Abstract
Mammary tumor–associated amyloidosis (MTAA) in dogs is characterized by amyloid deposition in the stroma of mammary adenoma or carcinoma; however, the amyloid precursor protein remains unknown. We attempted to identify an amyloid precursor protein and elucidated its etiology by characterizing 5 cases of canine MTAA. Proteomic analyses of amyloid extracts from formalin-fixed paraffin-embedded specimens revealed α-S1-casein (CASA1) as a prime candidate and showed the N-terminal truncation of canine CASA1. Both immunohistochemistry and immunoelectron microscopy showed that amyloid deposits or fibrils in MTAA cases were positive for CASA1. Reverse transcription-polymerase chain reaction and quantitative polymerase chain reaction revealed the complete mRNA sequence encoding CASA1, whose expression was significantly higher in the amyloid-positive group. The recombinant protein of the N-terminal–truncated canine CASA1 and the synthetic peptides derived from canine and human CASA1 formed amyloid-like fibrils
Amyloid has a cross-beta structure and is generated by the misfolding of precursor proteins. Amyloidosis is the general term for disorders characterized by amyloid deposition and can be either localized or systemic. Amyloidosis is classified by the specific protein deposition. To date, 42 types of amyloidosis have been identified in humans; however, there are only 11 types in animals. 5 Although comparative pathological studies using animal cases are effective for an integrated understanding of the pathogenesis of amyloidosis and for accurate risk assessment, 4 only a few types of animal amyloidosis have been recognized.
Most cases of primary breast amyloidosis in humans originate from immunoglobulin light-chain (AL) amyloidosis.18,34 Meanwhile, mammary AL amyloidosis is uncommon in animals.19,48 Cattle, rats, and dogs develop spherical amyloid bodies called corpora amylacea in the mammary gland lumen,3,31,40 and α-S2-casein has been identified as the precursor protein in cattle.26,42 In addition to α-S2-casein, the amyloidogenicity of other milk proteins, such as κ-casein, β-casein, α-lactalbumin, and β-lactoglobulin, has been identified
Mammary tumor–associated amyloidosis (MTAA) in dogs has been reported as a localized mammary amyloidosis.40,41,45 A casein protein has been implicated in the MTAA pathology; however, the amyloid precursor protein has not been identified conclusively.40,41 In this study, we performed a mass spectrometry–based proteomic analysis on 5 cases of canine MTAA and identified CASA1 as the amyloid precursor protein. Subsequently, we evaluated the amyloidogenicity of canine and human CASA1 via
Methods
Case Information and Histopathological Analyses
Tissues collected from 5 dogs with MTAA were analyzed (Supplemental Table S1). All dogs were female, 4 of which were spayed. Four were miniature dachshunds, and 1 was a Shiba dog. Their ages ranged from 9 years and 8 months to 13 years and 6 months, with an average age of 10 years and 11 months. All dogs were reared in households. The mammary masses were collected at private veterinary hospitals, fixed in formalin, and sent to the authors’ laboratory for pathological testing.
Formalin-fixed tissues were embedded in paraffin, cut into 3-μm sections, and subjected to hematoxylin and eosin staining, Congo red staining, and immunohistochemistry (IHC) analyses. Mammary gland tumors were histologically diagnosed by 2 Japanese College of Veterinary Pathologists–licensed veterinary pathologists (Murakami and Nomura) according to the classification described by Goldschmidt et al. 12 Amyloid deposits were identified as Congo red–stained materials showing yellow to green birefringence under polarized light.
Tissue Microdissection
Protein was extracted from formalin-fixed paraffin-embedded (FFPE) specimens of cases 1–5. Each specimen was cut into 8-µm sections, collected on polyethylene naphthalate membrane glass slides (Thermo Fisher Scientific, Tokyo, Japan), and stained with Congo red. Congo red–positive lesions (1,000,000 μm2) were manually dissected using 31-gauge needles (Becton, Dickinson and Company, Franklin Lakes, NJ) under a SMZ745 stereomicroscope (Nikon, Tokyo, Japan). The tissues dissected were collected in 40 µL of lysis buffer (containing 10-mM Tris, 1-mM ethylenediaminetetraacetic acid [EDTA], and 0.002% Zwittergent 3–16; Calbiochem, San Diego, California), boiled for 90 minutes, sonicated in a water bath for 90 minutes, and subjected to the following enzyme treatments.
Liquid Chromatography/Tandem Mass Spectrometry
The microdissected samples from cases 1–5 were divided into 2 equal portions; each sample was digested with 1.5 μg of trypsin (Mass Spectrometry Grade; FUJIFILM Wako, Osaka, Japan) or chymotrypsin (Mass Spectrometry Grade; Promega, Madison, Wisconsin) at 37°C for 6 hours, followed by reduction with dithiothreitol. Liquid chromatography/tandem mass spectrometry (LC/MS/MS) of the digested peptides was performed using high-performance LC interfaced with a mass spectrometer (LTQ Orbitrap XL; Thermo Fisher Scientific). The digested peptides were separated using an Inertsil ODS-3 column (φ1 mm × 100 mm, 3 μm; GL Sciences Inc., Tokyo, Japan) and eluted with a linear gradient of 2%–40% buffer B (100% acetonitrile and 0.1% formic acid) in buffer A (0.1% formic acid in water) at a flow rate of 0.05 mL/min. MS and MS/MS data were acquired using the data-dependent top 10 method.
MS/MS Ions Search
The resulting MS/MS data were collated with the theoretical fragmentation patterns of tryptic or chymotryptic peptide sequences of proteins registered in the NCBIprot database using the Mascot Server (Matrix Science Inc., Boston, Massachusetts). “Trypsin” and “chymotrypsin” were used as the in silico proteolytic enzymes. To search for possible chemical modifications, data were first subjected to an error tolerant search, and then oxidation of methionine and tyrosine, phosphorylation of serine and threonine, deamidation of asparagine and glutamine, and pyroglutamylation of glutamine and glutamic acid, which were significantly detected, were used as the in silico variable modifications. To identify exact fragment sites of the protein, MS/MS data were also analyzed using a database created by the authors (Supplemental Table S2); in silico proteolytic enzymes “semiTrypsin” and “none” in the Mascot Server’s drop-down list were selected to identify nontryptic and nonchymotryptic peptides, respectively. Statistically significant proteins (score > 40) and peptides (score > 20) were extracted using Mascot’s probability-based scoring algorithm.
Gene Sequencing Analysis
Because the amino acid sequences of canine CASA1 (cCASA1) registered in National Center for Biotechnology Information (NCBI; accession: XP_013974153.1) and UniProt (accession: J9P7W6) were computationally predicted, reverse transcription-polymerase chain reaction (RT-PCR) and sequencing were performed to determine the cCASA1 encoding gene sequence (
Real-Time RT-PCR Analysis
Real-time RT-PCR was performed to compare expression levels of the
Preparation of Recombinant Truncated cCASA1
A recombinant protein encompassing residues 81–183 of cCASA1, which was determined to be in the amyloidogenic region based on the tissue proteomic analysis, was prepared. The
Production of Anti-CASA1 Antiserum
Antisera were obtained by immunizing mice with recombinant cCASA181–183. Eight-week-old female BALB/c mice weighing 17–22 g (Japan SLC, Hamamatsu, Japan) were immunized intraperitoneally with 50 µg of cCASA181–183 with complete Freund’s adjuvant (Wako, Osaka, Japan) and boosted intraperitoneally using the same antigens with incomplete Freund’s adjuvant (Wako) on days 14, 28, and 39 after the first immunization. Mice were euthanized 3 days after the administration of the last booster, and whole blood was drawn to obtain serum. The study protocol described herein was approved by the Animal Care and Use Committee of the Tokyo University of Agriculture and Technology (approval number 30-83), and the animal experiments were performed according to the principles of the Declaration of Helsinki.
Immunohistochemistry
Anti-cCASA181–183 antiserum was used as the primary antibody. For antigen retrieval, specimens were autoclaved in 10-mM sodium citrate buffer (pH 6.0) at 121°C for 10 minutes prior to reaction with the primary antibody. Horseradish peroxidase–labeled polymer antimouse immunoglobulin antibodies (Dako, Carpinteria, California) were used as secondary antibodies. A diaminobenzidine-4HCl substrate kit (Liquid DAB + Substrate Chromogen System; Dako) and hematoxylin were used for color development and counterstaining, respectively. Normal mouse serum was used instead of the primary antibody for the negative control.
Ultrastructural Electron Microscopy
Transmission electron microscopy (TEM) and immunoelectron microscopy (IEM) of the formalin-fixed tissue of case 1 were performed as described previously. 24 IEM was performed using the anti-cCASA181–183 antiserum and a secondary antibody labeled with gold nanoparticles (EMGMHL10; BBI Solutions, Crumlin, UK). For the negative control, the primary antibody was omitted. Ultrathin sections with or without immunostaining were stained using EM stainer (Nisshin EM, Tokyo, Japan) and lead citrate and examined under a JEM-1400 electron microscope (JEOL, Tokyo, Japan) at 80 kV.
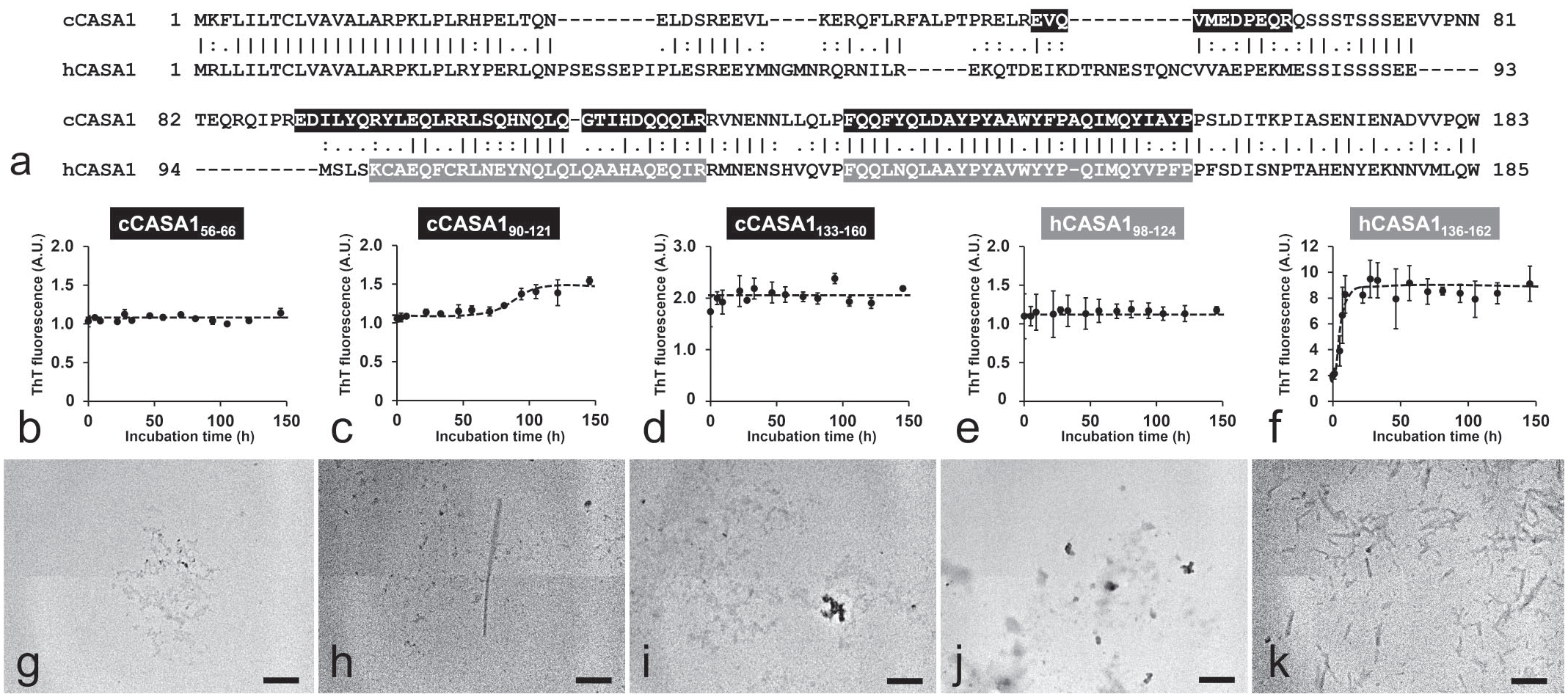
Pairwise Sequence Alignment of Canine and Human CASA1
Pairwise sequence alignment of cCASA1 and human CASA1 (hCASA1; UniProt accession no. P47710) was analyzed using the EMBOSS Needle. 25
Western Blot Analysis
Western blotting was performed on amyloid extracts of FFPE specimens from cases 1 to 3. Three 10-μm-thick sections were deparaffinized, suspended in 70% formic acid, and incubated at 20°C with agitation at 400 rpm. After 7 days of incubation, the soluble components were collected and evaporated. The dried pellets were dissolved in a sample buffer containing 6-M urea and subjected to SDS–PAGE followed by western blotting. cCASA181–183 was used as the positive control. Anti-cCASA181–183 antiserum and anti-mouse immunoglobulin G horseradish peroxidase-linked antibodies (GE Healthcare) were used as primary and secondary antibodies, respectively. Bands were detected using an ECL Prime detection kit (GE Healthcare) and visualized using a STAGE-5100 analyzer (AMZ System Science, Tokyo, Japan).
In Vitro Fibrillization of Recombinant Protein and Synthetic Peptide Fragments
cCASA181–183 was suspended at 1 mg/mL in F12 medium without phenol red (Research Institute for the Functional Peptides, Yamagata, Japan; pH 7.0) containing 0.02% sodium azide. The mixtures were incubated at 37°C with agitation at 800 rpm and were used for further assays.
Synthetic peptides (>98.0% purity) were obtained from GL Biochem (Shanghai, China). Three regions of cCASA1 (cCASA156–66, EVQVMEDPEQR; cCASA190–121, EDILYQR YLEQLRRLSQHNQLQGTIHDQQQLR; cCASA1133–160, FQQFYQLDAYPYAAWYFPAQIMQYIAYP) and 2 regions of hCASA1 (hCASA198–124, KCAEQFCRLNEYNQLQLQAA HAQEQIR; hCASA1136–162, FQQLNQLAAYPYAVWYYPQI MQYVPFP) were prepared. The preparation and concentration measurements of the peptide solutions were performed as described by Sawashita et al. 35 Each synthetic peptide was suspended in an F12 medium (100 µM) containing 5% dimethyl sulfoxide and 0.05% sodium azide, without phenol red. Mixtures without synthetic peptides were used as the negative control. The mixtures were incubated at 37°C with agitation at 800 rpm and were used for further assays.
Thioflavin T Binding Assay
After incubation, recombinant cCASA181–183 or the synthesized peptide mixtures were collected at arbitrary intervals. Then, the mixtures were adjusted to a total volume of 52 µL comprising 2 μL of mixtures and 50 μL of 1-μM thioflavin T (ThT)/50-mM glycine-NaOH buffer (pH 9.0). After briefly mixing the solutions at 20°C–25°C, ThT fluorescence intensities were measured using a fluorescence spectrophotometer (FT-8300; JASCO Corporation, Tokyo, Japan) at λex = 450 nm and λem = 482 nm. Each mixture was measured in triplicate, and the average was calculated.
Amyloid formation of the synthetic peptides was determined using Congo red staining. After 7 days of incubation, 10 μL of each reaction mixture was placed onto a glass slide and evaporated. The dried smears were stained with Congo red and examined using polarized microscopy.
Examination of In Vitro Amyloid Fibrils Using TEM
After 7 days of incubation (after incubation), each mixture was subjected to TEM analysis. cCASA181–183 before incubation was also analyzed. Aliquots of the suspensions were placed on 200-mesh copper grids supported by a Formvar-carbon film, blotted to remove excess water, and air-dried. The grids were stained with EM Stainer (Nisshin EM) and examined under a JEM-1400 electron microscope (JEOL) at 80 kV.
Prediction of Structure and Aggregation Propensity
The amino acid sequence of cCASA1, which was determined based on gene sequencing, and hCASA1 (UniProt accession: P47710) was analyzed using SignalP 4.0, 29 AmylPred 2, 44 PASTA 2.0, 46 and ZipperDB. 11
Statistical Analysis
Data were analyzed using Student’s
Results
Histopathological and Ultrastructural Characteristics of MTAA Cases
The following 5 cases were histopathologically diagnosed: case 1, carcinoma solid; case 2, adenoma simple; case 3, carcinoma solid; case 4, carcinoma tubulopapillary; case 5, adenoma simple (Supplemental Table S1). In all cases, moderate to severe amyloid deposits were observed as Congo red–positive amorphous lesions in the interstitium (Fig. 1a). In cases 2 and 5, nodular amyloid deposits replaced most tumor tissues, showing an amyloidoma-like lesion (Fig. 1b). Congo red–stained amyloid deposits showed yellow to green birefringence under polarized light as amyloid-specific reactions (Fig. 1a, b: inset). TEM of 1 sample revealed ultrastructurally dense fibrils, which were 7–10 nm in diameter, within the amyloid lesion (Supplemental Figure S2).
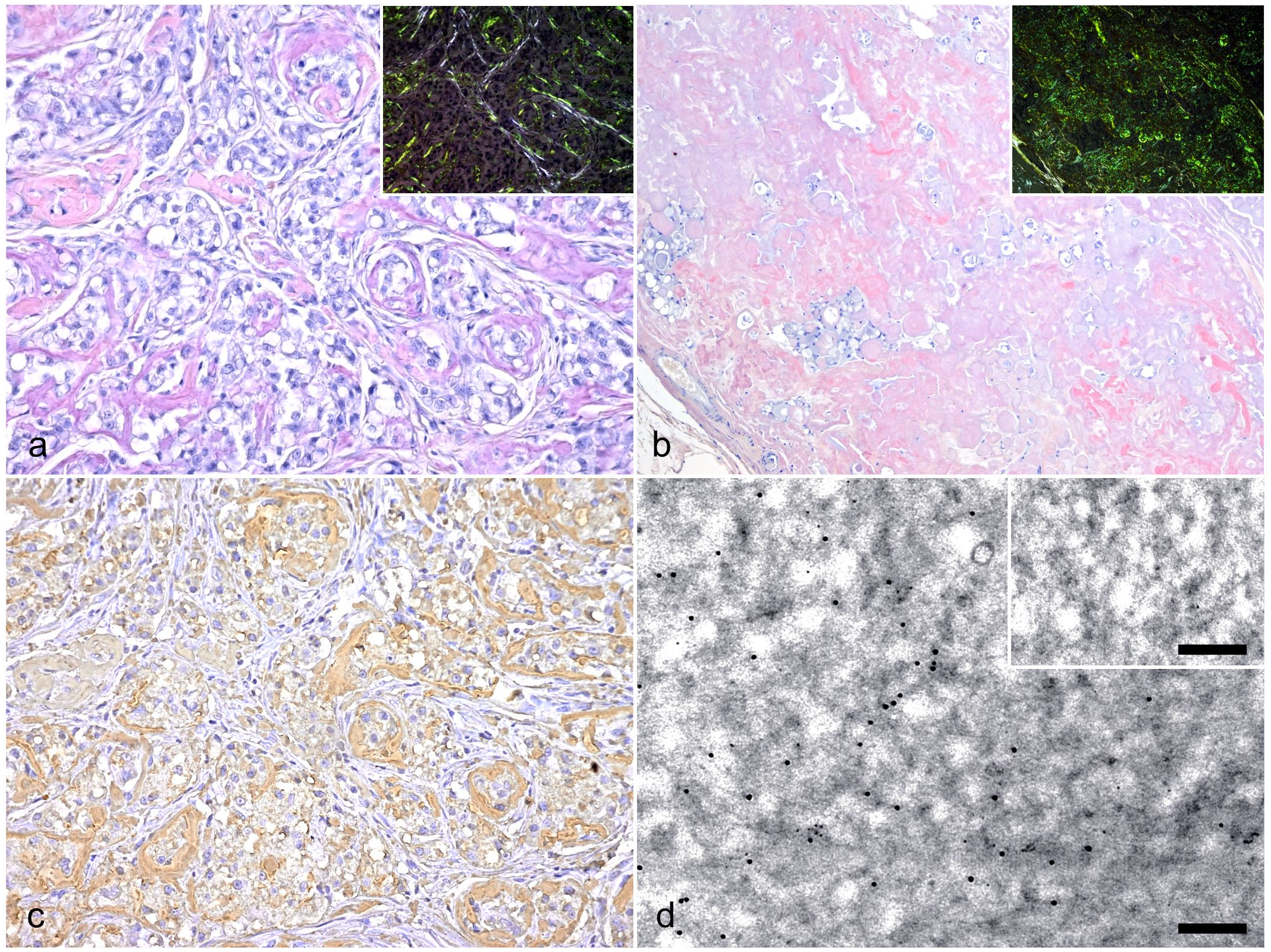
Histological features and immunological detection of amyloid deposits. (a, b) Amyloidosis, mammary gland, dog. Representative images of amyloid deposits stained with Congo red. Inset: same region viewed under polarized light. (a) Amyloid deposits were observed in the tumor stroma (case 3). (b) Nodular amyloid deposits replaced the tumor tissues (case 2). (c) Immunohistochemistry of serial section with Fig. 1a. Amyloid deposits immunolabeled with anti-cCASA181–183 antiserum (case 3). (d) Amyloid fibrils immunolabled with the anti-cCASA181–183 antiserum identified with transmission electron microscopy. Immunoreaction was visualized by 10-nm gold particle–labeled secondary antibodies. Few nonspecific immunoreactions were observed in the negative control (inset) (case 1). Bars, 200 nm.
Identification of CASA1 as an Amyloid Precursor Protein
Microdissected amyloid deposits were tryptic-digested and analyzed using LC-MS/MS to identify the major protein component. CASA1 was detected at prominently high levels in all cases (Table 1 and Supplemental Table S5). Apolipoprotein A-VI, apolipoprotein A-I, and lambda light chain, which have been reported to be amyloidogenic
Proteins identified from microdissected amyloid deposits.
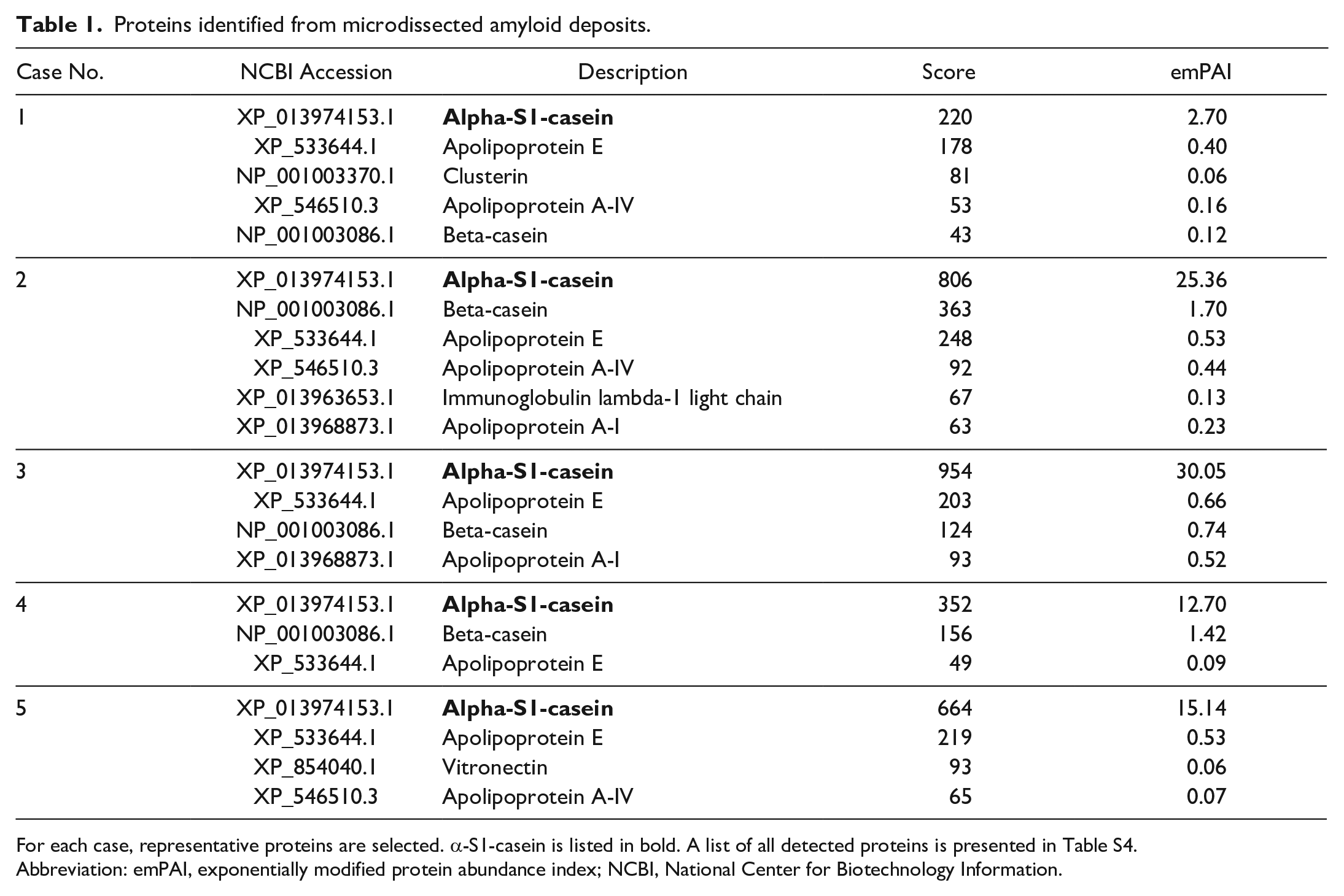
For each case, representative proteins are selected. α-S1-casein is listed in bold. A list of all detected proteins is presented in Table S4.
Abbreviation: emPAI, exponentially modified protein abundance index; NCBI, National Center for Biotechnology Information.
IHC analysis revealed that amyloid deposits were positive for anti-cCASA181–183 antiserum (Fig. 1c). IEM confirmed that amyloid fibrils were labeled with the anti-cCASA181–183 antiserum (Fig. 1d). In both IHC and IEM, similar positive reactions were not observed in the negative controls.
Gene Sequencing Analysis of cCASA1
Sequencing results on
Elucidation of the Amyloidogenic Mechanism of cCASA1
Real-time PCR using a primer set targeting the full
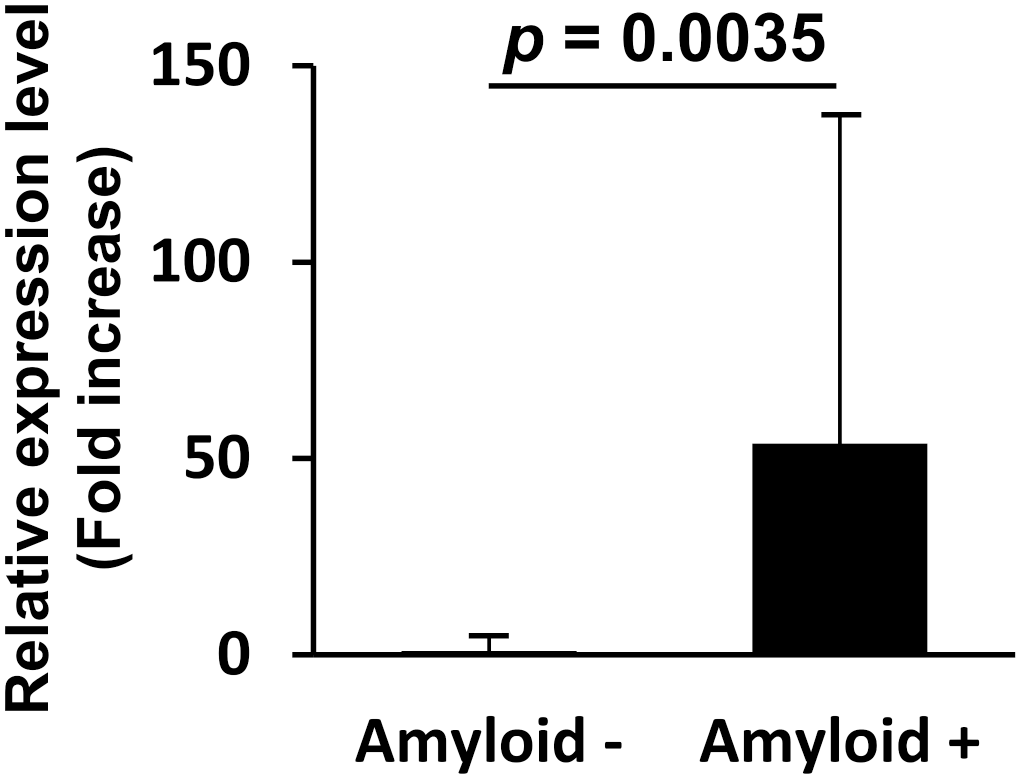
The average expression level of the newly discovered
To identify the amyloidogenic regions in CASA1, MS/MS data were again analyzed using a unique database containing 2 amino acid sequences translated from the newly identified
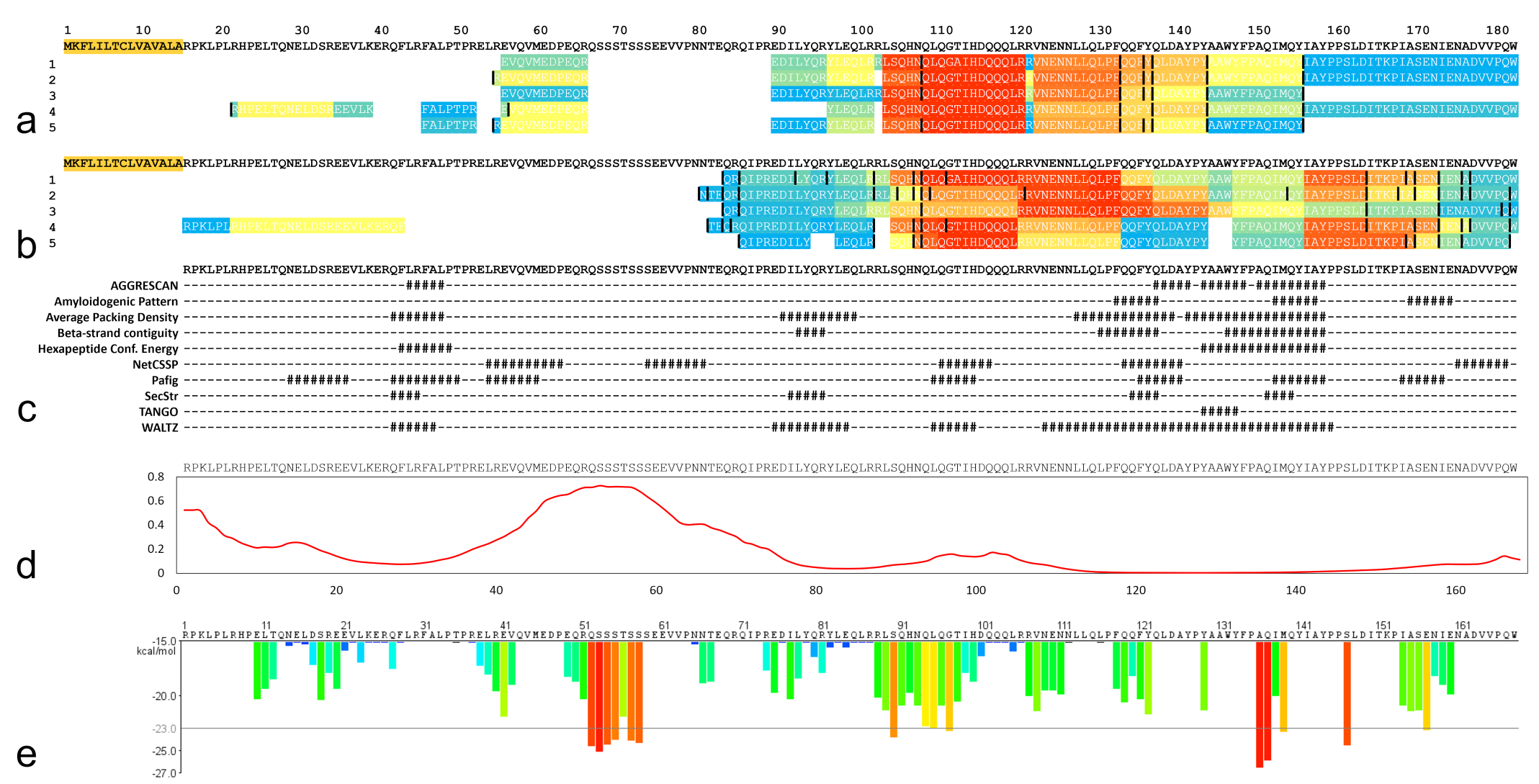
Amyloidogenic factors of cCASA1. (a, b) Tryptic peptides (a) and chymotryptic peptides (b) identified by mass spectrometry. The top row shows the amino acid sequence of CASA1 newly identified in this study. Sheer numbers on the left indicate case numbers. Yellow highlights indicate signal sequences. Each bar was colored according to the total number of peptides detected in 5 cases, with warmer colors (orange-red) indicating a higher number of significantly detected peptides. The maximum number of detections (a) with tryptic digestion (shown in red) was 35, 45, 49, 32, and 34 (cases 1–5, respectively) and (b) with chymotryptic digestion was 48, 53, 67, 34, and 28 (cases 1–5, respectively). Black bars indicate unpreferred truncation sites by the respective enzymes. (c) Prediction of the amyloidogenic potential of cCASA1 using AmylPred2. Regions with high amyloid propensity were indicated as #. (d) Disorder probability profile of cCASA1 based on PASTA2.0. The y-axis represents arbitrary units, with higher values representing domains that are predicted to have increased disorder. (e) Amyloid nucleating (zipper) sequence prediction for cCASA1 by ZipperDB. Bars are in warmer colors (orange-red) with energy below the indicated energetic threshold of −23 kcal/mol (gray line) and are predicted to form fibrils.
The locations of chemical modifications are shown in Supplemental Figure S3. In all samples, deamidation of asparagine and glutamine, oxidation of methionine, and pyroglutamylation of the N-terminal peptide fragment were frequently observed. Pyroglutamylation was observed frequently, especially in Gln84, Gln86, Glu90, Gln95, and Gln109, suggesting gradual fragmentation in these regions.
Of the 2 CASA1 sequences determined based on gene sequencing, CASA1 with Thr113 was detected in cases 2, 3, and 5; CASA1 with Ala113 in case 1; and both CASA1s were detected in case 4 (Supplemental Tables S6 and S7). These results support the genetic analysis results that CASA1 in case 4 is heterozygous for the p.Thr113Ala mutation; they also indicate that CASA1 in case 1 is homozygous for the p.Thr113Ala mutation. Furthermore, the N-terminal sequence (RPKLPLRHPELTQ) detected in case 4 is not present in the NCBI-registered CASA1 (XP_013974153.1). This supports that CASA1 in amyloid deposits is derived from the complete protein-encoding sequence of the canine
Western blotting of the extracted amyloids from 3 cases was performed to confirm the fragmentation of CASA1. Bands of approximately 13 kDa were identified in all 3 cases (Fig. 4), indicating the presence of truncated cCASA1 proteins.
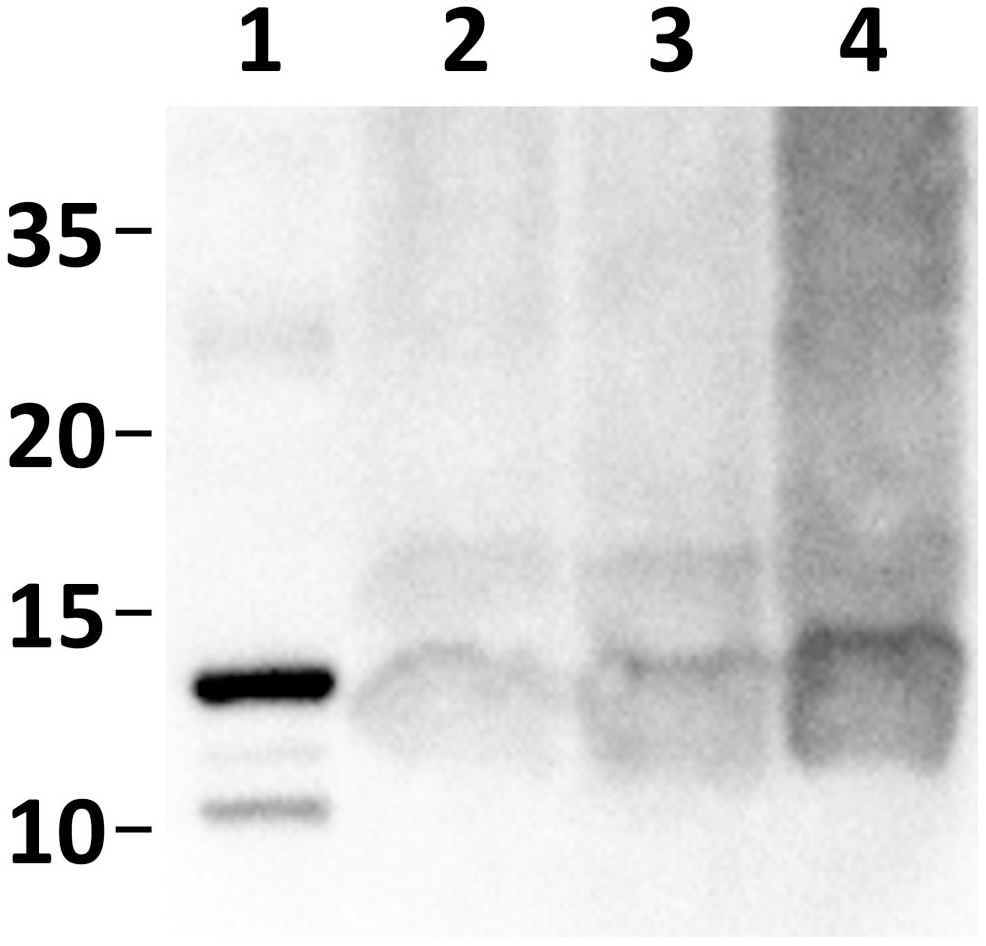
Representative western blot images using anti-cCASA181–183 antiserum. Bands around 13 kDa were observed in formic acid extracts from the formalin-fixed paraffin-embedded tissue of the amyloid-positive cases: lane 1, recombinant cCASA181–183; lane 2, extract from case 1; lane 3, extract from case 2; and lane 4, extract from case 3.
The amino acid sequence translated from the newly identified
AmylPred analysis 44 of hCASA1 predicted the presence of regions with amyloid-forming propensity, particularly in the C-terminal regions, similar to cCASA1 (Supplemental Figure S4). On the other hand, PASTA analysis 46 predicted lower disorderedness of the N-terminal sequence than cCASA1. ZipperDB analysis 11 indicated that hCASA1 has a broader range of multiple zipper sequences than cCASA1.
Demonstration of CASA1 Amyloidogenicity In Vitro
To demonstrate the amyloidogenicity of CASA1, the amyloidogenic potential of recombinant truncated cCASA1 was evaluated
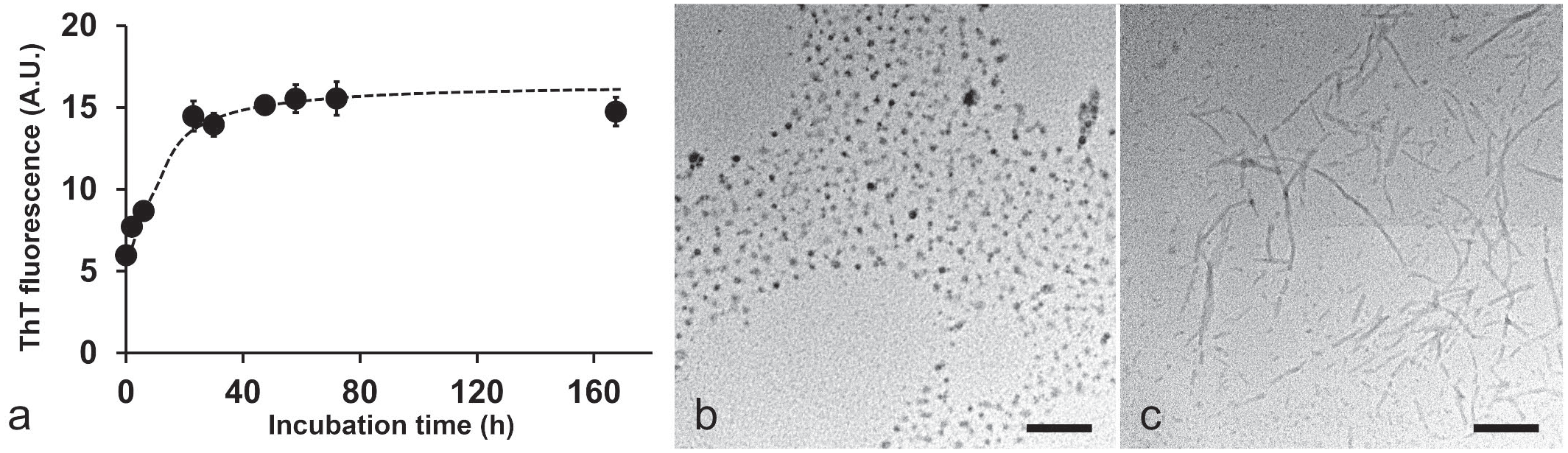

To identify the region with high amyloidogenic propensity within CASA1, the
Discussion
Based on mass spectrometry results, CASA1 was detected as a significant component of amyloid deposits in all 5 MTAA samples. Amyloid deposits were positive for CASA1 by IHC and IEM, and the recombinant C-terminal fragment and shorter synthesized peptides derived from CASA1 formed amyloid-like fibrils
We sequenced the complete
Mass spectrometry revealed that CASA1 was cleaved around Asn81–Arg85. Additionally, in amyloid deposition, peptides on the N-terminal side of CASA1 were less detectable than those on the C-terminal side. Western blot analysis results indicated that the ~13-kDa fragment is a major component of cCASA1-derived amyloids. Furthermore, we showed that N-terminal–truncated cCASA1 formed amyloid-like fibrils
AmylPred analysis indicated that the central and C-terminal regions of sequence (approximately Glu90-Tyr159) of cCASA1 had higher amyloidogenicity than the N-terminal region, with the exception of perhaps Gln42-Leu48. Indeed, in the ThT assay using synthetic peptides, cCASA190–121 exhibited amyloidogenicity under physiological conditions; however, cCASA156–66 did not form amyloid fibrils. PASTA analysis indicated several disordered regions in the N-terminal region of cCASA1. α-Synuclein is known to enhance amyloid aggregation by truncation of the C-terminal disordered region. 36 Similarly, the disordered N-terminal region of CASA1 may hinder amyloid formation of the full-length protein, with its proteolytic removal resulting in amyloidogenesis of the remaining truncated protein.
The mammary gland, which constantly secretes a high concentration of proteins, is at a high risk of conformational diseases.16,17 Various milk proteins, including α-S2-casein,
42
κ-casein,
43
β-casein,
28
α-lactalbumin,
30
and β-lactoglobulin,
13
have been reported to exhibit amyloidogenicity
In various types of amyloidosis, misfolded proteins exhibit cytotoxic properties. 23 For example, the milk protein κ-casein–derived protofibrils and fibrils are cytotoxic to PC-12 cells, 9 indicating that the protofibrils or fibrils formed during protein aggregation inherently have cytotoxic properties. We observed amyloidoma-like lesions in 2 of 5 MTAA cases. Amyloidoma, observed in AL amyloidosis, is a lesion characterized by nodular amyloid deposition without cell proliferation due to amyloid-induced cell death. 47 In addition, we propose that the cCASA1-derived amyloids associated with the amyloidoma-like lesions observed in this study are also cytotoxic, but additional studies are needed to prove this hypothesis. It should be noted that the amyloid deposits in this study occurred not in the lumen but in the tissue interstitium. In contrast, the mammary corpora amylacea of dogs, rats, and cattle, which likely contains casein-derived amyloids, mainly forms in the mammary glandular lumen;3,31,40 however, its cytotoxicity has not yet been reported. Corpora amylacea, also known as wasteosomes, is thought to sequester and detoxify the amyloid and waste products within a glycan structure. 32 In contrast, interstitial amyloid is less likely to be sequestered by other secretions and would be expected to result in amyloidoma-like lesions.
Identifying and characterizing diseases that widely target animals can contribute to a better understanding of their etiologies in humans. For example, in 1983, insulin-associated amyloidosis was reported as iatrogenic amyloidosis induced by continuous local insulin injection
38
because the
In conclusion, we demonstrated the amyloidogenicity of CASA1
Supplemental Material
sj-pdf-1-vet-10.1177_03009858221148511 – Supplemental material for Identification of novel amyloidosis in dogs: α-S1-casein acquires amyloidogenicity in mammary tumor by overexpression and N-terminal truncation
Supplemental material, sj-pdf-1-vet-10.1177_03009858221148511 for Identification of novel amyloidosis in dogs: α-S1-casein acquires amyloidogenicity in mammary tumor by overexpression and N-terminal truncation by Tomoaki Murakami, Toshisuke Kaku, Kaori Tsukakoshi, Susumu Iwaide, Yoshiyuki Itoh, Miki Hisada, Kohji Nomura, Rikako Kubo, Kazunori Ikebukuro, Yukiko Sassa-O’Brien and Fuyuki Kametani in Veterinary Pathology
Supplemental Material
sj-xlsx-2-vet-10.1177_03009858221148511 – Supplemental material for Identification of novel amyloidosis in dogs: α-S1-casein acquires amyloidogenicity in mammary tumor by overexpression and N-terminal truncation
Supplemental material, sj-xlsx-2-vet-10.1177_03009858221148511 for Identification of novel amyloidosis in dogs: α-S1-casein acquires amyloidogenicity in mammary tumor by overexpression and N-terminal truncation by Tomoaki Murakami, Toshisuke Kaku, Kaori Tsukakoshi, Susumu Iwaide, Yoshiyuki Itoh, Miki Hisada, Kohji Nomura, Rikako Kubo, Kazunori Ikebukuro, Yukiko Sassa-O’Brien and Fuyuki Kametani in Veterinary Pathology
Supplemental Material
sj-xlsx-3-vet-10.1177_03009858221148511 – Supplemental material for Identification of novel amyloidosis in dogs: α-S1-casein acquires amyloidogenicity in mammary tumor by overexpression and N-terminal truncation
Supplemental material, sj-xlsx-3-vet-10.1177_03009858221148511 for Identification of novel amyloidosis in dogs: α-S1-casein acquires amyloidogenicity in mammary tumor by overexpression and N-terminal truncation by Tomoaki Murakami, Toshisuke Kaku, Kaori Tsukakoshi, Susumu Iwaide, Yoshiyuki Itoh, Miki Hisada, Kohji Nomura, Rikako Kubo, Kazunori Ikebukuro, Yukiko Sassa-O’Brien and Fuyuki Kametani in Veterinary Pathology
Footnotes
Acknowledgements
We sincerely thank Dr Kazufumi Kawasako (Rakuno Gakuen University) for motivating us to initiate this study. We are greatly indebted to Prof. Keiichi Noguchi and Prof. Satoshi Nakaba (Tokyo University of Agriculture and Technology) for their technical assistance with the LC/MS/MS analysis and TEM analysis. We also greatly appreciate the helpful suggestions and comments of Prof. Naomi Hachiya (Tokyo Metropolitan Industrial Technology Research Institute) and Prof. Yuki Kobayashi (Nihon University).
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was supported by the JSPS KAKENHI (grant no. 20K15660); the Adaptable and Seamless Technology Transfer Program through target driven R&D (grant no. JPMJTM20CY) from the Japan Science and Technology Agency (JST); and the Program on Open Innovation Platform with Enterprises, Research Institute and Academia (OPERA) from JST.
Supplemental Material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
