Abstract
A comparative study was carried out on common and agile frogs (Rana temporaria and R. dalmatina) naturally infected with ranid herpesvirus 3 (RaHV3) and common toads (Bufo bufo) naturally infected with bufonid herpesvirus 1 (BfHV1) to investigate common pathogenetic pathways and molecular mechanisms based on macroscopic, microscopic, and ultrastructural pathology as well as evaluation of gene expression. Careful examination of the tissue changes, supported by in situ hybridization, at different stages of development in 6 frogs and 14 toads revealed that the skin lesions are likely transient, and part of a tissue cycle necessary for viral replication in the infected hosts. Transcriptomic analysis, carried out on 2 naturally infected and 2 naïve common frogs (Rana temporaria) and 2 naturally infected and 2 naïve common toads (Bufo bufo), revealed altered expression of genes involved in signaling and cell remodeling in diseased animals. Finally, virus transcriptomics revealed that both RaHV3 and BfHV1 had relatively high expression of a putative immunomodulating gene predicted to encode a decoy receptor for tumor necrosis factor in the skin of the infected hosts. Thus, the comparable lesions in infected frogs and toads appear to reflect a concerted epidermal and viral cycle, with presumptive involvement of signaling and gene remodeling host and immunomodulatory viral genes.
Amphibians are undergoing an unprecedented mass extinction around the globe with a dramatic loss in biodiversity. 8 It is becoming increasingly clear that infectious diseases are critical players in this complex process. 15 Over the last 4 years, we have discovered and characterized 2 new viruses, namely, Ranid herpesvirus 3 (RaHV3) and Bufonid herpesvirus 1 (BfHV1), associated with a proliferative skin disease in free-ranging common (Rana temporaria) and agile frogs (Rana dalmatina; RaHV3) and common toads (Bufo bufo; BfHV1), respectively. 2,23,24 These viral infections are associated with a similar proliferative epidermal lesion but infiltrating inflammatory cells are absent or minimal. RaHV3 and BfHV1 are invariably observed in the diseased skin of the affected frogs and toads, respectively. 2,11,23,24 The skin lesions vary greatly in severity, and their clinical relevance is not clear. However, considering the importance of the integument in amphibians physiology, 7 the infections might have significant effects.
RaHV3 and BfHV1 are structurally and phylogenetically related. Putative immunomodulatory genes are present in both viruses, possibly contributing to the very limited host cellular immune response. The diseases in frogs and toads appears to be seasonal, occurring at the end of hibernation (brumation) and during the mating season, suggesting that environmental, host physiological factors (such as hormonal status), and social interaction might also contribute to the occurrence of the skin lesions.
The similarities shared by these 2 distinct diseases include (1) similar lesions, (2) lack of a prominent leukocyte cell infiltrate, (3) presence of a herpesvirus, and (4) seasonal occurrence. This suggests similar pathogenesis of these diseases. Accordingly, the aim of this investigation was to gather preliminary data that could illuminate the molecular basis of the observed diseases and, ideally, matching them with the observed complementary morphological changes in the affected amphibian species.
Materials and Methods
Histopathology
All tissues used in this investigation were obtained from amphibians that were necropsied at the Centre for Fish and Wildlife Health (FIWI) at the University of Bern for diagnostic purposes. Accordingly, no experimental permit was necessary for obtaining these samples and to carry out this investigation.
Common frog (n = 4; Rana temporaria), agile frog (n = 2; Rana dalmatina), and common toad (n = 14; Bufo bufo) tissues were previously processed for routine diagnostic examination at the FIWI from 2014 to 2018 (Supplemental Material 1). The samples were fixed in 10% buffered formalin, processed, sectioned at 5 μm, and stained with hematoxylin and eosin according to standard protocols. One agile frogs was part of a previous study, 2 and the paraffin block used in this study was processed as described above.
Transmission Electron Microscopy
Samples of affected skin from 4 frogs and 4 toads, showing fully developed lesions and excellent tissue conservation (Supplemental Material 1), were fixed with 2.5% glutaraldehyde in 0.1 M cacodylate buffer, pH 7.4, and osmicated samples were dehydrated and then embedded in Epon (FLUKA). Ultrathin sections were double-stained with 0.5% uranyl acetate (Sigma Aldrich) and 3% lead citrate (Leica) and examined in a Philips CM12 transmission electron microscope (FEI). 23
In Situ Hybridization
Tissue blocks for in situ hybridization were selected from 1 frog and 1 toad naturally infected with RaHV3 and BfHV1 that had fully developed lesions, were positive by PCR [polymerase chain reaction], and had excellent tissue preservation, as well as 1 frog and 1 toad that were uninfected and had no detectable lesions, were PCR-negative, and had excellent tissues preservation (Supplemental Material 1). Sections (5-μm-thick) of formalin-fixed, paraffin-embedded tissue were deparaffinized in xylol and then treated according to the manufacturer’s instructions (RNAscope, ADC). The hybridization probes were designed to anneal to the DNA polymerase mRNA of RaHV3 and BfHV1, respectively (Cat. No. 563871-RaHV3; 563861-BfHV1; ADC).
Preparation of cDNA and Sequencing
Lesional skin samples were obtained from 2 common frogs and 2 common toads (Supplemental Material 1) naturally infected with RaHV3 and BfHV1, respectively, as confirmed by PCR and histopathology and with fully developed lesions. Skin samples from 2 uninfected common frogs and 2 common toads that were PCR-negative for RaHV3 or BfHV1 and did not have any lesions used as negative controls. Total RNA was extracted using the RNeasy kit (Qiagen) according to the manufacturer’s instructions and stored at −80°C until use. The total RNA extracted was pooled by species and condition as described below. One pool of total RNA from the 2 RaHV3-infected common frogs, one pool from the 2 RaHV3-uninfected common frogs, one pool from the 2 BfHV1-infected common toads, and one pool from the 2 BfHV1-uninfected common toads. Transcriptomic analysis was conducted using the Illumina platform (Fasteris SA) as follows. Following quality control and Ribozero treatment, cDNA libraries were prepared using the Illumina TruSeq Stranded Total RNA (Illumina) according to the manufacturer’s instructions. Four cDNA libraries were prepared from the pools described above that were derived from RaHV3-infected common frog tissues (GYB-11), BfHV1-infected common toad tissues (GYB-12), uninfected common frog tissues (GYB-30), and uninfected common toad tissues (GYB-31). The libraries were sequenced on an Illumina HiSeq 2500 to produce about 100 M raw paired-reads (PE) of 125 bases long per library. Following sequencing, adapters were removed using Trimmomatics, version 0.32, and the reads were filtered according to their quality (using a 4-base wide sliding window and cutting the reads when the average quality per base dropped below 5). Ribosomal RNA reads still present after Ribozero treatment were removed by mapping the reads with BWA version 0.7.5a on a database of rRNA sequences (downloaded from https://www.ncbi.nlm.nih.gov/nuccore; containing 2423 rRNA sequences) and by removing all mapped reads.
Host Transcriptomics
The main goal of the molecular investigation of this study was to compare the gene expression of RaHV3-infected and uninfected common frogs with BfHV1-infected and uninfected common toads. Accordingly, we prepared a common reference dataset based on Xenopus sp. data because this is a close and comparable reference for both species and because the Xenopus sp. genome and transcriptome are the most thoroughly characterized among amphibians. Consistently, we mapped all libraries using BWA version 0.7.5a on Xenla_9.1 (downloaded from Xenbase: ftp://ftp.xenbase.org/pub/Genomics/JGI/Xenla9). The selected reference comprised 402 502 scaffolds for a total of 2.76 Gbases. Annotations for Xenla_9.1 were downloaded from the publicly available repository: ftp://ftp.xenbase.org/pub/Genomics/JGI/Xenla9.1/1.8.3.2/. They comprised a total of 2 061 902 annotations, including 45 099 gene features. The number of reads mapping to the homologous genes was counted using the BEDtools version 2.21.0. The package DESeq2 was used to normalize the counts and compute the Log2 ratios between the infected and uninfected libraries. The 50 most differentially expressed genes (either up- or downregulated) from each species considered were selected (Supplemental Material 2). The biological functions of each of the identified transcripts were then determined by basic local alignment search tool (BLAST) using the gene IDs in Uniprot (https://www.uniprot.org). Additional protein repositories that were considered, although not systematically, included QuickGO (https://www.ebi.ac.uk/QuickGO) and GeneCards (https://www.genecards.org). For each of the genes identified, we determined the gene ontology (GO) molecular function and biological process associated. Additional information was obtained by BLAST analysis of the identified gene or predicted protein translation using Reactome (https://reactome.org; Supplemental Material 2).
Virus Transcriptomics
Adapter removal and reads filtering was carried out as described above for the host transcriptomics. Mapping was carried out using the reads after adapter removal. The reads from infected tissue of common frogs (GYB-11 library) were mapped on the RaHV3 genome (207’914 bp-Genbank NC_034618.1), whereas the reads from infected tissue of common toads (GYB-12 library) were mapped on the BfHV1 genome (158 250 bp-Genbank NC_040681.1). A total of 186 annotated genes for RaHV3 and 152 annotated genes for BfHV1 were considered. The number of reads overlapping each gene were counted using the BEDtools, version 2.21.0. In order to rank the expression of the viral genes, the total number of reads for each transcript was divided by the length of the transcript itself, obtaining the normalized value which could then be compared with all the other genes (Supplemental Material 3).
Results
Pathologic Findings in Frogs
Animals
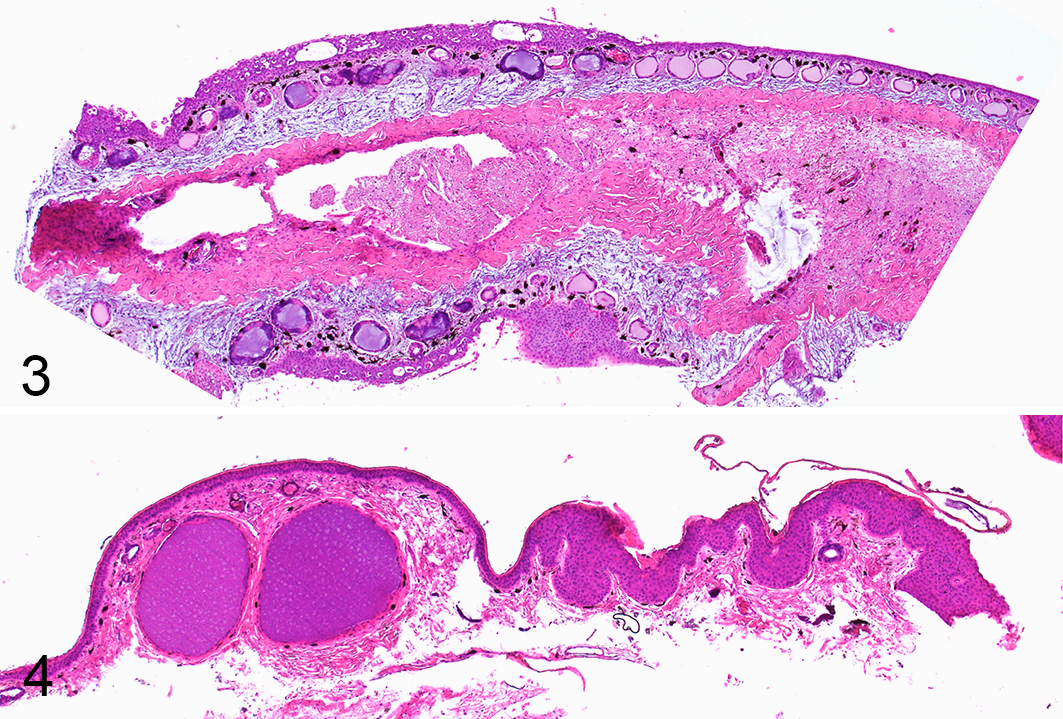
A total of 6 individuals (4 common frogs [Rana temporaria]; 2 agile frogs [Rana dalmatina]) collected during 4 different outbreaks in Switzerland (n = 3; 2015, 2017, and 2018) 23 and in Italy (n = 1; 1994), 2 were analyzed (Supplemental Material 1). All frogs included in this investigation were collected alive and showed variable but consistent macroscopic lesions of multifocal to coalescent, gray raised skin lesions of variable size (Fig. 1) and histological changes (Fig. 3) as described previously. 23
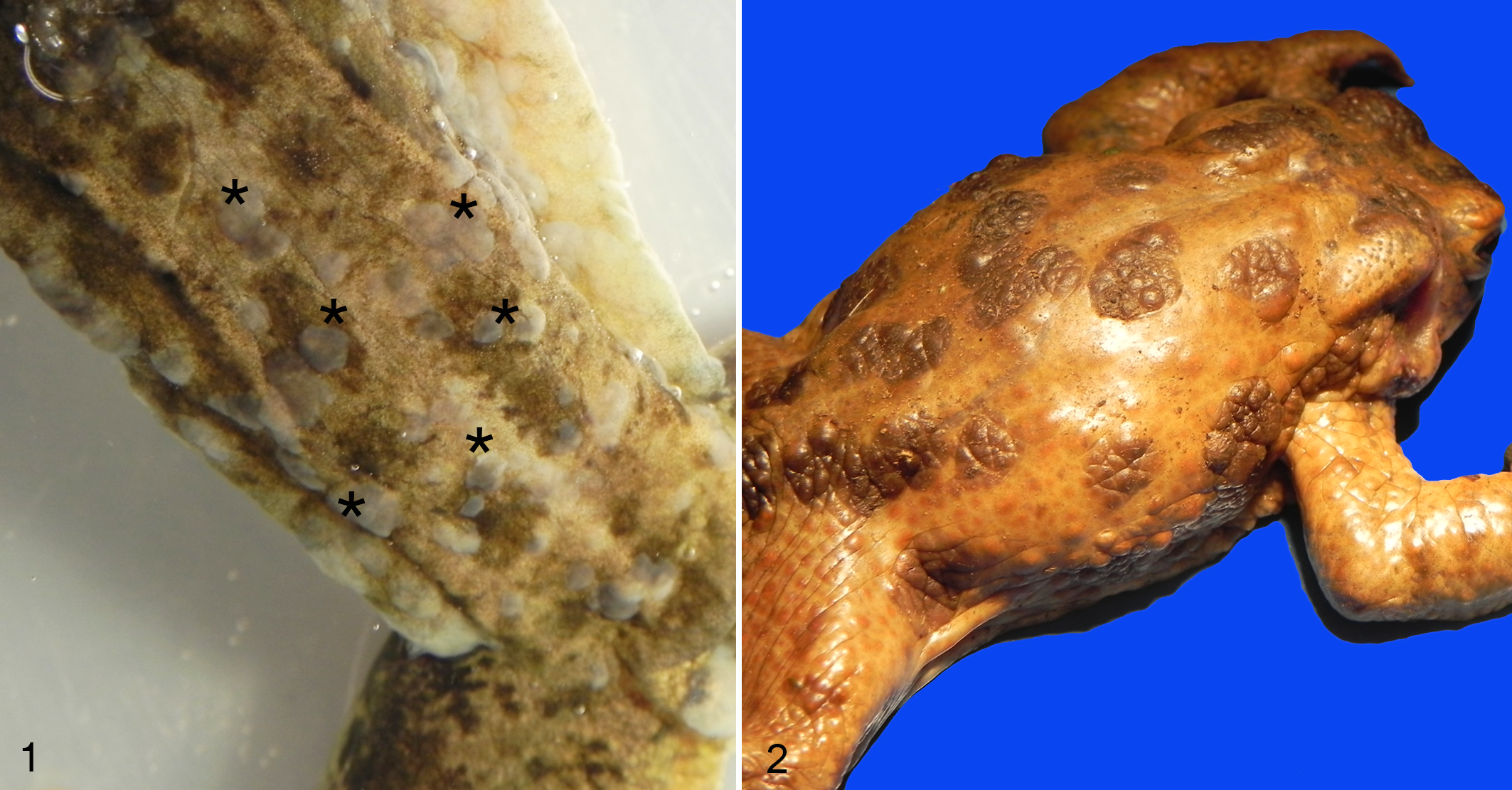
Herpesvirus-associated proliferative skin disease, skin.
Histological examination evaluated 3 major features of the lesions: (1) lesion development, (2) infiltration of immune cells (leukocytes), and (3) intranuclear inclusion bodies.
Lesion Development
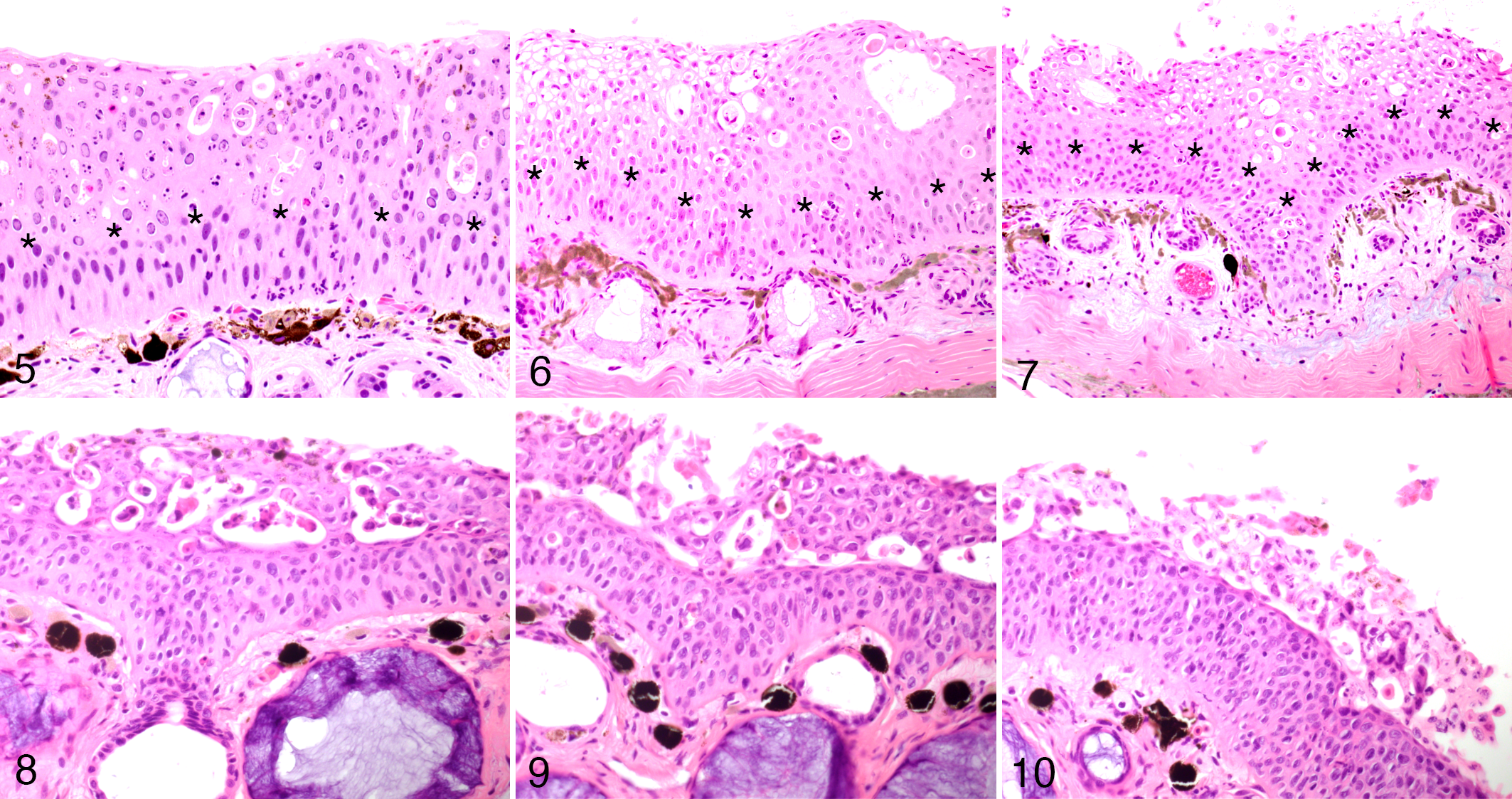
The skin lesions had differing morphology in the superficial and deeper layers of the epidermis. Specifically, these included (1) an exuberant superficial layer, which progressively underwent variably extensive vacuolization (Figs. 5–7), eventually followed by sloughing (Figs. 8–10), and (2) deeper or basilar and/or parabasilar layers, which remained relatively unaffected (Fig. 10). The demarcation between the 2 layers in the putatively early lesions corresponded to the “sloughing line” detected in individuals with putatively more advanced lesions (Figs. 5–10). Edema was commonly present, with severe distension of the stratum spongiosum of the dermis. Dermal glands had variable, but not systematic changes including, cell rounding and vacuolization, glandular epithelial cell necrosis, luminal ectasia, and occasional inflammatory infiltration.
Herpesvirus-associated proliferative skin disease, skin. Hematoxylin and eosin (HE).
Herpesvirus-associated proliferative skin disease, skin, common frog. Hematoxylin and eosin (HE). The figures depict the proposed progression of the histologic changes. In the putative early stages of disease, there is epidermal hyperplasia and superficial keratinocyte degeneration and necrosis (
Infiltration of immune cells. The cellular inflammatory infiltrate was very limited in the skin lesion of all individuals examined. Specifically, polymorphonucleated and few mononuclear cells were present in the basal and parabasal layers of the epidermis (exocytosis). In contrast, only few scattered inflammatory cells were present in the more superficial hyperplastic layers of the epidermis in the most severe lesions (Fig. 11). In the tissue samples with prominent vacuolization of the epidermis, there was a slight increase of mononuclear inflammatory cells within the dermis (Fig. 12). In contrast, in skin areas unaffected by epidermal hyperplasia and with no detectable or obvious viral presence, infiltrates of leukocytes were occasionally moderate to severe (Figs. 11, 12, insets).
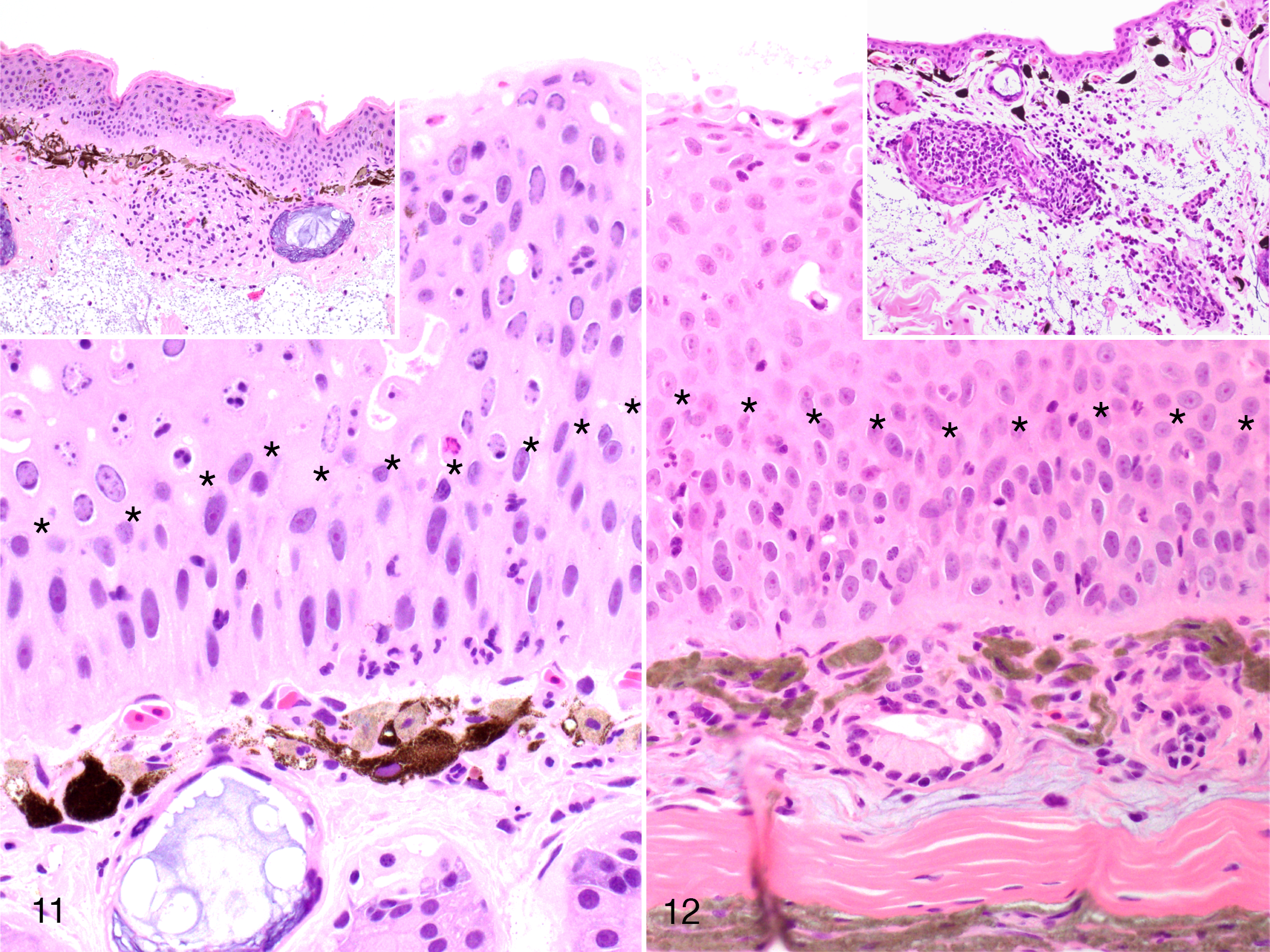
Herpesvirus-associated proliferative skin disease, skin, common frog. The epidermis is hyperplastic and viral inclusions are limited to the superficial layer (above the line of asterisks). An inconspicuous inflammatory infiltrate is limited to the deeper layers of the epidermis (below the line of asterisks) and slightly increased in the dermis. The scant inflammatory infiltrate in the epidermis contrasts with the robust dermal infiltrate in non-virus-infected areas (insets). Hematoxylin and eosin. Insets reprinted from Origgi et al, 2017, Veterinary Pathology, 23 with permission.
Inclusion Bodies
Intranuclear eosinophilic to amphophilic inclusions with marginalization of chromatin were detected in all the skin samples examined in the superficial layers of the hyperplastic epidermis (Fig. 13).
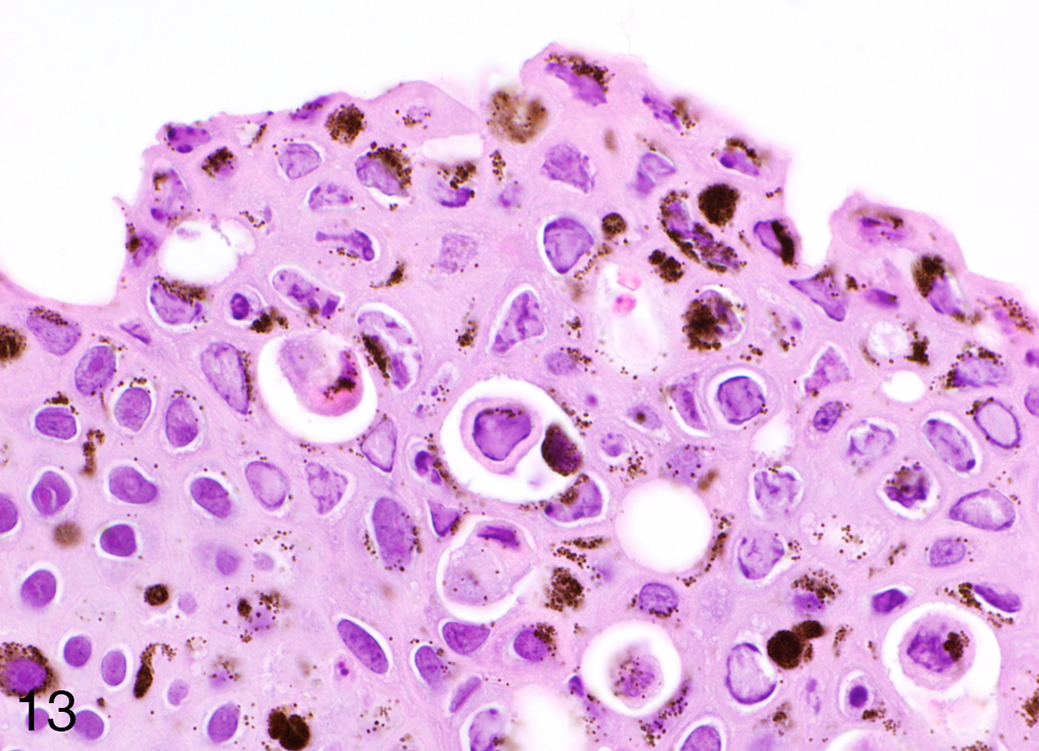
Herpesvirus-associated proliferative skin disease, skin, common frog. Numerous prominent intranuclear eosinophilic to amphophilic inclusions are scattered in the hyperplastic epidermis (HE).
Pathologic Findings in Toads
Animals
A total of 14 common toads (Bufo bufo) with gross cutaneous lesions were examined. The toads were part of 5 independent outbreaks occurring in Switzerland in 2014 (n = 1 individual), 2015 (n = 3), 2016 (n = 1), and 2017 (n = 9; Supplemental Material 1). 24 All the toads from 2014, 2015, and 2016 were collected alive; 8 of 9 toads collected in 2017 were found dead. Details concerning the specific outbreaks have been published previously. 24 All the toads examined showed the proliferative, multifocal to coalescent, brown skin lesions (Fig. 2) and histologic lesions (Fig. 4) as described previously. 24
Lesion Development
The lesion evolution in toads was limited to a transition from prominently thickened epidermis with similar morphology of the deep and superficial epithelial layers, to a similarly thickened epidermis but with a marked demarcation between the upper and lower hyperplastic layers. No vacuolization or obvious physical separation of the 2 layers could be observed (Figs. 14–17). Moderate to marked hyperkeratosis was present.
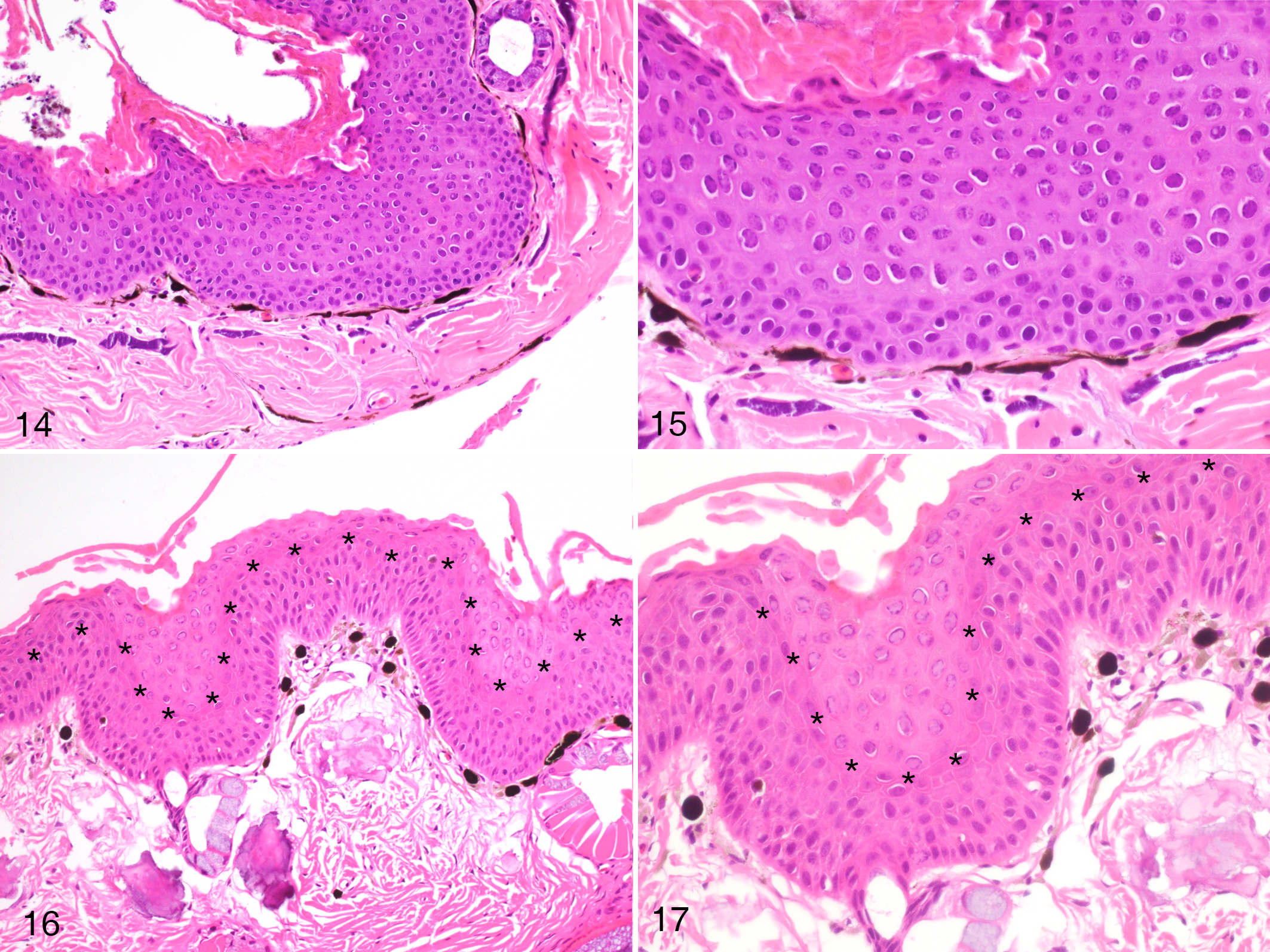
Herpesvirus-associated proliferative skin disease, skin, common toad. The epidermis is markedly thickened and moderately hyperplastic with numerous intranuclear inclusions. These are restricted to the superficial layer of the tissue. The line of asterisks is added to distinguish the superficial and deep layers (
Infiltration of Immune Cells
The cellular immune response was mild but evident in several sections. The leukocyte infiltrate was predominantly mononuclear and was present mainly at the dermal-epidermal junction and only occasionally infiltrated the basal portion of the epidermis in the more hyperplastic areas, and virtually absent from the superficial layers (Figs. 18–19).
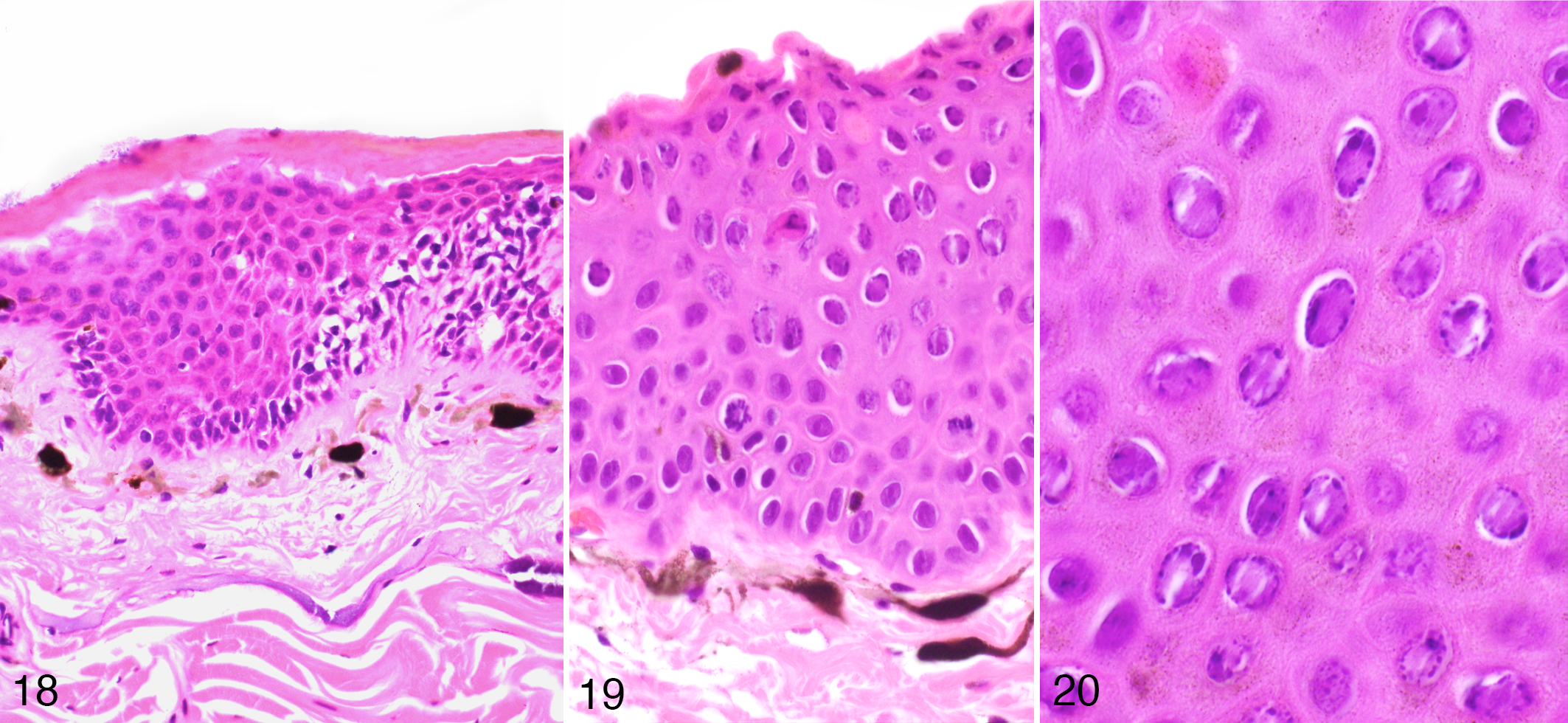
Herpesvirus-associated proliferative skin disease, skin, Common toad. Inflammatory cells are present within the dermis and the deeper epidermal layer, but are absent in the superficial layer where active viral replication occurs (as evidenced by inclusion bodies) (HE).
Inclusion Bodies
Intranuclear eosinophilic to amphophilic inclusions with marginalization of the chromatin were seen in all the BfHV1-infected toads in the upper layers of the hyperplastic epidermis (Fig. 20).
Comparison
The RaHV3- and BfHV1-associated had more defined lesion evolution pattern in frogs. Findings were suggestive of the following progression: (1) epidermal hyperplasia, (2) epidermal vacuolization, and (3) sloughing of the hyperplastic epidermal layer (Figs. 5–10). However, this staging was only partially observed in infected toads, and no detachment or sloughing of the exuberant hyperplastic epidermis was observed in any of the toad samples (Figs. 14–17). The cellular immune response was minimal to mild in both species, although slightly more accentuated in toads. Overall, the cellular inflammatory infiltrate was predominantly present in the deeper layers of the epidermis, corresponding to those layers where no viral inclusions were detected and which appeared to be less affected by the degenerative changes (vacuolation) associated with the viral infection. When present, the inflammatory cells were virtually all within the basal and parabasal layers of the hyperplastic epidermis, but not in the more superficial layers (Figs. 11, 12 and 18, 19). Inclusion bodies were slightly more abundant or more easily detectable in infected frogs compared to infected toads. Hyperkeratosis was observed in infected toads but not in infected frogs. Finally, degenerative glandular changes were observed in frogs, but not obvious in toads.
Ultrastructural Pathology
Ultrastructural changes in the skin of frogs infected with RaHV3 and of toads infected with BfHV1 at different stages of the disease indicated key steps of the presumptive viral cycle, including part of the virion maturation and transport from the nuclei to the extracellular space (Figs. 21–28).
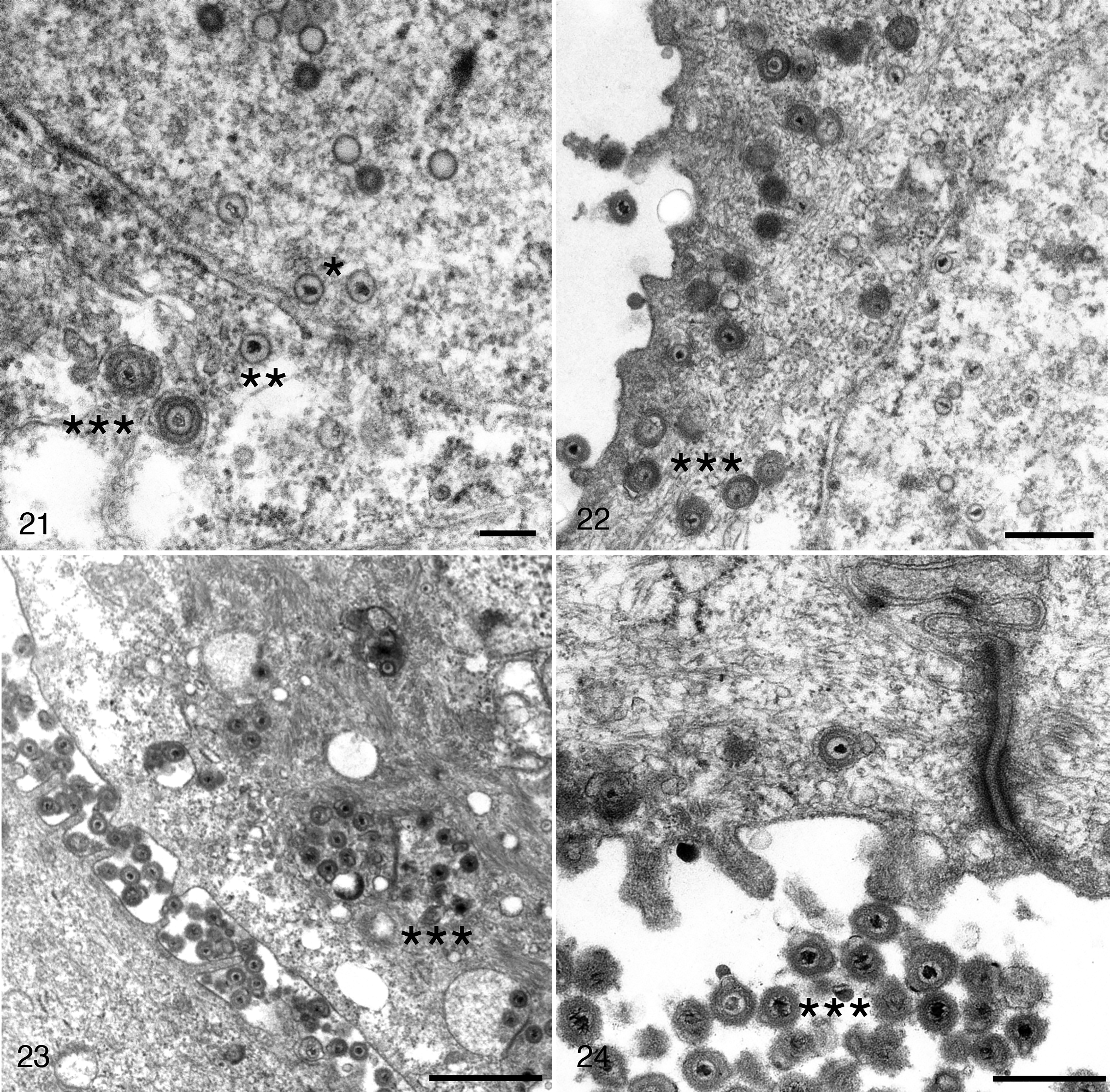
Herpesvirus-associated proliferative skin disease, skin, common frog. Transmission electron microscopy.
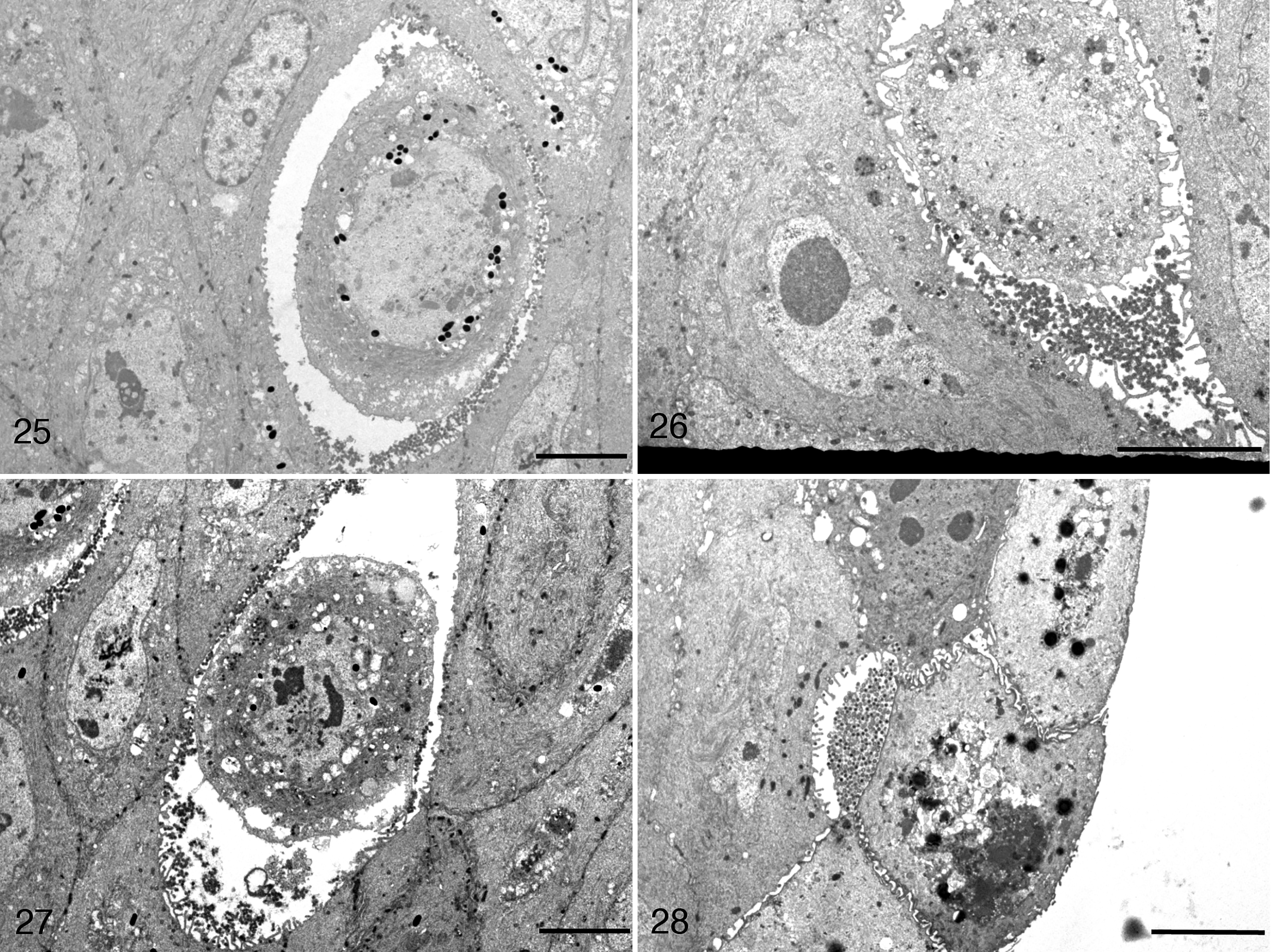
Scale bars: 5 μm. Viral particles accumulate within the intercellular space, which is expanded (Figs. 25, 26). Cell detachment (Fig. 27) and sloughing (Fig. 28) are also observed. Figure 28 reprinted from Origgi et al, 2017, Veterinary Pathology, 23 with permission.
Frogs
In RaHV3-infected frogs, naked capsid particles were present within the nuclei (Fig. 21). Occasionally, they were also present in the cytoplasm, in close proximity to the nuclear membrane (Fig. 21). However, in the cytoplasm, the majority of viral particles had additional layers consistent with a tegument and an envelope. Viral particles in proximity to the cell membrane were observed either as single particles or in variably dense clusters within large cytoplasmic vesicles, most consistent with Golgi vesicles (Figs. 22, 23). Large numbers of extracellular viral particles were collecting within expanded intercellular clefts (Figs. 23, 24). The accumulation of extracellular virions was associated with a progressive detachment of the infected cells from the surrounding keratinocytes. The severely vacuolated, rounded cells with poorly defined nuclear margins (necrotic) were seen sloughing together with both cell-free and cell-associated virions (Figs. 25–28).
Toads
In BfHV1-infected toad keratinocytes, the maturation pattern was similar to that observed in RaHV3-infected cells (Figs. 29–32), but with a few specificities. Naked capsids were present in the nuclei. Virions surrounded by an integument were present in the cytoplasm along with a presumptive temporary envelope within probable Golgi vesicles (Figs. 29, 30). No large vesicles containing multiple virions were observed in BfHV1-infected cells. However, a few single or more rarely double particles surrounded by a membrane were observed near the cell membrane (Fig. 30). Here, the single-virion vesicles appeared to fuse with the cell membrane, eventually releasing the virions into the extracellular space (Figs. 31, 32). Collapse of degenerating and necrotic cells was observed (Figs. 33–36). Large numbers of virions were present in the sloughing necrotic cells (Fig. 36).
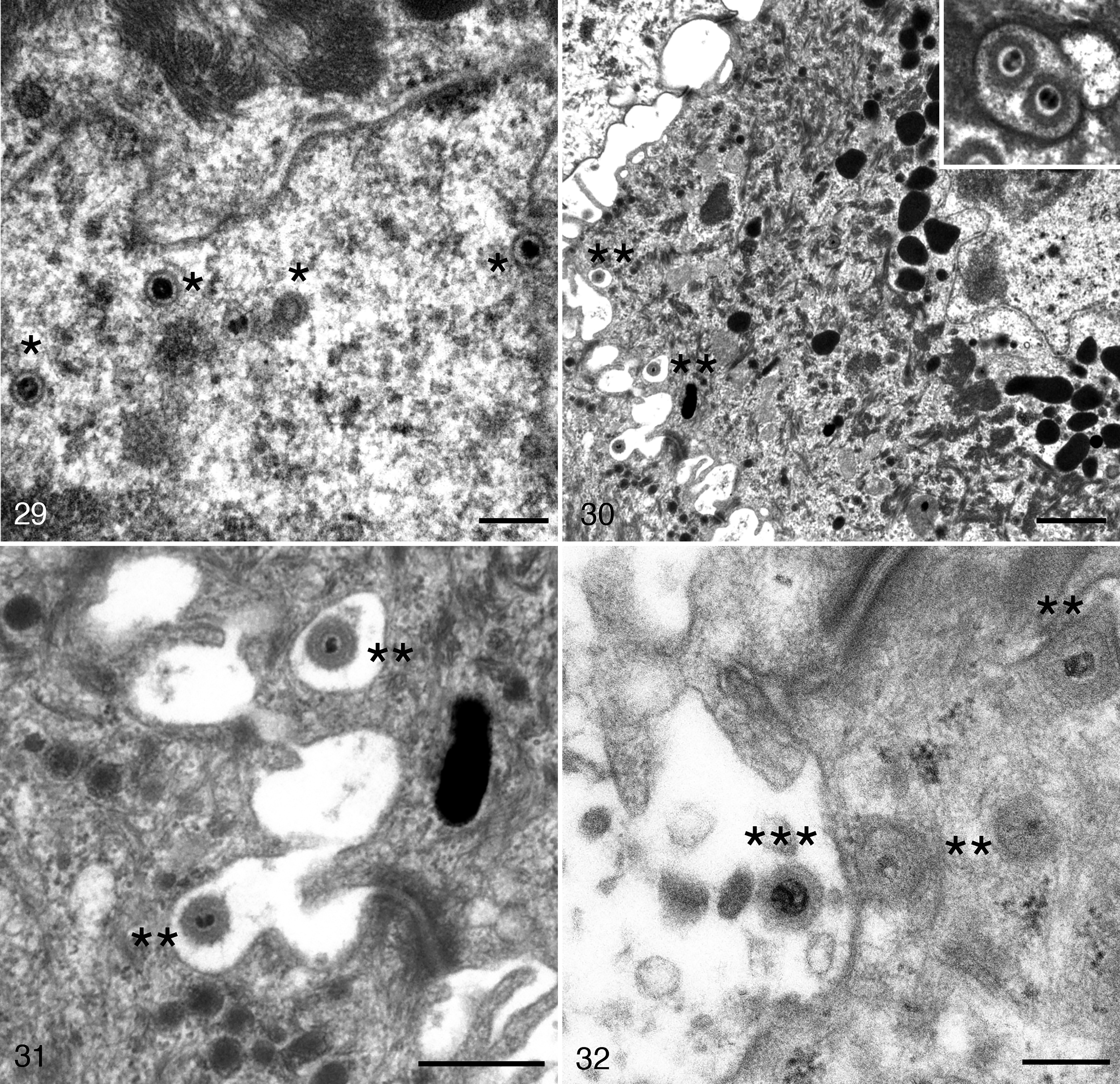
Herpesvirus-associated proliferative skin disease, integument, common toad. Transmission electron microscopy.
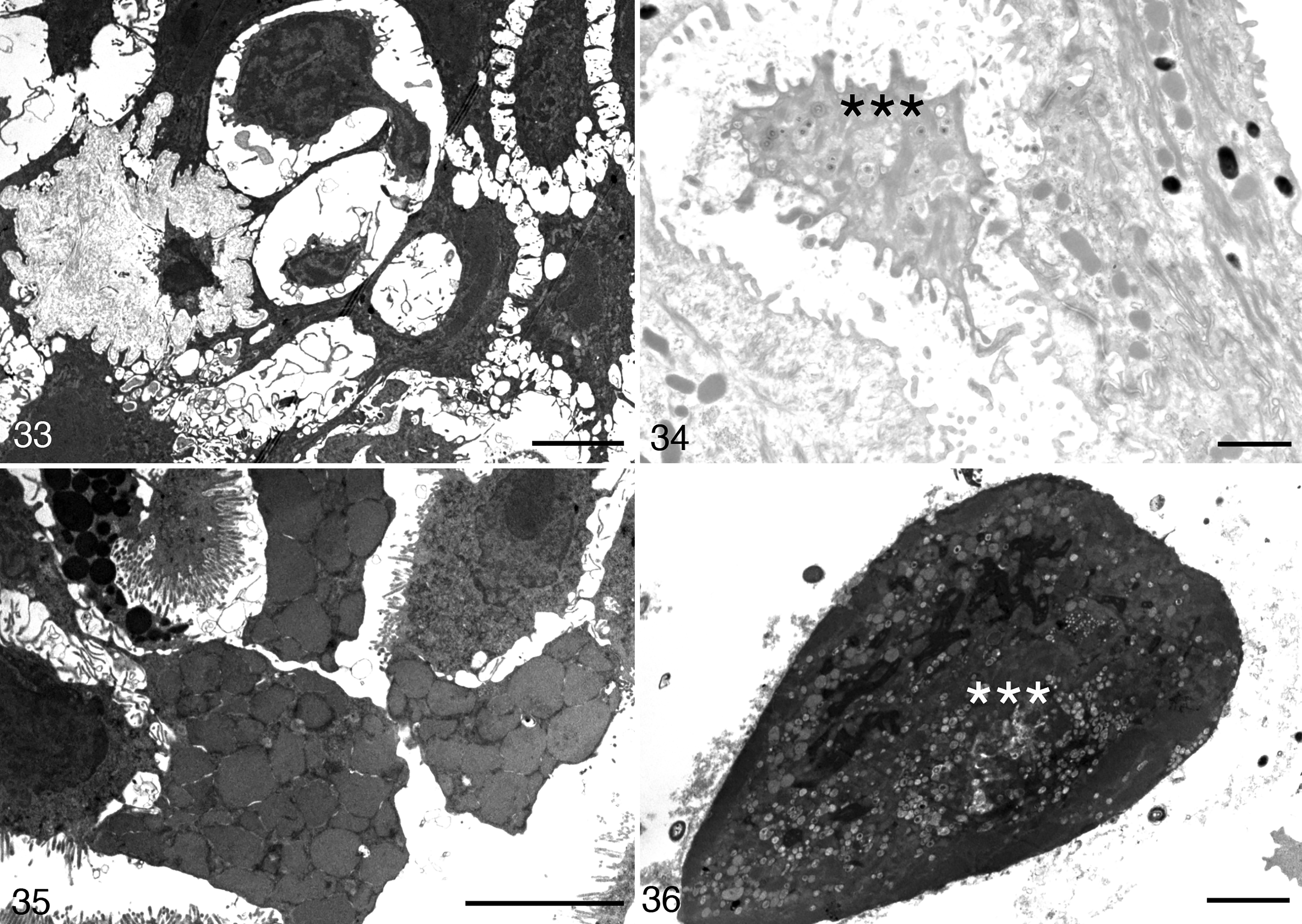
The degenerated keratinocytes are retracted and large intercellular spaces appear where few viral particles collect. Scale bars: Figure 33 = 5 μm; Figure 34 = 1 μm.
Comparison
Ultrastructural investigation revealed both similar and distinct features in virion maturation and egress. RaHV3 appeared to exploit 2 distinct systems of transporting maturing virions in which multiple virions and single virions, respectively, are putatively shuttled to the cell membrane in Golgi vesicles of variable size (Figs. 22, 23). In contrast, BfHV1 appeared to support a system in which single virions are shuttled within small vesicles before eventually fusing with the cell membrane and releasing the mature virions (Figs. 29–32). An obvious difference was the number of virions produced, which was remarkably higher with RaHV3 compared to BfHV1. Accordingly, the massive number of virions collecting in the extra-cellular spaces appears to be a specific feature of RaHV3 but not of BfHV1 (Figs. 21–36).
In Situ Hybridization
In situ hybridization revealed strong viral gene expression in the areas of epidermal hyperplasia both in affected frogs and toads (Figs. 37–42).
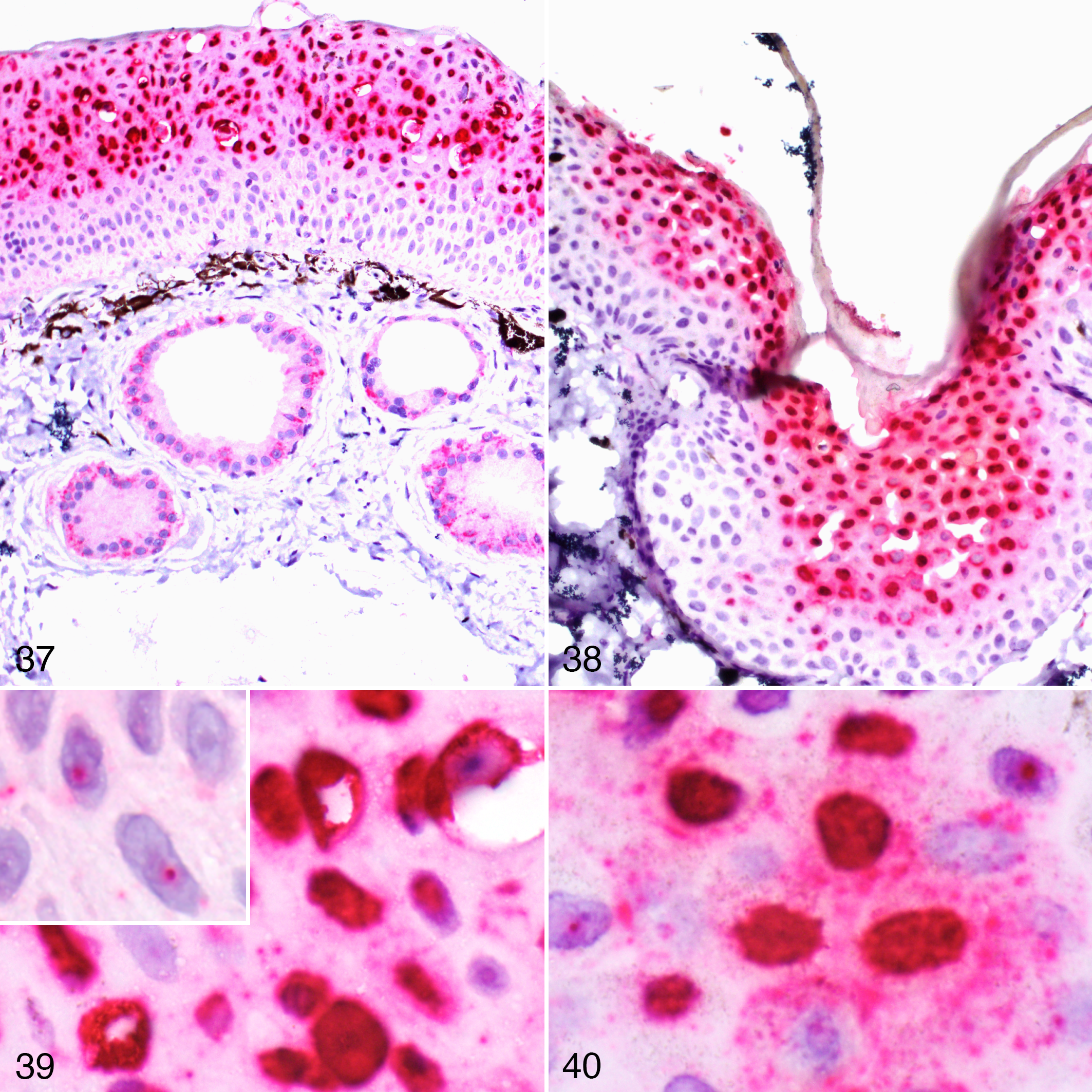
Herpesvirus-associated proliferative dermatitis, integument. In situ hybridization for herpesvirus mRNA.
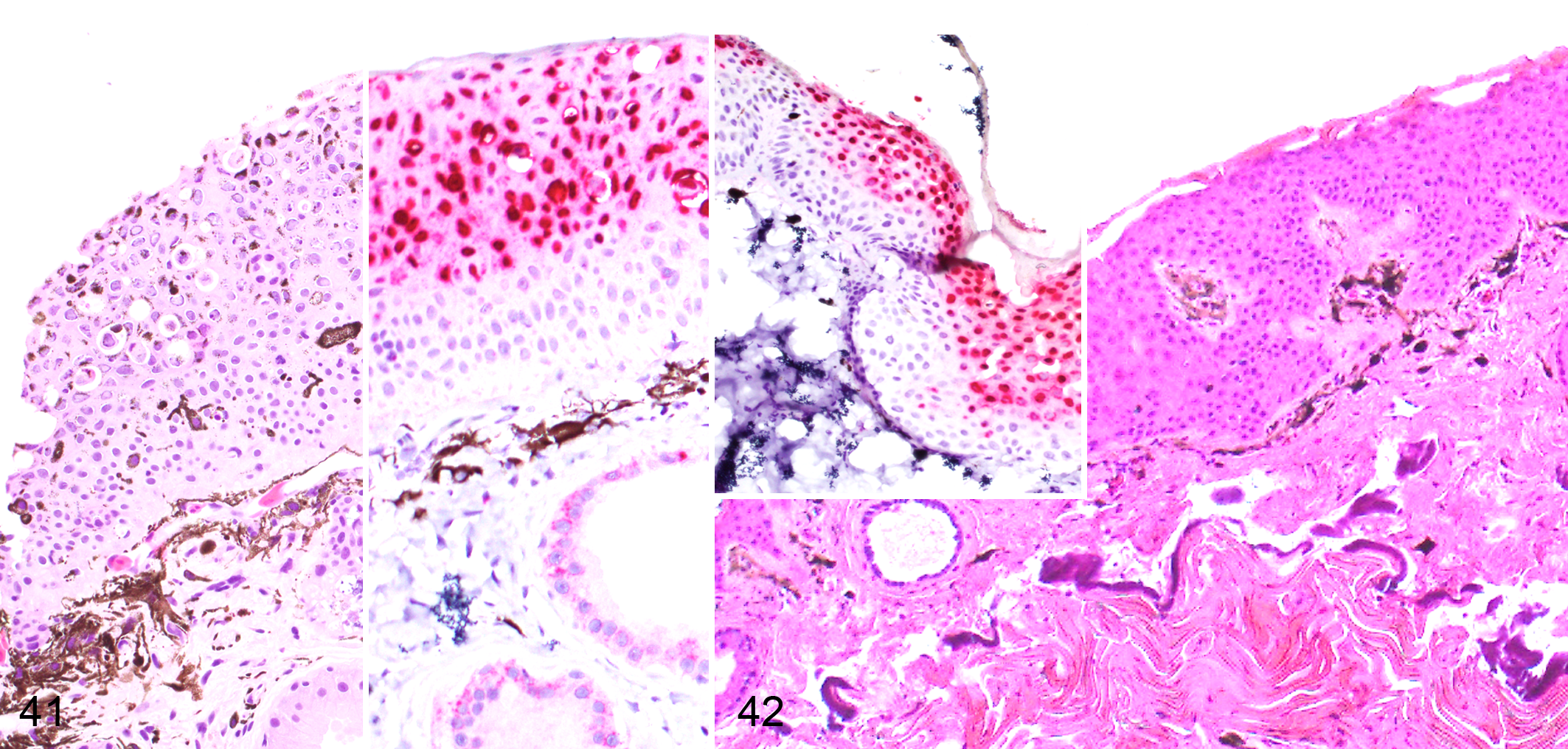
A composite figure to depict the proposed pathogenesis of herpesvirus-associated proliferative dermatitis in the common frog (Fig. 41) and common toad (Fig. 42). Viral gene expression appears to be confined to the superficial portion of the hyperplastic epidermis and this region also coincides with the distribution of viral inclusion bodies which will be progressively sloughed from the surface. In contrast, the deeper epidermal layer contains minimal or no viral nucleic acid, indicating a lack of viral replication. Hematoxylin and eosin. Insets: in situ hybridization for herpesvirus gene expression, red chromogen.
Frogs
In situ hybridization revealed that viral gene expression was limited to the upper portion of the hyperplastic epithelium. Rarely, the positive signal was present in a few keratinocytes within the deeper layers of the epidermis, but never in the basal layer (Figs. 37, 38, and 41). The signal was strongly nuclear with only occasional and limited involvement of the cytoplasm (Fig. 39). All of the most affected nuclei were in the upper half of the hyperplastic epidermis, whereas the few affected nuclei of the lower strata of the epidermis showed dense and sharp foci of positive signal within the nuclei, occasionally overlapping with the nucleolar regions (Fig. 39, inset). Multiple sharp foci of positive signals were also observed in the cytoplasm but limited to the upper portion of the epidermis. A light signal was observed in the cytoplasm of the keratinocytes, including those of the basal layer. However, this was interpreted as a nonspecific background because it was also observed in the negative control (uninfected skin; Fig. 37; data not shown). A prominent positive signal was observed in the cytoplasm of the epithelial cells of the mucous glands in the absence of a distinct nuclear stain (Fig. 37). Similarly, this signal was considered nonspecific because it was also observed using an irrelevant probe (O. ophiodiicola; data not shown).
Toads
In situ hybridization revealed strong viral gene expression in the upper portion of the hyperplastic epidermis. The signal was rarely seen in a few cells of the deeper layers of the epidermis but never in the basal layer (Fig. 38). The positive signal was mainly nuclear, with highly stained nuclei within the upper portion of the epidermis and small sharp foci in rare positive cells of the deeper layers (Fig. 38). A relatively strong signal with a patchy distribution was present in the cytoplasm of positive cells in BfHV1-infected toads, and these cells also had a dense network of positive cytoplasmic small cores and light filaments (Fig. 40). Occasionally, the entire cell was strongly positive (Fig. 40). A light signal was observed in the cytoplasm of the basal layer, which was interpreted as likely background on the basis of a similar signal level in the negative control (uninfected epidermis) in the same location (data not shown). No background signal was observed in the dermal glands of the infected toads (Figs. 37–42).
Comparison
In situ hybridization had similar positive signal distribution both in RaHV3-infected frogs and BfHV1-infected toads. In both species, the signal was confined to the upper portion of the hyperplastic epidermal layer, with rare positive cells in the lower epidermal layers. The positive signal was mainly nuclear in both species with a light and more inconsistent cytoplasmic pattern in toads. The labeling expanded in the nuclei from small and sharp cores, often corresponding to the nucleoli, to the whole nucleus. An evident compartmentalization of the lesions occurred in both RaHV3 infection in frogs and BfHV1 infection in toads (Figs. 37–42).
Transcriptomics
The amount of raw paired-end (PE) reads were comparable in all the libraries, with the exception of the GYB-31 library, corresponding to the BfHV1-free toad skin, which was approximately 30% higher than the others. However, the percentage of the mapped reads was relatively close across libraries (Supplemental Material 4). Similarly, the percentages of reads mapping to the 2 viral genomes were very similar, consistent with a proportional amount of viral transcription for both viruses at the time of sampling. No RaHV3 reads could be found in the tissues of the BfHV1-infected toads and no BfHV1 reads could be found in RaHV3-infected frogs, and neither RaHV3 nor BfHV1 reads could be found in the uninfected frog and toad samples (Supplemental Material 4).
Host Transcriptomics: Common Frogs
The transcriptomic analysis carried out on RaHV3-infected and uninfected common frogs allowed the identification of more than 45 000 transcripts. The highest Log2 ratio (upregulated genes) was equal to 1.007, whereas the lowest Log2 ratio (downregulated genes) was equal to −0.997. Given that no replicates were available and that the experimental samples were obtained by pooling the total RNA obtained from the skin of 2 uninfected and 2 RaHV3-infected common frogs, no significance threshold could be obtained. Accordingly, we adopted a highly conservative approach in evaluating our results. More specifically, we evaluated only the 50 most upregulated and the 50 most downregulated genes (out of the more than 45 000 examined). Furthermore, all selected genes had a Log2 ratio either higher than +0.9 (upregulated) or lower than −0.9 (downregulated; Supplemental Material 2). The biological functions determined for each gene identified 4 main functional groups including those involved with (1) signaling, (2) cell remodeling (cytoskeleton), (3) cell proliferation, and (4) immunity. Within the signaling group, we identified the following categories: (1) transduction, (2) RNA transcription/processing, (3) environment sensing, and (4) posttranslational modification. The genes classified in the cell remodeling (cytoskeleton) group were subdivided into 3 additional categories: (1) cell dynamics, (2) adhesion, and (3) intermediate filaments. The cell proliferation genes were divided into 2 categories: (1) DNA-binding molecules and (2) cell division (mitosis)-associated genes. Finally, the immunity genes included (1) antimicrobial, (2) innate immunity, and (3) adaptive immunity genes. A fifth gene group included all those genes for which no functional activity could be identified (nonclassified genes). The 50 most upregulated genes in RaHV3 infected common frogs included 20 signaling genes, 15 cell remodeling-associated genes, 3 cell replication-associated genes, 9 immune-associated genes, and 3 nonclassified genes (Supplemental Materials 2 and 5). Of the 50 most downregulated genes observed in RaHV3 infected frogs, 19 were signaling genes, 11 were cell remodeling genes, 4 were cell replication-associated genes, 6 were immune-associated genes, and for other 10 genes, the function could not be identified (Supplemental Materials 2 and 5).
Host Transcriptomics: Toads
A similar analysis was also carried out on BfHV1-infected and uninfected common toads. The identified transcripts showed a Log2 ratio ranging from 1.043 to −1.041, and we focused on the 50 most upregulated and the 50 most downregulated genes (with a Log2 ratio either higher than +0.9 [upregulated] or lower than −0.9 [downregulated]; Supplemental Materials 2 and 5). The same functional categories as for the frogs were identified for the toad genes and, accordingly, the 50 most upregulated genes clustered as follows: 22 signaling genes, 10 cell remodeling genes, 7 cell replication genes, 3 immunity-associated genes, and 8 nonclassified genes (Supplemental Materials 2 and 5). Of the 50 most downregulated genes, 23 were signaling genes, 6 were cell remodeling genes, 3 were cell replication genes, 8 were immune-associated genes, and 10 could not be identified (Supplemental Materials 2 and 5).
Comparison of Host Transcriptomics
The analysis of the upregulated and downregulated genes showed consistency across the 2 species and viral infections. In infected frogs and toads, the gene ranking revealed the signaling-associated genes to be the group of genes whose expression variation is most associated with viral infection and tissue changes (Supplemental Material 5). Cell (cytoskeleton) remodeling-associated genes ranked second, whereas immunity-associated genes ranked third and cell replication genes ranked fourth for infected frogs and toads (Supplemental Material 5). Immunity-associated genes were predominantly upregulated in infected frogs (9 vs 6) and predominantly downregulated in infected toads (8 vs 3). More specifically, frogs showed critically downregulated genes with antiviral activity including TRIM14- and Matrin 3-like, which are part of the innate defense against viruses (TRIM14) 29 and specifically against DNA viruses (Matrin 3) 22 such as herpesviruses (Supplemental Materials 2 and 5). Conversely, toads showed a predominant downregulation of antimicrobial and adaptive immunity-associated genes. Furthermore, 3 of the 4 overall most downregulated genes in infected toads were immune-associated. Specifically, these included the SLAM family member CD84 (a molecule critical for both adaptive and innate immunity) as first most downregulated, and mucin-5AC-like (a gel-forming glycoprotein relevant for skin protection) as second-most downregulated (Supplemental Material 2).
Involvement of Homologous Frog and Toad Genes
A closer comparative analysis of the upregulated and downregulated genes in infected frogs and toads revealed a small group (n = 7) of frog and toad homologous genes, which underwent the same proportional expression change in both species. More specifically, a total of 4 among the 50 most downregulated host genes in RaHV3-infected frogs were also highly downregulated in BfHV1-infected toads, although not included within the 50 most downregulated genes in this species. Similarly, 2 among the 50 most downregulated genes in BfHV1-infected toads were also strongly downregulated in RaHV3 infected frogs although not within the 50 most downregulated. Of these 7 genes, the ribosomal protein S15a (RPS15a) was consistently strongly downregulated in both species and it was the only gene out of the 7 genes identified that was included in the 50 most downregulated genes in both frogs and toads (Table 1 – boldface). Additionally, to RPS15a, this small group of homologous genes strongly downregulated in both infected frogs and toads included histone-encoding genes, a serine peptidase (spint2L), a microsomal glutathione S-transferase important for prostaglandin synthesis, a desmoplakin L-like gene critical for cell adhesion, and an L-aminopeptidase-like homologue (Table 1). 3 –6,16 –21,25,27,28,30 –34
Differentially Expressed Genes in the Skin of Frogs and Toads With Herpesvirus Infections.a
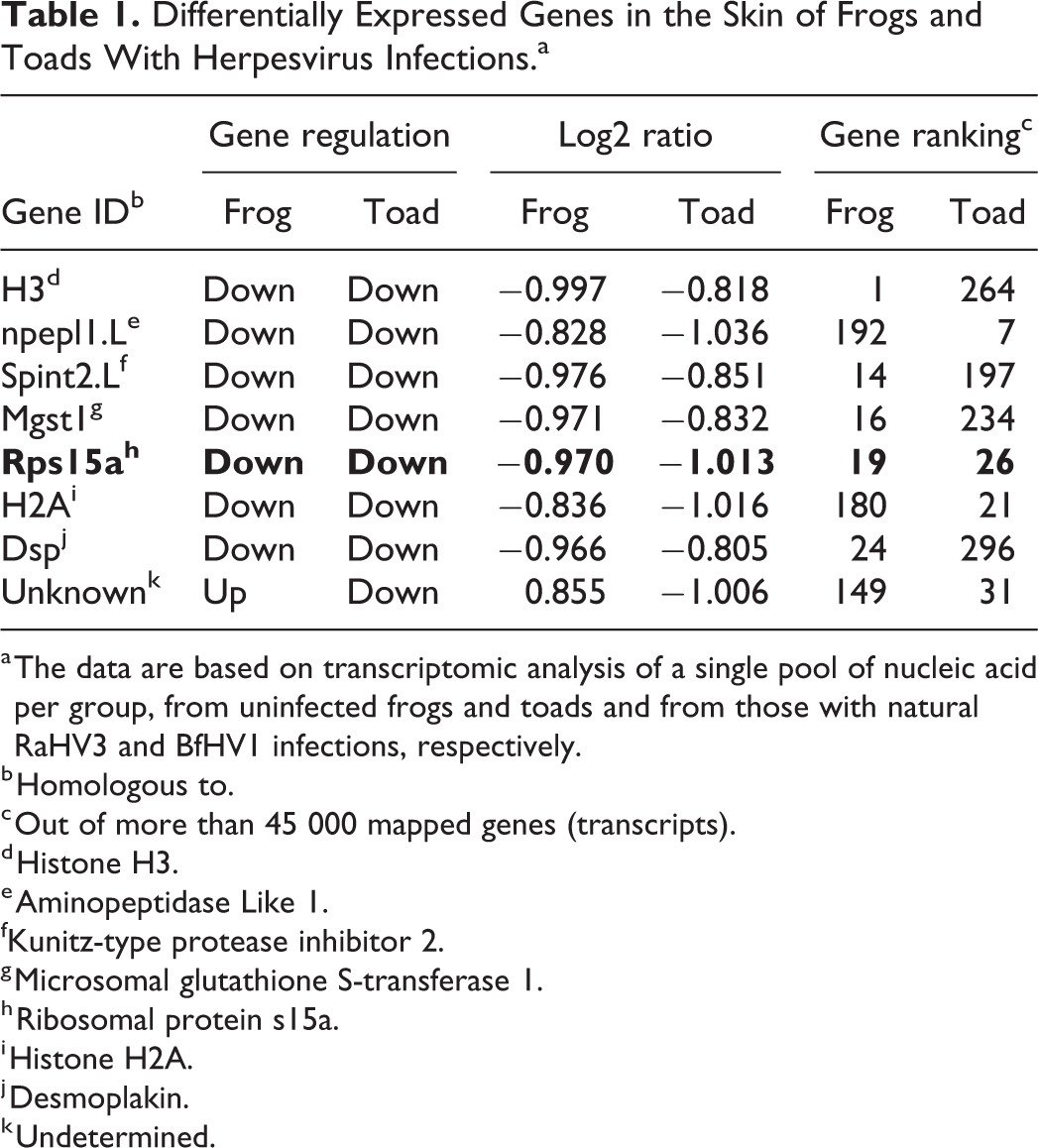
a The data are based on transcriptomic analysis of a single pool of nucleic acid per group, from uninfected frogs and toads and from those with natural RaHV3 and BfHV1 infections, respectively.
b Homologous to.
c Out of more than 45 000 mapped genes (transcripts).
d Histone H3.
e Aminopeptidase Like 1.
fKunitz-type protease inhibitor 2.
g Microsomal glutathione S-transferase 1.
h Ribosomal protein s15a.
i Histone H2A.
j Desmoplakin.
k Undetermined.
Viral Transcriptomics: RaHV3
RaHV3 viral reads aligned well with their annotated genomes (Genbank NC_034618) and confirmed their overall correctness. The tissue samples were collected from animals showing fully developed skin lesions. The normalized expression of the RaHV3 transcriptome showed a predominant cluster of C-type lectin-encoding genes among the most-expressed, with ORF124 being the most-expressed among them. Of the putative immunomodulatory genes, the 2 most-expressed were ORF22 and ORF23 encoding for a predicted apoptosis inhibitor and for a protein sharing strong similarities with the tumor necrosis factor receptor, respectively. ORF30 encoding for a B-22-like protein known to block T-cell activation, and ORF150 predicted to encode an IL-34-like protein known to enhance antiviral activity in macrophages, 1,12,13 recorded the highest number of reads (not normalized) among the predicted immunomodulatory genes. A RaHV3 gene predicted to encode for a protein resembling a well-known proto-oncogene (FOXM1) and cell proliferation-associated master transcriptional regulator of several human carcinomas 9 was also observed to be transcribed in infected frogs (Supplemental Material 3).
Viral Transcriptomics: BfHV1
BfHV1 viral reads aligned well with their annotated genome (Genbank MF143550.1) and confirmed their overall correctness. The normalized expression of the BfHV1 genome showed ORF19 and 20 (2 predicted C-type lectin proteins encoding genes) as the most expressed (Supplemental Material 3). Of the predicted immunomodulatory genes, the most expressed appeared to be ORF97, similar to a member of the TNF-receptor superfamily and homologous to RaHV3-ORF23.
Comparison of Viral Transcriptomics
Both the RaHV3 and BfHV1 transcriptome showed overlapping features including C-type lectin genes being the most expressed, and a TNF-R-like molecule as the most commonly expressed putative immunomodulatory gene. Interestingly, the BfHV1 transcriptome showed a drastic difference in the expression of the few top genes in comparison to the bulk of the others. Conversely, in the RaHV3 genome, the transition between the groups of the most expressed and least expressed genes was smoother (Supplemental Material 3). Relevant metabolic genes including the RaHV3 and BfHV1 homologs of RaHV1 ORF25, RaHV1 ORF26, RaHV1 ORF83, RaHV1 ORF40, major capsid protein, terminase, and RaHV1 ORF82 and RaHV1 ORF87 showed a similar expression ranking, consistent with similar roles and significance in their life cycles and, eventually, with their associated diseases. Additionally, in both the RaHV3 and BfHV1 transcriptome, the predicted metabolically relevant genes were transcribed at an obviously lower level than the most-expressed C-type lectin protein-encoding genes (Supplemental Material 3).
Discussion
The examination of multiple common frogs and toads infected with either RaHV3 or BfHV1, respectively, confirmed the findings originally described. 23,24 Additionally, the examination of individuals showing lesions at apparently different stages of development revealed undescribed aspects of the lesions that inform the pathogenesis in particular with respect to lesion development and the associated immune response. The skin lesions showed a striking compartmentalization, both in infected frogs and toads. Interestingly, the thickened hyperplastic epidermis could be easily subdivided into upper and deeper layers. The upper hyperplastic layer had the most severe changes including intranuclear inclusions, cellular vacuolization, degeneration, and necrosis. In contrast, the deeper layer of the hyperplastic epidermis showed a virtual absence of detectable tissue changes. Remarkably, the demarcation between upper and deeper layers corresponded to the “sloughing” line of the epidermis that was observed in infected frogs at a certain, probably advanced, stage of the disease. Although no sloughing was observed in the infected toads, a similar demarcation line was also obvious in this species, suggesting that a similar process could occur in toads. Accordingly, the failure to notice this process in toads might simply reflect the lack of specimens at that specific stage of lesion development.
The immunological features in infected frogs and toads reinforced the hypothesis of a compartmentalization of the lesions, with inflammatory cells almost completely confined to the deeper layers of the hyperplastic epidermis, paradoxically sparing the superficial layers where all the intranuclear inclusions—a proxy of viral activity—were observed.
Strikingly, in situ hybridization results paralleled the presumptive demarcation observed histologically, suggesting that the tissue compartmentalization mentioned above likely reflects a specific stage of the life cycle of the virus (replication and maturation) and of the associated lesions, with heavy viral activity in the superficial layers and no detectable virus in the deeper layers. The hybridization technique revealed that viral spread into the cell likely started from single sharp foci within the nuclei.
Ultrastructural pathology allowed us to unveil critical steps concerning virion maturations in infected frogs and toads, and surprising differences concerning the intracellular transport of the virions were observed. Our investigation suggested the existence of a system apparently able to process large number of particles in RaHV3-infected frogs in comparison with a relatively low number in BfHV1-infected toads. Interestingly, we observed few naked capsids in the cytoplasm of RaHV3 infected cells. These were observed in close proximity of the nuclei and they likely reflect the loss of the primary envelope after the egress from the perinuclear cisterna. 14
Transcriptomics was considered an important tool to partially dissect the pathogenesis of these viral diseases and to uncover preliminary data, although an important caveat for a correct interpretation of these data is the absence of replicates. Potentially, some transcriptomic differences between RaHV3-infected frogs and BfHV1-infected toads might have been influenced by the selected sampling and data collection strategy.
An interesting overlap was observed between the transcriptional regulation occurring in herpesvirus-infected frogs and toads with a high number of signaling and cell remodeling-associated genes, consistent with a likely articulated and presumptively coordinated proliferative and degenerative process that the affected skin of these amphibians is undergoing. Interestingly, 7 homologous gene identities were highly downregulated both in infected frogs and toads, consistent with a putative key role in the pathogenesis. The overall most downregulated of them in both frogs and toads was the RPS15a gene, a gene involved in carcinogenesis in humans and other mammals. 5,20,31,33 Interestingly, this gene appears to promote cancer when upregulated whereas its negative effect is blocked once its expression is suppressed. Differently, the other 6 homologous genes highly downregulated in both infected frogs and toads have been reported to be associated with carcinogenesis and metastasis proliferation when their expression is hampered.
Within the limits of the preliminary nature of the transcriptomic data, it is suggestive to observe that both host and viral genes (such as RaHV3-FOXM1) relevant for cell proliferation might be critically involved in the pathogenesis of the disease. Accordingly, cell proliferation, the first hallmark of the associated disease, seems to mirror a complex balance between host and, likely, viral gene regulation.
Similarly, transcriptomics was instrumental to better understand the immune-associated dynamics. The overall immune transcriptional changes revealed an apparently stronger downregulation in toads. This was mediated by the downregulation of genes putatively encoding for large spectrum antimicrobials, similar to what has been reported in chytrid infections in amphibians, 10 and the involvement of an overall higher number of genes, individually characterized by a broad activity spectrum such as CD84, a key molecule bridging innate and adaptive immunity. However, the lower number of downregulated immune genes in frogs might not necessarily reflect a less significant immunomodulation, but simply a more targeted one with involvement of genes known to be relevant in anti-DNA virus activity. 22,29 Finally, and in addition to the host immune-relevant molecules, an immunosuppression activity might be virus-mediated, as the presence of a strong transcription of the TNF receptor-like gene encoded by both RaHV3 and BfHV1 would indicate, within the limits of the preliminary nature of these data. Interestingly, TNF has been shown to be a key player in herpesvirus-associated pathology in fish 26 by mediating the development of a beneficial “fever,” instrumental for the host to fight the viral agent. The heavy production of a putative TNF receptor-decoy by both RaHV3 and BfHV1 indirectly suggests a relevant role of TNF in both these species. The remarkably poor cellular immunity observed in the epidermis might be secondary to a coordinated activity mediated on one side by viral factors and on the other side by a balance of the host’s upregulated and downregulated immunorelevant genes.
The observations of natural disease cases made in this investigation lead to a presumptive pathogenesis of RaHV3- and BfHV1-associated diseases in frogs and toads, respectively. Following viral infection, the epidermis would undergo a strong proliferative activity. The exuberant, newly formed epidermis would then provide the necessary substrate for the viral replication. The viral progeny would be released into the intercellular space, which would progressively expand to form variably extensive vacuoles, ending in cell degeneration and necrosis with sloughing of the upper portion of the hyperplastic epidermis resulting in the shedding of the newly produced virions. The basal layer would be spared, allowing restitution of the epidermis to its normal structure after the upper hyperplastic layers are sloughed. The transcriptomic data provided a solid foundation to this hypothesis highlighting that signaling and cell-remodeling genes are those most transcriptionally affected during the infection. We hypothesize that viral replication is mainly restricted to the superficial epidermis because there it is shielded from the host immune response both directly (viral immunomodulatory genes, such as TNFr-like molecules) and indirectly (specific upregulation or downregulation of immune-relevant host genes), explaining the striking compartmentalization of the lesions, including the distribution of the leukocyte infiltration, observed both in infected frogs and toads. Finally, the process would restart the following season with the reactivation of the herpesviruses from a presumptive latency, bringing the process to a full cycle (Fig. 43). This would imply a potential recurrent nature of the disease, which needs to be demonstrated together with the infectious route of the viruses and the overall clinical significance of RaHV3 and BfHV1 and their associated diseases for free-ranging amphibian populations. The understanding of these aspects is of great interest for the overall disease ecology of anuran herpesviruses and for a better understanding of the molecular mechanisms behind the proliferative nature of DNA virus-associated diseases.
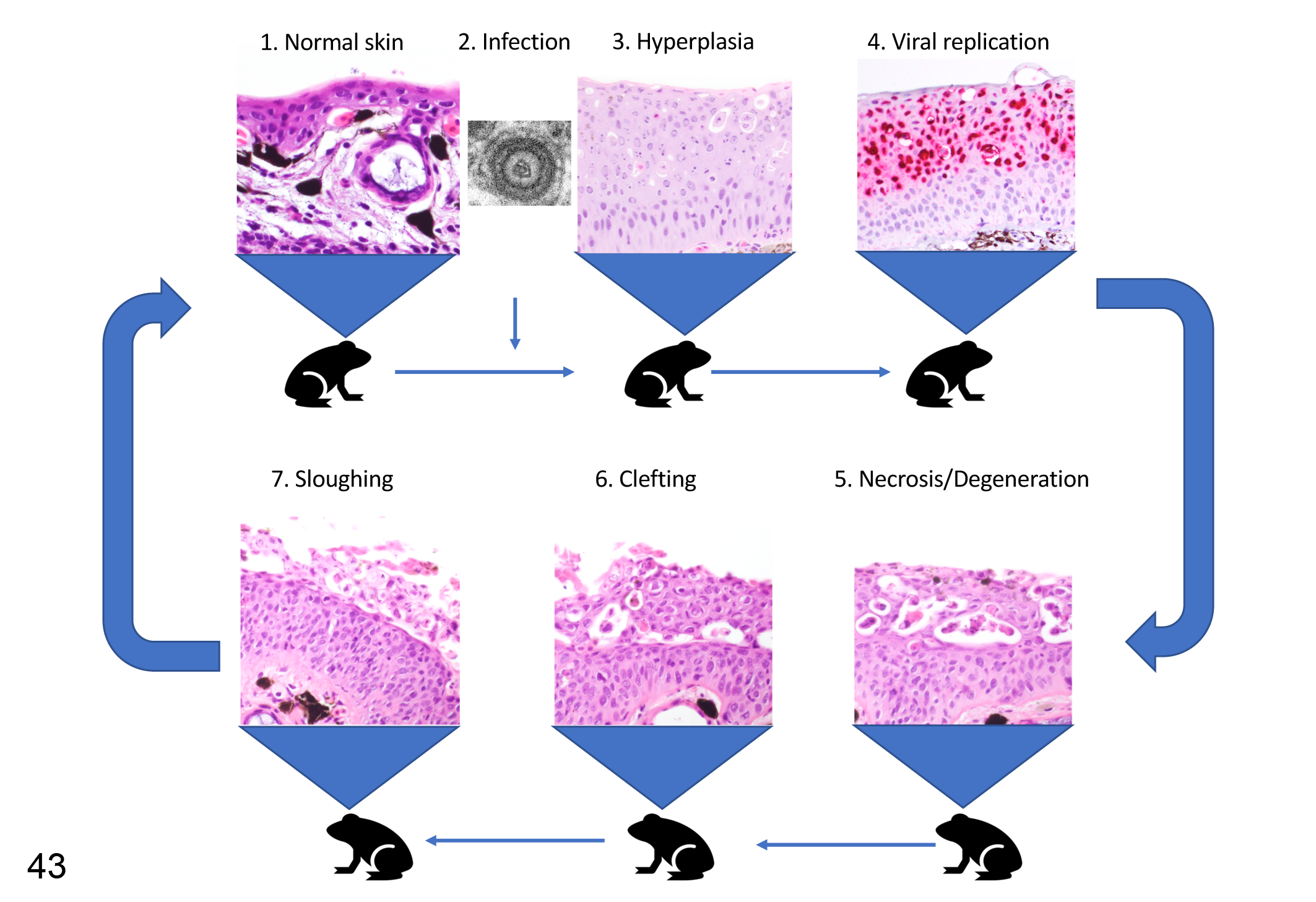
Proposed pathogenesis of herpesvirus-associated proliferative dermatitis. The skin (1, normal; hematoxylin and eosin [HE]) is infected with the herpesvirus (2, transmission electron microscopy) and subsequently undergoes epidermal hyperplasia (3, HE) with viral replication limited to the superficial layers (4, in situ hybridization for herpesvirus gene expression). Progressive epithelial degeneration and necrosis leads to formation of vacuoles in the superficial epidermis (5, HE), which enlarge to form a cleft between the superficial and deeper epidermal layers (6, HE) leading to sloughing of the superficial layer (7, HE). We expect that the affected skin would then progressively return to the thickness and morphology of normal skin (1). Reactivation of viral replication during the following season may then restart the pathologic process. These images are from RaHV3-infected common frogs, but the implications of the predicted pathogenesis are intended for both RaHV3 and BfHV1.
Supplemental Material
Supplemental Material, sj-pdf-1-vet-10.1177_03009858211006385 - Herpesvirus-Associated Proliferative Skin Disease in Frogs and Toads: Proposed Pathogenesis
Supplemental Material, sj-pdf-1-vet-10.1177_03009858211006385 for Herpesvirus-Associated Proliferative Skin Disease in Frogs and Toads: Proposed Pathogenesis by Francesco C. Origgi, Patricia Otten, Petra Lohmann, Ursula Sattler, Thomas Wahli, Antonio Lavazza, Veronique Gaschen and Michael H. Stoffel in Veterinary Pathology
Footnotes
Acknowledgements
This work was made possible by a number of people that I am indebted with, including the pathology residents, interns, doctoral students, assistants, and visiting students of the Centre for Fish and Wildlife Health (FIWI) of the University of Bern (Ezgi Akdesir, Roman Mayer, Gaia Moore Jones, Simone Pisano, Marco Tecilla, Ursula Teubenbacher), Professors Helmut Segner and Marie-Pierre Ryser for their constant support and the technical personnel of the FIWI. I am very grateful to Dr Benedikt Schmidt for his expertise, support, and collaboration. A special thanks to Dr Francis Cordillot and Dr Beatrice Werffeli for their special attention to these topics. Finally, I am in debt with all the amphibian volunteers and biologists in the field, who are the foundation of this whole investigation.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was partially supported by the Vontobel Foundation (a special thanks to Mrs Jeannette Frei for her help and support) and by the Federal Office of the Environment (FOEN; Grants R212-1468 and R414-1866).
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
