Abstract
Caryospora-like organisms (CLOs) form a clade of at least 11 genotypes of related coccidia that can cause epizootic mortality in marine turtles. The biology, transmission, host species range, and host cell tropism of these organisms are still largely unknown. The goal of this study was to characterize the host cell tropism, pathologic and ultrastructural features, and phylogeny associated with the first report of a mortality event due to CLO in the freshwater red-eared slider turtle (Trachemys scripta elegans). Sudden mortalities within a clutch of captive-raised red-eared slider hatchlings (n = 8) were recorded, and deceased animals had severe segmental to diffuse, transmural, fibrinonecrotic enterocolitis and multifocal to coalescing hepatic necrosis, among other lesions associated with numerous intracytoplasmic developing stages of intralesional coccidia. Among the different developmental stages, merozoites were ultrastructurally characterized by an apical complex. A pan-apicomplexan polymerase chain reaction (PCR) yielded a 347 bp-amplicon matching the Schellackia/Caryospora-like clade with 99.1% identity to the US3 strain from green sea turtles (Chelonia mydas) and 99.1% identity to Schellackia sp. Isolate OC116. Surviving hatchlings were treated with toltrazuril sulfone (ponazuril) but were subsequently euthanized due to the risk of spreading the parasite to other chelonids in the collection. The ponazuril-treated hatchlings (n = 4) had mild proliferative anterior enteritis, with few intraepithelial coccidia in one hatchling confirmed as CLO by PCR. This is the first report of Caryospora-like coccidiosis in non-cheloniid turtles, highlighting the relevance of this disease as an emerging highly pathogenic intestinal and extra-intestinal form of coccidiosis of turtles with potential cross-species infectivity.
Caryospora-like organisms (CLOs) form a clade of important emerging pathogens of marine turtles that cause epizootic mortality events associated with enterocolitis. These coccidia were first identified in the 1970s as the cause of an epizootic of enterocolitis in a group of maricultured green turtles (Chelonia mydas) in the Caribbean, and the organism was subsequently named Caryospora cheloniae.22,31 Morphologically similar coccidia were associated with epizootic mass mortality events in wild green turtles off the Australian coasts in 1991 and 20146,13 as well as off the southeastern United States coasts in 2014 and 2015. 34 Since then, sporadic cases have been reported in green loggerhead (Caretta caretta) and Kemp’s ridley turtles (Lepidochelys kempii) in the southeastern U.S. Atlantic coast, Gulf of Mexico, Central Pacific region (Hawaii), and Northeastern Brazil.8,34
The type species of Caryospora within the order Eucoccidioridae, family Eimeriidae is C. simplex, which uses snakes in the genus Vipera as definitive hosts and can use rodents as intermediate hosts. 21 There are at least 81 morphologically described species in the genus, 10 but of these, only five named species have available DNA sequence data, and these do not include the type species. Molecular phylogeny has revealed that they are paraphyletic, with at least 4 distinct non-sibling clades; C. bigemina, Caryospora-like coccidia from marine turtles, C. ernsti, and C. neofalconis. C. neofalconis actually clusters in the family Sarcocystidae rather than Eimeriidae, and recent evidence suggests they should be transferred to the genus Eumonospora. 7 As it is not possible to say which of the remaining clades contains the type species, and if therefore truly Caryospora, the term CLOs is used to describe these marine turtle coccidia until this issue can be resolved. The marine turtle CLOs consist of at least 2 distinct clades. One is closely related to Schellackia, from snake hosts, contains the strains most strongly associated with epizootics, and has only previously been reported in marine turtles in the family Cheloniidae. The other clade has been found in both Dermochelyidae (leatherback sea turtles) and Cheloniidae. 34 At this time, 11 genotypes of related coccidian organisms have been identified in stranded marine turtles in the United States, and 2 genotypes have been identified in Australian marine turtles. 34 Some authors have suggested that the marine turtle CLO should be transferred to the nearest monophyletic genus, Schellackia, found in squamate erythrocytes; 9 however, the ecologic niches and host ranges of the marine turtle CLOs appear to be distinct from Schellackia, and without knowing the phylogenetic position of C. simplex, it is not possible to make this determination.
Stranded CLO-infected marine turtles in the Northern hemisphere and Brazil exhibit signs of severe dehydration such as sunken eyes and gaunt neck, shoulder, and prefemoral regions due to coccidian-associated enterocolitis. Lesions include watery intestinal contents and ulcerative to proliferative heterophilic enterocolitis with intraluminal fibrin casts and numerous intracytoplasmic and central luminal coccidia in different developmental stages. Oocysts can be identified by fecal exam and, when sporulated, typically have a single sporocyst with 8 sporozoites, a morphologic feature consistent with Carysopora. 34 In addition to the intestinal lesions, marine turtles in Australia exhibit abnormal neurologic behavior associated with meningoencephalitis. Thyroiditis, nephritis, and pneumonia associated with extra-intestinal, intracytoplasmic coccidia have also been identified.6,13 To our knowledge, CLOs have only been, thus far, reported in marine turtles. The aim of our study was to describe the epidemiological, host cell tropism, and pathologic and ultrastructural features of the first reported outbreak of systemic CLOs in freshwater, red-eared slider turtles (Trachemys scripta elegans).
Materials and Methods
Description of the Outbreak, Animals Included in the Study, and Clinical History
In June of 2021, 8 red-eared slider eggs were collected from the wild near Baton Rouge, Louisiana, incubated, and hatched on commercial vermiculite. The hatchlings were housed together in a 60-gallon tank containing 20 gallons of water with rock and driftwood as substrate and provided with UVB and heat support. The hatchlings were fed a commercial pellet diet with commercial frozen-thawed Atlantic silverside fish (Menidia menidia). In October 2021, 4 of 8 hatchlings (Table 1) were sequentially found dead over a 3-week period and were submitted for postmortem examination. The remaining hatchlings (n = 4) were treated with a 60 mg/kg loading dose of toltrazuril sulfone (ponazuril, Merial, Inc., Duluth, GA, USA) followed by 20 mg/kg dose every 4 days for 48 days. 30 In December 2021, one treated hatchling was found dead and was submitted for histological evaluation. In January 2022, the remaining treated hatchlings (n = 3) were euthanized due to concerns for disease transmission to other chelonians in the collection and were submitted for postmortem examination (Table 1).
Red-eared slider cases (n = 8) submitted to the Louisiana Animal Disease Diagnostic Laboratory and their gross findings, pan-apicomplexan 18S rRNA PCR and bacteriological culture results.
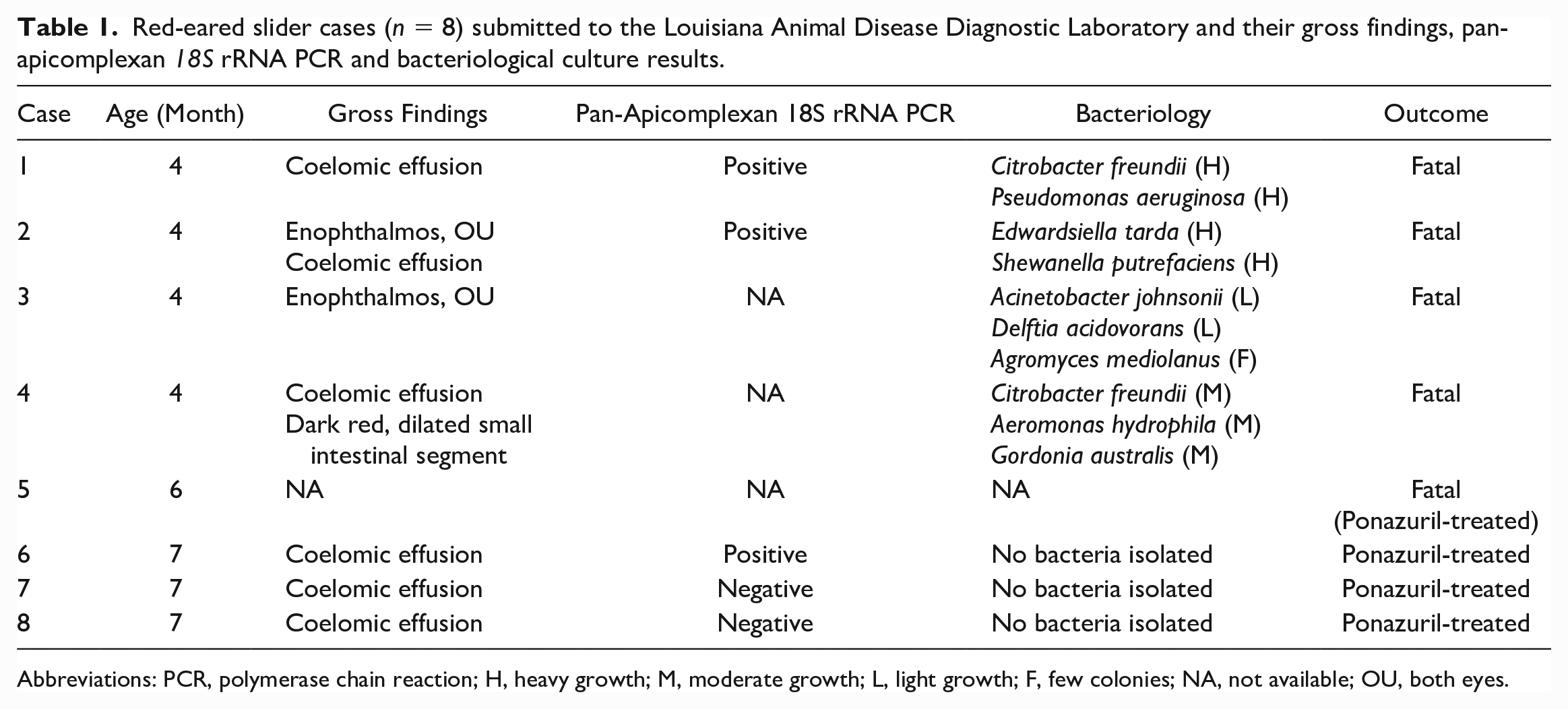
Abbreviations: PCR, polymerase chain reaction; H, heavy growth; M, moderate growth; L, light growth; F, few colonies; NA, not available; OU, both eyes.
Postmortem Examination and Sample Collection
Seven hatchlings (cases 1–4 and 6–8) were submitted fresh for full routine postmortem examinations. Coelomic swabs from all turtles were submitted for aerobic and Salmonella-specific bacteriological cultures (see below). Fresh samples of yolk sac remnants, lung, liver, and intestine were frozen at -80°C for ancillary testing, and the remaining specimens were fixed in 10% neutral buffered formalin. A segment of colon from case 4 was submitted for direct smear; however, no parasites were identified. Case 5 was submitted fixed in 10% neutral buffed formalin and, after routine decalcification, serial sections of the head and body were examined microscopically.
Histopathology
Sections of all major organs were routinely processed into paraffin blocks, sectioned at a 4 µm thickness, and stained with hematoxylin and eosin. The heads were decalcified in 15% formic acid until soft, and multiple serial transverse sections were obtained and similarly processed. Following standard methods, Gram, Fite-Faraco, and Steiner histochemical stains were performed on representative sections of liver, lung, and kidney from case 2. Based on lesion severity, the microscopic alterations were categorized as not present, minimal (<10% area affected), mild (10%–25% area affected), moderate (25%–50% area affected), marked (50%–75% area affected), or severe (>75% area affected).
Bacteriology
Coelomic swabs from all turtles were plated on commercial blood agar (tryptic soy agar with 5% sheep blood) and MacConkey agar plates (Remel, San Diego, CA, USA). Inoculated blood agar plates were aerobically incubated at 37°C with 5% CO2 for the first 24 h and without CO2 for the next 48 h. Inoculated MacConkey agar plates were incubated at 37°C for up to 72 h. Bacterial colonies were subsequently identified by matrix-assisted laser desorption/ionization—time of flight (MALDI-TOF) mass spectrometry (Biotyper Compass; Bruker). Salmonella-specific culture was performed following standard procedures.
Immunohistochemistry (IHC)
All tissues collected from cases 2 to 7 were evaluated by IHC using a cross-reactive rabbit polyclonal antiserum specific to Neospora caninum (anti-NC-1 tachyzoite). Tissues from case 1 were not subjected to IHC due to significant postmortem autolysis. Briefly, 4 μm sections of formalin-fixed, paraffin-embedded tissues were mounted on positively charged Superfrost® Plus slides (VWR, Radnor, PA, USA) and subjected to IHC using the automated BOND-RXm platform and the Polymer Refine Red Detection kit (Leica Biosystems, Buffalo Grove, IL, USA). Following automated deparaffinization, heat-induced epitope retrieval was performed using a ready-to-use ethylenediaminetetraacetic (EDTA)-based buffer (pH 9.0; Leica Biosystems) at 100°C for 20 minutes before incubation with anti-N. caninum antiserum diluted 1:1,000 in Primary Antibody Diluent (Leica Biosystems). Sections were then incubated with the primary antibody for 30 minutes at room temperature, followed by a polymer-labeled goat anti-rabbit IgG coupled with alkaline phosphatase (Leica Biosystems) for 30 minutes at room temperature. Fast Red was used as the chromogen (15 minutes), and counterstaining was performed with hematoxylin. Slides were mounted with a permanent mounting medium (Micromount®, Leica Biosystems). An N. caninum-infected bovine turbinate cell pellet 5 as well as brain from an aborted calf were used as positive controls, and tissues incubated without the primary antibody were used as negative controls.
Transmission Electron Microscopy
Liver and intestinal tissue samples from one deceased turtle (case 3) were transferred from 10% neutral-buffered formalin to a fixative composed of 2% glutaraldehyde in 0.1 M phosphate buffer, pH 7.25. Samples were fixed overnight. After washing several times in 0.1 M phosphate buffer, the tissues were post-fixed in 1% osmium tetroxide in phosphate buffer for 1.5 hours. The tissue samples were washed several times in deionized water and then placed in a 0.5% aqueous uranyl acetate enbloc for 1 hour. After several more washes in deionized water, the tissue samples were dehydrated in a series of ethanol solutions of increasing concentrations (30%, 50%, 75%, 95%, and 100%) and cleared in two changes of acetone and two changes of propylene oxide. The tissue samples were infiltrated with 2:1, 1:1, and 1:2 mixtures of propylene oxide and Mollenhauer’s Epon-Araldite plastic mixture 27 for 2 hours each; then two changes of 100% Epon-Araldite plastic for at least 2 hours each before embedding the tissues in flat embedding molds. The embedded samples were polymerized in a 70°C to -80°C oven overnight. 3 One micrometer sections from the polymerized blocks were obtained using a Reichert Ultracut S ultramicrotome. The sections were placed on glass slides and stained with 1% toluidine blue O in 1% sodium borate. The stained sections were evaluated, and areas of interest were chosen before trimming the corresponding block face for thin sectioning. Sixty nanometer sections were obtained and placed on 200-mesh copper grids. The grids were post stained with 2% aqueous uranyl acetate and Reynolds lead citrate. 32 Grids were viewed with a JEOL JEM-1011 transmission electron microscope at varying magnifications using an accelerating voltage of 80 KeV. Images were acquired using an AMT XR80M Wide-Angle Multi-Discipline Mid-Mount CCD Digital Camera with a resolution of 3296 x 2460 pixels.
Molecular Testing
Pooled fresh-frozen lung and liver samples from cases 1 and 2, jejunal segments from each of the ponazuril-treated turtles (cases 6–8), and a sample of the fed silverside fish were individually processed for molecular testing (polymerase chain reaction [PCR]) at the Zoological Medicine Diagnostic Laboratory (UF ZMDxL), College of Veterinary Medicine, University of Florida. DNA was extracted using the DNeasy Blood and Tissue Kit (Qiagen, Valencia, CA, USA) following the manufacturer’s instructions. The mitochondrial cytochrome c oxidase subunit I gene (cox1) was partially amplified using forward primer CaryCOIF2 (5′-GTCTATTCACTTGGGCTATTGTATT-3′) and reverse primer CaryCOIR (5′-CTGTAGGGATAGCAATCATCAT-3′) with protocols as previously described. 34 The 18S ribosomal RNA gene (18S rRNA) was partially amplified using forward primer 1135F (5′-ACYATAAACTATGCCRACTAGA-3′) and reverse primer 1503R (5′-CYTCCYTRCRTTARACACGCAA-3′) with protocols as previously described. 12 To obtain additional sequence from the pooled case 1 and 2, a region of Eimeriidae rRNA containing small subunit ribosomal RNA, internal transcribed spacer 1, 5.8S ribosomal RNA, internal transcribed spacer 2, and large subunit ribosomal RNA was amplified using novel pan-Eimeriidae forward primer EimITSF (5’-GATCGGAGGGTCCTGTGAA-3’) and reverse primer EimITSR2 (5’-GGACACGCGATTTTCACTTT-3’). To connect this sequence with the 18S rRNA sequence previously obtained, primers 1446F (5’-TAAATAGGGTCGGGAACCTCTC-3’) and 1898R (5’-GATCCTTCYGCAGGTTCAC-3’) were used with protocols as previously described. 16 To obtain further 5’ 18S rRNA, primers 18F (CTGGTTGATCCTGCCAGTAGTC) and 1145R (GCAGGAGAAGTCGAGAATGACA) were used with protocols as previously described. 16 For PCR reactions, we used Invitrogen Platinum Taq DNA polymerase (ThermoFisher Scientific, MA, USA). Amplicons of the expected size were submitted for Sanger sequencing in both directions (Azenta, South Plainfield, NJ, USA). The gene sequences were compared to those in GenBank using BLASTN. 1 In addition, PCR for ranaviruses, reoviruses, and Monocercomonadidae flagellates were also performed to rule out these agents at the UF ZMDxL.35,37 Sequences generated were submitted to GenBank under accession numbers OP739494 and OP739495.
Phylogenetic Analysis
Homologous 18S rRNA genes from other apicomplexan species were downloaded from GenBank and aligned to the sequences obtained from the index red-eared slider case. For an outgroup, Theileria ovis (GenBank accession # AY260172), a non-Conoidasida member of the phylum Apicomplexa was used. The predicted homologous nucleotide sequences were aligned using MAFFT. 19 Bayesian phylogenetic analyses of nucleotide alignments were performed using MrBayes 3.2.7 on the CIPRES server, with a general time reversible model, gamma distributed rate variation, and a proportion of invariant sites.26,33 The first 25% of 2,000,000 iterations were discarded as burn in.
Results
Postmortem and Histologic Findings
Gross lesions
The deceased and euthanized turtles had carapace lengths of 3.3 to 3.6 cm. Gross alterations, listed by case in Table 1, were therefore sometimes difficult to appreciate due to the small organ size. The gross alterations noted in the deceased turtles (Fig. 1a–c) included severe bilateral enophthalmos, gaunt neck and shoulders (Fig. 1a), and scant (<1 ml) to moderate (approximately 1–2 ml) coelomic effusion. In case 4, a segment of duodenum and proximal jejunum was dark red, dilated, and had multiple pinpoint, tan, smooth serosal plaques (Fig. 1b). The stomach and small intestines in this turtle contained dark red mucoid digesta. The turtles euthanized following ponazuril treatment had scant to moderate coelomic effusion (Fig. 1c).
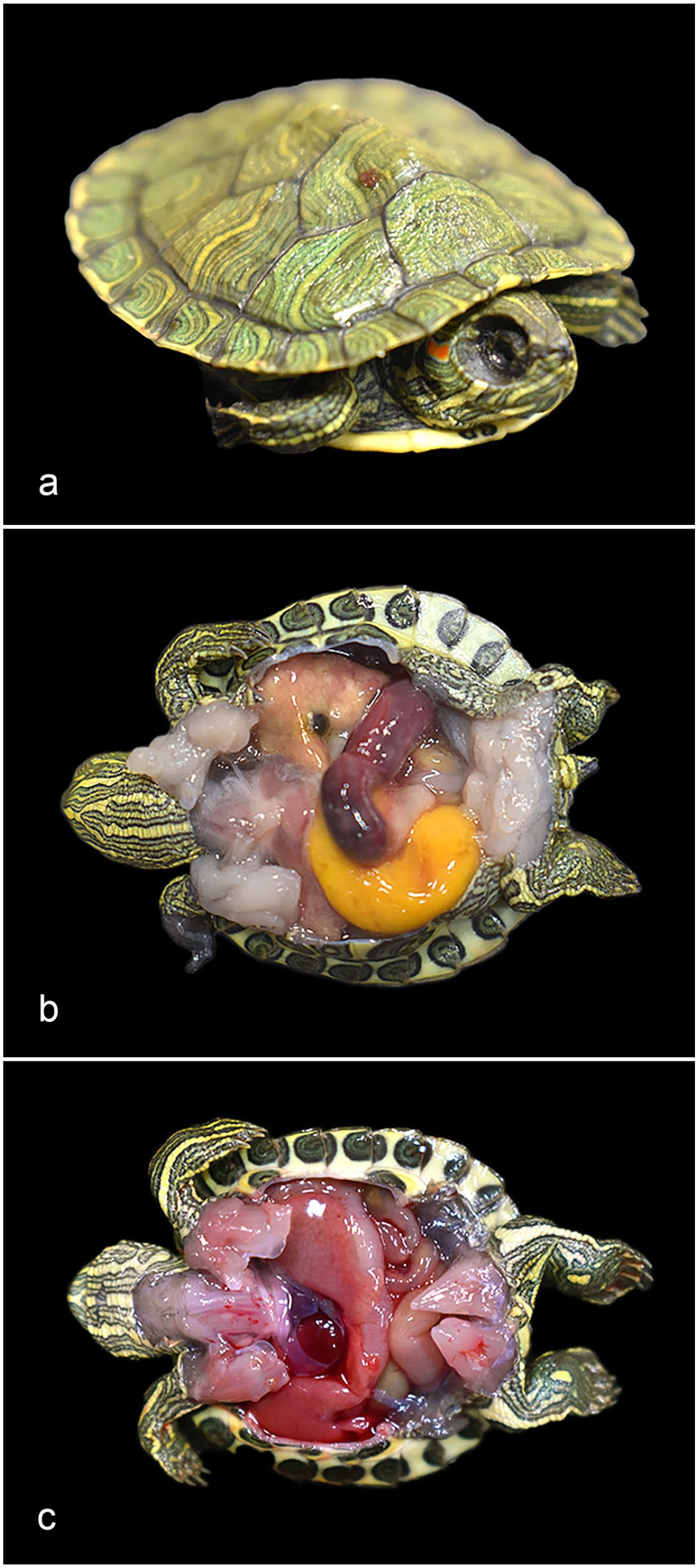
Caryospora-like organism (CLO) infection, red-eared slider turtles (Trachemys scripta elegans). (a) Fatal CLO infection, case 2. Severe enophthalmos. (b) Fatal CLO infection, case 4. Segmental dilation and hyperemia of the duodenum and proximal jejunum. (c) Post-ponazuril treatment, case 6. Grossly normal intestines for comparison, with scant coelomic effusion.
The histologic lesions, summarized in Table 2, were observed in multiple tissues. Common alterations across all deceased turtles mainly involved the gastrointestinal tract and liver. A representative whole slide image (WSI) is provided in the Supplemental Materials.
Histological findings associated with Caryospora-like coccidiosis in red-eared slider turtles (n = 8).
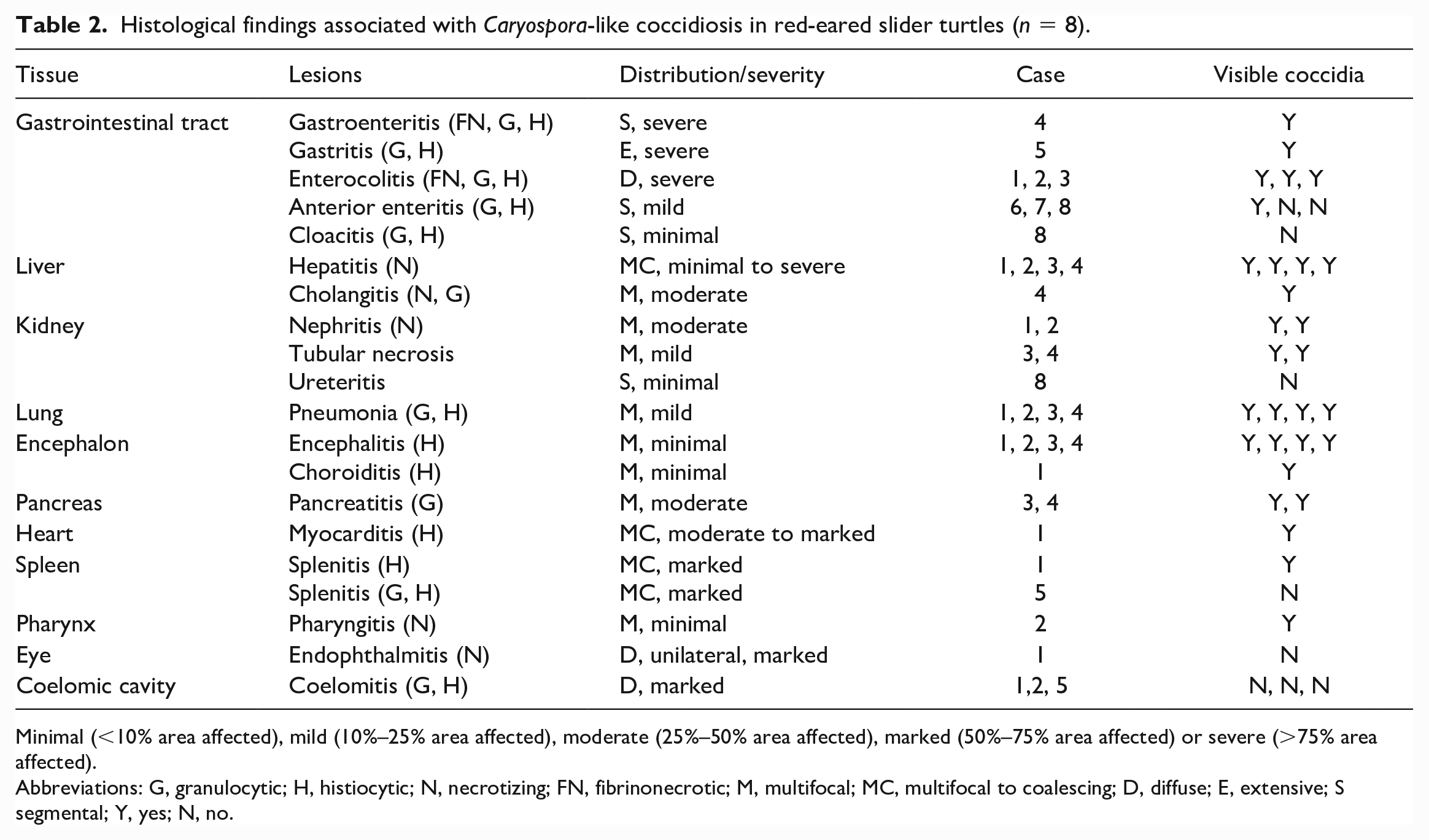
Minimal (<10% area affected), mild (10%–25% area affected), moderate (25%–50% area affected), marked (50%–75% area affected) or severe (>75% area affected).
Abbreviations: G, granulocytic; H, histiocytic; N, necrotizing; FN, fibrinonecrotic; M, multifocal; MC, multifocal to coalescing; D, diffuse; E, extensive; S segmental; Y, yes; N, no.
All deceased hatchlings (n = 4, cases 1–4) had severe segmental to diffuse, transmural, fibrinonecrotic, granulocytic, and histiocytic enterocolitis (Figs. 2a–f, 3a). These lesions were most severe in the duodenum and proximal jejunum and were characterized by transmural necrosis with complete loss of villi and intraluminal fibrin casts that were heavily colonized by mixed bacteria (Fig. 2c). Aborad segments of the jejunum had marked mucosal necrosis with villous blunting, crypt hyperplasia, and inflammatory infiltrates composed of heterophils and histiocytes (Fig. 2f). The colon had similar mucosal necrosis and inflammatory cell composition. All layers of the intestinal wall contained numerous intrahistiocytic and intraepithelial, intracytoplasmic coccidian organisms in various developmental stages (see below for details) as well as intraluminal, variably sporulated oocysts (Fig. 3a, b). The 3 ponazuril-treated and euthanized turtles had only mild segmental histiocytic and granulocytic anterior enteritis (Fig. 2b, e). Few similar intraepithelial coccidian stages were evident in 1 of the 3 ponazuril-treated turtles (Fig. 3c). One ponazuril-treated turtle (case 5) had extensive severe transmural granulomatous and heterophilic gastritis with rupture and intralesional pigmented fungal hyphae and intraluminal bacteria. Additionally, this turtle had severe histiocytic and heterophilic splenitis and coelomitis. Within the luminal debris of the stomach was a small cluster of 15 µm diameter, thin-walled, unsporulated oocysts.
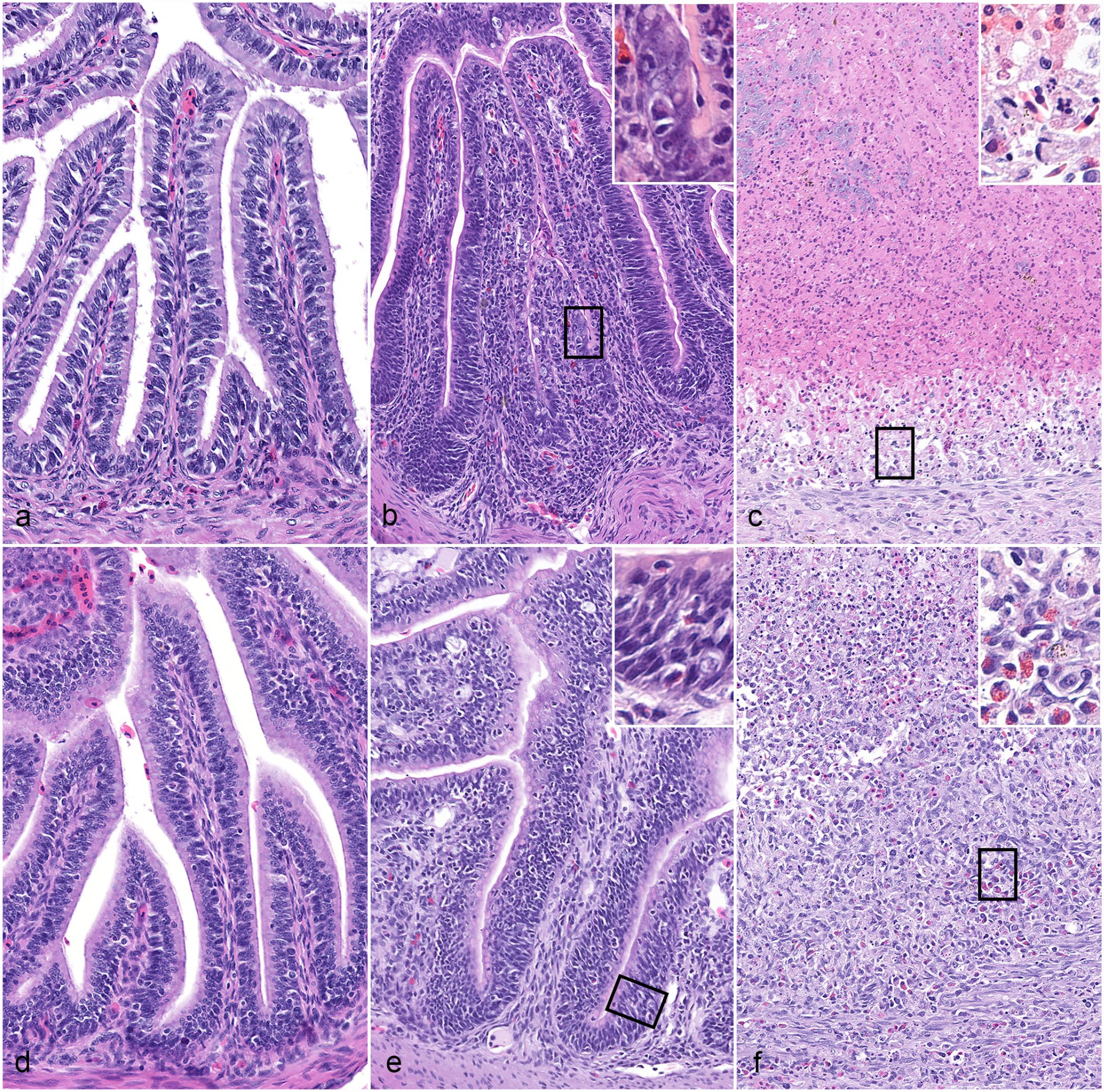
Comparison of CLO-associated duodenitis and jejunitis, red-eared sliders. Hematoxylin and eosin. (a-c) Duodenum. (a) Normal 1-month-old red-eared slider for comparison. (b) Post-ponazuril treatment hatchling, case 6. Mild segmental histiocytic, granulocytic and proliferative duodenitis (inset: magnified view of squared area). (c) Fatally infected hatchling, case 3. Severe fibrinonecrotic, histiocytic and granulocytic duodenitis (inset: magnified view of squared area) with complete loss of villi. (d-f) Jejunum. (d) Normal 1-month-old red-eared slider for comparison. (e) Post-ponazuril hatchling treatment, case 6. Mild segmental granulocytic and proliferative jejunitis (inset: magnified view of squared area). (f) Fatally infected hatchling, case 3. Marked fibrinonecrotic, histiocytic and granulocytic jejunitis (inset: magnified view of squared area) with severe villus blunting and ulceration.
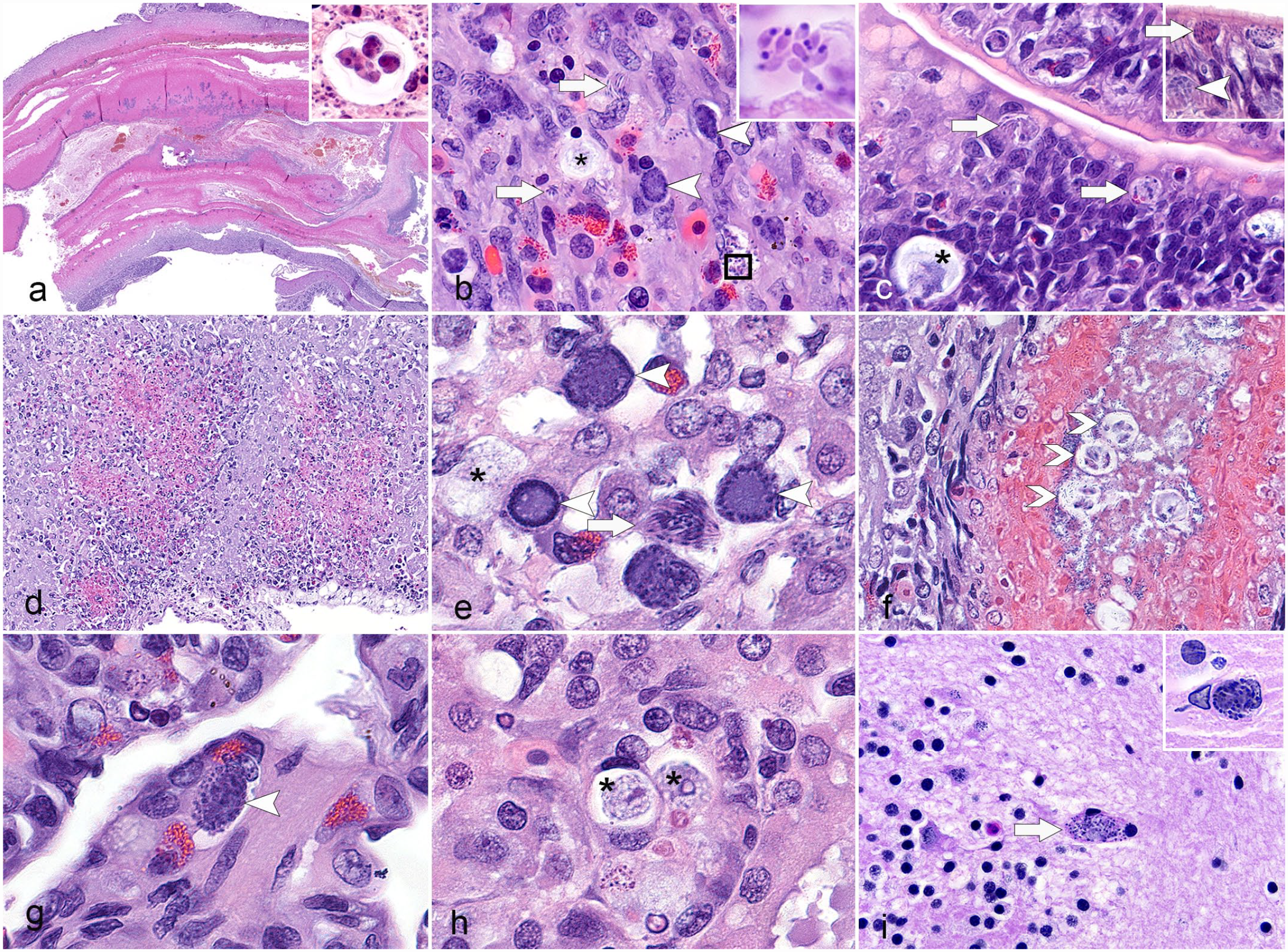
Systemic CLO infection, red-eared sliders. Hematoxylin and eosin. (a) Duodenum, fatally infected hatchling, case 4. Severe transmural necrotic enteritis with intraluminal fibrin casts and sporulated oocysts containing 4 paired sporozoites (inset). (b) Jejunum, fatally infected hatchling, case 4. Many presumably intrahistiocytic and intraepithelial meronts (arrow), gamonts (arrowhead), and unsporulated oocysts (asterisk) as well as extracellular merozoites (inset) throughout the intestinal wall. (c) Jejunum, post-ponazuril treatment, case 6. Few similar coccidian developmental stages (meronts [arrow], gamonts [arrowhead], and unsporulated oocysts [asterisk]) within hyperplastic epithelium. (d-f) Liver, fatally infected hatchling, case 4. (d) Random necrotic hepatitis. (e) Intrahepatocellular and intrahistiocytic gamonts (arrowhead), meront (arrow), and unsporulated oocyst (asterisk). (f) Fibrinonecrotic cholangitis with intraluminal sporulated oocysts (open arrowheads). (g) Lung, case 4. Interstitial granulocytic pneumonia with intralesional gamont (arrowhead). (h) Kidney, case 4. Intraepithelial and intratubular unsporulated oocysts (asterisks). (i) Brain, case 3 and 4 (inset). Rare perivascular intraglial meronts (arrow and inset).
Extraintestinal organs
Deceased hatchlings (n = 4; cases 1–4) had moderate to severe, multifocal to coalescing, necrotizing hepatitis with intralesional Gram-negative bacilli (Fig. 3d), intrahistiocytic and intrahepatocellular coccidian stages (Fig. 3e), fibrinoid vascular necrosis, and thrombosis. One turtle also had necrotizing cholangitis with intraluminal, variably sporulated oocysts (Fig. 3f). Various developmental stages of coccidia were also observed within the cytoplasm of pancreatic acinar cells, pneumocytes, and interstitial histiocytes in the lungs (Fig. 3g), renal tubular epithelium (Fig. 3h), ureteral epithelium, perivascular cerebral glial cells (Fig. 3i), and rarely within myocardial and splenic macrophages and pharyngeal epithelium. These coccidia were accompanied by variable necrosis and granulocytic and histiocytic inflammation within the different extraintestinal tissues. Additionally, two turtles had severe diffuse fibrinous and histiocytic coelomitis with intralesional Gram-negative bacilli and yolk material. One of the ponazuril-treated turtles had minimal histiocytic and heterophilic cloacitis and ureteritis; however, no extraintestinal organisms were microscopically identified in any of the 3 ponazuril-treated turtles. None of the special histochemical stains (Gram, Fite-Faraco, and Steiner) highlighted the coccidian organisms in the tissue sections.
Coccidia stages
Meronts measured up to 16 µm in diameter and were packed with numerous elongate merozoites that were 1 µm in diameter and had a round to ovoid polar nucleus (Fig. 3b). Extracellular individual merozoites had rounded and pointed ends with a round deeply basophilic structure within the pointed end (Fig. 3b, inset). Microgamonts measured up to 13 µm in diameter and had basophilic, granular cytoplasm with numerous round, central, deeply basophilic structures, while macrogamonts were rare, of similar size, and had peripheral deeply basophilic structures (Fig. 3e). Several unsporulated oocysts were scattered throughout all affected sites; they measured 15 µm x 12 µm and had a thick capsule and central pale eosinophilic vacuolated cytoplasm (Fig. 3h). Occasional round to ovoid variably sporulated oocysts containing 4 paired, 6 µm x 2 µm sporozoites were within the intestinal luminal debris and biliary tract (Fig. 3a).
Bacteriology
Multiple bacterial species were isolated from the coelomic swabs of deceased turtles, which are listed in Table 1. Selected isolates of potential concern, but were considered to be opportunistic, included Aeromonas hydrophila, Edwardsiella tarda, and Citrobacter freundii. Aerobic cultures of ponazuril-treated and subsequently euthanized turtles yielded no growth. Salmonella sp. cultures in all hatchlings were negative.
Immunohistochemical Findings
All tissues from 7/8 turtles (cases 2–8) were evaluated by IHC using a polyclonal antibody against N. caninum that immunolabeled all coccidian developmental stages including meronts, merozoites, microgametes, rare macrogametes, and unsporulated and sporulated oocysts (Fig. 4a–i). In the deceased turtles (n = 3), various coccidian stages were identified within enterocytes, fibroblasts, and histiocytes throughout the entire gastrointestinal tract (3/3; Fig. 4a–c); pancreatic acinar epithelium and inflammatory cells in the pancreatic interstitium (3/3); hepatocytes, Kupffer cells, and lumen of necrotic bile ducts in the liver (3/3, Fig. 4d–h); renal tubular epithelium (3/3); pneumocytes and interstitial pulmonary histiocytes within faveolar septa of the lungs (3/3); cerebral glia (3/3, Fig. 4i); cardiomyocytes (2/3); splenic macrophages (2/3); ureteral epithelium (1/2), macrophages within yolk sac remnants (1/3); mesentery (as free merozoites, 1/3); pharyngeal epithelium (1/3); and bone marrow precursors (1/3). In the treated turtles (n = 4), similar yet infrequent to rare coccidian stages were identified in the duodenal and jejunal epithelium (3/4 and 1/4, respectively; Fig. 4d–f), hepatocytes (2/4), pancreatic acinar epithelium (1/4), and pneumocytes and interstitial pulmonary macrophages (1/4). No organisms were identified within the examined thyroid or adrenal glands of the turtles evaluated nor in the tissues of the ponazuril-treated hatchling with fungal and bacterial gastritis and coelomitis (case 5). The cases and the affected tissue types are reported in Table 3.
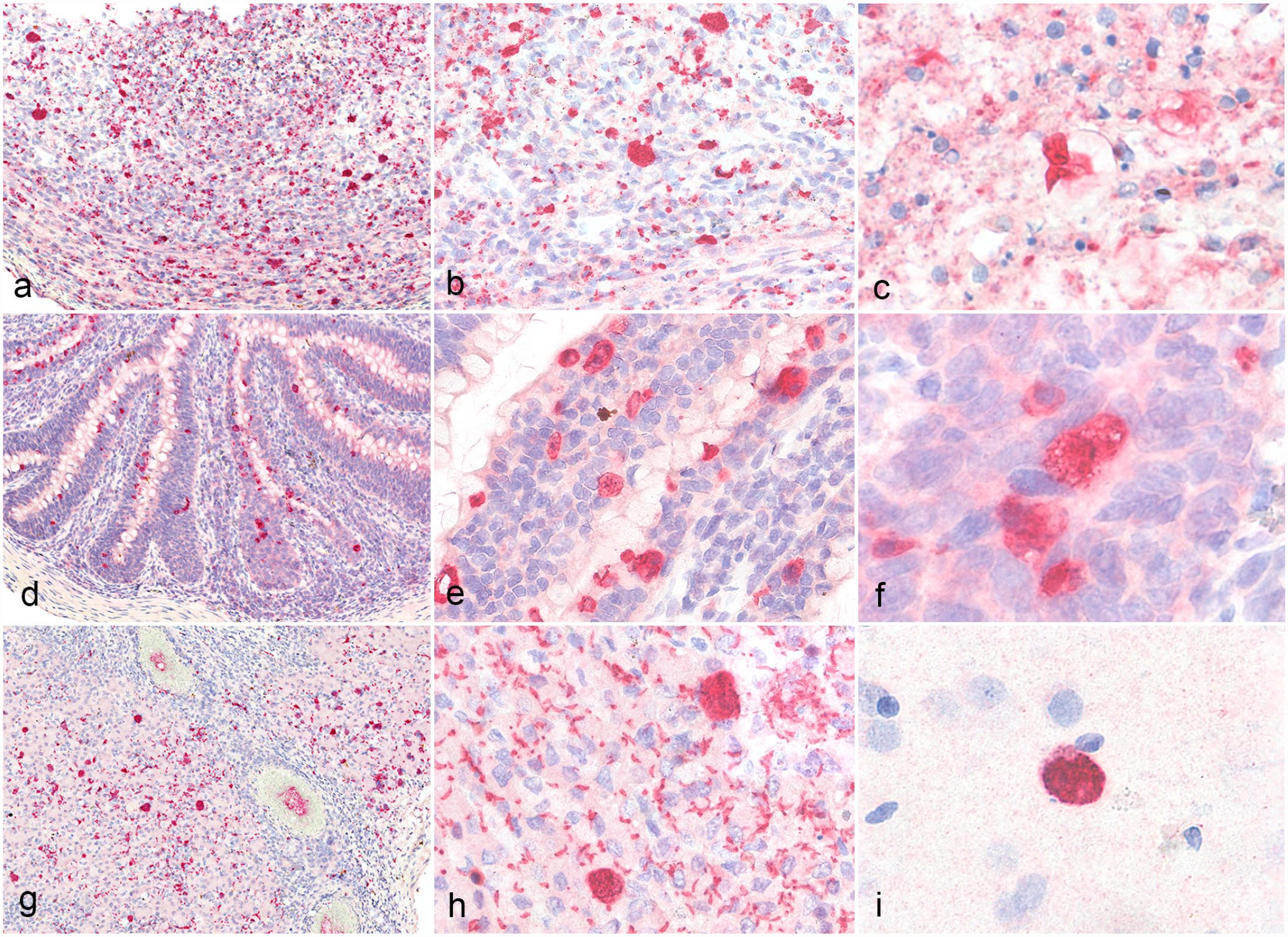
Systemic CLO infection, red-eared sliders. Immunohistochemistry using an anti-Neopsora caninum whole tachyzoite antibody. a-b) Jejunum, fatally infected hatchling, case 3. Numerous immunolabeled coccidian developmental stages transmurally throughout the intestine. c) Jejunum, fatally infected hatchling, case 4. Disintegrating sporulated oocysts with immunolabeled sporozoites. d-e) Jejunum, post-ponazuril treatment, case 6. Sporadic immunolabeled coccidian developmental stages confined to the lining enterocytes. f) Jejunum, post-ponazuril treatment, case 6. Immunolabeled meront with few extracellular merozoites. g-h) Liver, fatally infected hatchling, case 4. Numerous immunolabeled coccidian developmental stages within hepatocytes and macrophages as well as extracellular merozoites. i) Brain, fatally infected hatchling, case 4. Meront within the cerebrum.
Cases in which coccidian organisms, listed by tissue, were detected by immunohistochemistry using a cross-reactive anti-Neospora caninum polyclonal antibody (n = 7 turtles tested).
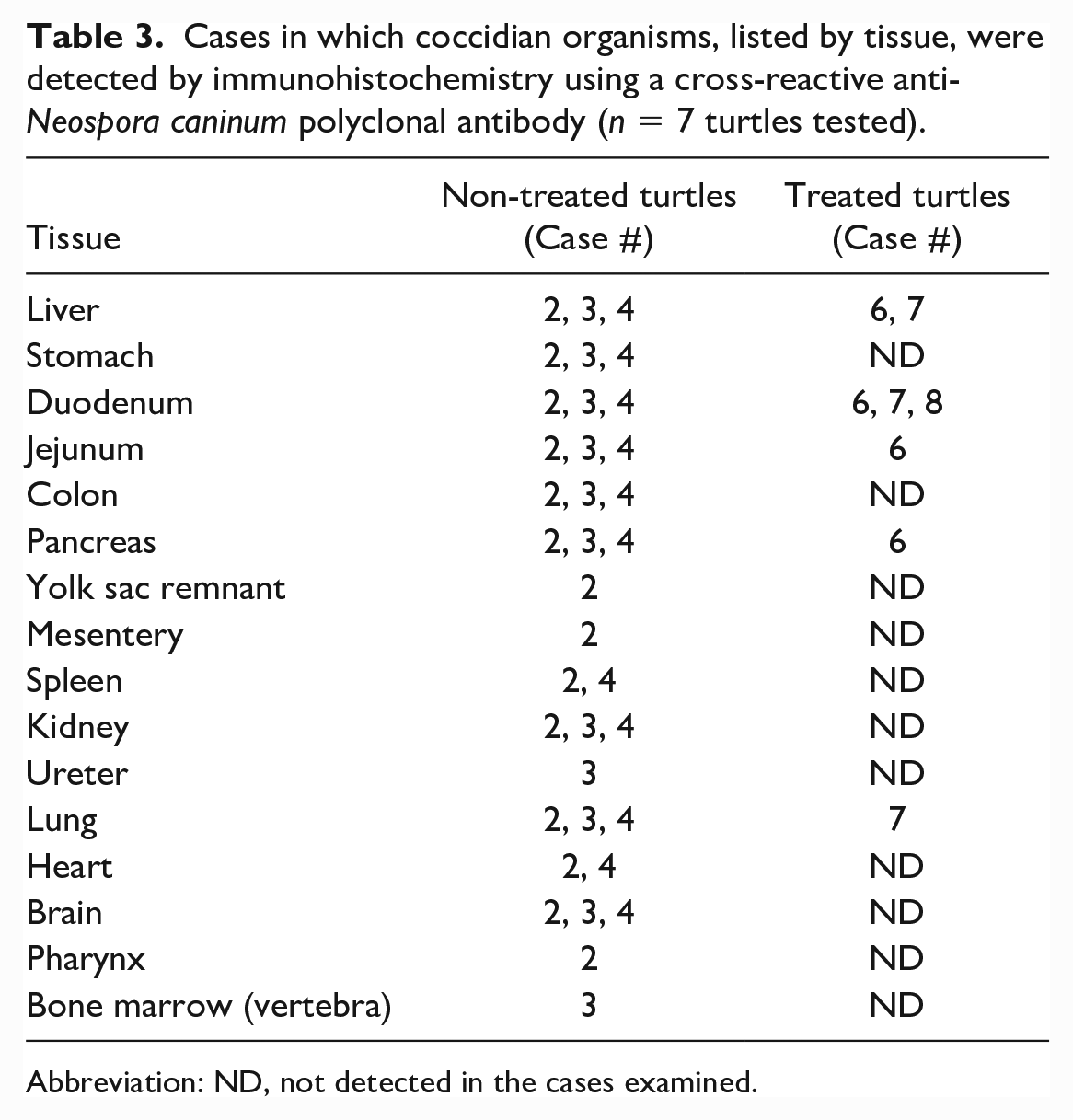
Abbreviation: ND, not detected in the cases examined.
Ultrastructural Features
Several apicomplexan developmental stages were identified in the intestinal and hepatic sections evaluated by transmission electron microscopy and were most numerous in the intestinal sections. Merogony stages (meronts) were the most frequently observed stage. Meronts ranged from 9 to 18 µm in diameter, were intracytoplasmic, surrounded by a parasitophorous vacuole, and contained a residual body along with 6 to 40 merozoites in various stages of maturation (Fig. 5a). Merozoites were 1 µm in diameter and ranged from 5.3 to 7 µm in length. Transections of merozoites revealed an apical complex composed of anterior conoid, micronemes, and rhoptries; a wall with a micropore; numerous microtubules; posterior refractile body and nucleus with nucleoli; and various cytoplasmic organelles such as amylopectin granules, ribosomes, and acidocalcisomes (Fig. 5b). 4
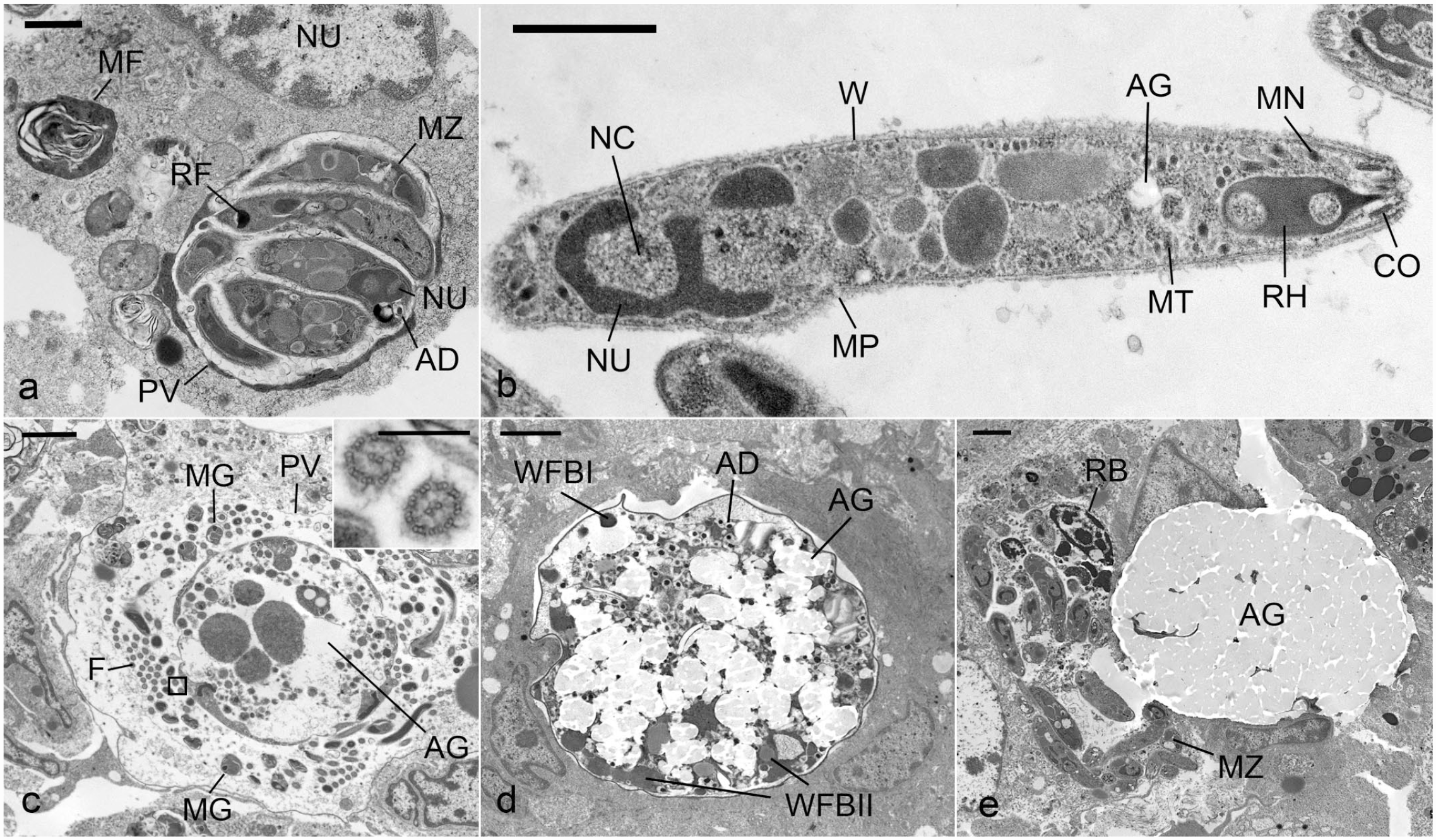
Systemic CLO infection, red-eared slider, case 3. Transmission electron microscopy. (a) Intestine. A degenerating enterocyte with cytoplasmic myelin figures (MF) and mature meront containing multiple merozoites (MZ) with a posterior nucleus (NU), refractile body (RF), and few acidocalcisomes (AD) that are surrounded by a parasitophorous vacuole (PV). Bar = 1 µm. (b) Liver. Extracellular merozoite with a conoid (CO), micronemes (MN), rhoptry (RH), wall (W), micropore (MP), microtubules (MT), amylopectin granules (AG), nucleus (NU), and nucleoli (NC). Bar = 0.8 µm. (c) Intestine. Intracellular mature microgamont, contained by a PV, with multiple peripheral microgametocytes (MG), flagella (F) with 9 peripheral double microtubules and 2 central microtubules (inset; bar = 300 nm), and amylopectin granules (AG). Bar = 2 µm. (d) Intestine. Developing zygote with peripherally located smaller wall forming bodies (WFB I) intermixed with larger WFBII, AG, and AD. Bar = 2 µm. (e) Intestine. Intracellular meront with MZs and a residual body (RB) and an adjacent intracellular unsporulated oocyst with amylopectin granules (AG). Bar = 2 µm.
Microgamonts were infrequent, ranged from 6.5 µm x 9 µm to 10 µm x 14 µm, and were bound by a similar parasitophorous vacuole (Fig. 5c). Mature microgamonts had central residual cytoplasm with amylopectin granules and multiple peripheral microgametes 29 with flagella that had 9 peripheral double microtubules and 2 central microtubules (Fig. 5c, inset). 36 Macrogametes were rare, histologically, and not observed by electron microscopy. Developing zygotes ranged from 7 to 10.5 µm in diameter and contained peripherally located, small electron-dense type 1 wall forming bodies (WFBI) intermixed with larger type 2 wall forming bodies (WFBII), 38 lipid, amylopectin granules, and various organelles (Fig. 5d). Several mature unsporulated oocysts measured 11 µm x 15 µm and were composed of predominately amylopectin granules (Fig. 5e). Ultrastructural features are summarized in Table 4.
Electron microscopy findings of various Caryospora-like organism developmental stages in the intestine and liver of a red-eared slider turtle (case 3).
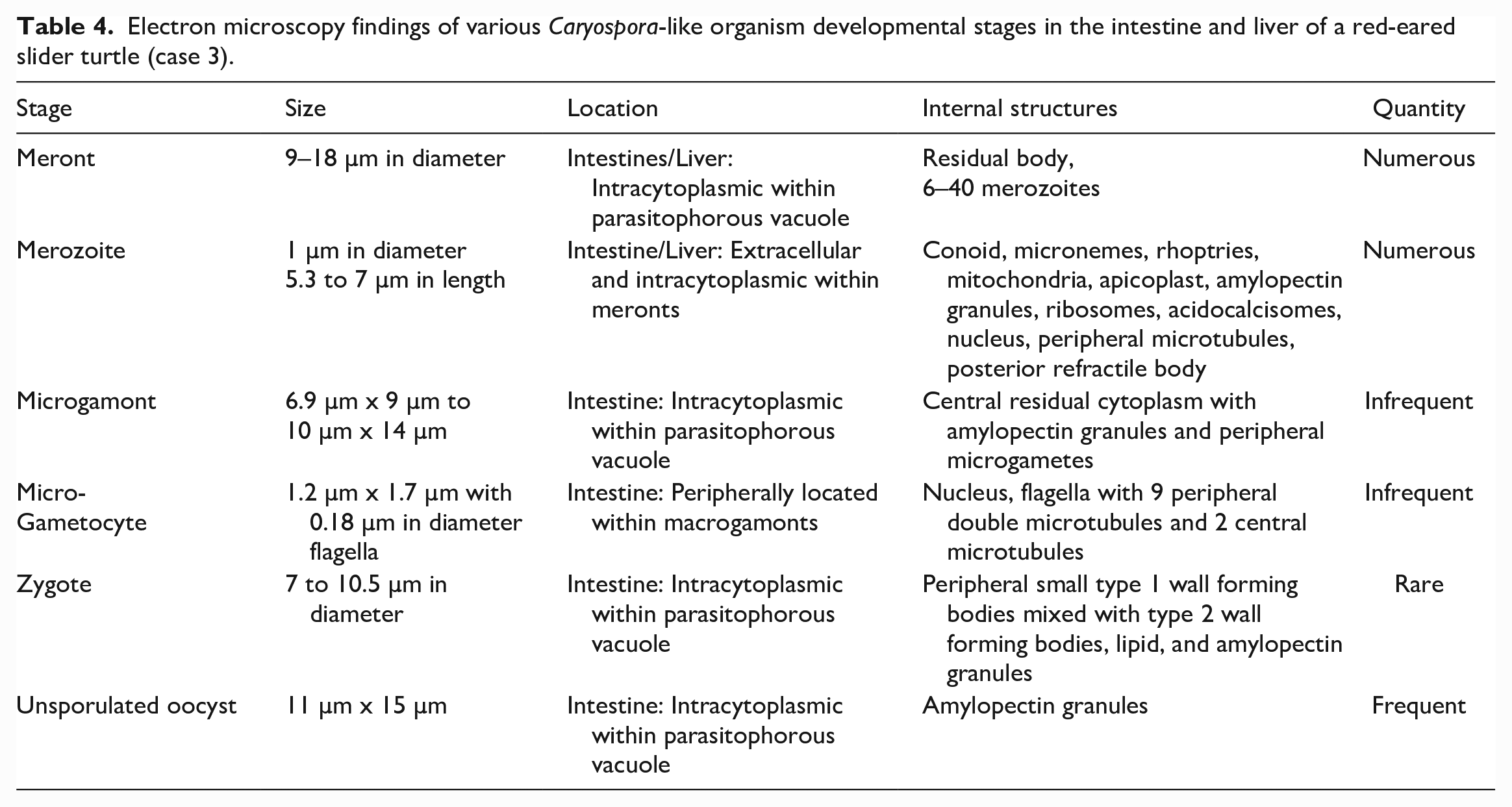

Infected cells showed several features of degeneration and necrosis. These included formation of myelin figures, nuclear pyknosis and karyorrhexis, mitochondrial swelling, and fragmentation of the cell membrane.
Molecular Testing
Initial amplification of the rRNA gene from the pooled liver and lung specimens (cases 1 and 2) resulted in a 347 bp product. BLASTN results indicated that the sequence obtained had highest identity to the Schellackia/Caryospora-like clade in the Eimeriidae, with 99.1% nucleotide identity to the US3 strain from green turtles (MN450817), as well as a 99.1% identity to Schellackia sp. isolate OC116 (MG775272.1). The jejunal sample of only one of the 3 ponazuril-treated turtles that were tested (cases 6–8) was positive for CLOs, the same turtle in which intestinal coccidian stages were histologically identified. The feeder silversides yielded a negative result for coccidia.
PCR of the mitochondrial cox1 gene resulted in a product of 355 base pairs after primers were edited out. The longer rRNA sequence with the additional PCR protocols was 2601 bp after assembly. Sequences were submitted to GenBank under accession numbers OP739494 and OP739495. PCR for ranaviruses, reoviruses, and Monocercomonadidae flagellates had negative results.
Phylogenetic Analysis
The Bayesian tree is shown in Fig. 6. This organism clustered with other members of the Eimeriidae, with a Bayesian posterior probability of 100%. Furthermore, this organism formed a clade with Eimeria/Isospora/Schellackia/Lankesterella/Caryospora and some of the Caryospora species (brackets 1–5, Fig. 6), with a Bayesian posterior probability of 100%. The most basal members of this clade (brackets 4 and 5, Fig. 6) all use poikilothermic hosts, including turtles, squamates, and anurans. Caryospora, Eimeria, and Isospora are all paraphyletic.
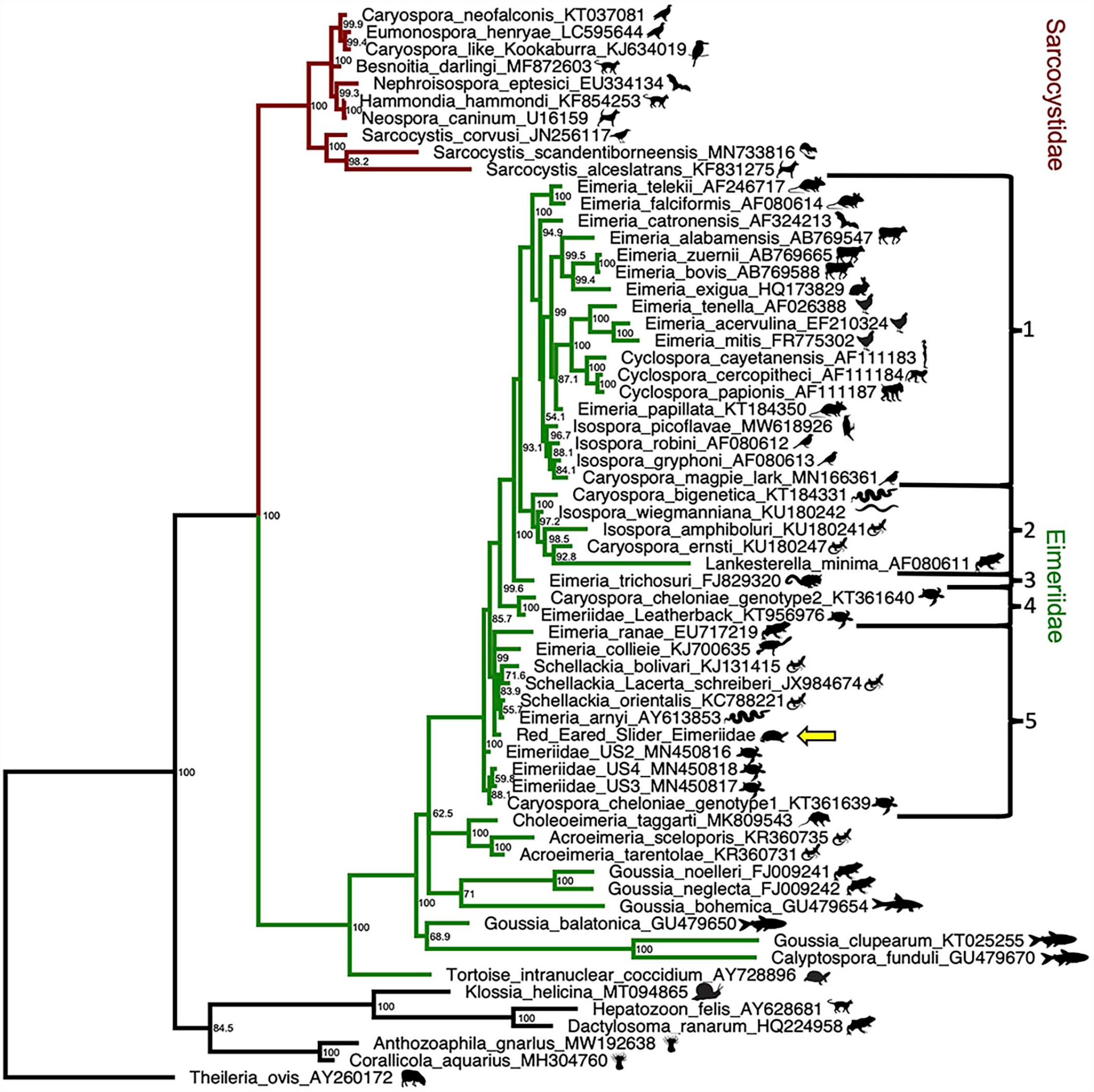
Phylogenetic analysis of 18S rRNA gene sequences from other apicomplexan species and the index red-eared slider case (yellow arrow). Theileria ovis (GenBank accession # AY260172), a non-Conoidasida member of the phylum Apicomplexa was used as an outgroup. Node support values are indicated. Brackets 1 through 5 represent different clades within Eimeriidae. Bayesian phylogenetic analyses of nucleotide alignments performed using MrBayes 3.2.7.
Discussion
To our knowledge, this is the first report of CLOs infecting freshwater non-cheloniid turtles, indicating potential cross-species infectivity and pathogen-host switching capability. Therefore, the aim of our study was to characterize the host cell tropism, pathologic and ultrastructural features, and phylogeny associated with CLOs in red-eared sliders. CLOs are emerging pathogens of marine turtles and a major cause of mass mortality-associated enterocolitis since the 1990s. During the 2014 Atlantic epizootic, there was a 36.1% increase in green turtle strandings and, of the stranded turtles that were positive for CLOs, 75% died while the remainder were released following treatment. 34 In this clutch of red-eared slider hatchlings, coccidia were identified immunohistochemically in 6 of the 7 tested turtles and were responsible for the death of 4 of 8 turtles (50% mortality). The interventional treatment attempted during the outbreak may have improved the survival rate. Similar to CLO infections in marine turtles in Australia, systemic CLO infection in this outbreak resulted in severe fibrinonecrotic enterocolitis as well as pneumonia, nephritis, and encephalitis. 6 This is in contrast to other CLO-infected marine turtles in the Southern Hemisphere, such as the most recently reported affected turtles from Northeastern Brazil, in which extra-intestinal lesions were not reported. 8 Additional affected organs and tissues of this clutch of red-eared sliders that have not been reported in marine turtles included the liver, biliary tract, stomach, pancreas, heart, spleen, mesentery, yolk sac remnant, pharynx, and bone marrow. Unlike some infected marine turtles,6,11 the examined thyroid and adrenal glands were unaffected, and no coccidian organisms were detected with immunohistochemistry in these endocrine tissues of the red-eared slider hatchlings. Similar coccidian tissue distribution as seen in the red-eared slider hatchlings was reported in 2 freshwater Indo-gangetic flap-shelled turtles (Lissemys punctata andersonii) with disseminated visceral coccidiosis. However, fibrinonecrotic enterocolitis and necrotizing hepatitis were not a feature in those cases, and the molecular characterization in those turtles revealed coinfections with 2 different Eimeria species. 14
Several treatments have been attempted in CLO-infected marine turtles including supportive care and the use of triazine anti-coccidials (ponazuril or toltrazuril),17,30 a group of anti-coccidials that are thought to target the apicoplast, and/or metronidazole. 24 Importantly, triazine treatments are reportedly effective at blocking intracellular propagation of the zoites, improving clinical disease.20,23 The 3 ponazuril-treated and subsequently euthanized hatchlings were clinically normal yet had mild proliferative anterior enteritis. Immunohistochemistry highlighted several organisms within the duodenum of these hatchlings and in the pancreas, liver, and lung of 1 hatchling (case 6), which also was PCR positive for CLOs. The presence of both intestinal and rare extraintestinal coccidia in these treated hatchlings suggests that, while this treatment regimen appears to be effective at controlling clinical disease, the organism is not completely cleared by it. The negative CLO PCRs in the 2 treated hatchlings in which only intestinal organisms were observed (cases 7 and 8) may be due to the low numbers and segmental distribution of organisms.
The phylogeny of much of the coccidia, especially Eimeriidae, is problematic and in need of revision. As found in previous analyses, Caryospora is paraphyletic, with some members, such as Caryospora neofalconis, not clustering in the Eimeriidae (Fig. 6). 7 It is possible that the Caryospora type species, C. simplex, may cluster with other Caryospora that use snakes as definitive hosts (bracket 2, Fig. 6), but data would be needed to prove this. Even though the CLOs of marine turtles are not monophyletic, Caryospora chelonae genotype 2 and the leatherback turtle coccidia form a supported clade (bracket 4 depicted in Fig. 6, 100% posterior probability) that is more closely related to a Eimeria/Isospora/Cyclospora/Caryospora/Lankesterella clade (brackets 1–3 depicted in Fig. 6, 99.6% posterior probability) with the majority of its members (brackets 1 and 3 depicted in Fig. 6) using warm-blooded hosts, rather than to other marine turtle CLOs (bracket 5, depicted in Fig. 6). Phylogenetic classification of coccidia based on sporulation patterns has proven problematic; there is significant homoplasy in sporulation patterns, and some of the Cystoisospora inconsistently undergo sporulation patterns that would cause them to be misidentified as Caryospora.15,18 Molecular data have shown that the presence or absence of Stieda bodies, the use of paratenic hosts, and host clade are all more phylogenetically informative than number of sporocysts or sporozoites, which was unfortunately used for classification prior to molecular data. 2
The phylogenetic resolution in the region where most turtle coccidia are found (bracket 5, Fig. 6) is not strong, with a large number of node support values <95%. It has been proposed that the CLOs of marine turtles may be classified within Schellackia, and some green turtle CLO sequences have been deposited in GenBank under this classification (accession # KY046254). Schellackia are intraerythrocytic coccidia which have been reported to cospeciate with their squamate hosts. 25 Our analysis also finds that Schellackia clusters with our red-eared slider CLO and some marine turtle CLOs (bracket 5, Fig. 6), although it is unclear whether they are the same clade or sister groups. Increased taxon and gene sampling are probably needed to resolve this. 28
The complete life cycles of CLOs and routes of transmission have yet to be fully elucidated. The global distribution in marine turtles and the genotype similarities between geographically separate epizootics has implicated interoceanic spread by transportation of infected hosts or oocysts. 34 Disease severity varies widely in marine turtles and it is not known if there are subclinical carriers that could serve as reservoirs. Despite a thorough investigation, how the clutch of red-eared sliders was exposed to this pathogen remains undetermined. The outbreak appeared to coincide with a new shipment of Atlantic silverside fish (Menidia menidia), but molecular testing of this batch of fish was negative for coccidia. Unlike the Sarcocystidae, members of the Eimeriidae have so far been found to have a 1-host life cycle, making a non-turtle reservoir less likely. While sporozoites from a marine turtle CLO outbreak have been shown to rapidly disintegrate within 2 to 3 days, indirectly implying that there is a need for an actively shedding host or reservoir for effective transmission of this coccidia, 13 no additional in-depth studies assessing transmissibility and environmental resistance have been performed for this genus. While other chelonids were housed in this facility, appropriate biosecurity measures were in place, and no other chelonids in the collection were affected. Given that this clutch was hatched within the facility and had no contact with other chelonids, transovarian transmission was also considered but could not be further investigated in this study. All fatally infected turtles had concurrent sepsis, likely secondary to the CLO-induced necrotic enterocolitis. The only ponazuril-treated hatchling that died had severe bacterial and fungal gastritis and coelomitis, but no associated coccidian organisms were identified by immunohistochemistry despite the small cluster of oocysts observed mixed with the intraluminal necrotic debris in the stomach in the hematoxylin and eosin-stained section. It is therefore unclear if the lesions responsible for the death of this turtle are directly related to previous CLO infection.
This is the first report of a CLO infection in a cohort of non-cheloniid turtles, and this report demonstrates this organism’s potential for cross species infectivity and considerable high pathogenicity. It is not known if this outbreak represents an isolated spillover event or reflects host broadening of this emerging disease of marine turtles. Further studies are needed to elucidate transmission, susceptible hosts, and potential reservoirs. Importantly, as this report demonstrates that hatchling red-eared sliders are susceptible to CLOs, this species could serve as an economically feasible model to study this important pathogen of marine turtles.
Supplemental Material
sj-pdf-1-vet-10.1177_03009858231179129 – Supplemental material for Systemic Caryospora-like coccidiosis in a clutch of hatchling red-eared slider turtles (Trachemys scripta elegans)
Supplemental material, sj-pdf-1-vet-10.1177_03009858231179129 for Systemic Caryospora-like coccidiosis in a clutch of hatchling red-eared slider turtles (Trachemys scripta elegans) by Naomi Falconnier, Michael Warshaw, James F. X. Wellehan, April L. Childress, Daniel K. Howe, Holly Taylor, Ingeborg M. Langohr, Mary B. Ard, Daniel B. Paulsen, Emi Sasaki, Maria S. Mitchell and Mariano Carossino in Veterinary Pathology
Footnotes
Acknowledgements
The authors would like to gratefully acknowledge Dr. Michael Garner, DVM, DACVP (Northwest ZooPath, Monroe, WA) for early consultation, his valuable input in this study and for critically reviewing this manuscript, and Dr. Eric Erwood, DVM (Louisiana State University) for assisting with tissue processing and evaluation. We would also like to kindly acknowledge the members of the Histology and Immunohistochemistry section at the Louisiana Animal Disease Diagnostic Laboratory (LADDL), School of Veterinary Medicine, Louisiana State University for their assistance. We would also like to acknowledge the Georgia Electron Microscopy laboratory at the University of Georgia for assistance with processing tissues and visualizing via transmission electron microscopy.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was partially supported by start-up funds from the School of Veterinary Medicine, Louisiana State University to Dr. Mariano Carossino (PG009641) and by funds generated by the Louisiana Animal Disease Diagnostic Laboratory (LADDL).
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
