Abstract
Hepatitis A virus (HAV) infects humans and nonhuman primates, typically causing an acute self-limited illness. Three HAV genotypes have been described so far for humans, and three genotypes have been described for nonhuman primates. We observed transiently elevated liver enzymes in Mauritius-origin laboratory-housed macaques in Germany and were not able to demonstrate an etiology including HAV by serology and polymerase chain reaction (PCR). HAV is a rare pathogen in cynomolgus macaques, and since all employees were routinely vaccinated against HAV, it was not a part of the routine vaccination and screening program. A deep sequencing approach identified a new HAV genotype (referred to as Simian_HAV_Macaca/Germany/Mue-1/2022) in blood samples from affected animals. This HAV was demonstrated by reverse transcription PCR in blood and liver and by in situ hybridization in liver, gall bladder, and septal ducts. A commercial vaccine was used to protect animals from liver enzyme elevation. The newly identified simian HAV genotype demonstrates 80% nucleotide sequence identity to other simian and human HAV genotypes. There was deeper divergence between Simian_HAV_Macaca/Germany/Mue-1/2022 and other previously described HAVs, including both human and simian viruses. In situ hybridization indicated persistence in the biliary epithelium up to 3 months after liver enzymes were elevated. Vaccination using a commercial vaccine against human HAV prevented reoccurrence of liver enzyme elevations. Because available assays for HAV did not detect this new HAV genotype, knowledge of its existence may ameliorate potential significant epidemiological and research implications in laboratories globally.
Cynomolgus macaques are commonly used for safety assessment of human biopharmaceuticals under development. Such nonclinical investigations typically involve the determination of indicators of liver injury, such as alanine aminotransferase (ALT) and glutamate dehydrogenase (GLDH). ALT and GLDH activities that exceeded the upper limit of the site-specific historical range were noted in vehicle-treated and naïve animals from a research facility in Germany.
Increases in ALT and GLDH activities in cynomolgus macaques can be caused by hepatotoxicity (such as inflicted by aflatoxins) but can also result from viral, bacterial, or parasitic infections. Relevant viral infections in cynomolgus macaques with a potential to cause hepatocellular necrosis are hepatitis A virus (HAV), B virus (macacine herpes virus 1), adenoviruses, and simian varicella virus (cercopithecine herpesvirus 9).12,28,26 Less likely viral etiologies include filoviruses (Marburg and Ebola Reston virus), Simian hemorrhagic fever virus, hepatitis B virus, hepatitis E virus, rhesus cytomegalovirus (macacine betaherpesvirus 3), and lymphocytic choriomeningitis virus.15,26 Hepatitis C virus and hepatitis D virus have not been associated with natural diseases in macaques.
Bacteria that can cause hepatic injury in cynomolgus monkeys include Mycobacterium tuberculosis and M. avium, Yersinia pseudotuberculosis and Y. enterocolitica, Francisella tularensis, and Burkholderia mallei.12,26 Parasites to be considered are trematodes, Cryptosporidium parvum, Toxoplasma gondii, Enterocytozoon bienusi, and Echinococcus. 26 Furthermore, elevated liver enzymes can be associated with hepatic amyloidosis or lipidosis in macaques.18,27
A root cause analysis was conducted with investigation into potential infectious and noninfectious causes. Because none of the investigations generated a positive result, we initiated a massive parallel sequencing approach, using serum samples from animals that were acutely affected by increased liver enzyme activities. This approach, also known as next-generation sequencing or deep sequencing, is a high-throughput sequencing approach that allows for the identification and characterization of low quantities of bacteria, fungi, parasites, protozoa, and viruses in a nontargeted and unbiased manner. 14 Unbiased massive parallel sequencing has been demonstrated to assist in detection and characterization of unidentified viral pathogens. 13 Combining massive parallel sequencing with sequence-independent single-primer amplification (SISPA) is especially useful when the nucleotide sequence of the desired molecule is both unknown and present in limited amounts. This approach yielded a previously unrecognized genotype of simian HAV.
HAV is a nonenveloped RNA virus in the family Picornaviridae. The host range for HAV is limited to man and several species of non-human primate. 4 The human HAV strains appear to have diverged from the simian HAV strains approximately 3,600 years ago.9,22 HAV is likely pervasive in many types of wild monkeys, but systematic data about its prevalence are lacking. HAV has been found in wild and captive monkeys of various origins, including Old World monkeys, New World monkeys, and great apes.6,7,8,17 Cynomolgus macaques that are experimentally infected with HAV demonstrate viral RNA in serum, saliva, and feces at 7 days postinoculation. 1 Biochemical and histological signs of infection are first seen at 15 days postinoculation, characterized by increased ALT and chronic periportal inflammation. 28 Infected animals demonstrate seroconversion with the appearance of anti-HAV IgM and IgG antibodies.16,28
This article presents the pattern of increased ALT and GLDH activities in captive cynomolgus macaques, presumably caused by this novel genotype of simian HAV, which evaded existing commercially available serologic and polymerase chain reaction (PCR) detection tests. We describe our root cause investigation and the subsequent discovery of this novel HAV. Finally, information is presented as to how to mitigate viral-induced effects in a laboratory setting.
Methods
Housing, Origin, and General Health of Animals
The facility in which this new genotype of HAV was discovered houses cynomolgus monkeys bred in Mauritius and Asia mainland. Prior to 2023, animals from Mauritius origin and animals from Asia origin were housed in separate rooms but without a strict separation of their housing areas. Animals are socially housed in groups of 2 to 20 in full compliance with the European Convention for the Protection of Vertebrate Animals used for Experimental and Other Scientific Purposes (ETS123) and Directive 2010/63/EU. Before entering the facility, all animals are screened for simian retro virus (serology and PCR), simian immunodeficiency virus (serology), and simian T-cell leukemia virus (serology). Animals from mainland Asia are additionally screened for macacine herpesvirus 1 (serology). In addition, all animals are screened for tuberculosis by a multiplexed fluorometric immunoassay and repeated tuberculin skin tests. Animals reported in this investigation were either allocated to a nonclinical safety study of a novel pharmaceutical or were colony animals (i.e. animals not yet allocated to an experiment). In case animals were allocated to a nonclinical safety study, only those animals treated with pharmaceutical excipients only (the nature of such vehicles cannot be disclosed, but none of them is known to cause liver enzyme elevations) were included in this report; animals administered a novel active pharmaceutical ingredient were excluded.
Biomarkers of Hepatocellular and Hepatobiliary Injury
Blood was collected into tubes without anticoagulant. The indicators for hepatocellular injury consisted of ALT, aspartate aminotransferase (AST), and GLDH activities, and the indicators for hepatobiliary injury consisted of gamma glutamyltransferase and alkaline phosphatase activities and total bilirubin concentration. These clinical chemistry endpoints were analyzed from serum by the Konelab system (Thermo Fisher Scientific Inc, Waltham, MA). The time points for blood collection for these endpoints varied between studies and animals. All data were entered into an electronic database (Pristima XD, Xybion, Princeton, NJ). Analysis and visualization were conducted using the TIBCO Spotfire platform (TIBCO Software Inc., Palo Alto, CA).
Histopathology
In 3 colony animals, a liver biopsy was collected for histopathologic evaluation immediately following the detection of increased ALT and GLDH activities. The liver tissue was fixed in 10% neutralbuffered formalin, paraffin blocks were prepared and sectioned, and sections were stained with hematoxylin and eosin. Microscopic examination was performed by a board-certified pathologist on at least 2 liver samples per animal.
Perl’s Prussian blue staining was conducted to demonstrate hemosiderin. Briefly, slides were dewaxed in xylene, immersed in industrial methylated spirit (IMS), rehydrated in 70% IMS, rinsed in deionized water, treated with Perls’ reagent for 10 minutes, washed in tap water for 10 minutes, stained in 1% neutral red for 2 minutes, washed in deionized water, dehydrated in 70% IMS and pure IMS, cleared in xylene, and mounted in synthetic resin.
Fouchet’s stain was conducted to demonstrate bile pigment. Briefly, slides were dewaxed in xylene, immersed in IMS, rehydrated in 70% IMS, rinsed in deionized water, stained in Fouchet’s reagent for 5 minutes, rinsed in deionized water, dehydrated in absolute alcohol, cleared in xylene, and mounted in synthetic resin.
Deep Sequencing
Blood samples from 6 animals that demonstrated elevated ALT and GLDH activities were used for massive parallel sequencing with a SISPA approach that was updated to sequence influenza viral RNA from clinical respiratory samples (Supplemental Figure S1).14,20
Sequencing read data were analyzed with a novel virus discovery pipeline. This pipeline uses scripts from the BBTools suite (http://sourceforge.net/projects/bbmap/) to mask host-specific reads (bbmask.sh), repair read pairing (repair.sh), and trim Illumina adapters (bbduk.sh). The pipeline then removes PhiX, human, and broad metagenomic reads with bbmap.sh, deduplicates reads with dedupe.sh, and merges paired and unpaired reads with bbmerge.sh. Merged trimmed reads are then assembled de novo with SPAdes (DOI: https://doi.org/10.1002/cpbi.102). After removing short and low-complexity contigs from the assembly (bbmask.sh and breformat.sh from BBtools), the pipeline classifies contigs with megablast, classifies unclassified contigs with blastn, and then outputs data for classified and unclassified contigs, DOI 1: https://doi.org/10.1186/1471-2105-10-421; DOI 2: https://doi.org/10.1016/S0022-2836,05)80360-2).
We visually inspected the SPAdes assemblies and generated a consensus sequence for the novel virus with Geneious Prime (Geneious Prime 2021.2.1). To account for excessive depths of coverage across many of the contigs, which may bias a consensus sequence, we used Geneious to reprocess NovaSeq reads by first trimming to remove adapters, low-quality sequences, and 19 bp from the 5' end of each sequence. Still in Geneious, we produced synthetic long reads by merging the trimmed reads with bbmerge and then mapped these reads to NCBI RefSeq # NC_001489, an HAV reference sequence, with the Geneious mapper. To ensure we detect novel virus reads, we required at least 100 bases of overlap between the reads and the reference sequence and tolerated up to 40% base mismatches. We then generated a prototype consensus sequence from the sample with the best coverage, corrected it by mapping reads from that sample onto its consensus using Multiple Sequence Comparison by Log-Expectation align, 10 transferred annotations from the human HAV NC_001489, trimmed low-quality ends, and translated the consensus sequence into amino acids. A GenBank formatted file of the 7490-bp consensus sequence (referred to as Simian_HAV_Macaca/Germany/Mue-1/2022), its annotations, and its translation is available at https://github.com/dholab/Mecklenburg-et-al-2022.
To compare Simian_HAV_Macaca/Germany/Mue-1/2022 with other Hepatovirus A sequences, we downloaded 19 complete genome sequences from Genbank (accessions KT819575, D00924, OK625565, MG181943, MH577309, MK829707, MN062164, MN832786, MN953474, MT181522, MT767252, NC_001489, NC_008250, DQ226541, EF067923, EF067924, EU140838, EU251188, and LC416594). We then used the Geneious Prime version 2022.2 Tree Builder to infer a Temura-Nei, neighbor-joining tree from Simian_HAV_Macaca/Germany/Mue-1/2022 together with the other Hepatovirus A sequences, with the possum HAV accession MG181943 as an outgroup. We also compared nucleotide and amino acid distances among this variety of HAVs with MUSCLE, version 3.8.425, with the default parameters implemented in Geneious Prime version 2022.2. Finally, we ran additional nucleotide sequence comparisons between Simian_HAV_Macaca/Germany/Mue-1/2022 and other simian HAV sequences, for which no complete genome is available: GenBank accession numbers M34084.1 (Simian HAV capsid protein VP1 strain PA21) and M59286.1 (HAV capsid proteins VP1, VP2, VP3, VP4, P2A of simian HAV strain Cy145).
Quantitative Reverse-Transcription PCR for the New Genotype of HAV
A quantitative reverse transcription PCR (qRT-PCR) method was developed to specifically detect Simian_HAV_Macaca/Germany/Mue-1/2022. Briefly, three sets of primers were designed: primer sets for qRT-PCR of Simian_HAV_Macaca/Germany/Mue-1/2022, one endpoint PCR primer pair that should detect human HAV and Simian_HAV_Macaca/Germany/Mue-1/2022 (and presumably some other divergent HAV), and one set of PCR primer pools that can be used for overlapping amplicon genome sequencing. Using the consensus sequence from the sample with the best coverage, we then generated primers in Geneious Prime. We set the primer tool to generate primers for between nucleotide 1000 and 6000 of the consensus to avoid ends, used the human mispriming library that is built into Geneious, and specified an optimal melting temperature of 60°C. To design endpoint PCR primers, we used a multiple sequence alignment method to align all sequences from the tree presented in Bennet et al. (2016), 6 and we generated a consensus sequence in Geneious Prime. 10 Finally, to design overlapping PCR primers for sequencing the whole genome of Simian_HAV_Macaca/Germany/Mue-1/2022, we used PrimalScheme with a target amplicon size of 1200 bases. 24 Primer pairs are summarized in the Supplemental Table S1.
For qRT-PCR, we extracted total RNA from serum samples (0.2 ml each) of 18 animals that showed elevated ALT and GDH activities and of 62 animals without liver enzyme elevation. In addition, we extracted total RNA from liver samples taken from 2 animals with elevated ALT and GLDH activities. RNA extraction methods and qRT-PCR assay conditions are summarized in the Supplemental Table S2.
In Situ Hybridization
Formalin-fixed and paraffin-embedded tissues from liver, gall bladder, and gastrointestinal tract were collected from 5 animals that had been assigned to the vehicle control group of a nonclinical study. Those animals had demonstrated liver enzyme elevations between 4 days and 11 weeks prior to necropsy.
For in situ hybridization, tissues were sectioned at 4–5 µm on Superfrost plus slides. In situ hybridization to detect Simian_HAV_Macaca/Germany/Mue-1/2022 as well as Mf-PPIB (positive control and tissue quality control) and DapB (negative control) genes was performed using reagents and equipment supplied by Advanced Cell Diagnostics (ACDBio, Hayward, CA) and Ventana Medical Systems (Roche, Tuscon, AZ). The in situ hybridization RNAscope probes were designed based on the Simian_HAV_Macaca/Germany/Mue-1/2022 sequence. The hybridization method followed protocols established by ACDBio and Ventana systems using a 3,3’-diaminobenzidine (DAB) chromogen. Briefly, sections were baked at 60°C for 60 minutes and used for hybridization. The deparaffinization and rehydration protocol was performed using a Sakura Tissue-Tek DR5 stainer with the following steps: 3 times xylene for 3 minutes each; 2 times 100% alcohol for 3 minutes; air dried for 5 minutes. Off-line manual pretreatment was conducted in 1X retrieval buffer at 98°C to 104°C for 15 minutes. Optimization was performed by first evaluating PPIB and DapB hybridization signals and subsequently using the same conditions for all slides. Following pretreatment, the slides were transferred to a Ventana Ultra autostainer to complete the in situ hybridization procedure including protease pretreatment; hybridization at 43°C for 2 hours followed by amplification; and detection with DAB chromogen and hematoxylin counterstain.
Vaccination
After the discovery of Simian_HAV_Macaca/Germany/Mue-1/2022, a vaccination program was initiated at the housing facility. For vaccination, a commercial vaccine against the inactivated human HAV HM175 strain, genotype IB (Havrix 720 Kinder, GSK) was used. Every newly arriving animal was vaccinated (0.5 ml per animal administered intramuscular) once within 14 days of arrival.
Results
Increased ALT and GLDH Activities
ALT and GLDH activities, which exceeded the upper limit of the site-specific historical range (range based on 2.5 and 97.5 percentiles for ALT males: 22–128 U/L based on data from 626 animals; ALT females: 22–160 U/L based on data from 532 animals; GLDH males: 7–64 U/L based on 555 animals; GLDH females: 8–59 U/L based on 460 animals), were identified in serum collected from Mauritius-origin cynomolgus macaques in the predose phase or from vehicle-treated control animals on nonclinical safety studies conducted between October 2020 and October 2021. ALT and GLDH activities in the affected animals were increased up to 11-fold (Fig. 1). These increases in ALT/GLDH activities were transient and only occurred once in any affected animal despite longitudinal monitoring. Increased ALT and GLDH activities did not correlate with clinical observations or macroscopic or microscopic findings in the liver at terminal necropsy, were infrequently associated with concurrently increased AST activity (only noted in 6% of the affected animals), and were unaccompanied by increases in indicators of hepatobiliary injury. Furthermore, no similarly increased ALT and GLDH activities were noted in cynomolgus monkeys of Asia mainland origin in studies concurrently conducted at the facility.
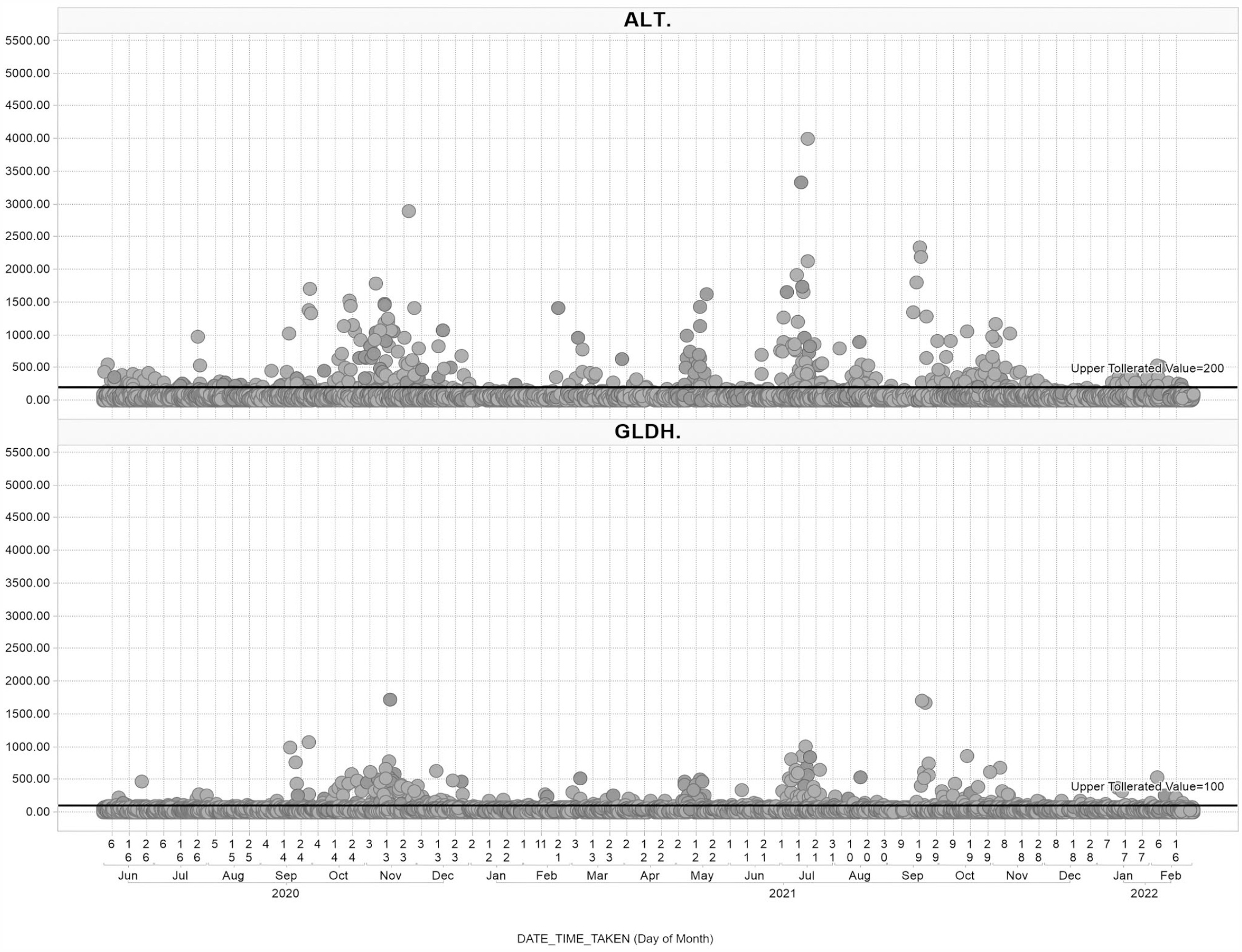
Scatter plot of alanine aminotransferase (ALT, U/l) and glutamate dehydrogenase (GLDH, U/l) values over time in 11915 (ALT) and 11436 (GLDH) blood samples from 3461 (ALT) and 3218 (GLDH) Mauritius-origin cynomolgus monkeys assigned to nonclinical safety studies at the test facility between June 2020 and February 2022. Animals were administered various test articles and vehicles. The upper tolerated value is shown by a black line (200 U/l for ALT and 100 U/l for GLDH). Each dot represents a measurement recorded by calendar date.
Screening of liver enzymes was initiated from Mauritius-origin animals that were not yet assigned to a study. Out of 3956 blood samples collected between December 2020 and March 2022, 144 samples (3.6%) demonstrated increased ALT and GLDH activities (Supplemental Figure S2). These were identified between March and November 2021. Sequential monitoring of the ALT and GLDH activities in affected animals indicated that the increased enzyme activities were transient and generally returned to values within the historical range within 6 weeks (Supplemental Figure S3).
Diagnostic Investigations
Root cause analysis was conducted with investigation into potential infectious causes including HAV, hepatitis B virus, hepatitis C virus, hepatitis D virus, and lymphocytic choriomeningitis virus. In addition, investigations were conducted for noninfectious causes such as aflatoxins and intoxication by detergents or leachables from toys that were provided for behavioral enrichment. Animals had not been moved or handled in any different way, and the housing environment had not been changed. A site-wide chart highlighting transfers from animals across the facility did not indicate any relationship to elevated liver enzymes. It was noted, though, that animals showed a liver enzyme elevation typically in the first 8 weeks after they had been introduced to the facility. Because none of the investigations generated a positive result, we initiated a massive parallel sequencing approach, using serum samples from animals that were acutely affected by increased liver enzyme activities. This approach yielded a previously unrecognized genotype of simian HAV.
Histopathology
To verify any histomorphological evidence of chronic periportal inflammation as described for HAV in cynomolgus macaques, 21 liver tissue was sampled from 3 colony animals shortly after the detection of increased ALT and GLDH activities. In all animals, microscopy was characterized by minimal to slight periportal mononuclear cell infiltrates (Fig. 2a) like what is occasionally seen in vehicle-treated macaques. In addition, very few degenerating hepatocytes or hepatocellular single-cell necrosis was seen in 2 animals (Fig. 2b), and minimal pigment in macrophages or Kupffer cells (Perl’s positive; Fouchet negative; data not shown). Minimal hepatocellular vacuolation was seen in 1 case.
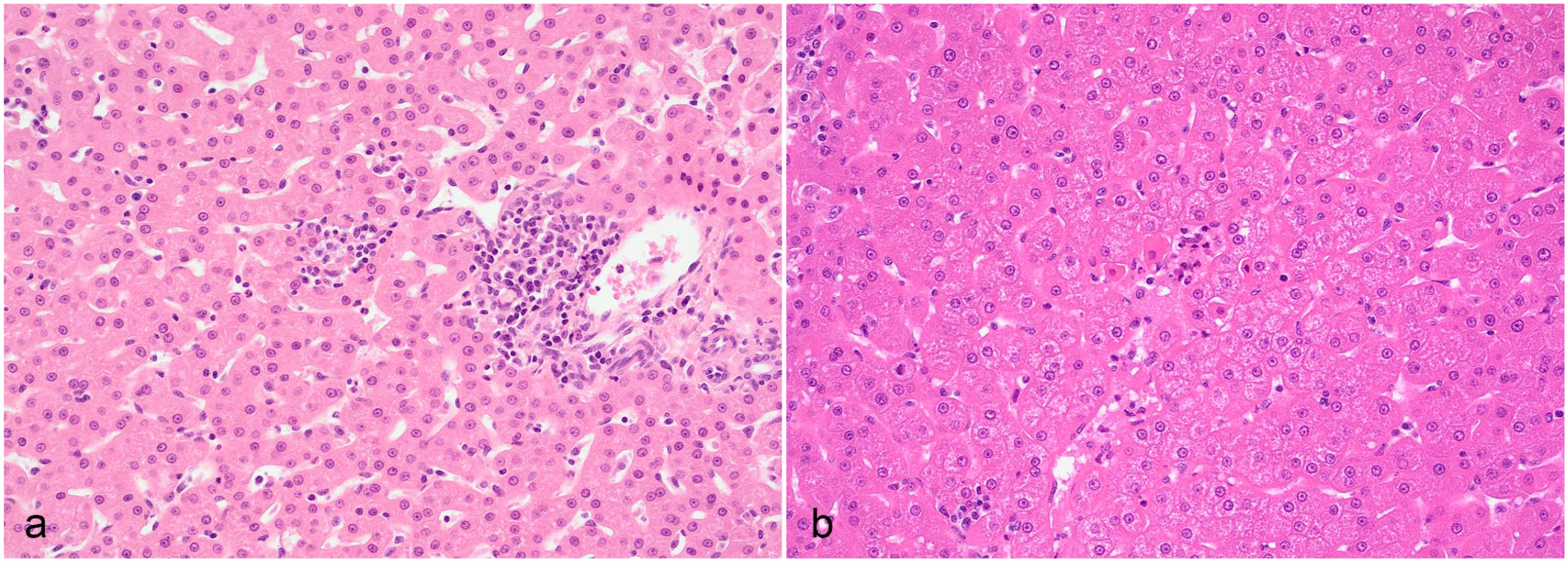
Histology of liver from a Mauritius-origin cynomolgus monkey with increased alanine aminotransferase and glutamate dehydrogenase enzyme activity. Hematoxylin and eosin. (a) Minimal mononuclear cell infiltrates in the periportal region with very few degenerating hepatocytes. (b) Single-cell necrosis of hepatocytes.
Deep Sequencing
Massive parallel sequencing yielded sequencing reads that matched a hepatitis A-like virus in 5 out of 6 samples. Consensus sequences of the viruses were virtually identical between samples. The identified nucleotide sequence of Simian_HAV_Macaca/Germany/Mue-1/2022 consists of 7490 bp, with only 80% identity to Simian HAV Cercopithecus/Kenya/AGM-27/1985 and to all human HAV sequences (Supplemental Figure S4). When only nucleotides presumably encoding for the viral proteins (VP) 1 and 2A (i.e. nucleotides 725–3243) of Simian_HAV_Macaca/Germany/Mue-1/2022 were used for comparison, 81.41% identity was found for the simian HAV strain PA21, and 85.99% identity was found for the simian HAV strain Cy-145 (Supplemental Table S2).
All samples with Simian_HAV_Macaca/Germany/Mue-1/2022 clustered closely on the same phylogenetic branch, consistent with each consensus sequence representing a single infecting lineage. The lineage was sister to all downloaded human and simian HAVs, indicating deeper divergence between Simian_HAV_Macaca/Germany/Mue-1/2022 and other previously described primate HAVs, including both human and simian viruses. Still, Simian_HAV_Macaca/Germany/Mue-1/2022 was in the same clade as other primate HAVs when duck and possum HAVs were included, both of which were in their own branches (Fig. 3).
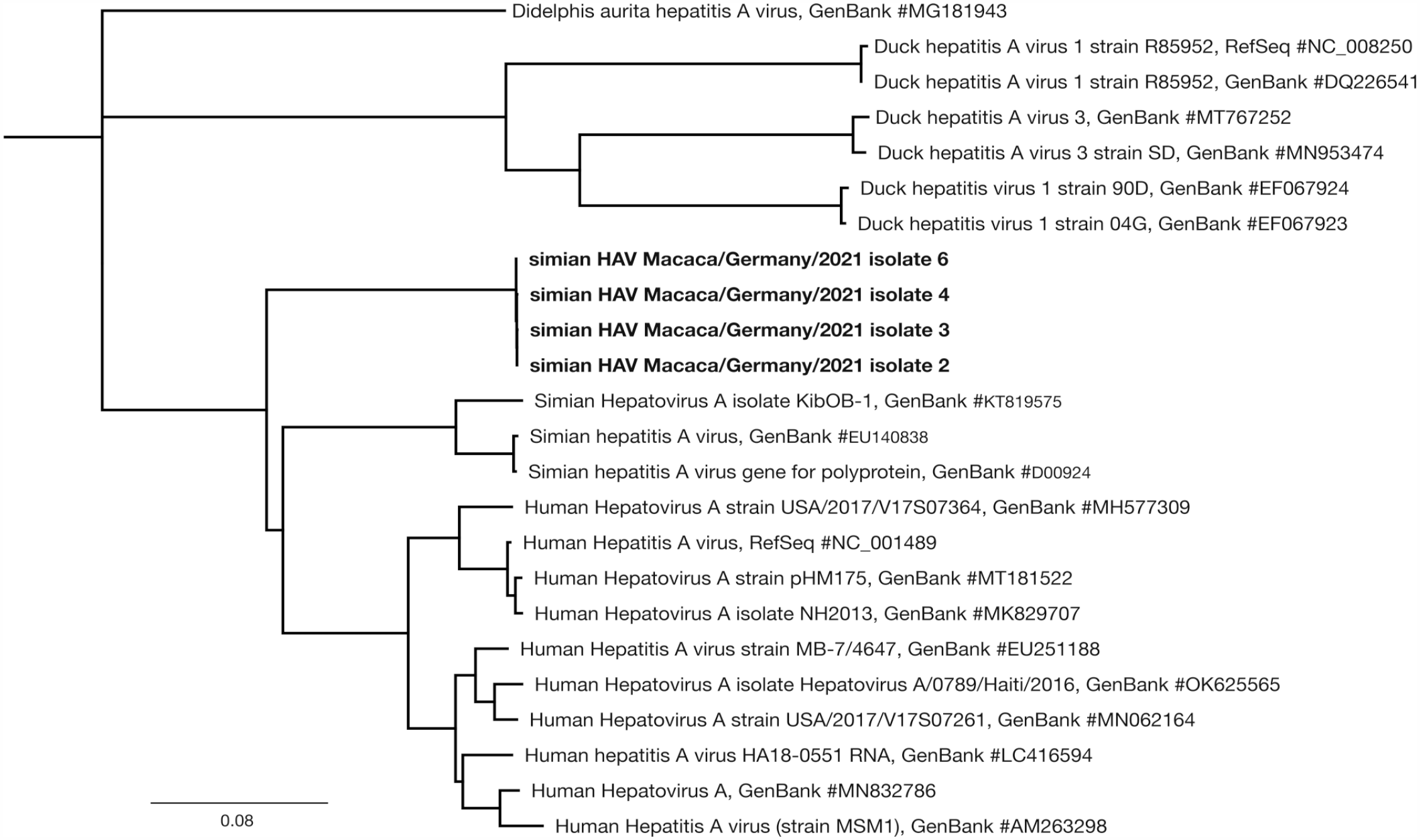
Phylogeny of 4 Simian_HAV_Macaca/Germany/Mue-1/2022 together with other hepatovirus A sequences, with the possum hepatitis A virus accession MG181943 as an outgroup.
At predicted amino acid level, the sequence of Simian_HAV_Macaca/Germany/Mue-1/2022 has 92.0% identity with the HAV polyprotein derived from olive baboons (GenBank accession no. ANJ65975.1), 91.97% identity with the simian HAV polyprotein derived from a captive rhesus monkey (GenBank accession no. ABX55994.1), and approximately 90% identity with numerous HAV polyprotein sequences from other HAV genotypes obtained from human patients (Supplemental Figure S5). A comparison of the predicted amino acid sequence specifically for VP1 (nucleotides 2203–3099) revealed a 98.66% identity with partial HAV polyprotein from HAV genotype IV (GenBank accession no. AAA45473.1) (Supplemental Figure S6). 23
Similar patterns were reflected in the nucleotide and amino acid distance matrices computed with MUSCLE. Simian_HAV_Macaca/Germany/Mue-1/2022 was a median of 80% nucleotide identity to previously described simian and human HAV sequences. Those previously described simian and human HAVs were more similar to each other than they were to Simian_HAV_Macaca/Germany/Mue-1/2022. The median nucleotide identity between previously described simian HAVs was 94.24% identical to other simian HAVs, while the median identity between the human HAVs was 91.01%. A distance matrix of translated open reading frames showed similar identities.
Diagnostic PCR
Because conventional serology and PCR assays for human HAV had not detected the new HAV genotype, we developed a quantitative RT-PCR method specific for Simian_HAV_Macaca/Germany/Mue-1/2022. This method was used on serum samples from 80 animals and on liver tissue samples from 2 animals. The method detected Simian_HAV_Macaca/Germany/Mue-1/2022 in all 18 serum samples and in both liver tissue samples from animals with increased ALT and GLDH activities. The 62 serum samples from animals without increased ALT and GLDH activities were negative.
In Situ Hybridization
In situ hybridization detected Simian_HAV_Macaca/Germany/Mue-1/2022 in liver tissues from 5 out of 5 animals. Signal was present to varying degrees within hepatocytes, sinusoidal cells, and biliary epithelium (Fig. 4). In addition, the Simian_HAV_Macaca/Germany/Mue-1/2022 signal was observed in the gut-associated lymphoid tissue of 1 animal with a high hepatic viral load. In 3 animals euthanized 4 days after increases in ALT and GLDH were detected, signal was observed in hepatocytes, sinusoidal lining cells including Kupffer cells, and biliary epithelial cells in the liver. The in situ hybridization signal in the liver correlated with the ALT elevation. The strongest signal was seen in the animal where ALT was elevated 13 times above baseline, followed by the animal with 11-fold elevation, and the animal with 6-fold elevation.
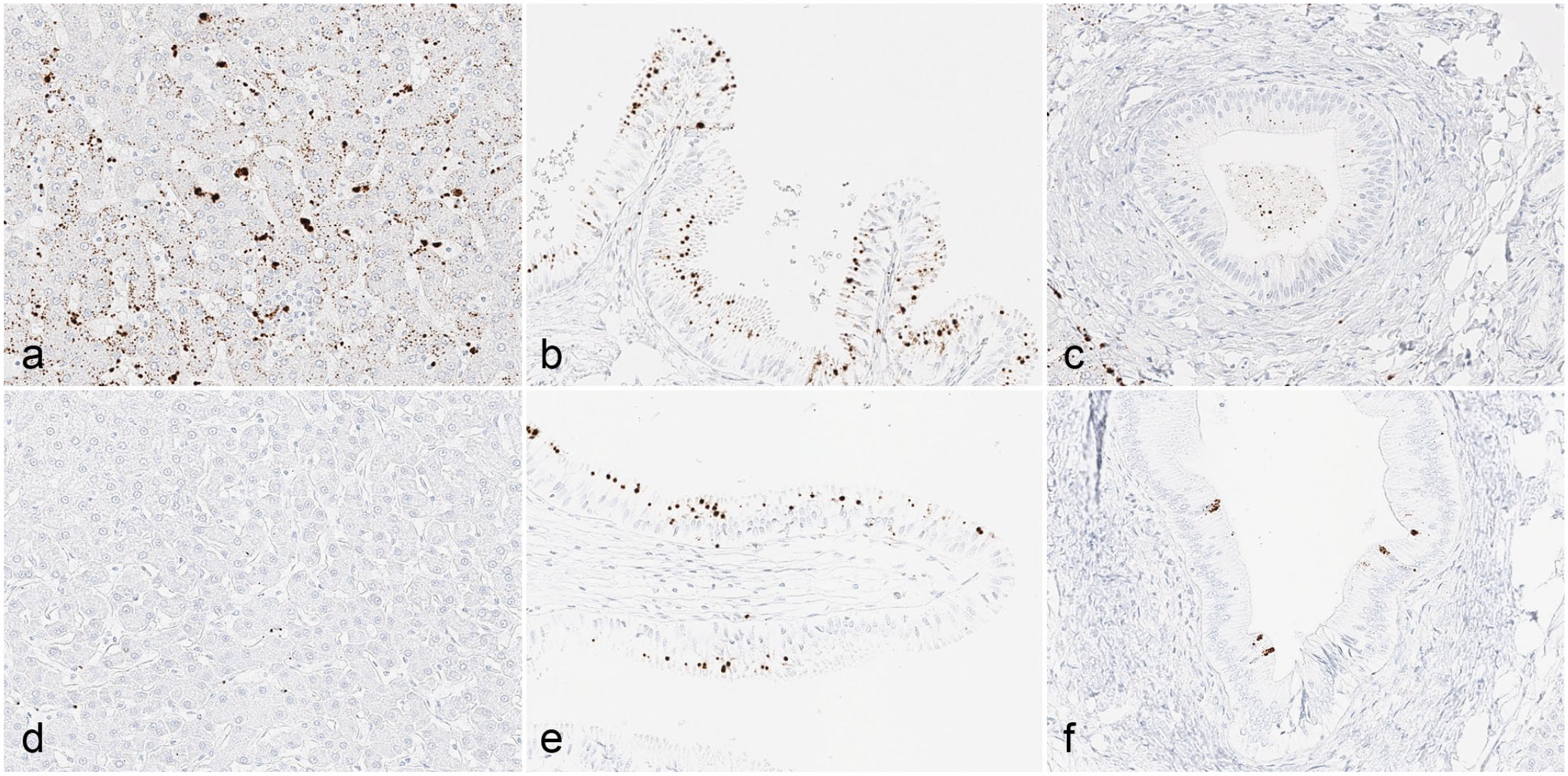
In situ hybridization for Simian_HAV_Macaca/Germany/Mue-1/2022 in Mauritius-origin cynomolgus monkey tissues from (a, d) the liver, (b, e) gall bladder, and (c, f) hepatic septal ducts from animals euthanized 4 days (a, b, c) and 11 weeks (d, e, f) after the detection of elevations in alanine aminotransferase and glutamate dehydrogenase.
In 2 animals euthanized 11 weeks after increases in ALT and GDLH activities were detected, signal was largely absent from the liver and not present in hepatocytes. Despite an absence of the Simian_HAV_Macaca/Germany/Mue-1/2022 signal in hepatocytes of these animals, signal was observed in gall bladder epithelium, suggesting the biliary epithelium may represent a reservoir of infection and shedding after resolution of hepatocellular infection.
Vaccination
After identification of Simian_HAV_Macaca/Germany/Mue-1/2022, we initiated a vaccination program using a commercial vaccine against the human HAV HM175 strain. All animals newly arriving to the test facility (both of Mauritius and Asia mainland origin) were vaccinated within 14 days of arrival. In addition, all colony animals housed at the facility but not yet allocated to an experiment were also vaccinated. The vaccination program started in August 2021. Since its initiation, newly arriving animals have been monitored with respect to liver enzyme activities. In all vaccinated animals, ALT and GLDH activities have remained within the reference range (Supplemental Figure S2).
Discussion
HAV is a known cause of hepatitis and elevated liver enzyme activities in captive cynomolgus monkeys. 28 However, disease outbreaks in captive monkeys are rarely reported. At the facility where this new genotype of HAV was discovered, no such disease outbreak had occurred for more than 40 years. Animals entering the facility were not routinely screened for HAV, since HAV was not perceived as a commonly encountered pathogen and since all employees were routinely vaccinated against human HAV.
When increased ALT and GLDH activities were identified, HAV was the main differential diagnosis. Appropriate diagnostic tests (PCR and serology) were initiated but ultimately failed to detect Simian_HAV_Macaca/Germany/Mue-1/2022. Despite broad-ranging diagnostic investigations, no etiology could be identified. Therefore, a massive parallel sequencing approach was employed. This approach allows identification and characterization of bacteria, fungi, parasites, protozoa, and viruses without prior knowledge of a specific pathogen.
The approach revealed a previously unreported HAV genotype with less than 80% whole-genome nucleotide sequence identity to any previously reported human or simian HAV. Highest nucleotide sequence identity was detected for nucleotides 725–3243, encoding for HAV capsid proteins VP1, VP2, VP3, VP4, and P2A, from Simian HAV Macaca/Philippines/CY-145/1988, which has been classified as a genotype IV hepatitis virus. 25 This sequence was derived from a cynomolgus macaque that was imported into the United States from the Philippines and that had serological as well as histopathological evidence of hepatitis. 23 The authors did not attempt to isolate and further characterize this new HAV in cell culture.
HAV is a small nonenveloped ancient virus with a long evolutionary history that is distinct from other picornaviruses. The HAV genome is a single-stranded, positive-sense RNA molecule, approximately 7500 nucleotides in length, that serves directly as messenger RNA for translation of HAV-encoded proteins. 21 Like other picornaviruses, HAV shows a high mutation rate, resulting in the emergence of a genetically diverse cloud of mutants. 9 Three HAV genotypes have been described for humans (genotypes I–III), and three genotypes have been described for nonhuman primates (genotypes IV–VI). Opposed to the human HAV genotypes, those in nonhuman primates are not well characterized. Genotype IV was described in a cynomolgus macaque from the Philippines. 23 Genotype V was first described in an African vervet monkey from Kenya and was also found in a rhesus monkey from India and an olive baboon from Uganda.3,6,5,29 Genotype VI was found in a cynomolgus macaque from Indonesia. 2 In addition, genotype IIIA HAV was discovered in an owl monkey from Panama. 7
The HAV genome contains an open reading frame encoding a large polypeptide with multiple cleavage sites. It is generally agreed that the polyprotein is processed to 4 structural capsid proteins (VP1–VP4) and 7 nonstructural proteins (2ABC, 3ABCD). Processing is performed by the proteinase encoded in the 3C region. The host humoral response against the capsid protein represents the primary weapon to control the infection. Capsid proteins VP1 and VP3 are major antibody-binding sites.9,11 While there is good evidence that most human strains of HAV are closely related antigenically, simian strains have significant antigenic differences from human HAV strains. 19 The newly discovered Simian_HAV_Macaca/Germany/Mue-1/2022 is sufficiently different from known human and simian HAVs, such that diagnostic methods that are directed against human HAV would not detect this new simian genotype. This was confirmed by directly comparing the genomic sequence of Simian_HAV_Macaca/Germany/Mue-1/2022 against the PCR primer sequences from the commercially available PCR test used to detect human HAV.
Despite the difference between Simian_HAV_Macaca/Germany/Mue-1/2022 and human HAV, we successfully vaccinated animals with a commercial vaccine against inactivated human HAV genotype IB. This vaccination program was able to prevent further occurrence of increased ALT and GLDH activities.
It is interesting to note that Simian_HAV_Macaca/Germany/Mue-1/2022 only caused increased liver enzyme activities in Mauritius-origin cynomolgus macaques. It is possible that Mauritius-origin monkeys are more sensitive to factors causing liver enzyme elevation, as Mauritius-origin cynomolgus macaques generally show a 20%–40% higher liver enzyme activity than Asia mainland-origin animals. 30 It is also possible that Simian_HAV_Macaca/Germany/Mue-1/2022 circulates in cynomolgus monkeys from Asia generating immunity of Asia-origin animals opposed to HAV-naïve Mauritius-origin macaques. Further investigations would be needed to determine the epidemiology of this new genotype HAV in cynomolgus monkeys.
We conclude that Simian_HAV_Macaca/Germany/Mue-1/2022 is present in cynomolgus macaques and that it poses a potential risk particularly for Mauritius-origin captive monkeys that are intended for use in scientific purposes. To prevent infection and associated elevations in liver enzyme activity, it is recommended to vaccinate animals. A commercial vaccine against human HAV has demonstrated efficacy in this matter.
Supplemental Material
sj-pdf-1-vet-10.1177_03009858231209691 – Supplemental material for A new genotype of hepatitis A virus causing transient liver enzyme elevations in Mauritius-origin laboratory-housed Macaca fascicularis
Supplemental material, sj-pdf-1-vet-10.1177_03009858231209691 for A new genotype of hepatitis A virus causing transient liver enzyme elevations in Mauritius-origin laboratory-housed Macaca fascicularis by Lars Mecklenburg, Rebecca Ducore, Molly Boyle, Andrew Newell, Laura Boone, Joerg Luft, Annette Romeike, Ann-Kathrin Haverkamp, Keith Mansfield, Kelley A. Penraat, JJ Baczenas, Nick Minor, Shelby L. O’Connor and David H. O’Connor in Veterinary Pathology
Footnotes
Acknowledgements
The authors acknowledge and highly appreciate the work of Dr. Artur Kaul and Dr. Annette Dammert from German Primate Center in Goettingen, Germany, for establishing and conducting qRT-PCR assays for Simian_HAV_Macaca/Germany/Mue-1/2022. They also acknowledge the contribution from Jakob Fritz who assisted in data analysis and from David Dahlhaus who formerly worked for Labcorp and participated in the root cause analysis.
Authors’ Contribution
LM, RD, MB, KM, and DHO designed the experiments. LB, JL, KAP, and SLO contributed to the experimental design. AN performed data visualization. AR and A-KH performed histologic evaluations. JJB and NM performed genetic analysis. The manuscript was written by LM with contribution from the other authors.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
