Abstract
Mycobacterium ulcerans ecovar Liflandii (MuLiflandii) was identified as the causative agent of mycobacteriosis in a research colony of Zaire dwarf clawed frogs (Hymenochirus boettgeri) at the University of Michigan. Clinical presentation included lethargy, generalized septicemia, cutaneous granulomas, coelomic effusion, and acute mortality. Identification of the mycobacterial species was based on molecular, microbiological, and histopathologic characteristics. These findings indicate that MuLiflandii is a primary cause of morbidity and mortality in Zaire dwarf clawed frogs and should be considered in the differential diagnosis of sepsis and coelomic effusion in amphibians. Mycobacterial speciation is important given the variability in pathogenesis within the family Mycobacteriaceae and the implications for both animal and human health as potential zoonoses. The Zaire dwarf clawed frog is a species common in the pet trade, and these findings provide consideration for this pathogen as a potentially important public health concern. This is the first report of MuLiflandii infection in the genus Hymenochirus and illustrates the diagnostic challenges of differentiating among both mycolactone-producing mycobacteria and Mycobacterium marinum. Furthermore, we demonstrate the utility of environmental sampling for this pathogen within the tank system, suggesting this mode of sampling could replace the need for direct frog surveillance.
Keywords
Mycobacterium spp. are aerobic, nonmotile pleomorphic bacilli in the order Actinomycetales and have a broad host range. There is a wide array of mycobacterial species in freshwater and marine environments, many of which are nonpathogenic. However, among those that are pathogenic, Mycobacterium marinum is particularly important because of its extremely broad host range, pathogenicity, and relatively high zoonotic potential. Another related pathogenic species, Mycobacterium ulcerans, is a water-borne, nontuberculous mycobacterium endemic to tropical regions of Africa, Australia, Central America, and eastern Asia. It is associated with necrotizing skin lesions (Buruli or Bairnsdale ulcers) in otherwise healthy people. 38
Nontuberculous mycobacteria present a significant concern for aquatic species, both in the laboratory setting and pet trade, as a cause of ulcerative dermatitis and septicemia. Trott et al 38 characterized a lethal outbreak of M. liflandii in Western clawed frogs (Xenopus tropicalis), in which animals presented with cutaneous granulomas, coelomic distension, and sepsis. Later, Suykerbuyk et al 35 identified a fatal outbreak of M. liflandii in a European colony of the same species and suggested the potential for Mycobacterial spp. transmission through the international pet trade. Godfrey et al 10 identified Mycobacteria liflandii in a colony of Xenopus laevis in the research laboratory setting, demonstrating pathogenicity in multiple anuran species.
Following its initial characterization, Mycobacterium liflandii was later phylogenetically identified as M. ulcerans-like, and in 2013, the classification M. ulcerans ecovar Liflandii (MuLiflandii) was coined based on whole-genome sequencing. 36 Outbreaks of fatal disease caused by M. ulcerans have been reported in anurans 36 as well as fish, 44 and given the reclassification of the organism, it is possible that other outbreaks may have actually been caused by MuLiflandii rather than M. ulcerans.
This report documents the first outbreak of disease due to MuLiflandii in African dwarf clawed frogs (Hymenochirus boettgeri). Historically, this frog species has been utilized as a laboratory animal model in the study of limb regeneration, 11 molecular ecology and evolution,18,19 antimicrobial properties of peptides,21,41 mechanisms of prey capture, 4 and chytridiomycosis research. 29 This species is physically smaller than other frogs, which may contribute to its historic popularity in the global pet trade as well as its more recent use as a research model. 21 Due to its relatively new introduction into the laboratory setting, the disease status and pathobiology of this species are largely unknown but are presumed to be like other aquatic anuran laboratory species.
This study highlights the importance of MuLiflandii as a pathogen in the laboratory setting, and the implications on global transmission and epidemiology of the disease in anuran species. Not only is the dwarf clawed frog a popular species in the global pet trade and a useful model in research settings, it has been introduced nonindigenously to the wild in southeastern Florida, likely as a result of the international pet trade.19,27 Historically, Hymenochirus sp. have been associated with outbreaks of salmonellosis, another potential zoonosis, 23 emphasizing the importance of characterizing disease susceptibility for this species. Finally, we highlight the importance of proper Mycobacterium speciation to understand the relevance of these organisms in disease pathogenesis of aquatic species, and the potential zoonotic implications for humans caring for these animals.
Materials and Methods
Animals and Husbandry
All animals were housed in an AAALAC-accredited facility (University of Michigan, Ann Arbor, MI) and under an approved Institutional Animal Care and Use Committee protocol designed to study chytridiomycosis in anurans. The colony consisted of approximately 300 individuals housed at about 2 frogs per liter of water, fed a mixture of blackworms (Lumbriculus variegatus) and bloodworms (Chironomidae), and free of disease for more than 19 months before clinical signs first occurred. At delivery, frogs were quarantined in static tanks and underwent gradual heat treatment to eradicate chytrid fungus before combining into a single recirculating system (Techniplast, West Chester, Pennsylvania). Gradual heat treatment spanned 3 weeks, starting with a week-long increase from ambient temperature to 30°C, maintenance at 30°C for the second week, and a gradual decrease to ambient temperature over the third week. At breeding age, the frogs were temporarily maintained in static tanks. The recirculating system was maintained at 28°C, and water quality parameters (pH, nitrates, nitrites, and ammonia) were assessed weekly (API; Mars Fishcare, Chalfont, Pennsylvania).
Necropsy and Histopathology
Clinically affected frogs were humanely euthanized by submersion in tricaine methanesulfonate (MS222) at a concentration greater than 250 mg/L for 10 minutes. A total of 8 frogs were collected for evaluation at necropsy. The coelomic cavity and buccal mucosa were aseptically sampled for microbiological analysis, followed by immersion fixation of the carcass in 10% neutral-buffered formalin for 12 to 24 hours. Following fixation, each frog was decalcified in Immunocal (StatLab, Columia, Maryland) for 5 to 8 hours. The body of each animal was bisected longitudinally, the limbs were removed, and hemisections of body and the limbs were placed in tissue cassettes for histopathologic examination by the In Vivo Animal Core facility in the Unit for Laboratory Animal Medicine at the University of Michigan by routine methods. Tissues were processed in an automated tissue processor (TissueTek VIP; Sakura Finetek USA, Torrance, California), embedded in paraffin, sectioned at 4 μm, and stained with hematoxylin and eosin by routine histological methods on an automated histostainer (Leica ST5010 XL; Leica Biosystems, Buffalo Grove, Illinois). Gram staining was performed using alcoholic crystal violet-oxalate stain followed by aqueous Lugol iodine solution, decolorized in acetone, and stained with Twort stain (Newcomer Supply, Middleton, Wisconsin). Ziehl-Neelsen acid-fast staining was performed using Ziehl-Neelsen carbol-fuchsin followed by decolorization in acid alcohols and then staining with methylene blue working solution (Rowley Biochemical, Danvers, Massachusetts). Grocott methenamine silver stain was performed using oxidation in 4% aqueous chromic acid then staining with methenamine silver solution, toning with 0.2% gold chloride, and counterstaining with Light Green Working Solution (Rowley Biochemical).
Bacterial Culture and Identification
The coelomic cavities of 3 frogs displaying clinical signs were opened and swabbed for general bacterial culture. The swabs were cultured on BBL trypticase soy agar plates with 5% sheep blood (TSA II; Becton Dickinson, Franklin Lakes NJ), BBL chocolate II agar (Becton Dickinson), and either BBL CDC 5% sheep blood agar with phenylethyl alcohol (PEA; Becton Dickinson, Franklin Lakes NJ) or tryptone yeast extract salts (TYES) agar, which was prepared following a published formulation. 13 Blood agar plates and TYES agar plates were incubated for 5 days at 22°C in room air, PEA and chocolate agar were incubated for 5 days at 35°C in an atmosphere of 6% CO2 and additional blood agar plates were incubated for 5 days at 35°C in an atmosphere of 6% CO2 and/or anaerobically at 35°C.
Aseptically collected coelomic tissues, excluding the gastrointestinal tract, were collected from 2 clinically affected animals and submitted to the IDEXX BioAnalytics microbiology laboratory for Mycobacterium-specific culture and identification. Multiple culture medias were employed to target fastidious Mycobacterium spp., including Middlebrook 7H11 agar (Thermo Fisher Scientific, Ann Arbor MI), BBL Lowenstein-Jensen agar (Becton Dickinson, Franklin Lakes NJ), BBL Mueller Hinton II agar (Becton Dickinson), and BBL MGIT mycobacterial growth indicator tubes supplemented with oleic acid, bovine albumin, sodium chloride, dextrose, and catalase (MGIT OADC Enrichment, Becton Dickinson), as well as amphotericin B, azlocillin, nalidixic acid, polymyxin B, and trimethoprim (MGIT PANTA Antibiotic Mixture; Beckon Dickinson). Mycobacterial cultures were incubated in an atmosphere of 7% CO2 at 30°C.
Morphologically distinct isolated colony types were harvested for proteomic analysis using a direct transfer method described previously. 25 Harvested bacteria on an etched target were overlaid with 1 µL of a saturated matrix solution of α-cyano-4-hydroxycinnamic acid in 50% acetonitrile and 2.5% trifluoroacetic acid (HCCA, Bruker Daltonics, Billerica, Massachusetts) and air dried prior to evaluation by matrix-assisted laser desorption/ionization time-of-flight (MALDI-TOF) mass spectrometry using a commercially available mass spectrometer (Microflex, Bruker Daltonics) and flexControl software (Bruker Daltonics). Identification of isolated bacteria was based on automated comparisons of the collected spectra for each colony type with integrated reference spectral databases by MALDI BioTyper software (Bruker Daltonics).
Molecular Analysis
A coelomic swab from a clinically infected frog was initially submitted to a commercial veterinary diagnostic laboratory for real-time polymerase chain reaction (PCR) analysis, and the extracted total nucleic acids were returned to the University of Michigan upon request. Previously extracted total nucleic acids, live feed samples, environmental samples, visceral tissues, and coelomic swabs were subsequently submitted to IDEXX BioAnalytics (Columbia, Missouri) for additional molecular analysis. Environmental samples included detritus from a static tank containing clinical frogs, feed samples, and recirculating system filter samples containing biofilm. Feed samples included both whole blackworms and frozen whole bloodworms. Blackworms are purchased commercially and stored at 5°C with water. Frozen bloodworms are purchased commercially, stored frozen, and defrosted before use. Proper personal protective equipment was utilized while handling feed. Total nucleic acids were extracted from frog tissues and all other sample types using a commercially available platform (NucleoMag VET; Macherey Nagel, Bethlehem, Pennsylvania). All real-time PCR assays tests were based on the IDEXX BioAnalytics proprietary service platform (IDEXX Laboratories, Westbrook, Maine). Hydrolysis probe–based real-time PCR assays used a commercially available master mix (LightCycler 480 Probes Master; Roche Applied Science, Indianapolis, Indiana) and a commercially available instrument (LightCycler 480; Roche Applied Science). Positive and negative controls were performed as well as a multiplexed hydrolysis-probe-based real-time PCR assay, targeting a eukaryotic gene (18S rRNA) and bacterial gene (16S rRNA) to demonstrate recovery of amplifiable nucleic acids and the absence of PCR inhibition. Conventional PCR to amplify sequences from an isolated Mycobacterium sp. colony used proprietary sequencing primers that flank the targets for a genus-wide Mycobacterium spp. assay and an assay targeting Mycobacterium ulcerans species, including MuLiflandii—referred to as Mycobacterium ulcerans/liflandii. Polymerase chain reaction products were submitted for Sanger sequencing (GENEWIZ, South Plainfield, New Jersey).
Several additional loci were amplified by conventional PCR for Sanger sequencing (GENEWIZ), including the enoyl reductase (ER) domain of the mycolactone polyketide synthase A gene (mlsA), insertion sequence 2404 (IS2404), malate synthase gene (glcB), fibronectin-binding protein A gene (fbpA), exported repetitive protein gene (Erp), and the 16S ribosomal RNA gene. Multiple attempts were made to amplify insertion sequence 2606 (IS2606) using 2 published primer sets.15,20,34,43 Sequencing primers and cycling conditions are described in Table 1. The resulting PCR products from all conventional PCR reactions were submitted for Sanger sequencing (GENEWIZ) and assembled into contigs with Sequencher software (Gene Codes Corporation). Primer binding sequences and regions of poor-quality sequence were trimmed from the ends. BLAST software (National Center for Biotechnology Information, http://www.ncbi.nlm.nih.gov) was used to compare contig sequences with sequences deposited in GenBank.
PCR primers used for detection of Mycobacteria ulcerans ecovar Liflandii (MuLiflandii).
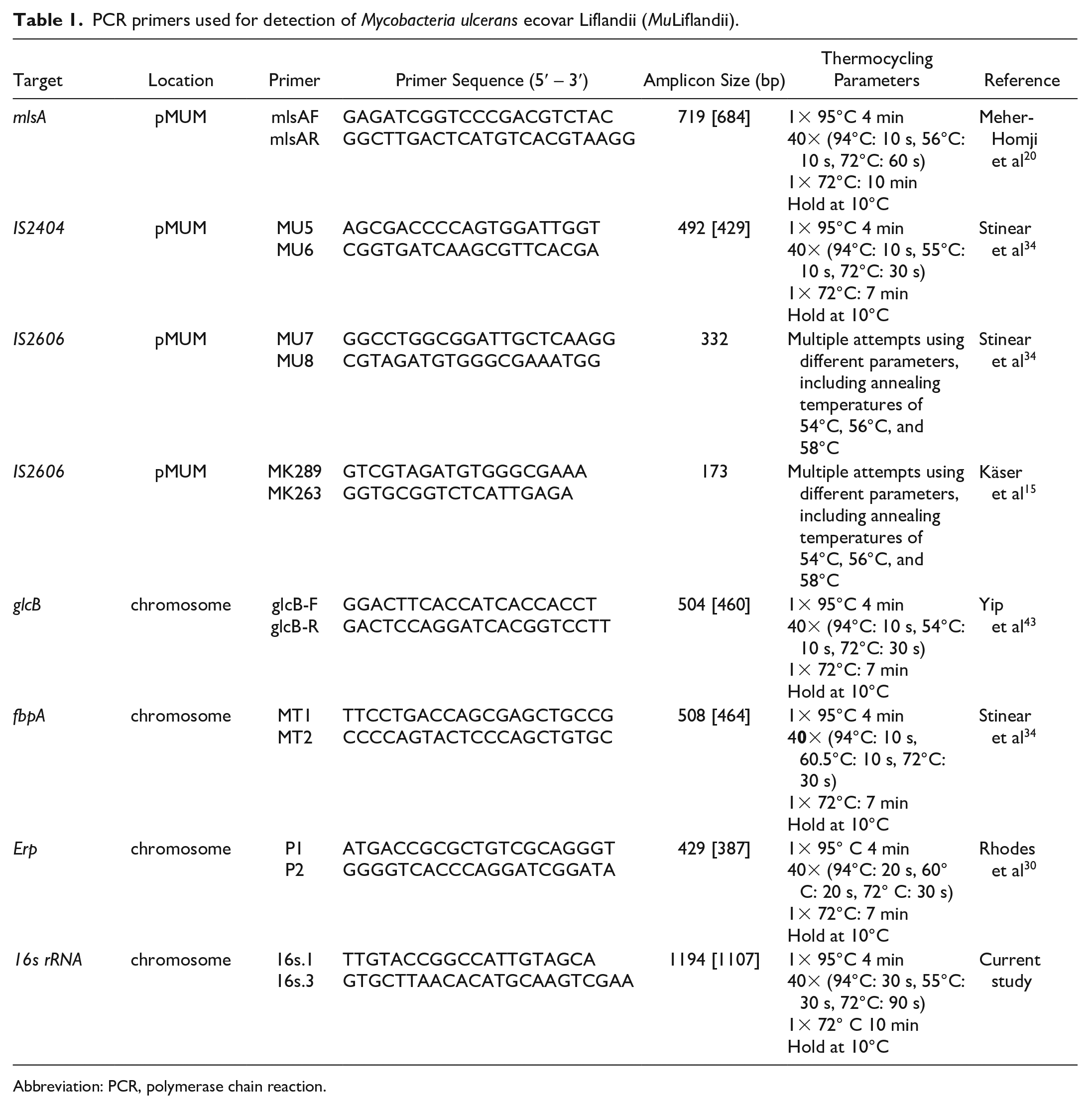
Abbreviation: PCR, polymerase chain reaction.
Histological lesion distribution in clinically affected frogs infected with M. ulcerans ecovar Liflandii (MuLiflandii).
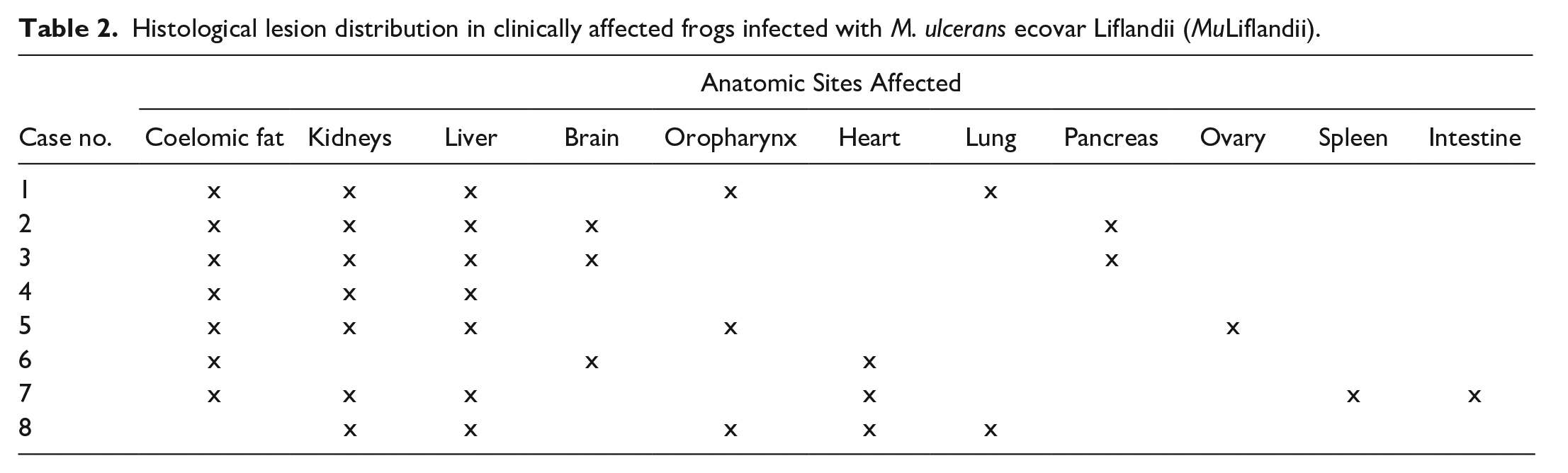

Results
Clinical Presentation, Necropsy, and Histopathology
Over a 5-month period, a colony of approximately 300 frogs from static and recirculating tanks experienced periodic mortality and morbidity (approximately 4 frogs per month) characterized by erythema of the inguinal region and the buccal mucosa, and dull mentation. Initial clinically impacted frogs were from stock purchased from the vendors. However, impacted frogs eventually included individuals born at our facility, housed in both static and recirculating systems. Clinically affected frogs appeared to be in adequate body condition as evidenced by sufficient adipose stores and skeletal muscling. Gross findings in affected frogs included oral and buccal erythema, cutaneous hyperemia, petechiae, ecchymoses, and coelomic distension with serous effusion (Fig. 1). In one frog, there was a focal, irregular, pale cutaneous lesion along the mandible, consistent with a granuloma.
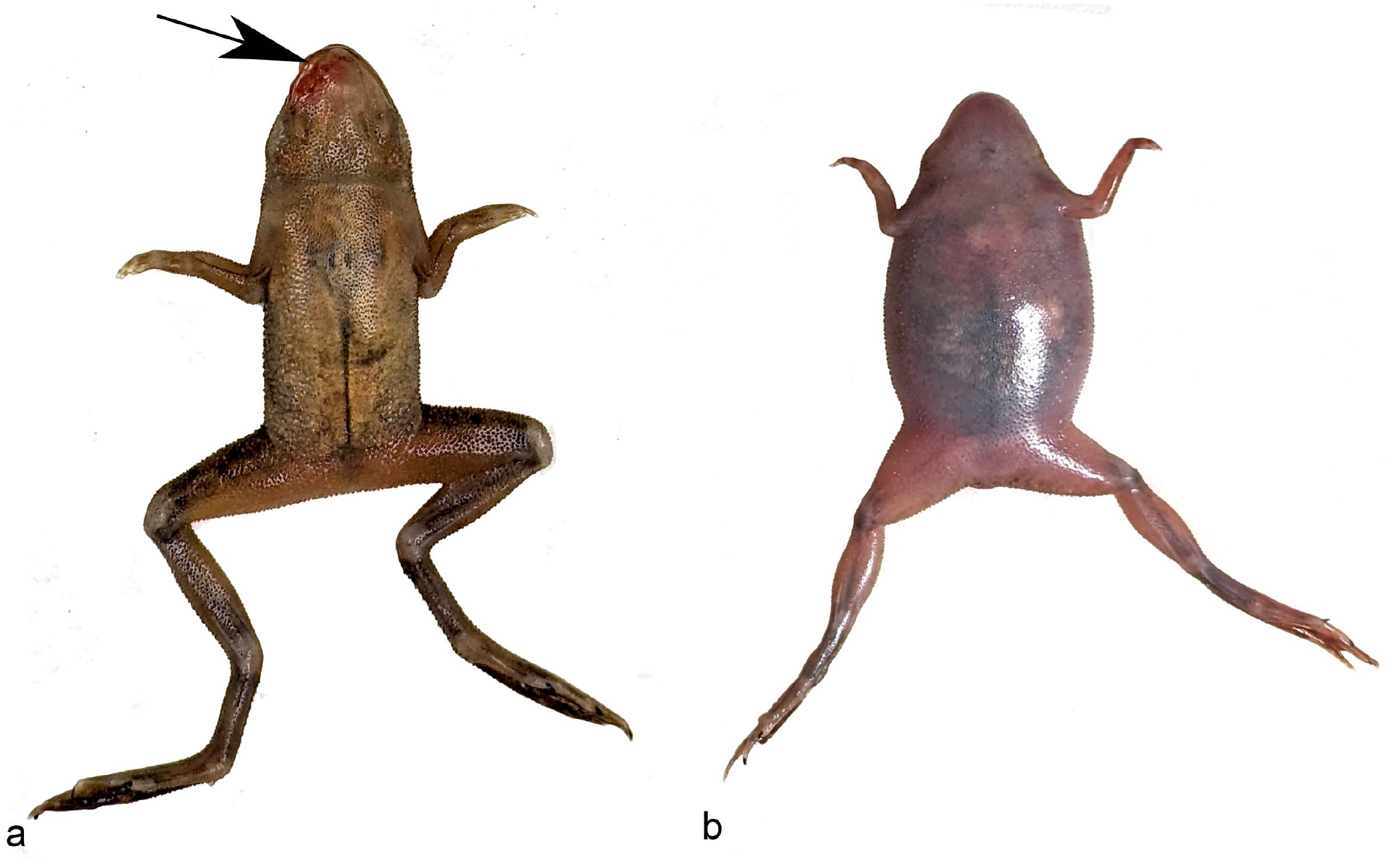

Systemic MuLiflandii infection, African dwarf frog. Affected frogs presented with (a) oral or cutaneous hyperemia (arrow), and/or (b) coelomic distension with effusion.
Histologically, lesions in affected frogs were observed in multiple organs, but appeared predominantly in the coelomic cavity as coelomic fat necrosis, with local extension to the serosal surfaces and parenchyma of other coelomic organs (Fig. 2a, b), including the kidney, liver, spleen, pancreas, intestines, and ovary, but also systemic spread to distant organs, including the brain. Necrotizing lesions in the coelomic cavity and associated organs were characterized by loss of cellular and tissue architecture, hypereosinophilia, nuclear pyknosis, karyolysis, and karyorrhexis, and were accompanied by generally minimal to mild inflammatory cell infiltrates composed predominantly of lymphocytes, histiocytes, and few heterophils. In several animals, there was multifocal mucosal necrosis of the bronchial epithelium (Fig. 3a), oral cavity, and oropharynx characterized by effacement and loss of surface epithelium, which was replaced by amorphous to granular eosinophilic necrotic debris, occasionally admixed with histiocytes and fewer lymphocytes that extended into the submucosal layers. In sections of liver (Fig. 3b), spleen, and kidney, there were often random, multifocal inflammatory infiltrates of lymphocytes and histiocytes without the presence of necrosis. In one animal, there was necrosis of the middle ear epithelium, which extended into the underlying tissues (Fig. 3c). In the brain of 3 animals, there was variable necrosis of the neuropil centrally within the cerebrum (Fig. 3d) or accompanied by mild to moderate inflammatory infiltrates affecting the meninges. As evidenced with Ziehl-Neelsen acid-fast staining, zones of necrosis contained myriad, extracellular, slender acid-fast bacilli (Figs. 2c, 3, insets), while foci of purely lymphohistiocytic inflammation contained relatively few extracellular and intrahistiocytic organisms (Fig. 3b, inset). Bacterial rods were gram-positive (Fig. 3e) and positive on Grocott methenamine silver stain (Fig. 3f).
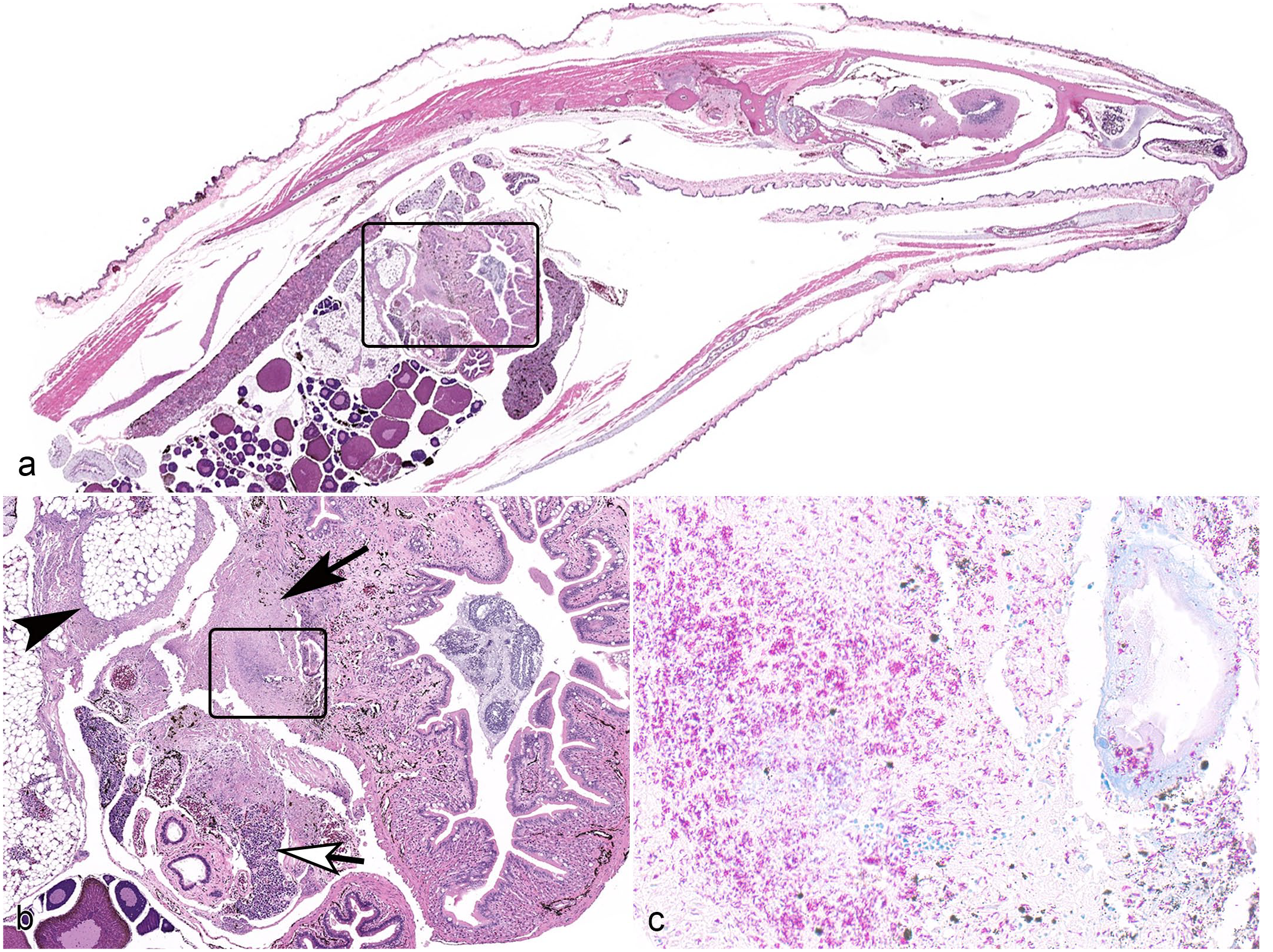
Systemic MuLiflandii infection, African dwarf frog. (a) Sagittal section of a frog with coelomic inflammation (black box). Hematoxylin and eosin (HE). (b) Higher power magnification of the box in (a) demonstrating involvement of the pancreas (white arrow), coelomic fat (black arrowhead), and serosal surfaces of the intestinal tract (black arrow). HE. (c) Higher power photomicrograph of black box in (b) demonstrating myriad acid-fast bacilli within necrotic lesions. Ziehl-Neelsen acid-fast stain.
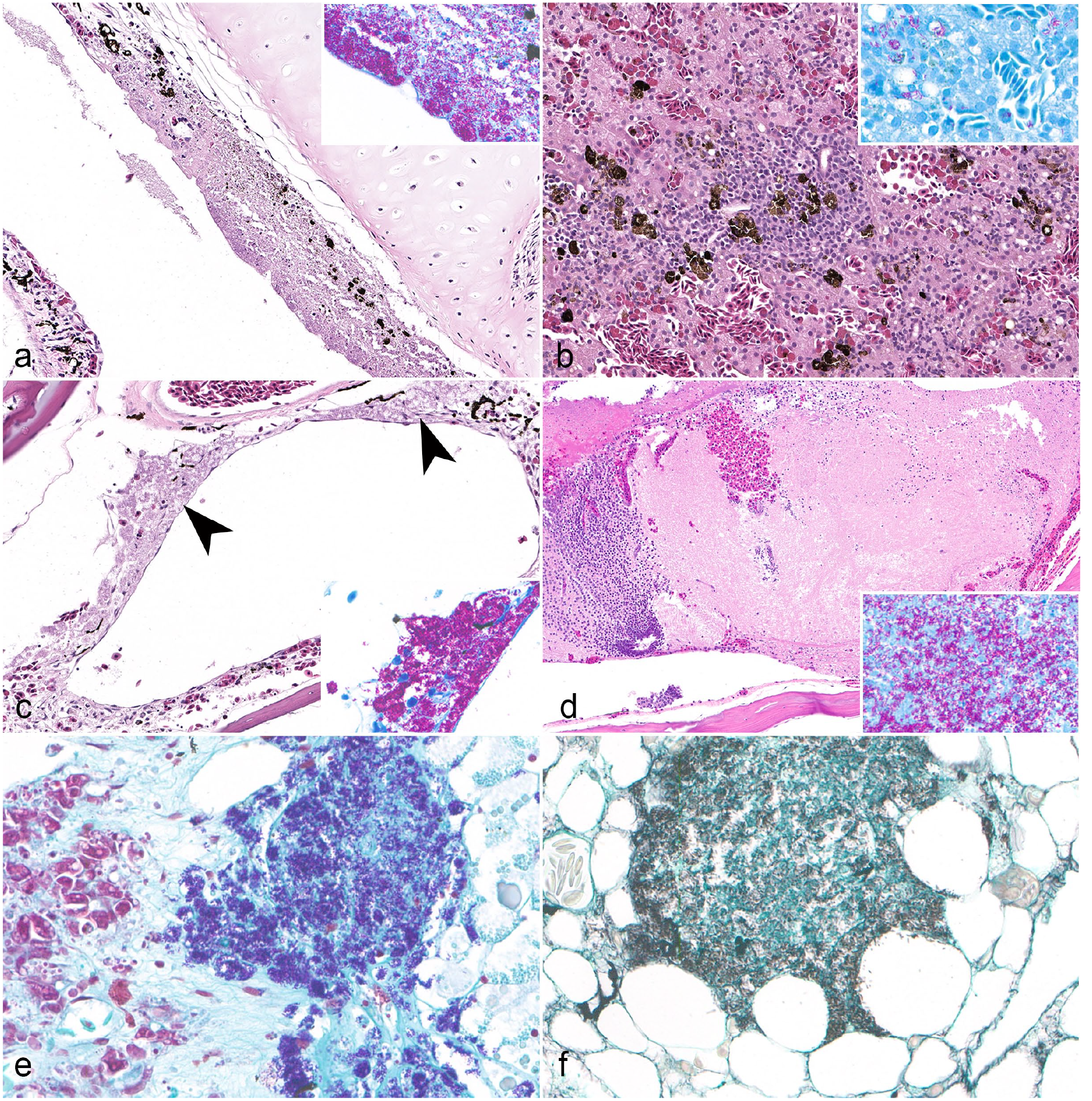
Systemic MuLifandii infection, African dwarf frog. (a) Full-thickness necrosis of the bronchial epithelium and underlying submucosa. Hematoxylin and eosin (HE). Inset, myriad acid-fast bacteria within necrotic debris. Ziehl-Neelsen acid fast (AF)). (b) Focal mononuclear to granulomatous inflammatory infiltrates within the liver (HE), associated with small numbers of acid-fast bacilli (inset, AF). (c) Areas of necrosis (arrowheads) within the middle ear characterized by hypereosinophilia, loss of cellular architecture, and cellular debris (HE), accompanied by large numbers of acid-fast bacilli (inset, AF). (d) Locally extensive necrosis within the cerebrum (HE), with myriad acid-fast bacilli present (inset, AF). (e) Bacteria within areas of necrosis were gram-positive (Hucker-Twort Gram stain) and (f) strongly positive on Grocott methenamine silver staining.
Bacterial Culture and Identification
A variety of commensal species and opportunistic pathogens were isolated from the initial coelomic swab, including Aeromonas hydrophila, Chryseobacterium arthrospharae, C. indologenes, Lysobacter sp., Mycobacterioides (basonym Mycobacterium) sp., Paenibacillus thiaminolyticus, and unidentified gram-negative rods and cocci.
From the Mycobacterium-specific cultures, small, rough irregular colonies with undulate margins morphologically consistent with Mycobacterium sp. were isolated and further evaluated by MALDI-TOF mass spectrometry. This isolate grew well at 30°C similar to M. ulcerans and M. marinum, but in contrast to Mycobacterium pseudoshottsii, a very closely related species that exhibits either no growth or very minimal growth at 30°C. 30 The collected mass spectra most closely matched the database spectra for Mycobacterium marinum, although with such a low score that the organism could only be identified to the genus level. Importantly, spectra from MuLiflandii were not represented in the available spectral databases.
Molecular Analysis
The initial submission of a coelomic swab to the first commercial diagnostic laboratory was reported positive for Mycobacterium marinum by real-time PCR. Subsequently, the previously extracted total nucleic acids tested positive for MuLiflandii and negative for M. marinum at IDEXX BioAnalytics.
Real-time PCR analysis of coelomic swabs collected from clinically affected frogs tested positive in all animals for MuLiflandii and uniformly negative for other pathogens, including Batrachochytrium dendrobatidis, Mycobacteroides (Mycobacterium) chelonae, Mycobacterium marinum, Pseudocapillaroides xenopi, and ranavirus (Table 3). A recirculating filter sample and an environmental detritus sample also tested positive for MuLiflandii and negative for M. marinum by real-time PCR, and live feed samples tested negative for both MuLiflandii and M. marinum by real-time PCR (Table 3).
Results of microbiological culture, real-time PCR analysis, and sequencing of samples for Mycobacterium spp.
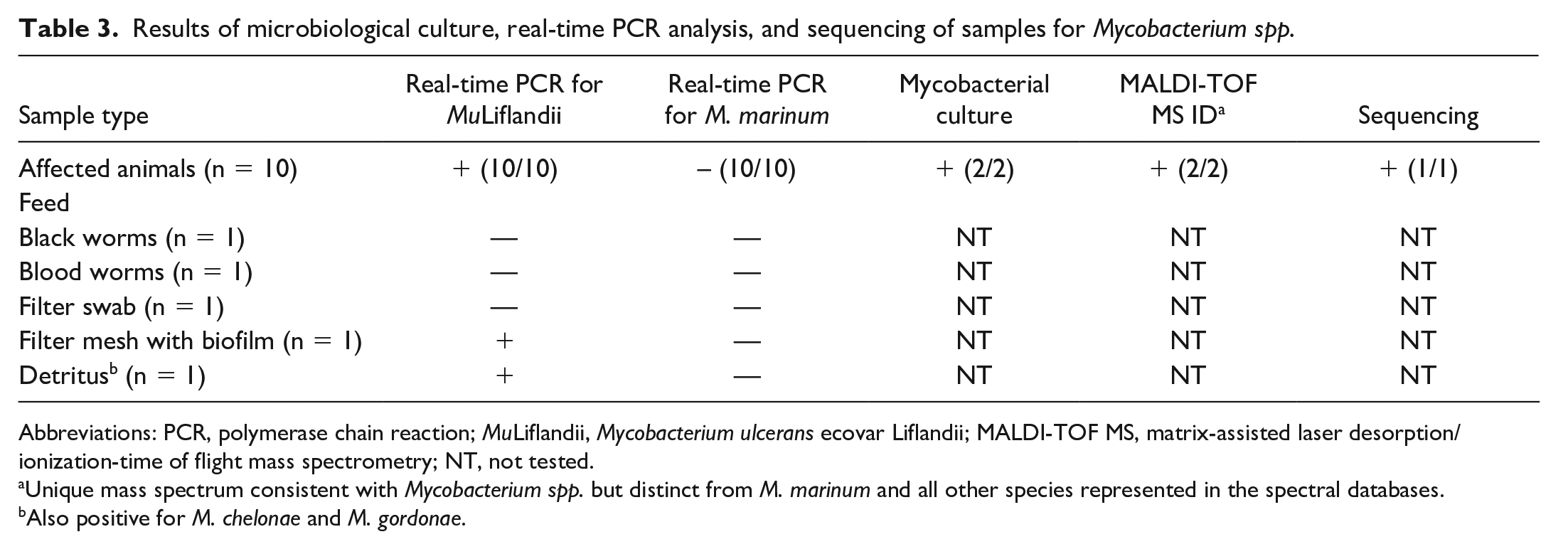
Abbreviations: PCR, polymerase chain reaction; MuLiflandii, Mycobacterium ulcerans ecovar Liflandii; MALDI-TOF MS, matrix-assisted laser desorption/ionization-time of flight mass spectrometry; NT, not tested.
Unique mass spectrum consistent with Mycobacterium spp. but distinct from M. marinum and all other species represented in the spectral databases.
Also positive for M. chelonae and M. gordonae.
Confirmatory sequence analysis across the targets for the diagnostic real-time PCR assays supported the positive real-time PCR test results for MuLiflandii. The gene segment surrounding the target for the genus-wide Mycobacterium spp. assay displayed 100% identity over 350 bases to MuLiflandii strain ASM001 (GenBank Accession CP023138.1) and differed from MuLiflandii strain 128FXT, other M. ulcerans strains, and some M. marinum strains by only 1 or 2 mismatched bases. The gene segment surrounding the target for the Mycobacterium ulcerans/liflandii assay displayed 100% identity over 221 bases to MuLiflandii, other M. ulcerans strains, and M. pseudoshottsii. The closest M. marinum sequence shared only 73.81% identity over 84/234 (35%) of the bases searched and the rest of the sequence did not align at all with any M. marinum sequences.
Taken together, sequence analysis was consistent with identification of the isolated Mycobacterium sp. as MuLiflandii. The ER domain of mlsA is required for production of the mycolactone core and is contained within a contiguous 110 kb cluster of genes for mycolactone biosynthesis. 22 The mlsA sequence displayed 100% identity to MuLiflandii, M. ulcerans, and M. pseudoshottsii with 684 matching bases out of 684 searched but did not align with any M. marinum sequences. IS2404 is present in M. ulcerans, including MuLiflandii and M. pseudoshottsii, but is not present in M. marinum. 33 The IS2404 sequence from the isolated Mycobacterium sp. displayed 99.3% identity to M. ulcerans, including MuLiflandii strains 128FXT and ASM001, and M. pseudoshottsii with 426 matching bases out of 429 searched. The IS2404 sequence did not align with any M. marinum sequences except for one sequence from M. marinum DL240490. This particular strain is very slowly growing, mycolactone F-producing, scotochromogenic strain isolated from cultured sea bass (Dicentrarchus labrax) that carries a pMUM plasmid with 99.9% identity to M. pseudoshottsii over 89,597 bases, is incapable of growth at 30°C, 28 displays phenotypic similarities to M. pseudoshottsii, and may have been misidentified as M. marinum. 30 The glcB sequence showed 100% identity over 460 bases to MuLiflandii strains 128FXT and ASM001, M. ulcerans strain BS123, and M. pseudoshottsii. The closest M. marinum sequence differed by a single base mismatch. The fbpA sequence displayed 100% identity over 464 bases to MuLiflandii strains 128FXT and ASM001 and M. ulcerans strain BS123. The closest M. marinum strain differed by a single base; however, the mismatch in this gene is the T-for-C transition at a polymorphic site that has been reported to serve as a key discriminator between M. marinum and M. ulcerans.33,43 The Erp sequence displayed 100% identity over 387 bases to MuLiflandii strain ASM001, and less than 96.7% identity to all other species. The 16S rRNA gene from the isolated Mycobacterium sp. displayed 100% identity over 1107 bases to MuLiflandii strain ASM001, another MuLiflandii isolate, Mycobacterium sp. CSTy-8.29, and a sequence identified and deposited in GenBank (AF456239.1) as M. marinum prior to the initial description of MuLiflandii. This sequence and Mycobacterium sp. CSTy-8.29 were nearly identical and when the entire sequences of these accessions were searched using BLAST, they displayed the greatest identity to MuLiflandii (99.74%-99.93%). IS2606 was not detected in the isolate using 2 published primer sets.15,34 IS2606 is present in M. pseudoshottsii, absent in M. marinum, 43 present in high copy numbers in human M. ulcerans isolates (eg, 91 copies in M. ulcerans Agy99), 36 and can be present either in low copy numbers (eg, 4 copies) 36 or completely absent 16 in MuLiflandii.
Discussion
This report documents the first outbreak of disease due to MuLiflandii in African dwarf clawed frogs. The disease presentation due to this organism is similar to other bacterial septicemias that clinically present as cutaneous hyperemia and/or ulceration (ie, “red leg”), but has some interesting characteristics that differentiate it clinically and histologically from other mycobacterial species. First, lesions due to MuLiflandii in this case series were decidedly necrotizing rather than predominantly granulomatous. Although mycobacterial disease in anurans is not always observed as discrete granulomas, historically granulomas are an indication of this disease in multiple species.3,24 There was a lack of clear and predominant granulomatous inflammation, including epithelioid macrophages and multinucleate giant cells, in these animals; rather, a predominantly necrotizing process was evident as previously reported in Xenopus spp. 9 The necrotizing character of these lesions has been reported in Xenopus tropicalis frogs with MuLiflandii infection, but disease in these frogs was also associated with more prominent tuberculoid lesions, most commonly affecting the spleen, among other organs, and was less commonly disseminated than in this series. 9 Similar to cases in Xenopus frogs, large numbers of acid-fast bacteria were prominent in areas of necrosis. The necrotizing lesions associated with MuLiflandii infections in anurans is thought to be due to production of mycolactone E, which has potent cytopathic effects leading to tissue necrosis,9,10 and the lack of multinucleated giant cell formation, possibly due to an inherent defect or deficiency in fusion of macrophages in this species. 9
Second, while the disease observed was multisystemic in nature, as often is the case with mycobacterial disease, 24 there was predominant involvement of coelomic fat, with what appeared to be secondary lesions along serosal surfaces as well as other organs. This is not necessarily unusual considering the coelomic cavity allows for local spread of infection within a contained space, but the consistent involvement of coelomic fat was an interesting feature in this group of animals. Given the variable presence of lesions in the oral cavity, oropharynx, middle ear, and bronchial epithelium, the route of exposure was assumed to be through these sites secondary to environmental water contamination, but an obvious route for coelomic inflammation was not apparent. It is likely that defects in these areas, or even translocation through the gastrointestinal tract allowed for entry of the bacteria into the tissues and resulted in local and systemic spread.
Of the observed clinical signs, coelomic distension was of particular note. Effusion within the coelomic cavity, subcutaneous lymphatics, or both is a common nonspecific finding in anurans with multiple potential differentials. 24 In captive anurans, the most common cause of coelomic distension is chronic renal disease, 24 but this clinical presentation may also be caused by a variety of conditions, including cutaneous lesions, vasculitis, cardiac insufficiency, systemic bacterial or viral infections, or parasitic diseases. 24 Noninfectious causes may include acquired polycystic nephropathy, hypocalcemia, hypoproteinemia, gastrointestinal disease, heart failure, renal toxicity, and congenital anomalies of lymph hearts. 24 In laboratory African clawed frogs, a common cause of coelomic distension is ovarian hyperstimulation syndrome, due to their use in reproductive studies. 24 In this study, coelomic distension was determined to be of an infectious nature, due to systemic infection with MuLiflandii, and therefore should be part of a differential list that includes other mycobacteria as potential etiologies. In our experience with the current study, we have found that parenchymal abdominal organs can be collected in situ for PCR or bacterial culture with good results, due to the small nature of this species. However, it is important to avoid collection of the gastrointestinal tract, or inadvertent contamination of tissues with gut contents, as gram-negative gut bacteria will outgrow and overtake mycobacteria in traditional media, leading to misleading results and misdiagnosis. The most reliable, routine method of detection of mycobacteria is by PCR and acid-fast staining in histological sections. However, speciation is problematic, and interpretation of PCR results may prove difficult with current commonly offered commercial tests. Therefore, the most definitive way to detect and speciate MuLiflandii is through mycobacterial culture and sequencing of virulence factors, laboratory techniques that are not available on a routine basis. These findings may support the interest of future work to enhance commercially available tests for proper speciation, especially as some mycobacterial species show clear zoonotic potential while the zoonotic potential of MuLiflandii remains unknown.
Epizootic M. ulcerans infections resulting in high mortality have been reported in research anuran colonies.35,38 Given the eventual reclassification of the organism, it is possible that outbreaks may have actually been caused by MuLiflandii rather than M. ulcerans. In one report involving Xenopus tropicalis, infection began with the formation of cutaneous lesions in a few individuals, which progressively spread between tanks. Depopulation was performed, but additional animals experienced cutaneous lesions and generalized septicemia a year later, 38 suggesting environmental contamination and persistence of the organism.
It has been suggested that MuLiflandii is the product of an intermediate degree of reductive evolution between the Agy99 strain of M. ulcerans and M strain of M. marinum. Like M. pseudoshottsii and other lineages of M. ulcerans, MuLiflandii carries the pMUM plasmid and produces mycolactone, a polyketide toxin that is key to the pathogenesis of M. ulcerans. Pathogenic human isolates of M. ulcerans produce mycolactone A/B, C, and D, 31 whereas MuLiflandii produces mycolactone E.2,22 Some authors have suggested that this bacterium could be niche adapted as a frog pathogen, fulfilling environmental pressures and giving the ecotype specification, “ecovar.” 36 Comparisons have demonstrated at least 3 discrete lineages of mycolactone-producing mycobacteria, with the frog pathogen MuLiflandii stemming from M. ulcerans isolates originating from Africa and Australia. 36 To the author’s knowledge, the zoonotic potential of MuLiflandii is currently unknown, and thus speciation is critical as M. ulcerans is associated with direct human disease.
Human infection with Mycobacterium ulcerans is associated with the formation of Buruli ulcers, a chronic, ulcerative cutaneous lesion characterized by toxin-mediated necrosis of adipose-rich subcutaneous tissue. Unrecognized or incompletely treated Buruli ulcers can lead to permanent location-dependent cosmetic or functional disabilities due to tissue destruction, scarring, and contracture formation. 42 M. ulcerans has been recognized by the World Health Organization as a potential pathogen of concern in humans. 1 It is endemic in at least 33 countries, spans various ecosystems and climates (mostly tropical), has been identified in aquatic insects, and has been shown to infect other species experimentally. 17 However, both the reservoir and mode of transmission between environmental M. ulcerans and people are currently unknown. There is evidence that mosquitoes may transmit the disease in southern Australia, suggesting a potential vector to humans.20,42
With regard to Mycobacterium marinum infection in humans, this organism is associated with focal cutaneous granulomas arising from the site of exposure. These lesions are sometimes colloquially referred to as “fish-tank granulomas” as transmission is classically associated with exposure of abraded skin to contaminated fish or water. Cutaneous manifestations of disease are described as focal papulonodular, verrucose, and/or ulcerated granulomas; however, dissemination of the bacteria through the lymphatic system has also been reported.3,14,37 Similarly, infection in leopard frogs with M. marinum has been associated with noncaseous granulomas when immunosuppressed. 33 While MuLiflandii has not yet been implicated in human disease or definitive spread between animals and humans demonstrated, the potential for human infection or zoonotic spread, particularly in immunocompromised individuals, should be considered given its similarities to other mycobacteria, such as M. ulcerans or marinum.
Preliminary PCR testing of coelomic swabs from affected animals was positive for M. marinum. However, subsequent PCR speciation at a separate laboratory indicated the presence of M. ulcerans (later confirmed by sequence analysis as MuLiflandii) and was repeatedly negative for M. marinum. Several possibilities were considered for this discrepancy. First, commercially available PCR assays often cannot differentiate between M. marinum and M. ulcerans or MuLiflandii. In addition, commercially available spectral databases do not typically include spectra for MuLiflandii and may only include spectra from human M. ulcerans isolates or M. marinum. Clusters of mycobacterial species are phylogenetically closely related, including M. marinum, M. ulcerans, M. shottsii, and M. pseudoshottsii. Some authors have suggested that all mycolactone-producing mycobacteria (M. ulcerans, MuLiflandii, M. pseudoshottsii, and M. marinum DL240490) should be categorized as M. ulcerans given the similarities, including bacteria previously called M. “liflandii.” 26 However, this potentially oversimplifies the broad host and environmental adaptations demonstrated by mycolactone-producing mycobacteria, which is of particular concern for biosecurity within laboratory settings which may house multiple aquatic species in contact with people. Because the zoonotic potential of MuLiflandii is unknown and most commercial tests cannot currently differentiate between similar species, further study of these organisms is needed along with development of new diagnostic tools. In our hands, we found that molecular evaluation of coelomic swabs and histology, particularly of coelomic fat, appeared to be the most efficacious diagnostics currently available.
General aerobic bacterial culture isolated a variety of commensal and opportunistic pathogens, including Chryseobacterium spp., Aeromonas spp., Lysobacter spp., and Paenibacillus spp. Chryseobacterium spp. specifically, C. indologenes and C. meningosepticum, have been associated with disseminated disease in aquatic laboratory anurans, including the African clawed frog (Xenopus laevis). 23 Aeromonas hydrophilia (among others) has been implicated in opportunistic bacterial dermatosepticemia (“red leg”). These commensal and opportunistic bacteria are present ubiquitously in the aquatic environment, and are typically normal inhabitants of amphibian skin and gastrointestinal systems.12,23 In addition, bacilli within areas of inflammation and necrosis were acid-fast positive, gram-positive, and Grocott methenamine silver-positive, consistent with the morphology and histochemical staining pattern of mycobacteria.5,9,24,38 As culture results for these organisms were inconsistent between individuals, often only reported as being few organisms, and numerous acid-fast bacteria were demonstrated within the lesions, the presence of these organisms on bacterial culture was considered incidental and unrelated to the cause of morbidity and mortality in these animals.
Environmental sampling indicated that detectable amounts of nucleic acids associated with MuLiflandii were present in the environment and harbored within the aquaria filter during infection. While buccal swabs from symptomatic and asymptomatic animals were consistently negative for mycobacterial agents, coelomic swabs from symptomatic frogs were consistently positive for MuLiflandii by real-time PCR. Detritus from tanks harboring symptomatic animals was also positive for MuLiflandii by real-time PCR, as well as M. chelonae and M. gordonae, illustrating the presence of multiple mycobacterial species as part of the normal milieu of the tank environment. These observations suggest that detritus, filter biofilm, and coelomic swabs may be effective ways to test for the presence of MuLiflandii.
Environmental sampling has been shown to be an effective way to screen for pathogenic mycobacteria in zebrafish, even at low numbers, 6 and environmental samples have been used to detect other pathogens in anurans. 8 Ideally, environmental screening would be an appropriate colony health surveillance method, as it reduces animal usage. MuLiflandii was detectable by PCR in environmental samples, suggesting the potential for simple and effective surveillance. Nucleic acid detection by PCR is preferable for demonstrating mycobacteria over other methods, such as bacterial culture, because culture of mycobacteria is notoriously difficult, often requiring specialized culture media and lengthy incubation periods. Furthermore, gastrointestinal bacteria and environmental commensals may interfere with the growth of mycobacterial species in culture, so aseptic technique during collection of culture material is critical to an accurate diagnosis.
In this study, the examined animals were all clinically symptomatic, moribund, or found deceased. Clinically unaffected frogs were not examined, so the possibility of the presence of asymptomatic carriers as persistent sources of infection in this colony is unknown. However, following the outbreak of morbidity and mortality in this colony, there have been no subsequent reported symptomatic cases. This suggests that the remaining animals have become resistant to infection, are subclinical carriers, or have cleared the infection completely.
These animals were acquired as adults from a biologic supplier with an unknown pathogen exposure and disease status history prior to acquisition. Acquisition of frogs from multiple vendors is common and can be a source of asymptomatic carriers of disease or the spread of pathogens among laboratory facilities. Environmental testing, quarantine status, and clinical progression suggest that these animals likely harbored the bacteria and were asymptomatic upon arrival, with eventual dissemination and multisystemic effects occurring late in the course of disease.
Mycobacteria are often found incidentally in asymptomatic laboratory fish and anurans. The impact of such infections is potentially confounding to research data and provides a nidus of infection for susceptible animals. Therefore, proper identification and eradication must be performed to prevent disease outbreaks, maintain proper animal welfare, and ensure and protect quality research data. Previous reports have utilized sodium hypochlorite (bleach) for disinfection following disease outbreaks, with variable effectiveness at controlling subsequent infections in other colonies at later dates. 38 This might suggest that bleach is potentially adequate in environmental decontamination, while asymptomatic carriers might propagate ongoing disease. In addition, it has been suggested that MuLiflandii is niche-adapted and has the potential to rapidly evolve, thus potentially impacting the efficacy of environmental decontamination. 36 Furthermore, vertical transmission of Mycobacterium spp. has been reported in viviparous fish. 7 Similarly, mycobacteria have been noted on the chorion of zebrafish (Danio rerio), and thus sodium hypochlorite surface disinfection is routinely used to limit transmission. 39 This and previous studies have demonstrated ovarian involvement in mycobacterial infection of anurans. 35 In other anurans, such as Xenopus laevis, surface disinfection protocols have been explored, notedly using ethanol. 40 However, to the author’s knowledge surface disinfection is not routine in Hymenochirus spp., leaving this as a potential conduit for colony infection. Further studies on the vertical transmission of Mycobacterium spp. and disinfection for eradication of these organisms are warranted.
There is recent concern for a progressive decline in wild frog populations globally. While multifactorial, emerging infectious diseases in susceptible species are an important contributor. For example, the spread of Batrachochytrium dendrobatidis, the cause of chytridiomycosis, from minimally affected populations in Asia to other continents has been potentially facilitated by humans. 32 Hymenochirus spp. and other anuran species are globally traded as pets and may harbor and spread diseases of great concern within the international frog trade. MuLiflandii has been associated with the international spread of disease to laboratory species such as Xenopus tropicalis; 35 therefore, the implications of this disease in a variety of anurans should not be understated, for the global pet trade or the research laboratory environment.
Conclusion
In this study, we report the first colony outbreak of septicemia caused by MuLiflandii in a research colony of Zaire dwarf clawed frogs. This species is common in the global pet trade and is increasingly used as a laboratory animal model; therefore, proper early identification and treatment of such pathogens are critical for overall animal health and welfare, colony and ecosystem maintenance, and preservation of research data. Furthermore, although the zoonotic potential of this pathogen is unknown, given its phylogenetic relationship with other human mycobacterial pathogens, proper identification of this organism is important for those individuals caring for these animals as pets or working with research colony animals, particularly if those individuals are immunocompromised.
Footnotes
Acknowledgements
The authors would like to thank the In Vivo Animal Core (IVAC), Unit for Laboratory Animal Medicine (ULAM) for pathology and histology expertise, Kyle Willenburg for technical assistance with PCR and sequence analysis, and Joann Young and Joe Myers for technical assistance with mycobacterial culture and mass spectrometry.
Author Contributions
The manuscript was written by M.B. and M.J.H. with contributions from the other authors; M.B. and J.K. performed the clinical evaluations; M.B. performed necropsies; M.J.H. performed histological evaluations; M.J.C. performed molecular analysis of specimens; M.J.H., J.K., and M.B. contributed to the experimental design.
Declaration of Conflicting Interests
The author(s) declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: M.J.C. is an employee of IDEXX BioAnalytics, a division of IDEXX Laboratories, Inc., a company that provides veterinary diagnostics.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
