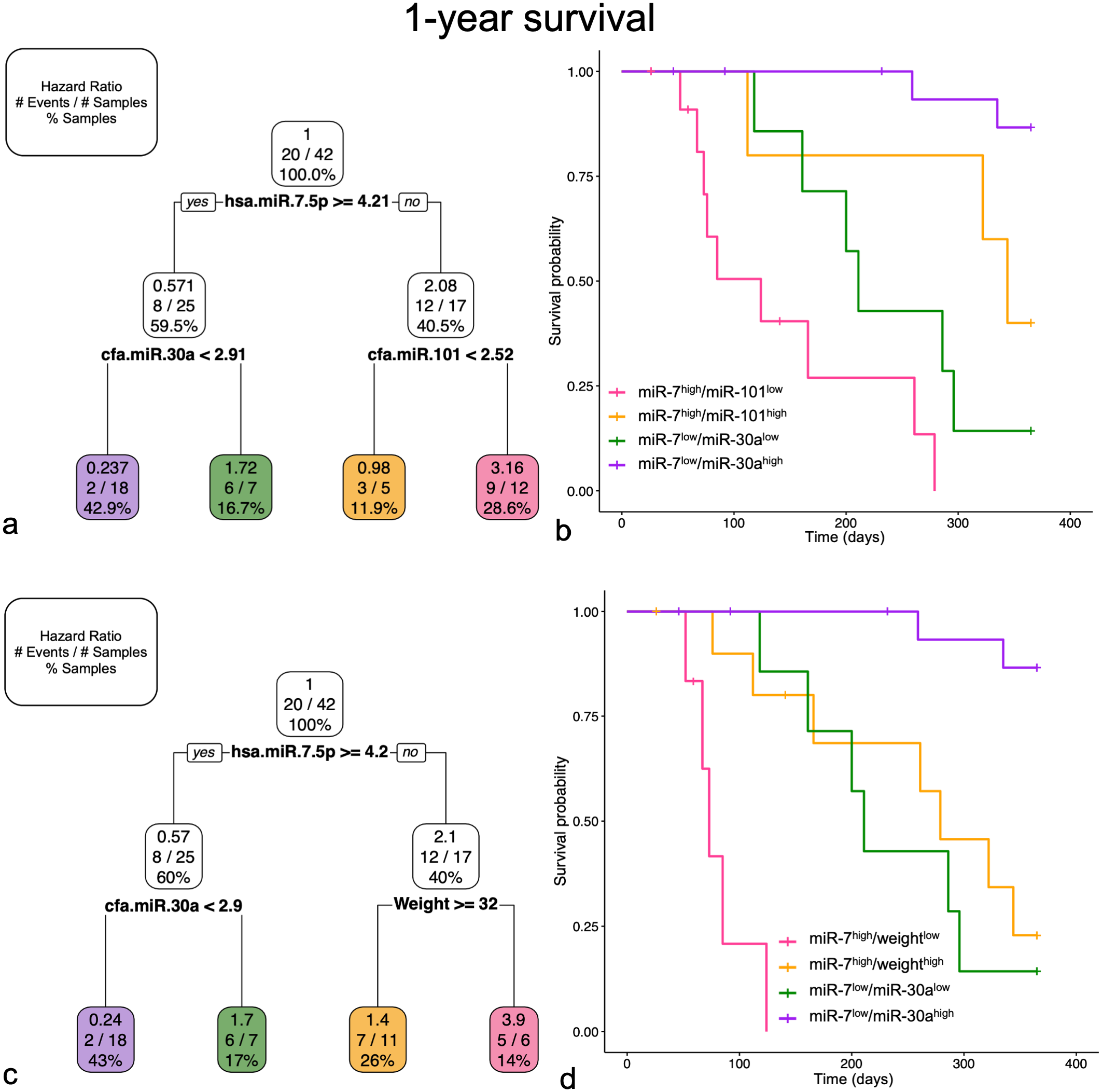
Multiple miRNA (a and b) and multiple variable (c and d) models for 1-year survival. Selected groups in both graphs are represented by corresponding miRNA profile and color. Pruned decision tree (a and c) for 1-year classified samples into 4 groups with different hazard ratios (HRs). Each step includes the HR, number of events defined by death prior to 1-year out of the total number of samples, and percentage samples in each group. Note that decision values are normalized Ct. A “yes” determination is always to the left. Kaplan-Meier survival curve for each of the groups (b and d). (a) Pruned decision tree only including miRNAs selected miR-7-5p, miR-30a, and miR-101. (b) In the Kaplan-Meier survival curve for the multiple miRNA model, the probability of 1-year survival for miR-7low/miR-30ahigh is significantly better compared to miR-7low/miR-30alow (P < .001) and miR-7high/miR-101low (P < .001). The other group comparisons are not statistically significantly different. (c) Pruned decision tree including multiple variables selected miR-7-5p, miR-30a, and weight (greater or equal to 32 kg). (d) In the Kaplan-Meier survival curve for the multiple variable model, the probability of 1-year survival for miR-7low/miR-30ahigh is significantly better compared to all other groups (P < .01). The probability of 1-year survival for miR-7high/weightlow is significantly worse compared to all other groups (P < .01). The other group comparisons are not statistically significantly different.
MicroRNA Expression in Matched Tissue and Plasma to Predict Clinical Outcome Parameters
Twenty-two miRNAs were previously investigated in OSA plasma samples and subsequently in matched primary OSA tissue samples from 15 dogs. 27 The normalized Ct values between the tissue and plasma samples were compared. Using a non-parametric 2-tailed Spearman correlation test, only miR-885-5p was considered statistically significant (r = −0.5607, 95% CI = −0.8386 to −0.05131; P < .05). MiR-16-5p (r = −0.4679, 95% CI = −0.7968 to 0.07505), miR-145-5p (r = −0.4464, 95% CI = −0.7867 to 0.1019), and miR-1271 (r = −0.4607, 95% CI = −0.7935 to 0.08410) were approaching statistical significance (P < .1). All had negative correlations.
Fifteen dogs with matched primary OSA tissue miRNA and plasma miRNA values were considered in multiple miRNA models to predict OS, DFI, and 1-year survival (Supplemental Figure S1). For OS (Supplemental Figure S1a), tissue miR-542high (HR = 2.35; MD = 0.56; OS = 121 days) had a decreased probability of survival, but this was only significantly different from tissue miR-542-3plow/plasma miR-195low (HR = 0.84; MD = 0.14; OS = 537.5 days). Tissue miR-542-3plow/plasma miR-195high (HR = 0.27; MD = 0.48; OS = <50% of cases experienced an event) only included 3 cases, which may have impacted the ability to determine statistical significance. Plasma miRNAs miR-19a and miR-195 predicted DFI (Supplemental Figure S1b). Plasma miR-19alow (HR = 2.55; MD = 0.59) had a shorter DFI (112 days), which was significantly different from plasma miR-19ahigh/plasma miR-195low (HR = 0.91; MD = 0.18; DFI = 406 days). The last group, plasma miR-19ahigh/plasma miR-195high (HR = 0.26; MD = 0.49; DFI = <50% of cases experienced an event), again only included 3 cases and was not considered significantly different from the other groups. Finally, for 1-year survival (Supplemental Figure S1c), all 3 groups were significantly different from each other. Tissue miR-19ahigh/plasma miR-19alow had the worst probability of 1-year survival (HR = 2.73; MD = 0.81). Tissue miR-19ahigh/plasma miR-19ahigh represented an intermediate probability of 1-year survival (HR = 1.32; MD = 0.54), while tissue miR-19alow had the best probability of 1-year survival (HR = 0.21; MD = 0.23), with all non-censored dogs reaching 1 year.
In Silico MicroRNA Targets
TargetScan and miRDB databases were searched for each miRNA shown in Figs. 1, 2. The top 5 targets for each miRNA in each database were recorded and compared (Supplemental Table S10). Genes determined to be one of the top 5 targets in both databases were reported. For miR-9-5p, targets included ONECUT2 and POU2F1. For both miR-196a-5p and miR-196b, HOXA7 was reported. MiR-196a-5p additionally had HOXC8 and NR6A1. HECTD2 was reported in both databases for miR-221 and similarly FOXJ3 for miR-708-5p. GALNT3 was reported for miR-885-5p and B3GNT5 for miR-30a. Three common genes were reported amongst the databases for miR-145-5p, including FSCN1, ABHD17C, and FLI1. Finally, genes OSR1 and PSMB8 were reported gene targets for miR-451a, and HSPB8 was reported for miR-126-5p. MiR-378a-3p, miR-542-3p, and miR-30c-5p had no gene targets in common in the top 5 selections between the 2 databases.
Discussion
Multiple individual tissue miRNAs in this study were able to separate this OSA population into long and short DFI and long and short OS groups, as well as differentiate between those surviving beyond 1 year or not (Supplemental Tables S6–8). Eleven of these individual miRNAs were similarly predictive across all clinical outcome measures. This contrasts with other clinical parameters, including location and serum ALP activity, which did not predict clinical outcome in this study (Supplemental Table S9). As the outcome prediction for canine OSA has not changed in decades, this represents a potential advancement for clinical decision-making.4,21,34,36,38
Multiple tissue miRNA models separated patients into 4 different groups for DFI, OS, and 1-year survival (Figs. 4–6). These refined the stratification of patients compared to single miRNAs. For example, in the miR-7-5plow/high miR-30ahigh expression group, only 2 of 18 cases died prior to 1 year. In contrast, 8 of 25 cases included in the low expression group of miR-7-5p alone died prior to 1 year. For DFI, miR-16-5p alone included 9 cases in the group with a poor prognosis, whereas the multiple miRNA models, which included miR-16-5p, miR-451a, and miR-885-5p, refined the poor prognosis group and included 20 cases (Fig. 4). The miR-16-5plow, miR-16-5phigh/miR-451ahigh, and miR-16-5phigh/miR-451alow/miR-885-5plow groups were similar and not significantly different. MiRNAs were the strongest variables when combined with other clinical parameters. In multiple variable analyses, only miRNAs were selected for DFI and OS (Figs. 4, 5). Weight was previously correlated to progressive disease and 1-year survival in other studies. 34 Weight (≥ or <32 kg) was selected by our model of 1-year survival in multiple variable analyses along with miR-7-5p and miR-30a (Figs. 6c, d). Weight was used to distinguish between dogs with high expression of miR-7-5p and resulted in miR-7-5phigh/weightlow having a decreased likelihood of achieving 1-year survival. Based on our results, tissue miRNA profiles were successfully able to stratify patients for DFI, OS, and 1-year survival. Dailey et al 11 previously investigated multiple miRNA signatures (3-miRNA expression-based risk score) to distinguish between patients with what they described as having “good” and “poor” prognosis. Their models were only used to separate their population into 2 groups. In contrast to this study, other clinical parameters in combination with miRNAs were not used to determine their risk score. In their multivariate analyses, their risk score and proximal humeral location were both associated with clinical outcome.
Thirteen miRNAs had similar levels in both primary and pulmonary metastatic OSA tissues but were different compared to normal lung tissue (Figs. 2, 3). The miRNAs with decreased expression in OSA tissues compared to normal lung tissue had overlapping normalized Ct values. MiRNAs with increased expression in primary OSA tissues compared to normal lung tissue demonstrated the greatest separation of normalized Ct values. MiR-9-5p and miR-196a-5p had no overlap in expression between cases and controls. In fact, only 1 lung sample had a Ct value <35.00 across both miRNAs. MiR-9-5p has high expression in canine OSA tumor tissue and cell lines and is implicated in cell invasion and migration. 14 One of the target genes for miR-9-5p is ONECUT2, which is involved in multiple hallmarks of cancer across various human cancers. 43 This supports the role of miR-9-5p as an oncomiR. In addition, POU2F1 was determined to be a target of miR-9-5p. POU2F1 interacts with NPR3 and the PI3K/AKT pathway, which regulates cell growth in human OSA cell lines. 25 The findings here also support the massive upregulation of miR-196a-5p in canine OSA, at least compared to normal lung tissue. MiR-196a-5p is consistently down-regulated in human OSA tissue compared to normal bone; however, there is much larger variability in canine OSA tissue with many cases having massive upregulation compared to normal bone in 1 study. 29 MiR-196b was expressed in all OSA samples, except for 1 primary and 1 pulmonary metastasis sample, and showed no expression in any normal lung samples. Multiple presumed targets of miR-196a-5p and miR-196b are HOX genes, which are clustered alongside these miRNAs.24,42 These miRNAs and NR6A1 were described as a master regulator of HOX genes, which are normally involved in embryologic development.7,42 HOX genes are known to be involved in cancer development and progression.13,24 The mature miRNA sequence of miR-196a-5p and miR-196b only differs by 1 nucleotide. 8 Although the sequences are very similar, and these 2 miRNAs have a similar pattern overall, there are differences in the expression data in this study supporting sufficient specificity of the primers. These miRNAs, which are expressed in both primary and metastatic OSA tissue and not in normal lung tissue, along with their presumed gene targets, are likely important to disease progression and could represent promising novel therapeutic targets for treating microscopic disease.
The investigation of both primary and metastatic tissue allows for a greater understanding of how tumor cells adapt to a new tissue/organ microenvironment. The analysis of whole tissues by RT-qPCR allows investigation of the miRNA profile of the tumor cells and the tumor microenvironment. MiR-206 was not differentially expressed between normal lung tissue and pulmonary metastases but was differentially expressed compared to the primary OSA tissues. In addition, 2 miRNAs were differentially expressed across all 3 groups (Fig. 1; miR-200c-3p, miR-214-3p). Determining whether these miRNAs are reflective of the tumor microenvironment or are important in disease progression and metastases is more challenging.
MiR-200c-3p had the highest expression in normal lung tissues (Ct <26.50) and is enriched in lung tissue and other epithelial tissues but not in bone. 19 Pulmonary metastases had intermediate expression levels of miR-200c-3p (Ct values from 25.43 to 35.00), compared to the other 2 groups, with 2 samples having no expression. Most primary OSA samples had no expression or had very low expression. This suggests this miRNA is important in lung tissue and may be involved in the growth and establishment of pulmonary metastases. Berlanga et al 3 proposed that miR-200c is upregulated in OSA cells present in the lung to promote their integration into this new tumor microenvironment. One caveat in the use of whole tissue is the inability to determine with exact certainty the precise composition of the tissue despite robust quality control practices. An alternative explanation for increased miR-200c-3p in the pulmonary metastatic samples is the presence of variable amounts of normal lung tissue. If the 2 metastasis samples that had no expression of miR-200c-3p were composed entirely of tumor cells, this could explain their similarity to primary OSA samples in this regard. Methods that use formalin-fixed, paraffin-embedded tissue sections, such as in situ hybridization or miRNA isolation from formalin-fixed paraffin-embedded tissues for RT-qPCR could allow more precise targeting and evaluation of tumor-only areas within affected lungs.
In multiple studies, miR-214-3p levels in plasma of dogs with appendicular OSA receiving standard-of-care therapy reflected clinical outcome.16,17,27 High expression of this miRNA was correlated to a shorter DFI and OS, and decreased likelihood of 1-year survival. In contrast to plasma, high expression of miR-214-3p in primary OSA tissue predicted a longer DFI and OS and greater likelihood of 1-year survival. This inverse relationship between plasma and tissue expression of this miRNA and clinical outcome parameters poses additional questions into the mechanisms that may be involved. Further investigation into the cellular source of miR-214-3p production, such as by in situ hybridization, is warranted as greater evidence mounts for its importance in canine OSA. One hypothesis for its relationship to clinical outcome parameters is the release of this miRNA into circulation by both primary and metastatic tumors. In tissue, miR-214-3p had the highest expression in primary OSA cases, followed by pulmonary metastases and the lowest expression in normal lung tissue. This supports this hypothesis. MiR-214-3p holds great promise as a prognostic biomarker for canine OSA but may not be suitable as a therapeutic target, as normal lung tissue also has relatively high expression (Ct values <29.00).
The data in this study comparing matched samples (tissue and plasma) in canine OSA remains novel in the canine literature. Fifteen cases had matched plasma and primary OSA tissue samples, and correlations were evaluated. Only miR-885-5p had a significant correlation. Craig et al 10 only found 1 miRNA (miR-143) that had a significant correlation in plasma and lymph node aspirates from healthy dogs and 7 and 8 miRNAs that correlated between the 2 sample types in dogs with B-cell and T-cell lymphoma, respectively. This could be due to the low number of matched pairs or the small subset of miRNAs investigated. Unfortunately, direct fold-change comparisons are not possible due to the substantially different methods, particularly with regard to sample input and normalization strategies between plasma and tissue. Finally, only linear relationships were investigated in this study, and there may be a nonlinear relationship between plasma and tissue expression.
A single biomarker would be cost-efficient and convenient, but patients and OSA are complex, and this may not be a realistic solution. In our multiple variable models, miRNAs predominated in determining clinical outcome, and miRNAs have proven to be important compared to other clinical variables in other canine OSA studies.11,17 As standard-of-care therapy involves amputation and histology to confirm the diagnosis, tissue is regularly available and plasma is readily obtained pre-operatively. Within our matched samples here, multiple miRNA models selected both tissue and plasma levels of miRNAs to predict OS and 1-year survival. Interestingly, the 1-year survival model chose miR-19a-3p expression in both the plasma and tissue. These results support the utility of miRNA expression in patient-matched samples to predict clinical outcome, and further investigation is warranted in a larger population of patient-matched samples.
The normal lung tissue group only included 5 samples. This is an important limitation of the study, and results utilizing this group, although strong for some miRNAs, should be interpreted with caution. A larger study population to account for sex and other factors is required to confirm these results. Unfortunately, due to study timeline and case load restrictions, only 1 female dog was able to be included in this group. To the author’s knowledge, no studies have been performed in canine lung tissue to determine if sex differences may exist in this tissue specifically. The female lung sample was an outlier for miR-29b-3p, -126-5p, -150-5p, and -223-3p, so these miRNAs in particular may be further interpreted with caution. Sex differences were investigated in the primary OSA tissue group, as this represented a greater cohort of animals to more accurately investigate whether sex may be a relevant factor in miRNA expression in tissues. No miRNAs were statistically significantly different when cases were separated into male or female groups (Supplementary Table S4). The overall weight range in the normal lung was also lower than is typical of dogs with OSA.
An additional limitation of this study is that non-pulmonary sites of metastases were not investigated. Since miR-9-5p, miR-196a-5p, and miR-196b have limited expression in normal lung tissue, these findings cannot necessarily be extrapolated to other tissues in the body, which might have higher background expression. Likewise, metastases to other locations may not have high expression of these miRNAs. Inclusion of a greater number of pulmonary metastases, and those from additional locations, combined with normal tissues, would be useful to confirm these findings. Whether treatment, such as radiation, influences miRNA expression in the pulmonary metastases was also not evaluated, as it was varied and so the groups would be too small. The prognostic relevance of these miRNAs can only be applied to dogs diagnosed with OSA without clinically detectable metastatic disease receiving standard-of-care therapy, as these were the inclusion criteria for this study. Subtype, grade, and other histologic features were not evaluated in this study to determine if miRNAs alone or in combination with clinical parameters can predict prognosis without histology. Future studies should investigate if there is a relationship between histologic features and miRNAs and whether combining histology and miRNA expression can better predict clinical outcomes. In addition, only select miRNAs were investigated, and there may be others that have similar or better predictive value. Finally, whole tissue offers insight into the overall tumor and its microenvironment, but the proportion of tumor tissue and normal surrounding tissue cannot be accurately quantified. Further investigation with formalin-fixed paraffin-embedded tissues can offer more targeted tumor area selection, and methods such as in situ hybridization can provide direct visualization of miRNA expression for further understanding the role of miRNAs in canine OSA.
Conclusion
MiRNAs are differentially expressed between primary OSA, pulmonary metastases, and normal lung tissues. MiRNAs such as miR-9-5p, miR-196a-5p, and miR-196b had limited expression in normal lung but relatively consistent expression in OSA tissues, making them important potential therapeutic targets. Multiple miRNA models are better able to stratify patients than single miRNAs and successfully identify both short- and long-term survivors. Further investigation is required to better understand the relationship of miRNAs within the tumor microenvironment and how multiple sample types, such as tissue and plasma, used together can better predict clinical outcome with miRNAs.
Supplemental Material
sj-pdf-1-vet-10.1177_03009858251356603 – Supplemental material for Differential expression of miRNAs in primary canine appendicular osteosarcoma tissue and pulmonary metastases
Supplemental material, sj-pdf-1-vet-10.1177_03009858251356603 for Differential expression of miRNAs in primary canine appendicular osteosarcoma tissue and pulmonary metastases by Latasha Ludwig, Heather Treleaven, Roger Moorehead, Robert A. Foster, R. Ayesha Ali, R. Darren Wood and Geoffrey A. Wood in Veterinary Pathology
Footnotes
Acknowledgements
The authors would like to acknowledge Dr Matthew A. Vermey, Grace Kim, Dr Melissa Prickaerts, and David Marom for their technical assistance. As well as past members of the G. Wood laboratory, including Dr Courtney Schott, for collection of tissue samples, for the tissue archive from which these samples were accessed, and clinical outcome information.
Author Contributions
All authors contributed to advisement on the experimental design. LL performed and/or supervised the experimental protocols. Statistical analyses were performed by HT with advisement from RAA, GAW, and LL. The manuscript was written by LL, HT, and GAW with contributions from the other authors.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was funded by OVC Pet Trust (055845) and NSERC (RGPIN-2020-06472) to GAW and Vanier Canadian Graduate Scholarship (NSERC) and OVC Fellowship to LL.
Supplemental Material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
