Abstract
Shuni virus (SHUV), an orthobunyavirus of the Simbu serogroup, was initially isolated in Nigeria in the 1960s, further detected in other African countries and in the Middle East, and is now endemic in Israel. Transmitted by blood-sucking insects, SHUV infection is associated with neurological disease in cattle and horses, and with abortion, stillbirth, or the birth of malformed offspring in ruminants. Surveillance studies also indicated a zoonotic potential. This study aimed to test the susceptibility of the well-characterized interferon (IFN)-α/β receptor knock-out mouse model (Ifnar−/−), to identify target cells, and to describe the neuropathological features. Ifnar−/−mice were subcutaneously infected with two different SHUV strains, including a strain isolated from the brain of a heifer showing neurological signs. The second strain represented a natural deletion mutant exhibiting a loss of function of the S-segment–encoded nonstructural protein NSs, which counteracts the host’s IFN response. Here it is shown that Ifnar−/−mice are susceptible to both SHUV strains and can develop fatal disease. Histological examination confirmed meningoencephalomyelitis in mice as described in cattle with natural and experimental infections. RNA in situ hybridization was applied using RNA Scope™ for SHUV detection. Target cells identified included neurons and astrocytes, as well as macrophages in the spleen and gut-associated lymphoid tissue. Thus, this mouse model is particularly beneficial for the evaluation of virulence determinants in the pathogenesis of SHUV infection in animals.
Shuni virus (SHUV) is an insect-transmitted orthobunyavirus of the Simbu serogroup within the family Peribunyaviridae. 3 SHUV was initially isolated in Nigeria in the 1960s and subsequently detected in other African countries. 1 The first cases outside the African continent were noticed in Israel in 2014. By now, SHUV disease is endemic in this country.5,7
The tri-segmented RNA genome of SHUV encodes 6 proteins. The large genomic segment (L-segment) encodes the viral RNA-dependent RNA polymerase. The M-segment encodes the nonstructural protein NSm and the surface glycoproteins Gn and Gc, which are involved in viral attachment, cell fusion, and the induction of neutralizing antibodies. The small segment (S-segment) encodes for the nucleocapsid protein (N) and the nonstructural protein NSs in two overlapping reading frames.2,21,24 Like for other orthobunyaviruses, the NSs represents a major virulence factor in vertebrate hosts, as this protein counteracts the shut-off of host-cell protein synthesis and the induction of interferon (IFN).2,8,25
SHUV has been associated with neurological disease in cattle and horses, and was also detected in other domestic and wildlife species.6,18,23 In addition, an infection of naïve ruminants during gestation may lead to abortion, stillbirth, or the birth of malformed lambs and calves.5,6 A zoonotic potential was suspected since SHUV antibodies have been found in veterinarians in South Africa and became even more likely after SHUV RNA detection in cerebrospinal fluid from human neurologic disease cases.13,22
During the recent disease outbreaks in Israel, SHUV was isolated from brains of animals with both clinical presentations, namely encephalitis in cattle (strain 2162/16) and abnormal course of gestation in a sheep (strain 2504/3/14).5,6,8 The SHUV strain 2504/3/14 originated from a malformed, aborted sheep fetus and harbored mutations in the S-segment gene that encodes the nonstructural protein NSs, leading to a loss of the NSs function. 8 In vitro data suggested that this NSs-truncated SHUV 2504/3/14 did not prevent the activation of the host’s innate immune IFN response anymore. 8
We recently demonstrated that experimental infection of cattle with the “encephalitis strain” SHUV 2162/16 leads to short-term viremia and seroconversion. Five out of 6 animals showed elevated body temperature, but neurological signs were not recorded within 21 days. 17 However, as reported for naturally infected animals, 6 the cattle developed nonsuppurative meningoencephalitis, and SHUV RNA was identified intralesionally by RNA in situ hybridization (ISH). 17 In clear contrast, the NSs-truncated “malformation strain” 2504/3/14 did not lead to viremia. Seroconversion was found at very low titers in only 1 out of 6 animals, indicating that this natural mutant represents a dead-end artifact that is not fit for the usual transmission cycle. 17
To date, no information is available on the neuropathological features of SHUV infection in humans. This highlights the importance of using well-characterized animal models to reveal clues to the potential pathogenesis of this disease not only in humans, but also in domestic animals. Studies using large animals are complicated by the high costs and the limited availability of adequate high-containment housing for laboratory animals. However, these obstacles could be mitigated by small animal models as soon as suitable species are identified. Research on orthobunyaviruses has significantly benefited from the introduction of the IFN-α/β receptor knock-out (Ifnar−/−) mice, which were shown to be highly susceptible to the Simbu serogroup viruses Schmallenberg virus (SBV) and Sabo virus.19,28 In contrast to these viruses, the pathogenesis of SHUV has not yet been characterized in a mouse model in detail. Not yet fully immunocompetent newborn wild-type mice have been used for virus isolation, 5 but by using intracerebral infection an inoculation route was selected that does not resemble the natural route of infection. Thus, we investigated whether the Ifnar−/− mouse model is also applicable to study SHUV infection following subcutaneous inoculation and whether this mouse model mirrors the alterations seen in cattle.
Materials and Methods
Virus
Two virus strains originating from Israel were used. The “encephalitis strain” SHUV 2162/16 was isolated on Vero cells from the brain homogenate of a heifer showing neurological signs. 6 The “malformation strain” SHUV 2504/3/14 expresses a truncated NSs protein due to a stop codon in the NSs open reading frame and was obtained from the brain of a malformed, aborted sheep fetus. Isolation was performed using suckling mice and Vero cells (for details see Golender et al5,6). Both isolates were passaged once each on Culicoides sonorensis cells (KC cells, L1062, collection of cell lines in veterinary medicine (CCLV), Friedrich-Loeffler-Institut (FLI; Greifswald-Insel Riems, Germany), and baby hamster kidney cells (BHK 21 cells, L0164 CCLV). The KC cells were grown in Schneider’s Insect Medium at 28°C and the BHK 21 cells in Minimum Essential Medium at 37°C.
Animal Experiment
The experimental protocol was evaluated by a state ethics commission and approved by the competent authority (permission number: LALLF M-VTSD/7221.3–2-010/18).
Twenty-six Ifnar−/− mice on a C57BL/6 genetic background (B6-IfnartmAgt) were obtained from the specific pathogen-free breeding unit of the FLI. The age of the mice on the day of inoculation was 35–38 weeks. Twenty-four Ifnar−/−mice were allocated to 4 groups with 6 animals per group. Mice of both sexes were distributed evenly among the groups and all animals were kept in single-ventilated type III mouse cages. Animal housing and all handling took place under biosafety level (BSL) 3** conditions. Animals were inoculated subcutaneously into the scruff of the neck with 104 (low-dose group) or 105 (high-dose group) 50% tissue culture infectious doses of either SHUV/2504/3/14 or SHUV/2162/16. The two remaining mice were injected with phosphate-buffered saline and kept as controls (environmental control). During the entire study, all animals were offered water and rodent pellets ad libitum, and were checked for clinical signs and weighed daily by veterinarians. A clinical scoring system comprising body weight, behavior, the appearance of neurological signs, and alterations of eyes or respiration was applied. Up to 3 points were awarded for each category as follows: (1) body weight: 0 points for ≥100% of the weight of the starting day, 1 point for 90–100% of the weight of the starting day, 2 points for 80–90%, and 3 points for 75–80%; (2) behavior: 0 points for active animals that are interested in their environment and show normal grooming behavior, 1 point for slowed movement, 2 points for lethargy and unkempt hair, and 3 points for stupor, hunched back, and ruffled hair; (3) neurological signs: 0 points for no indication of a neurological disease, 1 point for hyperactivity, 2 points for head tilt, and 3 points for ataxia and tremor; (4) eyes and respiration: 0 points for shiny eyes and normal breathing, 1 point for dull eyes, 2 points for conjunctivitis and/or sticky eyes, and 3 points for breath sounds and/or tachypnea. The points awarded per day and animal were added up. A clinical score higher than 10 points, >25% body weight loss since the starting day, and predefined clinical signs such as paralysis of the limbs or dyspnea were defined as endpoint criteria. Animals reaching the endpoint criteria were euthanized immediately and subjected to necropsy. All remaining mice were euthanized after 21 days.
Whole blood samples were collected at 3 and 7 days post inoculation (dpi). At necropsy, serum, whole blood (ethylenediamine tetraacetic acid [EDTA]), and tissue samples were collected, including heart, lung, liver, kidney, urinary bladder, spleen, jejunum, colon, central nervous system (CNS) including cervical spinal cord and brain (half brain, sagittal section), skeletal muscle, and gonads.
Serology
The sera collected at necropsy were analyzed for SHUV-specific antibodies by a previously described multispecies ELISA. 26
Real-Time Reverse Transcription-Polymerase Chain Reaction (RT-PCR)
Tissue samples were homogenized in 1 ml of Modified Eagle Medium using a TissueLyzer instrument (Qiagen, Hilden). Total RNA from 100 µl of the tissue homogenates, 20 µl of the sera, or 20 µl of the whole blood samples were extracted using the KingFisher 96 Flex purification system (ThermoFisher Scientific, Braunschweig) in combination with the NucleoMag VET kit (Macherey-Nagel, Düren) according to the manufacturer’s instructions. The extracts were subsequently tested for SHUV RNA using an S-segment–based generic Simbu serogroup real-time RT-PCR. 7 As positive control, RNA extracted from culture-grown SHUV was included in every PCR run. To control for efficient RNA extraction and amplification, thereby avoiding false negative results, an internal control based on the beta-actin gene was additionally tested. 20
Histopathology, Immunohistochemistry, and RNA ISH
Tissue samples including heart, lung, liver, kidney, urinary bladder, spleen, jejunum, colon, CNS, skeletal muscle, and gonads were collected and fixed in 10% neutral-buffered formalin and trimmed for paraffin embedding. Brain samples were trimmed in a sagittal plane and spleen samples were trimmed in cross section. All tissue samples were cut at 2-3-μm-thick sections and stained with hematoxylin and eosin (HE) for light microscopical examination. On consecutive CNS sections, conventional staining protocols were applied, including luxol fast blue/cresyl violet for detection of myelin sheaths and Nissl substance, von Kossa stain to show mineralization, and Prussian blue reaction for demonstration of ferric iron, indicating hemosiderin. Consecutive slides of the CNS were processed for immunohistochemistry (IHC). Briefly, sections were mounted on adhesive glass slides, dewaxed in xylene, followed by rehydration in descending graded alcohols. Endogenous peroxidase was quenched with 3% hydrogen peroxide in distilled water for 10 minutes at room temperature. Antigen heat retrieval was performed in a steamer for 20 minutes followed by a cooling period. Nonspecific antibody binding was blocked by pure goat normal serum for 30 minutes at room temperature. T-cell and microglia/macrophage markers were applied for all animals. B-cell and astrocyte markers were used for uninfected animals and those with identified meningoencephalomyelitis (details given in Table 1). Immunolabeling was visualized by 3-amino-9-ethylcarbazole substrate (AEC, Dako, Agilent, Santa Clara, California) producing a red-brown signal and sections were counter-stained with Mayer’s hematoxylin. Archived mouse tissue sections were used as positive controls for specificity, including spleen (cluster of differentiation 3 [CD3], paired box protein 5 [PAX5]) and brain (glial fibrillary acidic protein [GFAP], ionized calcium-binding adapter molecule 1 [Iba-1]) sections. As negative control, a normal rabbit serum was used at a dilution of 1:100.
Primary antibodies used for IHC and applications.
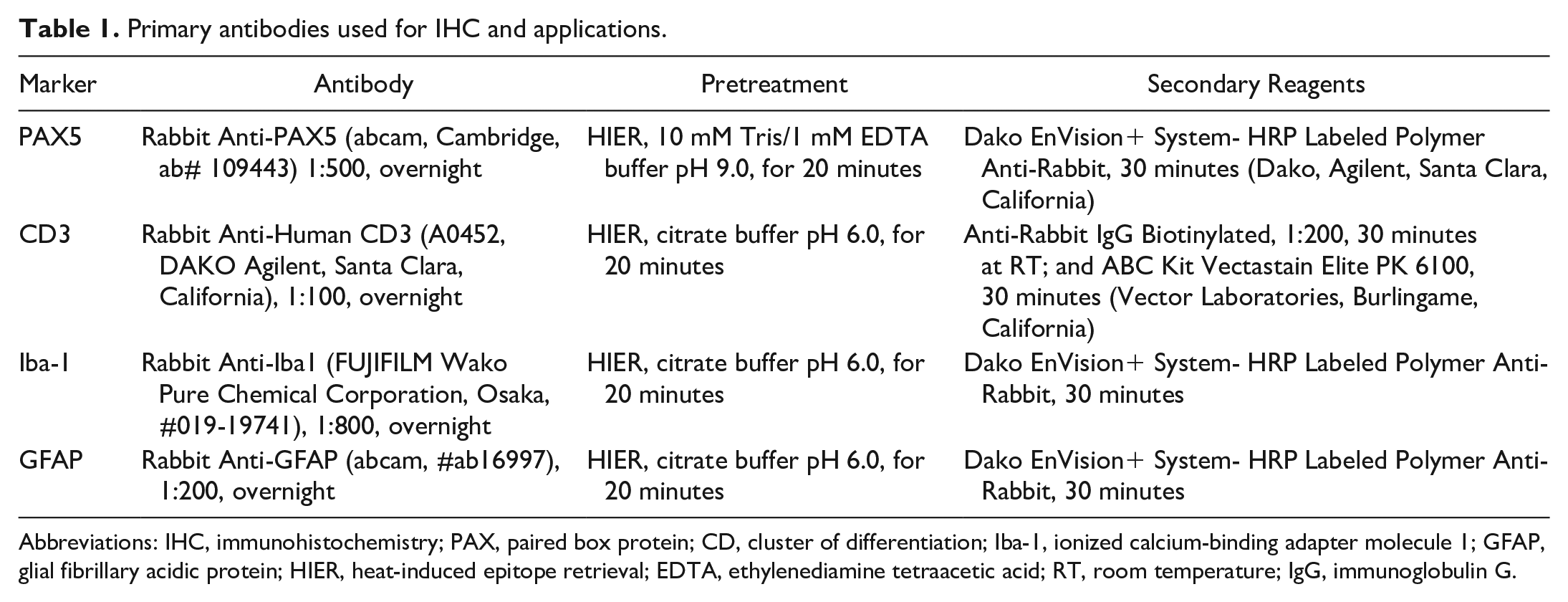

Abbreviations: IHC, immunohistochemistry; PAX, paired box protein; CD, cluster of differentiation; Iba-1, ionized calcium-binding adapter molecule 1; GFAP, glial fibrillary acidic protein; HIER, heat-induced epitope retrieval; EDTA, ethylenediamine tetraacetic acid; RT, room temperature; IgG, immunoglobulin G.
RNA ISH for the detection of SHUV RNA was performed on all tissues of selected animals based on PCR results (animals tested positive with SHUV PCR with quantification cycle (Cq) values below 30 in at least one tissue). The RNAScope™ 2-5 HD Reagent Kit-Red (ACD, Advanced Cell Diagnostics, Newark, California) was applied according to the manufacturer’s instructions. For RNA ISH, RNAScope™ probes were custom-designed for the nucleocapsid protein. As technical assay controls, a positive control probe (peptidylprolyl isomerase B) and a negative control probe (dihydrodipicolinate reductase) were included.
All sides were scanned using a Hamamatsu S60 scanner, and evaluation was done using the NDPview.2 plus software (Version 2.8.24, Hamamatsu Photonics K.K., Iwata City).
Evaluation and Scoring of Changes
Evaluation and interpretation were performed by a board-certified pathologist (DiplECVP) in a masked fashion using the postexamination masking method. 12
HE-stained sections of all tissues were evaluated and described. In particular, for the sagittal brain and spinal cord cross sections, meningeal infiltrates, perivascular cuffing, microglial nodules, neuronal or glial cell degeneration, and necrosis were recorded as “present” or “absent.” Parenchymal mineralization, hemosiderin deposition as well as demyelination, and loss of Nissl substance were also recorded as absent or present. In cross sections of the spleen, the number of secondary follicles was counted in 1.5 mm2 fields. Extramedullary hematopoiesis in the spleen, as well as tingible body macrophages in follicular germinal centers of the spleen, and gut-associated lymphoid tissue (GALT) were described in comparison with control animals. As two different dose levels and virus strains were used, a semiquantitative scoring was initially applied (see Supplemental Material), but did not identify differences between groups. Thus, the focus was laid on describing the pathological changes.
Following RNA ISH, the target cell was identified based on the morphology. The virus RNA distribution of labeled cells in the spleen, intestinal GALT, cerebrum, diencephalon, brain stem, cerebellum (arbor vitae, granular layer, Purkinje cell layer, molecular layer), and spinal cord (gray matter, white matter) was recorded as focal to oligofocal (up to 3 foci), multifocal (>3 foci, area positive <80%), coalescing to diffuse (area positive >80%).
Results
Clinical Findings
After inoculation, all animals were monitored daily for body weight loss and clinical signs (Fig. 1). SHUV 2504/3/14 infection led to body weight loss compared with the weight on the starting day in 2 out of 6 mice after low-dose infection. One animal was euthanized at 6 dpi due to slowed down movement and body weight loss (about 10% within 1 day). No clinical signs apart from marginal weight loss were reported in the remaining animals of this group. High-dose infection resulted in body weight loss in 1 out of 6 mice and an additional animal was euthanized at 9 dpi because of slowed down movement and body weight loss.
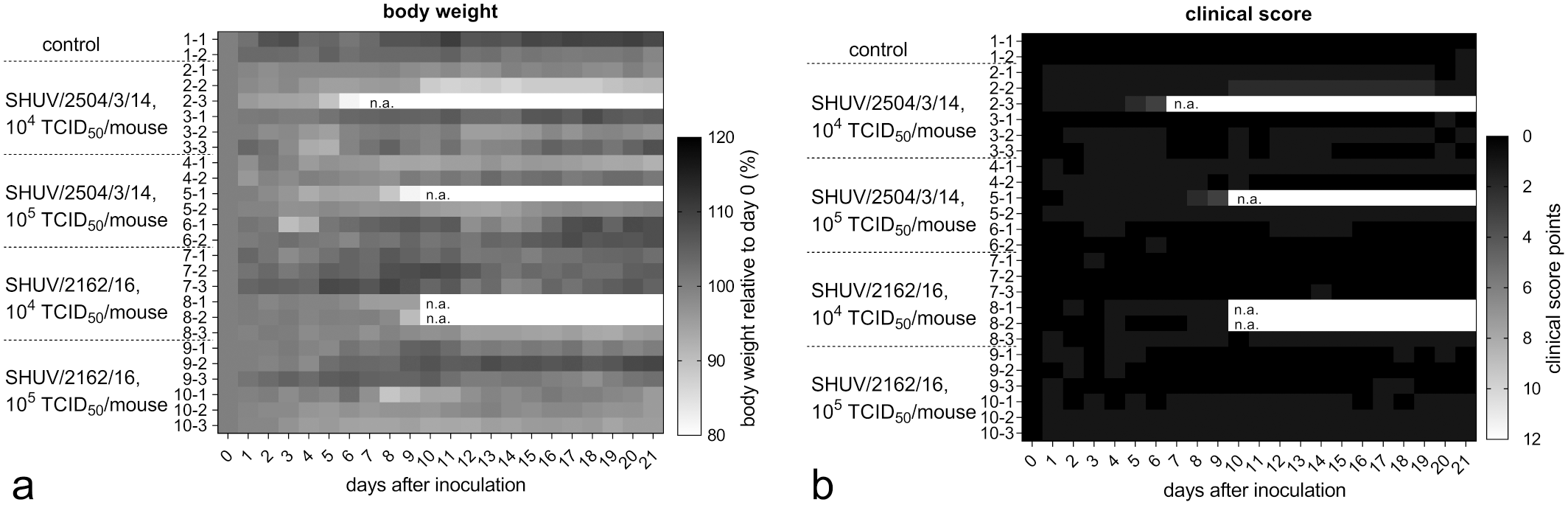
Body weight and clinical score of Ifnar−/− mice infected with Shuni virus, heat map. (a) Independent of the Shuni virus strain and infectious dose used, individual animals showed body weight loss. (b) Clinical scores reflect that no clinical signs were visible except for slowed down movement and body weight loss, scores never exceeded 3 points for any animal.
Low-dose infection with SHUV 2162/16 was associated with body weight loss of about 5% to a maximum of 10% in 3 out of 6 mice (Fig. 1a). Two of these animals were found dead at 10 dpi. High-dose infection led to weight loss in 3 out of 6 mice; no further clinical signs were recorded.
The clinical score did not exceed 3 points in any animal at any time point (Fig. 1b).
Viral RNA Detection and Serology
SHUV RNA was detected by real-time RT-PCR in serum, whole blood, and tissue samples of all groups except the uninfected control mice (Fig. 2). All SHUV-inoculated animals tested positive in at least one sample during the study. SHUV RNA was found in sera collected at 3 dpi in all inoculated mice with exception of animal No. 8-2, and Cq values ranged from about 28 to 31. No differences were seen between both virus strains and dose levels. Viral RNA was also found at 7 dpi in all but 3 animals. In specimens collected at 21 dpi from 20 infected mice, viral RNA was most consistently found not only in the spleen (Cq values 34–38, 9 infected mice) and jejunum (Cq values 32–37, 3 infected mice), but also in the heart, skeletal muscle, spinal cord, and brain (Fig. 2). However, only spinal cord and brain samples yielded lower Cq values compared with full blood samples (Fig. 2). In animals euthanized prematurely, viral RNA was found in a broad spectrum of tissues; however, only the Cq values in the brain (Cq values 16–19) and spinal cord samples (Cq values 16–22) markedly undercut those of the serum or blood (Fig. 2). By ELISA, SHUV-specific antibodies could be detected in one mouse 9 days after high-dose SHUV 2504/3/14 and in one animal 10 days after low-dose SHUV 2162/16 infection, and in all inoculated animals surviving until the end of the study (Fig. 3). The uninfected control animals remained seronegative.
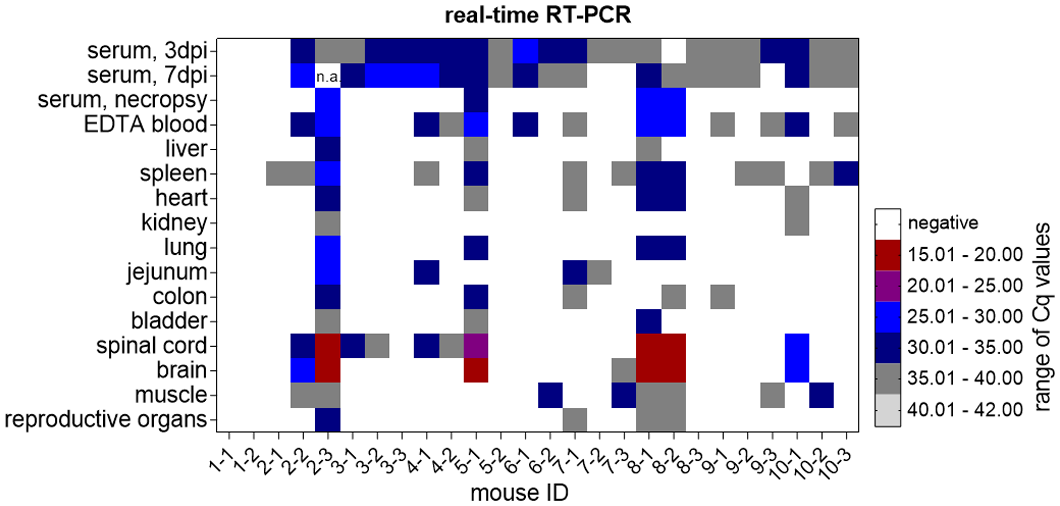
Viral RNA detection in serum, EDTA blood, and tissues of Ifnar−/−mice infected with SHUV reveals low quantification cycle (Cq) values in samples of clinically diseased animals (animal No. 2-3, 5-1, 8-1, 8-2), independent of the strain used for infection, heat map.
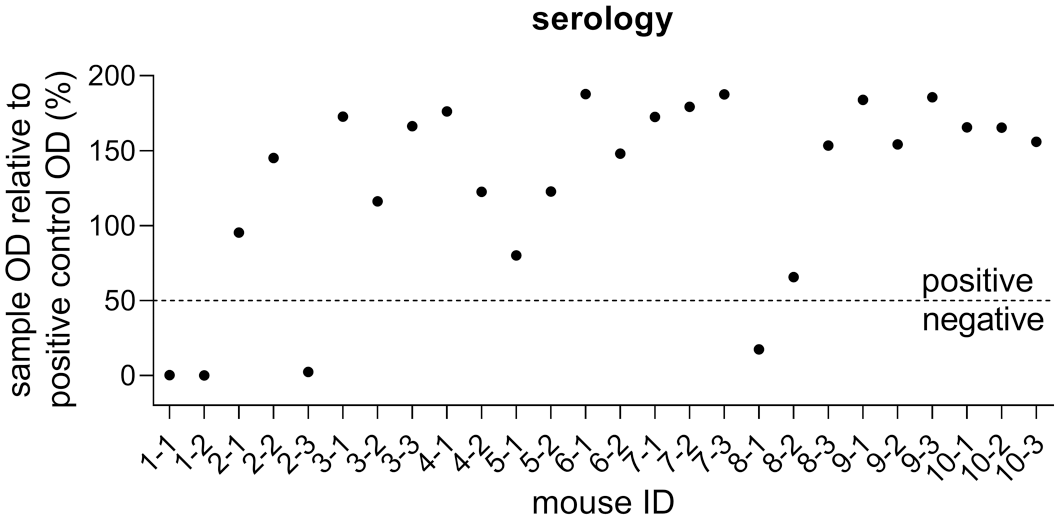
Serological data of Ifnar−/− mice infected with SHUV. A specific antibody response is detectable in all SHUV-inoculated animals with exception of the prematurely lost mice 2-3 and 8-1, while both uninoculated control animals remained seronegative.
Histopathology
Uninfected control animals did not show any histological changes in the heart, lung, liver, kidney, urinary bladder, jejunum, colon, skeletal muscle, gonads, and CNS. Cross sections of control mouse spleens contained up to 2 secondary follicles per 1.5 mm2, minimal extramedullary hematopoiesis, and minimal numbers of tingible body macrophages in germinal centers, all interpreted to be within normal limits. After SHUV infection, no histological changes were found in the heart, lung, liver, kidney, urinary bladder, jejunum, colon, skeletal muscle, and gonads of any mouse.
In the CNS, a range of pathological changes was found after infection in a small number of animals (Table 2). In total, 6 out of 24 mice exhibited histological alterations, affecting 1 or 2 animals per group. No correlation was found between the virus strain and the infectious dose with the presence of histologic changes.
Scoring of histopathologic findings in the CNS of control and SHUV-infected Ifnar−/− mice.
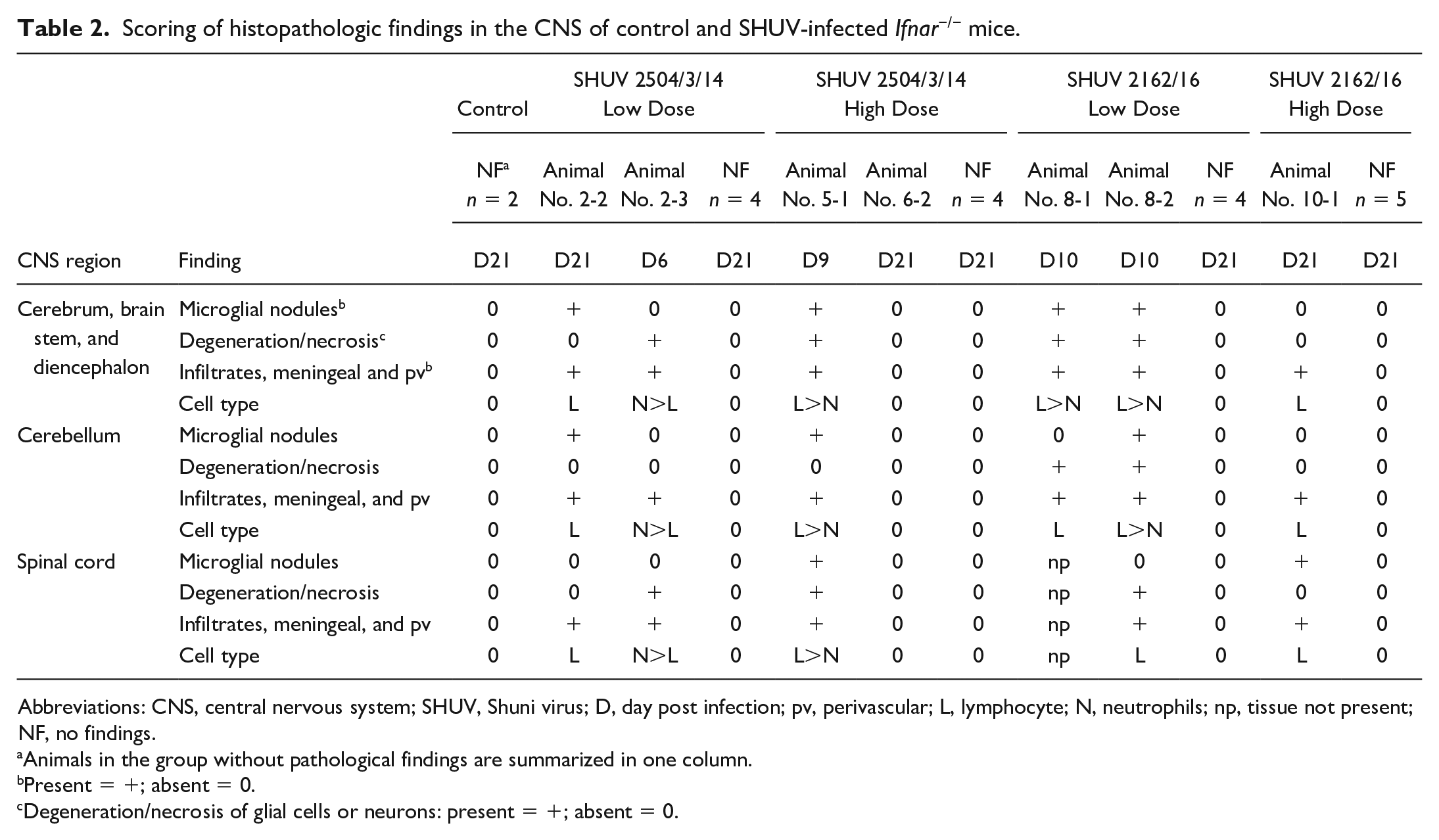
Abbreviations: CNS, central nervous system; SHUV, Shuni virus; D, day post infection; pv, perivascular; L, lymphocyte; N, neutrophils; np, tissue not present; NF, no findings.
Animals in the group without pathological findings are summarized in one column.
Present = +; absent = 0.
Degeneration/necrosis of glial cells or neurons: present = +; absent = 0.
Affected mice had meningeal and perivascular inflammatory infiltrates that never exceeded 4 cell layers in thickness in the brain and spinal cord (Table 2). Neutrophils constituted the dominant population of infiltrates at day 6 (1 animal, Fig. 4a). A higher proportion of lymphocytes and fewer neutrophils were present at 9 and 10 dpi (3 animals). At day 21, only lymphocytes could be identified (2 animals, Fig. 4b). Further, microglial nodules were found at day 9 and later (Table 2, Fig. 4c). The infection also led to neuronal as well as glial cell degeneration and necrosis, including satellitosis and neuronophagia (Fig. 4d), which were detectable in animals euthanized at 6, 9, and 10 dpi (Table 2). Pathological changes were not restricted to a particular region in the brain or spinal cord. However, the most severely affected area was located in the brain stem in the majority of animals and thus selected for the initial semiquantitative scoring of the cerebrum, brain stem, and diencephalon (Supplemental Table S1). Hemosiderin deposition, mineralization, demyelination, or loss of Nissl substance were not found in affected brain and spinal cord sections (details on special stains not shown).
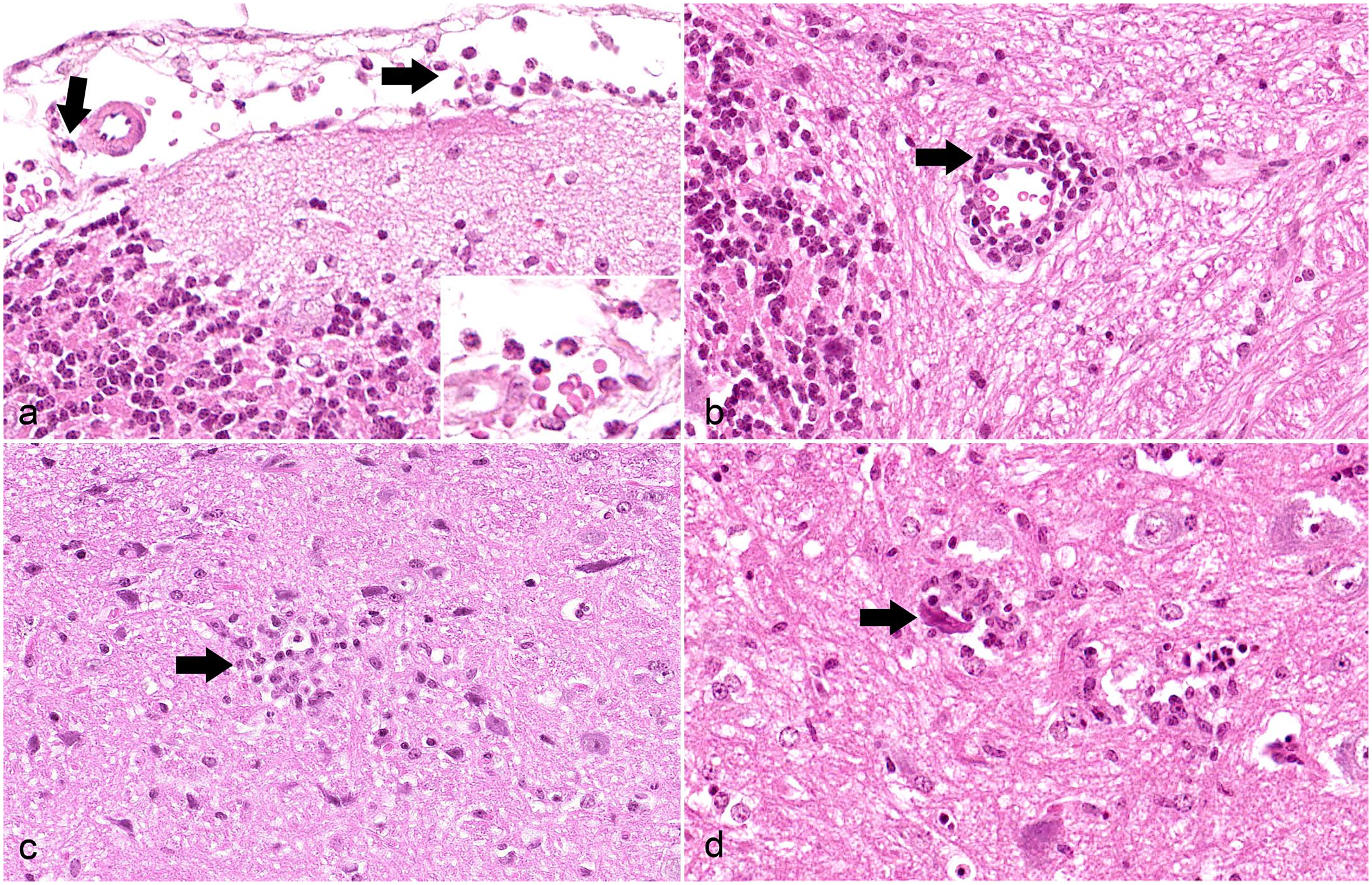
Shuni virus infection, Ifnar−/− mice. Hematoxylin and eosin. (a) Cerebellum. There are mainly neutrophils (arrows) infiltrating the meninges, animal No. 2-3, 6 dpi, inset: higher magnification. (b) Cerebellum. The perivascular space is expanded by lymphocytes (arrow), animal No. 2-2, 21 dpi. (c) Brain stem. Microglial nodule (arrow), animal No. 2-2, 21 dpi. (d) Spinal cord. The degenerate neuron is shrunken and hypereosinophilic (arrow) surrounded by glial cells indicating neuronophagia, animal No. 5-1, 9 dpi.
Regardless of the virus strain and infectious dose used, SHUV infection led to an increased number (≥4) of secondary follicles in the spleen of 20 out of 24 animals, with up to a maximum of 9 secondary follicles per 1.5 mm2. Extramedullary hematopoiesis in the spleen and tingible body macrophages in the spleen and GALT were slightly increased in comparison with controls in up to 2 animals per group (see Supplemental Table S2 for details). In addition, 1 animal showed focal, minimal lymphoid necrosis with few neutrophilic infiltrates in the GALT 6 days after SHUV 2504/3/14 low-dose infection.
Immunohistochemistry
To identify subtle infiltrates and microglial reactions that might have been missed by HE-based examination, we evaluated the CNS of all animals for CD3 and Iba-1–positive cells. B-cell and astrocyte markers were used for uninfected animals and those with identified meningoencephalomyelitis. Immunohistochemical findings in the CNS for CD3, Iba-1, and GFAP IHC are summarized in detail in Supplemental Table S1. Again, no correlation was found between the virus strain and the infectious dose used. In detail, meningeal infiltrates and perivascular cuffs in the brain and spinal cord of the 6 meningoencephalomyelitis-affected mice were composed of CD3-positive T-cells (Fig. 5a). Many Iba-1–labeled cells were also detected in the infiltrates, corresponding with vessel-associated, resident microglia and/or infiltrating macrophages (Fig. 5b). One high-dose SHUV 2504/3/14-infected mouse lacked identifiable lesions in the HE sections, but showed perivascular Iba-1 labeling in the cerebrum. Affected animals also showed CD3-positive T-cells and Iba-1–labeled resident microglia and/or infiltrating macrophages throughout the CNS parenchyma (Fig. 5a, b). As described for the histopathological changes, the most severely affected area was located in the brain stem in the majority of animals. The detection of scattered PAX5-positive B-cells was restricted to 1 animal euthanized 21 days after low-dose SHUV 2504/3/14 infection. Regarding indications for astrocyte reactivity, only 2 animals euthanized at 9 or 10 dpi and 2 mice euthanized at 21 dpi presented with diffuse GFAP labeling in the CNS, indicating a potential reactive state. However, GFAP-positive cells had a slender cell shape (Supplemental Table S1, Fig. 5c, d).
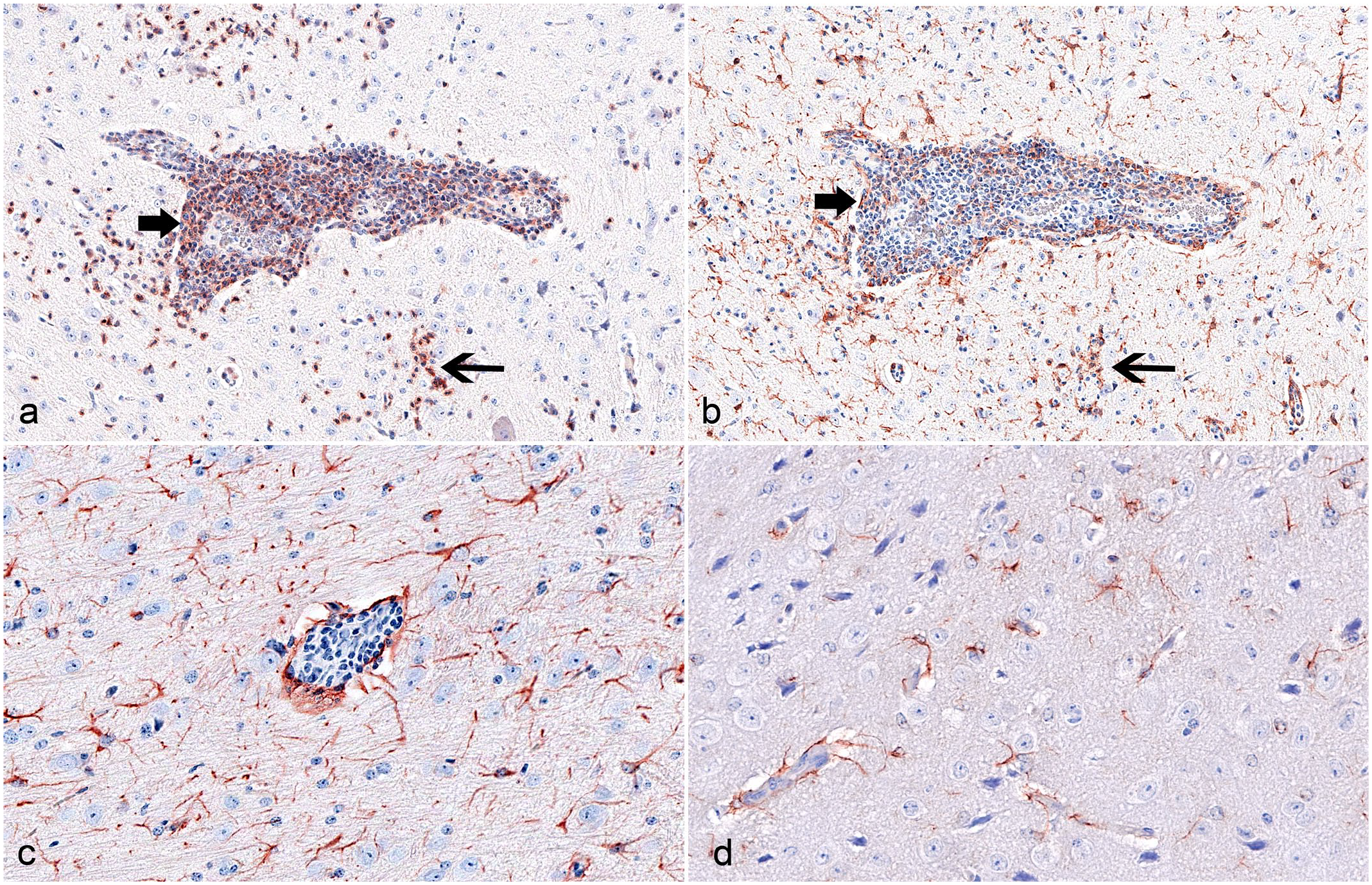
Shuni virus infection, Ifnar−/− mice. Immunohistochemistry for CD3, Iba-1, and GFAP in the brain stem. (a) The affected area contains numerous CD3-positive T-cell infiltrates in the perivascular space (bold arrow) and in the parenchyma (slender arrow), animal No. 2-2, 21 dpi. (b) Consecutive section to (a). There are numerous Iba-1–labeled cells in the perivascular space (bold arrow) and in the parenchyma (slender arrow). (c) There are numerous GFAP-labeled, long and slender-shaped astrocytes, animal No. 2-2, 21 dpi. (d) For comparison, fewer GFAP-positive astrocytes showing slender shape, uninfected control animal.
RNA ISH
RNA ISH was performed on all tissues from animals that tested SHUV PCR positive with Cq values below 30 in at least one tissue (Table 3). SHUV RNA was abundantly detected in the CNS in all 4 animals affected by meningoencephalomyelitis and euthanized or found dead at 6, 9, or 10 dpi. In the CNS, labeled cells, morphologically consistent with neurons and astrocytes, were found in the brain and cervical spinal cord (Fig. 6a–c). Viral RNA was rarely detected in single cells of the GALT and spleen after low-dose SHUV 2504/3/14 and SHUV 2162/16 infection; these were morphologically consistent with macrophages (Fig. 6d). Viral RNA was most consistently and abundantly found in the brain stem and diencephalon (Table 3). The detection of viral RNA in the CNS was associated with inflammatory infiltrates; however, in a few areas, viral RNA was found without evidence of lymphocytic infiltrates. Furthermore, 2 animals euthanized on day 21 had meningoencephalitis without viral RNA detection. Viral RNA labeling (Fig. 7a) was associated with neuronal degeneration, satellitosis, neuronophagia, and necrosis of glial cells (Fig. 7b), as well as presence of numerous, plump Iba-1–labeled resident microglial cells and/or infiltrating macrophages (Fig. 7c). Astrocytes in affected areas were slender (Fig. 7d). Details are given in Table 3.
RNA in situ hybridization for SHUV distribution in the CNS, spleen, and MALT of infected Ifnar−/− mice.
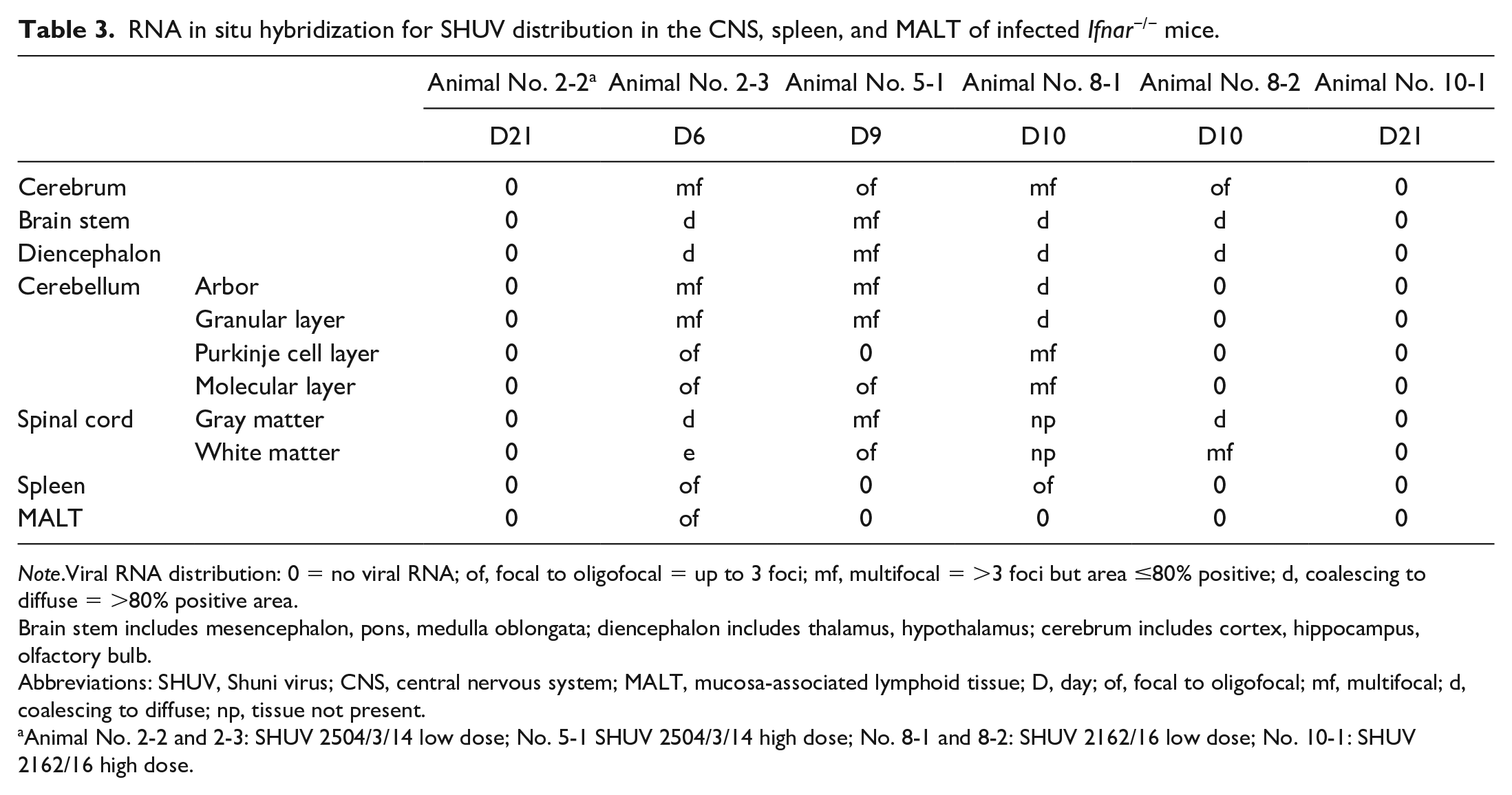
Note.Viral RNA distribution: 0 = no viral RNA; of, focal to oligofocal = up to 3 foci; mf, multifocal = >3 foci but area ≤80% positive; d, coalescing to diffuse = >80% positive area.
Brain stem includes mesencephalon, pons, medulla oblongata; diencephalon includes thalamus, hypothalamus; cerebrum includes cortex, hippocampus, olfactory bulb.
Abbreviations: SHUV, Shuni virus; CNS, central nervous system; MALT, mucosa-associated lymphoid tissue; D, day; of, focal to oligofocal; mf, multifocal; d, coalescing to diffuse; np, tissue not present.
Animal No. 2-2 and 2-3: SHUV 2504/3/14 low dose; No. 5-1 SHUV 2504/3/14 high dose; No. 8-1 and 8-2: SHUV 2162/16 low dose; No. 10-1: SHUV 2162/16 high dose.
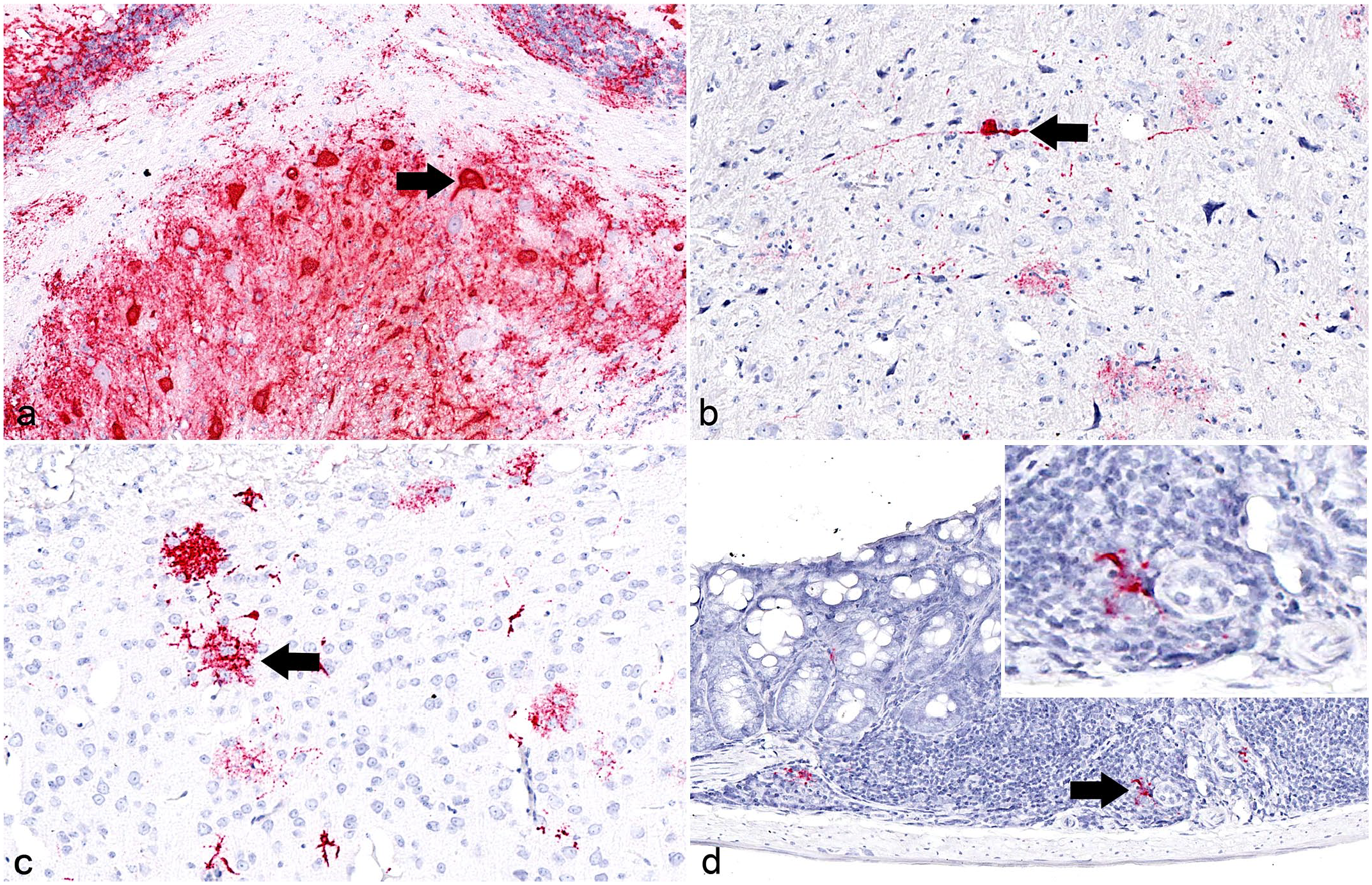
SHUV infection, Ifnar−/− mice. RNA in situ hybridization for detection of SHUV RNA in the brain and large GALT. RNAScope™ probes designed for nucleocapsid protein gene detection (red). (a) Brain, cerebellum. SHUV RNA is detected in neurons (arrow), here mainly in the arbor vitae but also in all layers of the cerebellar gray matter, animal No. 8-1, 10 dpi. (b) Brain, thalamus. Viral RNA is found in neuronal processes (arrow), animal No. 5-1, 9 dpi. (c) Brain, cortex. SHUV RNA detection in cells, morphologically consistent with astrocytes (arrow) of the larger pyramidal cell layer, animal No. 8-1, 10 dpi. (d) Large intestine, GALT. Viral RNA is found in the lymphoid tissue in cells, morphologically consistent with macrophages (arrow), animal No. 2-3, 6 dpi. Inset: higher magnification.
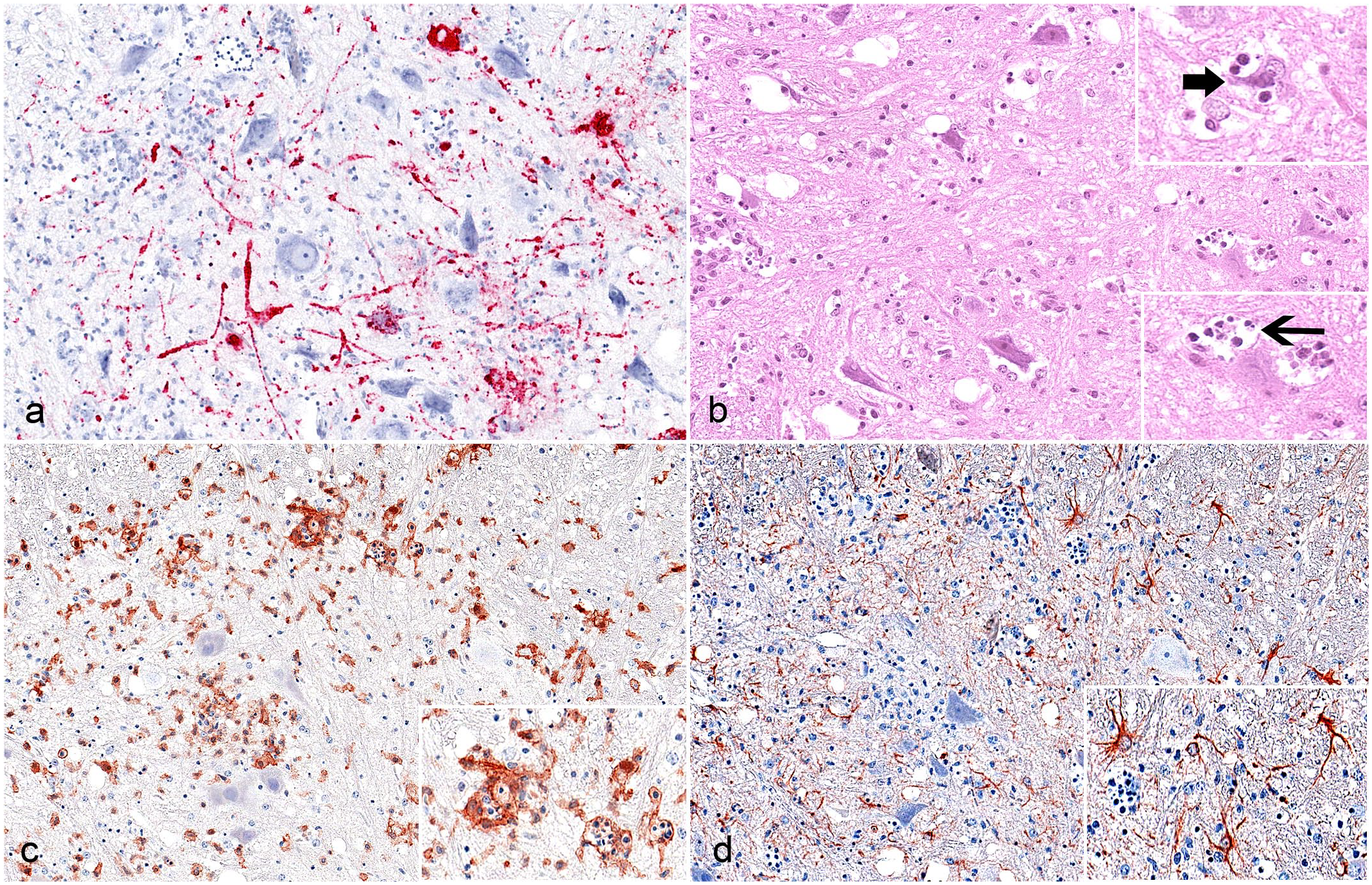
SHUV infection, Ifnar−/− mice, associated changes, cervical spinal cord, animal No. 5-1, 9 days post inoculation. (a) RNA in situ hybridization–based viral RNA detection for nucleocapsid protein gene detection (red) yielded abundant SHUV RNA. (b) A shrunken, degenerate neuron (upper inset, bold arrow) with satellite cells, indicating neuronophagia. Several glial cells are hypereosinophilic and/or show karyopyknosis and karyorrhexis (lower inset, slender arrow). Hematoxylin and eosin. (c) There are numerous Iba-1–labeled resident microglia/infiltrating macrophages exhibiting a plump cell shape. IHC. (d) There are numerous GFAP-labeled, slender-shaped astrocytes. Inset: higher magnification. IHC.
Discussion
Ifnar−/− mice are used as a small animal model for studying the pathogenesis of several orthobunyaviruses, as the neuropathology in these animals appears to be similar to that found in the natural hosts.19,28 Therefore, we have explored the potential of this animal system for studies of SHUV infection as well. Two different virus dose levels were selected and interestingly, they did not result in clear differences in the parameters of body weight, clinical findings, viral RNA detection, serology, or pathology in this mouse model.
As reported for experimentally SHUV-infected cattle, 17 the overall clinical picture was inconspicuous in most inoculated mice; however, 4 out of 24 Ifnar-/- mice developed fatal disease and 6 out of 24 mice developed a meningoencephalomyelitis. For Peribunyaviridae, a lethality rate below 100% is not uncommon, as Oropouche virus infection led to a lethality rate of 85% in 1-day-old BALB/c mice, and SBV yielded a lethality rate of about 20% in Ifnar−/− mice.15,27
The histopathologic changes in the CNS of experimentally SHUV-infected mice resembled those found in cattle after natural and experimental infection,6,17 showing meningeal infiltrates and perivascular cuffs, dominated by either neutrophils or CD3-positive T-cells. In addition, many Iba-1–labeled cells were found perivascularly, corresponding with vessel-associated, resident microglia and/or infiltrating macrophages. Further, numerous T-cells and Iba-1–labeled cells were also found throughout the CNS parenchyma, the latter indicating microgliosis. Rarely, PAX5-positive B-cells were identified. In cattle, “gliosis” was described after natural, but not experimental SHUV infection and astrogliosis was found after SBV infection.6,9,17 In the present study, we aimed to get an indication for astrocyte reactivity by exploring the distribution pattern and the morphology following GFAP labeling of astrocytes. In few mice affected by meningoencephalitis, GFAP labeling was found diffusely in the CNS indicating reactivity, but the cell shape remained slender. At this point, it should be noted that GFAP upregulation and hypertrophy are useful, but are now regarded as insufficient markers for astrocyte reactivity that need to be complemented by additional markers. 4 Thus, the interpretation of the reactive state of astrocytes is limited and needs further evaluation. To detect parenchymal mineralization, hemosiderin deposition, as well as demyelination and loss of Nissl substance, special stains were applied on brain slides. In contrast to infection with the closely related SBV, hemorrhage, mineralization, demyelination, and loss of Nissl substance were not found in the brain.6,9,17
RNA Scope™–based SHUV detection in infected Ifnar−/− mouse tissues demonstrated abundant viral RNA in neurons and astrocytes at 6, 9, and 10 dpi. By contrast, in cattle the viral RNA detection was restricted to rare, individual neuronal cell processes. This difference between species is most likely related to the fact that cattle were evaluated only at 21 dpi and viral RNA detection was limited. 17 Data for earlier time points after infection are not yet available for cattle. In turn, most of the mice were likewise investigated as late as 21 dpi in the present study and either no or only marginal levels of viral RNA could be detected in those animals. It is not known whether mice that did not show meningoencephalomyelitis or viral RNA in the CNS at 21 dpi were able to completely prevent neuroinvasion or effectively cleared the virus. Localized foci of inflammation and viral RNA may have been missed and thus underestimated the neuroinvasive capacity of SHUV in the small animal model. Interestingly, RNA ISH identified viral RNA in macrophages of the spleen and GALT of mice, the latter associated with focal, minimal lymphoid necrosis and neutrophilic infiltrates in the GALT in one animal. Additional changes in lymphoid tissues were nonspecific, with increased numbers of secondary follicles in the spleen in most infected animals, and increased extramedullary hematopoiesis and more tingible body macrophages in a few mice. Extramedullary hematopoiesis can increase in response to inflammation, and increased tingible body macrophages are a distinctive although nonspecific feature of follicular hyperplasia. Remarkably, viral RNA was also detected in SHUV-infected cattle by PCR in the spleen and intestine. 17 The phenomenon of viral RNA detection in lymphoid tissues has been previously reported for other viruses closely related to SHUV, such as SBV. After experimental SBV inoculation, viral RNA could be detected in cattle and sheep several weeks after infection in the lymphoid tissues, most often in the mesenteric lymph nodes.28 –30
To date, the mechanisms for brain invasion are not known for SHUV. Studies on the related Oropouche virus indicate that the virus enters the CNS in mice by at least 3 different routes: by the neural route, by crossing the blood-brain barrier, and by the Trojan horse mechanism using infected leukocytes and monocytes.14,16 In the present study, the brain stem was the most severely affected area in the majority of animals showing meningoencephalitis. Viral RNA was most consistently and most abundantly found in the spinal cord, brain stem, and diencephalon. Further, viral RNA was detected in macrophages in the GALT and spleen in mice by ISH. In cattle, SHUV RNA was found by PCR in the intestine and spleen. 17 Also in cattle, inflammation was most consistently found in the periductal matter of the cerebral aqueduct, where viral RNA was detected by ISH. 17 Thus, our findings indicate that SHUV may also take several routes for neuroinvasion, a phenomenon that needs to be investigated in more detail in the future.
As expected, both viruses, the “encephalitis” SHUV 2162/16 and the “malformation” strain SHUV 2504/3/14, did not differ in any parameter tested in our mouse model lacking a functional IFN-α/β receptor. Comparable results have already been obtained using other NSs-deficient mutant orthobunyaviruses including SBV. 10 Therefore, the effect of NSs on the pathogenesis in the natural host cannot be evaluated using the Ifnar-/- mouse model. However, these mice represent a suitable model to study the neuropathogenicity and the impact of virulence determinants beyond NSs, as the postmortem lesions mirror those seen in cattle. Studies on several viruses have shown the suitability of this model to test vaccines and to study the adaptive, humoral, and cellular immune responses, comparable with those achieved in the immunocompetent model. 11 In concrete terms, Ifnar-/- mice can be used to study: (1) the mechanism for neuroinvasion, (2) the early infection phase to clarify a potential prevention of neuroinvasion versus effective clearing, (3) astrocyte reactivity, and (4) the role of macrophages in the early and late phase of infection. However, the generation of a serially passaged, mouse-adapted SHUV with increased and/or more consistent virulence could allow a broader range of investigations, especially in immunocompetent mouse models.
The present study highlights the importance of histopathological evaluation and the value of techniques that enable lesion-associated pathogen detection. The latter is the prerequisite to distinguish indirect immunopathological processes from direct lesions triggered by infection. SHUV RNA has been detected only recently in the cerebrospinal fluid from human patients with neurologic disease. 13 However, target cells and neuropathology have not been described so far for humans. Well-characterized animal models may reveal clues to the potential pathogenesis of this disease in humans. Further, SHUV should be considered as etiologic differential diagnoses for meningoencephalomyelitis not only for wildlife and domestic animals but also for humans in endemically affected regions.
In conclusion, we show that Ifnar−/− mice are susceptible to SHUV infection and can develop fatal disease. The histopathological changes correspond to the findings described for cattle. Thus, our mouse model is particularly beneficial for evaluating virulence determinants in the pathogenesis. The current study provides first insights into the pathogenesis, but a deeper understanding of the virulence determinants and pathogenesis is urgently needed, especially as a further spread of the insect-transmitted pathogen into hitherto unaffected regions should be expected.
Supplemental Material
sj-pdf-1-vet-10.1177_03009858231155402 – Supplemental material for Characterization of experimental Shuni virus infection in the mouse
Supplemental material, sj-pdf-1-vet-10.1177_03009858231155402 for Characterization of experimental Shuni virus infection in the mouse by Angele Breithaupt, Franziska Sick, Natalia Golender, Martin Beer and Kerstin Wernike in Veterinary Pathology
Footnotes
Acknowledgements
We thank Bianka Hillmann, Silvia Schuparis, and Gabriele Czerwinski for their excellent technical assistance and the animal caretakers for their dedicated work.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was funded by the German Federal Ministry of Food and Agriculture (BMEL) through the Federal Office for Agriculture and Food (BLE), grant number (281B101816).
Supplemental Material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
