Abstract
Nine distinct papillomaviruses (Lambdapapillomavirus) have been described in domestic and nondomestic cats, but not in cheetahs. These viruses have been associated with cutaneous papillomas or plaques, bowenoid in situ carcinomas, feline cutaneous squamous cell carcinomas (SCC), feline sarcoids, and oral (often sublingual) papillomas. Fourteen cheetahs from the AfriCat foundation (Namibia) and one from the Ann van Dyk Cheetah center (South Africa) presented with sublingual lesions reminiscent of sublingual papillomas. Two animals were biopsied and the histopathology revealed benign proliferative epithelial lesions with prominent thickening of the overlying squamous epithelium. Throughout the squamous epithelial layers were cells with nuclear enlargement, irregularity of the nuclear membranes and cell contours, focal hyperchromasia of the nuclei, and perinuclear halos, reminiscent of a virus-associated process as seen in papillomavirus infections. Thirteen more cheetahs were sampled and the tissue snap frozen for molecular characterization. Amplification and sequencing of the papillomavirus L1, E6, E7, and E1 gene regions was achieved with modified primers. Maximum likelihood phylogenetic analyses revealed all 15 cheetah papilloma samples were 99.99% genetically similar and closely related to, but genetically distinct from any known felinepapillomaviruses. All cheetahs were FIV and FeLV negative. The results suggest the samples identified in this study can be considered a previously undescribed or novel feline papillomavirus and the authors propose “Acinonyx jubatus papillomavirus type 1” (AjPV-1), within the Lambdapapillomavirus 1 genus (Family: Papillomaviridae).
Papillomaviruses belong to the family Papillomaviridae and consist of approximately 30 to 35 genera.2,15 These double-stranded, circular-genome, DNA viruses are usually species-specific, and may be limited to certain tissues and areas of the body, specifically the epithelial cells of the skin; squamous epithelium of the mouth, anus, and female genitals; and the columnar epithelium of the cervix.2,26 However, detection of human papillomavirus (HPV) in oral squamous cell carcinomas (SCC) of domestic cats and bovine papillomavirus (BPV) in sarcoids of lions suggests cross-species transmission that challenges this dogma. 15
Traditional classifications of these viruses have used the conserved L1 gene, and sequences differing by more than 10% from any known papillomavirus type are considered a new type. 2 This choice is supported by a large body of current research, where most new isolates have genomic differences larger than 10%. 2 Currently, any variation within the L1 gene between 2 viruses of between 10% and 40% constitutes a different viral type and more than 40% difference, a different genus. 15 The predominant papillomavirus types have been described in humans (approximately 170), however, additional genera, species, and types may be described as more research on papillomaviruses in animals is conducted. 15 At least 9 distinct feline papillomaviruses (FPV) have been described by full genome sequencing. 15 Five of these were described in domestic cats 16 and a further 4 described from nondomestic felids.1,18,25 An in-depth description of these viruses in a previous study, based on detection of the E6, E7, E1, L1, and L2 oncogene regions, showed that all belonged to the lambdapapillomavirus genus. 15
In domestic cats (Felis catus), FPV most commonly causes skin lesions which can include cutaneous papillomas (CP),5,14 cutaneous viral plaques (CVP), 27 bowenoid in situ carcinomas (BISC), 10 feline cutaneous squamous cell carcinomas (SCC), 13 and feline sarcoids 21 while oral papillomas (often sublingual) have also been described.12,26 Oral SCC is a common tumor in cats but currently there is very little evidence that FPV plays a pertinent role in their development, which differs from known associations between papillomaviruses and SCC in humans.8,15,22 In nondomestic felids, FPVs have been implicated in the formation of cutaneous plaques and cutaneous SCC in snow leopards (Panthera uncia), sarcoids in African lions (Panthera leo) and ventral tongue papillomas in snow leopards, bobcats (Lynx rufus), Asiatic lions (P. leo persica), clouded leopards (Neofelis nebulosa), a Florida panther (Puma concolor coryi), a Canadian lynx (Lynx canadensis), and an Asiatic desert cat (Felis silvestris ornata).1,17,28 BISCs have not yet been described in any nondomesticated felid and the potential role of FPV in the development of oral SCC in snow leopards is uncertain. 15 There have been very few reports of FPV in non-domestic felids in southern Africa. In a study defining the manifestations of feline immunodeficiency virus in 2 African lion populations, oral papillomas were described and their link to FIV questioned. 20 A previous report by the authors described the partial molecular characterization of oral FPV in cheetahs and 1 lion in southern Africa.3,23 Here, we expand on the initial description in cheetahs and report on the possible long-term effects of this infection.
Materials and Methods
Samples were collected from cheetahs originating from a captive breeding facility in South Africa (The Ann van Dyk Cheetah center [S 25° 40’ 25.1” E 27° 55’ 25.4”]), and a rescue sanctuary in Namibia (AfriCat Foundation [S 20º 51’ 59” E 16º 38’ 22”]). The author (GS) was involved with annual health examination at these facilities during 2002 to 2016. 23 Any cheetah with sublingual papillomas had a small biopsy taken from the affected area. The papillomas on the ventral aspect of the tongue consisted of flattened small prominent papillomatous lesions, 2 to 4 mm in diameter (Fig. 1A, B). The initial biopsies (2005) were immersed in 10% buffered formalin after collection. All subsequent samples were kept in liquid nitrogen upon collection, and thereafter stored at -80°C as soon as they were brought to the laboratory. The project was approved by the National Zoological Gardens of South Africa’s Research and Ethics Committee (Project no. P11/07).
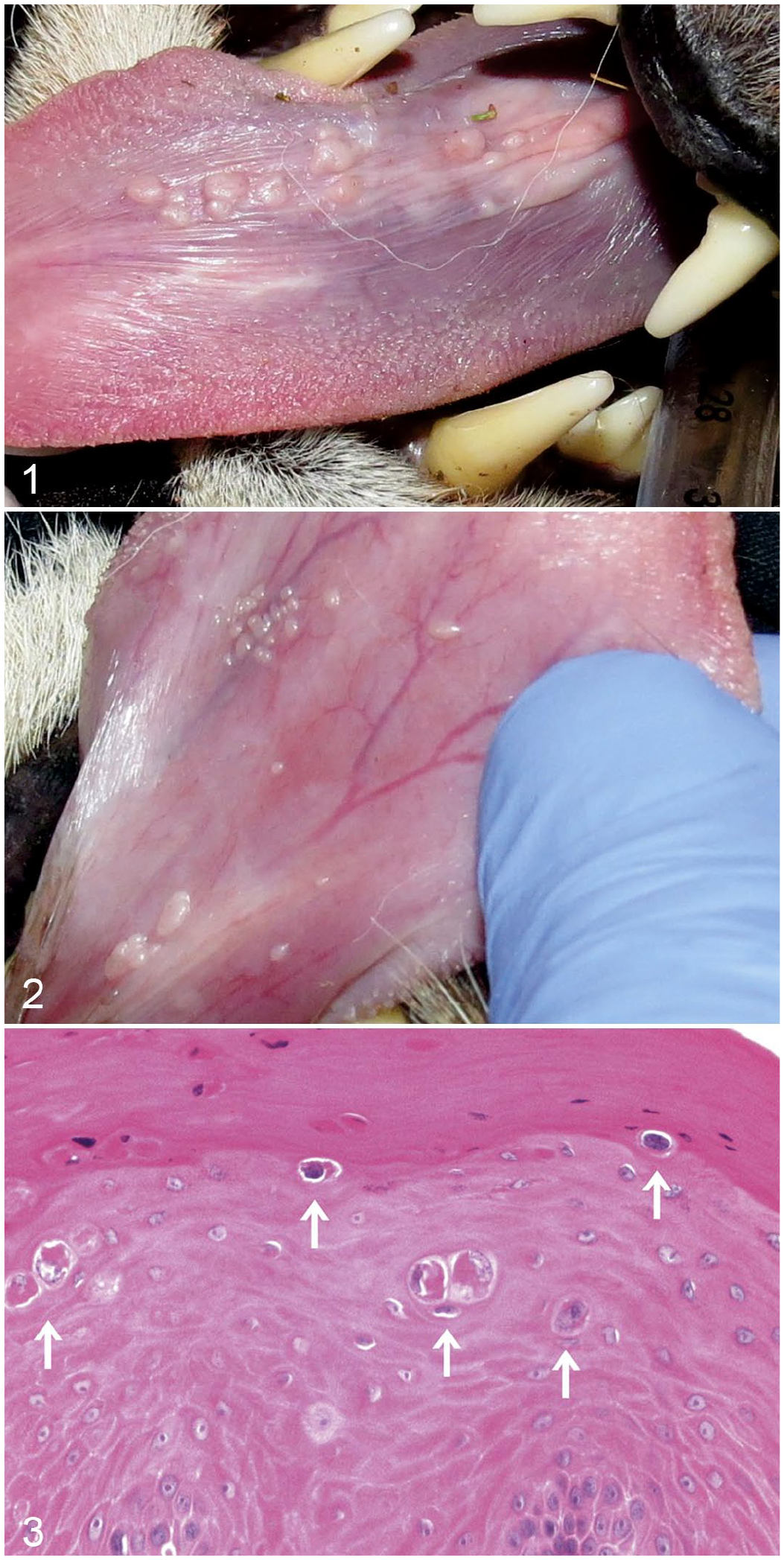
Papillomas, tongue, cheetah. Papillomatous lesions form discrete or coalescing nodules on the ventral surface of the tongue.
The oral proliferative tissue was embedded in paraffin blocks whereafter sections, 3 to 4 µm thick, were cut from them and stained with haematoxylin and eosin according to standard histology laboratory protocols.
Molecular Detection
A total of 15 samples from 13 individuals were submitted to the National Zoological Gardens of South Africa in Pretoria for detection, identification, and molecular characterisation of any papillomaviruses present in the biopsied samples (Supplemental Table S1). DNA was extracted using either the ZR Genomic DNATM Tissue MiniPrep kit (Zymo Research, California, USA) or the Epicenter MasterPure ™ Complete DNA and RNA purification kit (Whitehead Scientific, Gauteng, South Africa) according to the manufacturer’s tissue isolation protocol.
Molecular detection of papillomaviruses by amplification of partial L1, E5, E6, and E7 viral oncogenes using previously published primers6,7,9,11 was unsuccessful. Therefore, primers for the L1, E6, E7, and E1 gene regions were modified (Table 1) 19 by either changing the length, target region, and/or replacement of degenerate sites by incorporation of deoxyinosine (I). Polymerase chain reaction (PCR) was performed on all 15 samples using the Ampliqon Red PCR Mastermix (Lasec SA, PTY LTD) according to the following protocol:
Primer sets and gene regions used in this study.
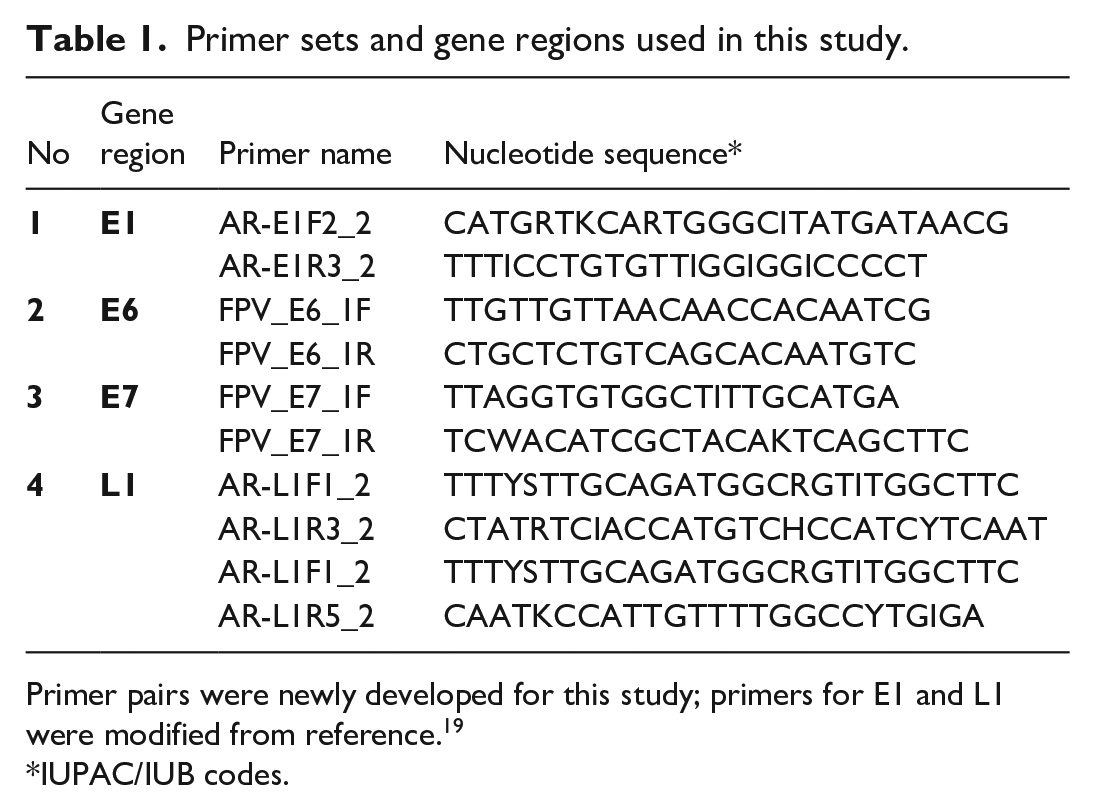

Primer pairs were newly developed for this study; primers for E1 and L1 were modified from reference. 19
IUPAC/IUB codes.
PCR of all samples was carried out in 25 μl reaction volumes comprised of 12.5 µl of 2 x Ampliqon Red Mastermix, 1 µl of each forward and reverse primer (10 pmol concentration), 1 µl of genomic DNA template (50 ng/ul concentration), and 9.5 µl nuclease free water. Final cycling conditions after optimization (by touchdown PCR) were as follows: Initial denaturation at 94°C for 5 minutes; 35 cycles of denaturation at 94°C for 30 seconds, annealing at 52°C (E6, and E7 primers) or 58°C (E1 and L1 primers) for 50 seconds and extension at 72°C for 1 minute. A final extension step of 72°C for 10 minutes concluded the cycling. Contamination was minimized by doing PCR setup in a DNA-free laminar flow hood, while reactions were monitored by the inclusion of a negative control, where the DNA sample was replaced with 1 µl of nuclease free water, for each gene region. Thereafter, agarose gel electrophoresis of each amplification product (5 µl) was performed on a 2% agarose gel in 1 × TAE buffer at 135 V for 30 minutes and visualized on the GelDoc EZ imaging system (Bio-Rad). All positive products were purified using Exonuclease I and FastAP (ThermoFisher, Massachusetts, United States). Cycle sequencing of the positive PCR products was performed using the BigDye Terminator version 3.1 Cycle Sequencing Kit (Applied Biosystems, California, USA) according to the manufacturer’s protocol. Sequencing products were purified using the BigDye Xterminator sequencing purification kit (ThermoFisher) and sequenced in both the forward and reverse directions on an ABI 3130 genetic analyzer (Applied Biosystems, California, USA). The raw sequence data were analyzed using the Applied Biosystems Sequencing Analysis software version 6. PCR results for each gene region are shown in Supplemental Table S1.
Phylogenetic Analyses
All sequences were initially analyzed using BLAST against the National Center for Biotechnology Information (NCBI) database to confirm each sequence identity as papillomavirus. Thereafter, the BLASTn program was used to search the NCBI database for closely related papillomavirus sequences for all gene regions amplified in this study. Relevant reference sequences were downloaded from the NCBI GenBank database into the Molecular Evolutionary Genetics Analysis (MEGA X) 24 software and separate sequence alignments were created for each gene region amplified in this study, except for the E6 and E7 gene regions which were combined and analyzed together.
Depending on availability, either Etapapillomavirus, Betapapillomavirus, or Bovine papillomavirus sequences were selected from GenBank and included as an out-group for the 3 gene regions. Phylogenetic analysis was performed on a 39-sample dataset for the L1 gene (15 sample isolates and 24 NCBI reference sequences), a 29-sample dataset for the combined E6 and E7 gene (15 sample isolates and 14 NCBI reference sequences), and a 34-sample dataset for the E1 gene (15 sample isolates and 18 NCBI reference strains) dataset. Sequence datasets were aligned with the ClustalW algorithm in MEGAX, 24 while all phylogenetic analyses were performed using the Maximum Likelihood model in MEGA X, with 100 bootstrap replications. The best fit model of evolution for each dataset was also determined in MEGA X under the AIC and were as follows: for the L1 gene region, the T92+G+I model was selected with G = 1.07; for the combined E6 and E7 dataset the K2+G+I model was employed with G = 3.79, and finally for the E1 dataset the T92+G+I model with G = 0.94 was selected.
Genetic distances between the cheetahpapillomavirus clade identified in this study and the feline and caninepapillomavirus clades were calculated under the maximum composite likelihood distance estimation in MEGA X. All sequences generated in this study were submitted to Genbank and accession numbers are listed in Supplemental Table S1.
Additional Tests
Serum samples were available from 11 affected animals and were tested for feline leukemia (FeLV) and feline immunodeficiency (FIV) virus infections. A commercially available test kit (Antigen rapid FIV Ab/FeLV Ag Test Kit, Bionote Inc, Republic of Korea) was used to test for FIV antibodies and the FeLV antigen according to the manufacturer’s specifications.
The data analyzed for this study are available from the authors.
Results
Fifteen cheetahs presented with sublingual lesions reminiscent of sublingual papillomas, from a total of 256 cheetahs seen at 2 facilities during the study period (specifically, 14 of the 92 seen at AfriCat Foundation, and 1 of the 164 seen at the Ann van Dyk Cheetah Center). The first 2 cases were seen in 2005 (1 at each facility). Subsequently, tissue from 13 additional cases were submitted for molecular analysis (Supplemental Table S1), including 14 samples from 12 cheetahs from AfriCat Foundation and 1 from the Ann van Dyk Cheetah center.
The lesions consisted of multiple proliferative exophytic nodules on a large area of the sublingual mucosa (Fig. 1). The nodules often appeared to coalesce to form larger flat lesions (Fig. 2). Histologically, samples from the first 2 cases (2005) had a benign proliferative epithelial lesion with prominent thickening of the stratified squamous epithelium, and scattered cells within the epithelium had nuclear enlargement, irregularity of the nuclear membranes and cell contours, focal hyperchromasia of the nuclei, and perinuclear halos (Fig. 3).
Amplification results for each gene region are shown in Supplemental Table S1. Maximum likelihood analyses were conducted and phylogenetic trees constructed (Figs. 4-6). Data from the 3 datasets yielded the same results, and a feline clade and canine clade were distinguishable with good bootstrap support (70%-100%). All cheetah papilloma sequences from this study clustered together with 99.99% genetic similarity across all 3 gene regions with very good bootstrap support (99%-100%), and these sequences were closely related to but genetically distinct from those of other known feline papillomaviruses. Genetic distance estimations revealed a 63.1% to 74% genetic similarity between the cheetah papilloma and feline papillomas for the 3 gene regions, while data showed a 35.9% to 54.3% similarity between the feline and canine clades.
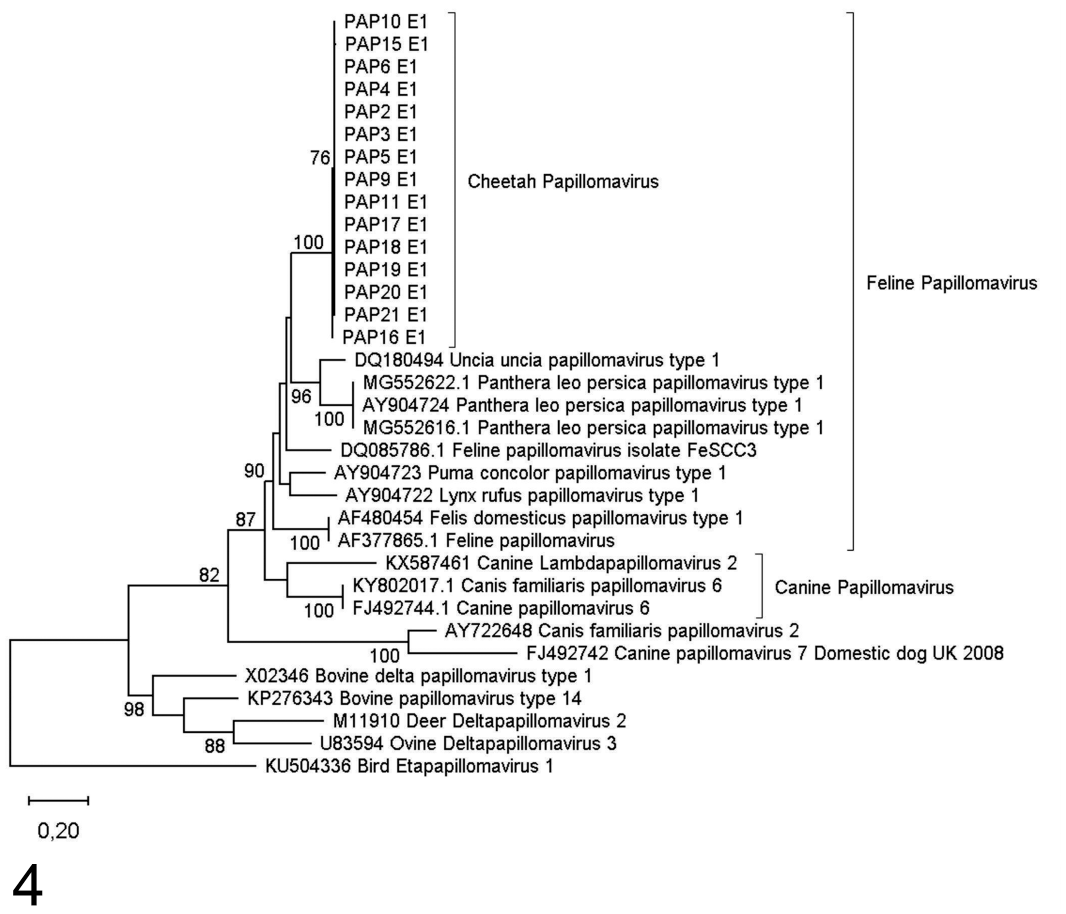
Maximum likelihood phylogenetic tree based on 946 bp of the L1 gene region for cheetah papillomaviruses identified from southern Africa. Bootstrap support indices above 70% are indicated on branches.
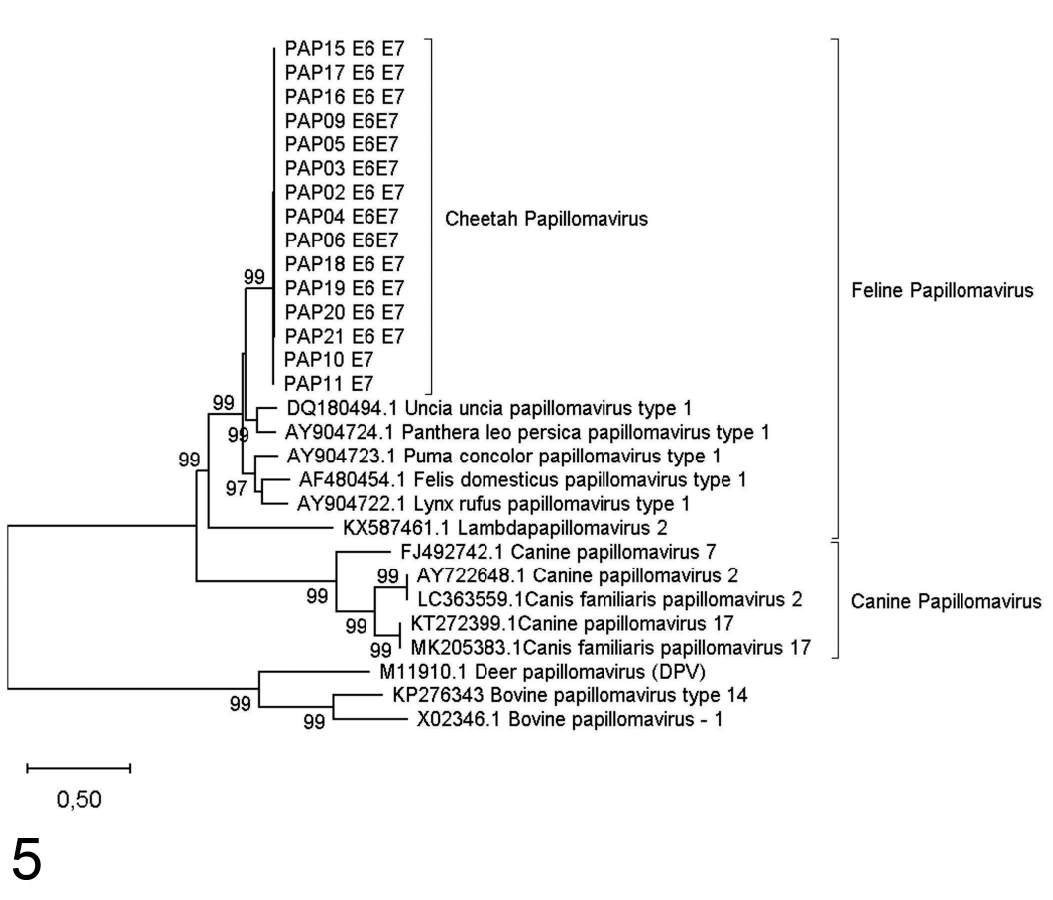
Maximum likelihood phylogenetic tree based on 911 bp of the combined E6 and E7 gene regions for cheetah papillomaviruses identified from southern Africa. Bootstrap support indices above 70% are indicated on branches.
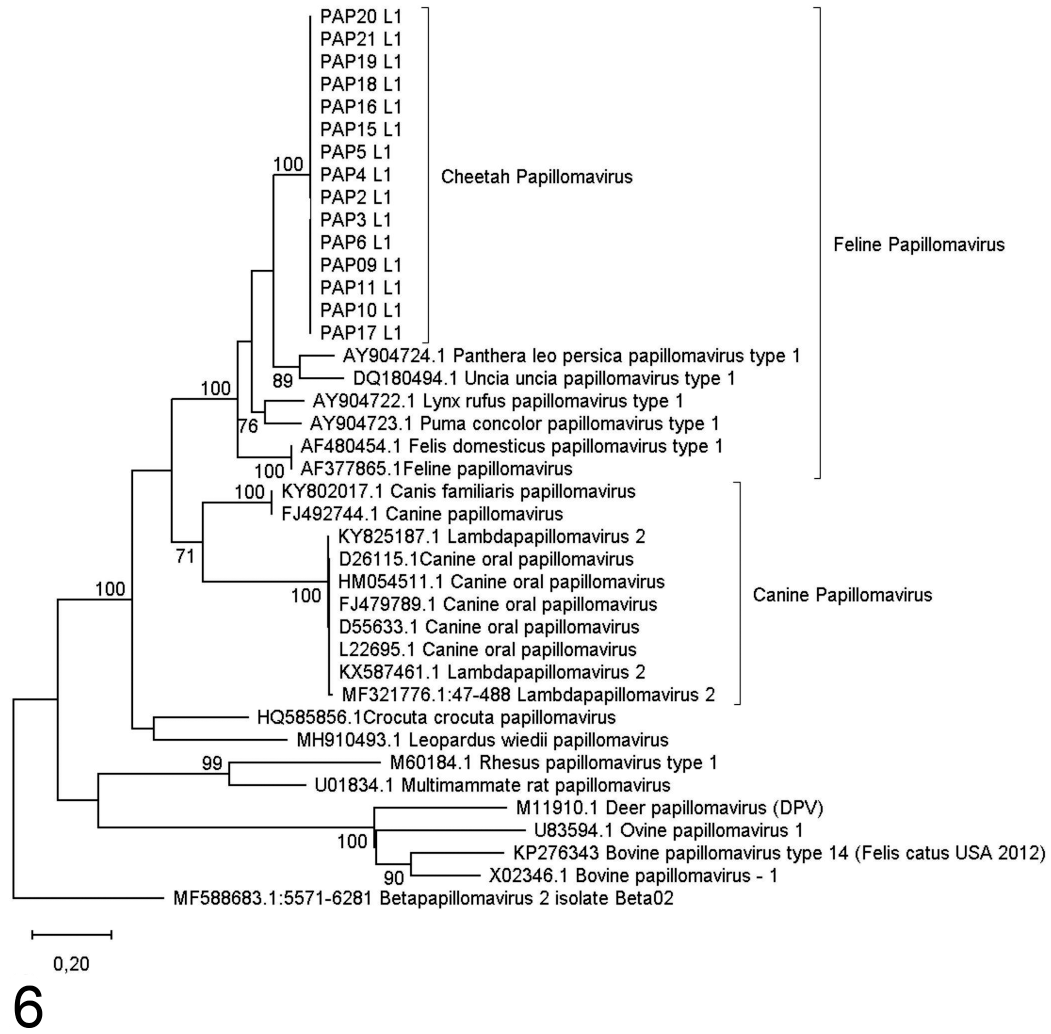
Maximum likelihood phylogenetic tree based on 372 bp of the E1 gene region for cheetah papillomaviruses identified from southern Africa. Bootstrap support indices above 70% are indicated on branches.
The results of all the samples tested for the FeLV and FIV viruses were negative.
Discussion
Papillomavirus infections in domestic cats were first diagnosed in 1990. 4 Since then, at least 5 different feline papillomaviruses have been identified in domestic cats and 1 non-feline papillomavirus (bovine papillomavirus-14) may also infect them. 16 Papillomaviruses infect stratified squamous epithelium and in domestic cats is most often identified in the skin and oral mucosa. Although most cats are infected with the FcaPV2 virus, it rarely is of any consequence. Even so, 6 distinct diseases have been associated with feline papillomaviruses and BPV-14 infection in domestic cats. 16
In non-domestic cats, to date, 5 different diseases have been attributed to papillomavirus infection including cutaneous plaques, cutaneous squamous cell carcinoma, ventral tongue papillomas, sarcoids and oral/sublingual SCC.1,17,20,28 The histological features of the samples in this study were reminiscent of papillomavirus infections, and presence of papillomavirus was confirmed by molecular diagnosis. Although causality cannot be confirmed without additional work, the detection of a novel species of feline papillomavirus in lesions consistent with papillomavirus infection warrants further attention. None the cheetahs with typical tongue papillomas showed any associated clinical signs. After removal of these lesions, they did recur but with no ill effect to the patients. Papillomas always recurred in the same individuals and never spread to other cheetahs on the facilities, including cheetahs sharing their enclosures. None of the lesions ever progressed to squamous cell carcinoma in the 12 years that we followed the cheetahs.
Sequence data suggest that papillomaviruses may be widespread among the captive felid community, with related strains in several captive wild felids from the United States1,11,17,28 and southern Africa (this study). Samples tested in this study were distinct from domestic canid and felid papillomavirus reference sequences in the NCBI nucleotide database, and all generated sequences were within the Feline papillomavirus cluster in the Lambdapapillomavirus 1 genus. Phylogenetic analysis showed that all papillomavirus sequences detected from cheetahs in this study differed from those of other FPVs in the NCBI nucleotide database by 26% to 36% for the L1, E6&7, and E1 gene regions. Therefore, the virus identified in this study can be considered as a previously undescribed or novel feline papillomavirus and the authors propose the designation “Acinonyx jubatus papillomavirus type 1” (AjPV-1), within the Lambdapapillomavirus 1 genus in the family Papillomaviridae.
Unfortunately, the lack of reference sequence data for papillomaviruses isolated from wildlife, and particularly from southern African wildlife, has hindered definitive identification and classification of these viruses. An increased awareness and effort to characterize these viruses in wildlife will provide valuable information to assess the true diversity of these viruses in wild felids in southern Africa.
All 13 cases tested were negative for feline immunodeficiency virus and feline leukemia virus using a commercially available kit. In African lions, a definitive association of sublingual papillomas with FIV has not been established; 20 however, in the current study, there was no association between the presence of cheetah papillomavirus in sublingual samples and FIV.
A genetically unique papillomavirus that is proposed as a novel species of feline papillomavirus (designated as Acinonyx jubatus papillomavirus type 1; AjPV-1) was detected from sublingual papillomas biopsied from cheetahs from the Ann van Dyk Cheetah Center in South Africa as well as the AfriCat Foundation in Namibia. The lesions had the typical plaque-like appearance that was originally identified in ventral tongue papillomas of snow leopards.1,11
Supplemental Material
sj-pdf-1-vpt-10.1177_03009858221109610 – Supplemental material for Sublingual papillomas of cheetahs in southern Africa
Supplemental material, sj-pdf-1-vpt-10.1177_03009858221109610 for Sublingual papillomas of cheetahs in southern Africa by Gerhard Steenkamp, Adrian S. W. Tordiffe, Essa Suleman, Almero Oosthuizen, Helene Brettschneider and Sonja C. Boy in Veterinary Pathology
Footnotes
Acknowledgements
We would like to thank the team at the Ann van Dyk Cheetah Center and the AfriCat Foundation (Namibia) for their dedication and assistance in collecting the material as well as Ms Estelle Mayhew for her assistance in preparing the figures.
Supplemental material for this article is available online.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
