Abstract
Lymphoma is among the most common cancer in dogs. Diffuse large B-cell lymphoma (DLBCL) is the predominant type, accounting for up to half of all cases. Definitive diagnosis of DLBCL relies on cytologic evaluation with immunophenotyping, or histopathology and immunohistochemistry when needed. A rapid and specific molecular test aiding in the diagnosis could be beneficial. Noncoding microRNAs (miRNAs) are regulators of gene expression involved in a variety of cellular processes, including cell differentiation, cell cycle progression, and apoptosis. Not surprisingly, miRNA expression is aberrant in diseases such as cancers. Their high stability and abundance in tissues make them promising biomarkers for diagnosing and monitoring diseases. This study aimed to identify miRNA signatures of DLBCL to develop ancillary molecular diagnostic tools. miRNA was isolated from formalin-fixed, paraffin-embedded lymph node tissue from 22 DLBCL and 14 nonneoplastic controls. Relative gene expression of 8 tumor-regulating miRNAs was achieved by RT-qPCR (reverse transcriptase quantitative polymerase chain reaction). The results showed downregulation of the let-7 family of miRNAs and miR-155, whereas miR-34a was upregulated in DLBCL compared to the controls. We demonstrated that the combination of expression levels of miR-34a and let-7f or of let-7b and let-7f achieved 100% differentiation between DLBCL and controls. Furthermore, let-7f alone discriminated DLBCL from nonneoplastic tissue in 97% of cases. Our results represent one step forward in search of a rapid and accurate ancillary diagnostic test for DLBCL in dogs.
Lymphoma is a heterogeneous disease that represents malignant transformation of lymphocyte subtypes at different stages of development. Lymphoma is the most common spontaneous malignant tumor in dogs, affecting all breeds as well as mixed-breed dogs. 37,47 Diffuse large B-cell lymphoma (DLBCL) is the most common type of lymphoma in dogs, comprises approximately 38% to 48% of cases, 29,34,47 and generally behaves aggressively. 4,21,29 Most dogs diagnosed with lymphoma will ultimately die of their cancer, as current therapies are not curative. The majority of the DLBCL cases can be diagnosed by cytology, but this cannot differentiate the subtype, which is essential for predicting behavior and determining adequate treatment. 4 Despite advances in molecular characterization of cancers, the current approach for subtyping lymphoma relies on histopathologic evaluation of a biopsy, which is an invasive procedure and can take up to 2 weeks to produce comprehensive diagnostic results. Given the heterogeneous nature of lymphoma subtypes and differences in prognosis, additional immunophenotyping through immunocytochemistry (IHC), flow cytometry, and polymerase chain reaction (PCR) for antigen receptor rearrangements (PARR) may be required. Thus, molecular biomarkers for accurate and rapid diagnosis of DLBCL are highly desirable.
MicroRNAs (miRNAs) have shown the possibility of being useful biomarkers for rapid diagnosis and prognostic analysis from tissue and body fluids. 7,24,26,27,43,50 MicroRNA research is relatively new with the first publication in 2002, which reported the downregulation and deletions of microRNAs in chronic lymphocytic leukemia in humans. 9 This discovery led to an increased interest in the diagnostic and therapeutic potential of microRNA-based biomarkers in cancer and other diseases. 8
Noncoding miRNAs are posttranscriptional regulators of gene expression of various cellular processes, including cell differentiation, cell cycle progression, and programmed cell death. 20,22 Thus, they also play an essential role in carcinogenesis as they have either oncogenic or tumor suppressor functions. 1,14 Furthermore, miRNAs can circulate in association with protein complexes or small vesicles (exosomes) and lipoproteins, making them stable in solid tissues and body fluids despite the presence of RNases. 38 They are also resistant to boiling, desiccation, and several freeze-thaw cycles, and can be reliably extracted from formalin-fixed, paraffin-embedded (FFPE) tissues. 15
miRNAs play a critical role in physiologic lymphocyte development and differentiation. 5 In humans, studies on the miRNomes of normal lymphocytes showed that mature B-cells, memory cells, and cells from the germinal centers have distinct miRNA profiles, and the expression of most miRNAs is specific to the stage of differentiation. 5 As expected, several studies have shown that the malignant counterparts to normal B-cells—DLBCL, chronic lymphocytic leukemia, follicular lymphoma, and mantle cell lymphoma—also have distinct miRNA signatures. 5
Evidence of miRNA dysregulation in dogs has been demonstrated, where miR-17-5p and miR-181a were upregulated in T-cell and B-cell lymphoma, respectively. 31,46 Studies with cell lines showed the upregulation of miR-19a+b, an inhibitor of apoptosis in T-cells, and downregulation of miR-218, a tumor suppressor in B cell lines. 46 Differentially expressed miRNAs were also identified in the serum of dogs with lymphoma by RT-PCR (reverse transcriptase-polymerase chain reaction). Of the 5 miRNAs analyzed, 4 (let-7b, miR-233, miR-92a, and miR-25) were downregulated, whereas miR-423a was upregulated compared to controls. 17 More recently, a larger study confirmed the dysregulation of miRNAs in canine lymphoma, and showed upregulation of the miR-181 family and downregulation of miR-29b and miR-150 were associated with decreased therapy response and shorter survival, while the upregulation of miR-155 and miR-222 were negatively associated with outcome in both B and T cell lymphoma. 13 Thus, this study aimed to identify miRNA signatures of DLBCL to be used to develop ancillary diagnostic tools and serve as a core for future studies of miRNA regulation in DLBCL.
Material and Methods
Animals
This retrospective study used 22 FFPE archival lymph node biopsy specimens collected from privately owned dogs presented to the Purdue University Veterinary Teaching Hospital (PUVTH), West Lafayette, IN, from 2006 to 2016, which were sampled at the time of diagnosis of primary nodal DLBCL. 12 Except for 2 emergency visits, all dogs were referred to the PUVTH Oncology Department for confirmation, further testing, and/or treatment of lymphoma. Surgical biopsies from affected lymph nodes included incisional wedge biopsies or entire lymph node removal. DLBCL was confirmed by histopathology and immunohistochemistry (positive for at least one of CD79a, Pax-5, or CD20, and negative for CD3). Breed, age, sex, presenting complaint, and pathological data from these dogs are shown in Supplemental Table S1. The controls used in the study were 14 archived FFPE, nonneoplastic lymph nodes from dogs used for teaching that had no significant hematologic or biochemical changes (based on evaluation by a board-certified clinical pathologist, APS). After selection of cases and controls, all histologic sections were reevaluated by a board-certified pathologist (Ramos-Vara). The samples available for this project were collected with informed consent from the dogs’ owners and were approved under the animal use protocols PACUC#1211000780 and PACUC#1111000308.
miRNA Extraction and Reverse Transcriptase Quantitative PCR
Total RNA was extracted from one 20-µm scroll from the paraffin blocks of the 36 FFPE lymph nodes using the miRNeasy FFPE kit (QIAGEN Inc) according to the manufacturer’s specifications. Quantity and quality of total RNA were assessed by capillary electrophoresis (Bioanalyzer, Agilent Technologies) for 8 samples and UV spectrophotometry (NanoDrop, ThermoFisher Scientific) for all samples.
First-strand cDNA was synthesized by reverse transcription using miScript II RT Kit (QIAGEN) using 500 ng of miRNA, according to the manufacturer’s specifications. Reverse transcriptase quantitative PCR (RT-qPCR) was performed using the miScript SYBR Green PCR Kit (QIAGEN) on a QuantStudio3 PCR system (ThermoFisher Scientific) using RNU6B as a normalizer. The samples were run in triplicate. Relative expression levels were calculated by the 2−ΔΔCt method. 5 Primer assays for canine miRNAs, let-7a, let-7b, let-7c, let-7e, let-7f, let-7g, miR-155, miR-34a, and the normalizer RNU6B were acquired from QIAGEN. No-template controls (NTC) were used in all plates.
Statistical Analysis
All analyses were performed using R version 3.4.1. Analyses had 2 goals. First, we sought to determine which of the 8 miRNAs analyzed were differentially expressed in DLBCL cases versus controls, and to quantify the differences in mean expression between the 2 groups. Second, we conducted a discriminant analysis to identify a combination of miRNAs that might be used to efficiently classify individuals as DLBCL-positive or -negative.
In the first analysis, we used a permutation-based multivariate analysis of variance (MANOVA) to screen for overall differences in mean miRNA expression between the case (DLBCL) and control groups, followed by Welch 2-sample t tests to examine differences in the fold-change in expression of each miRNA. MANOVA provides a global test for differences among 2 or more groups in a multivariate response (ie, relative expression of 8 miRNAs). In comparison to a series of univariate ANOVAs, initial screening by MANOVA minimizes the risk of false positives due to unnecessary multiple testing, and may also reveal multivariate differences among groups that cannot be detected by the univariate methods alone.
MANOVA assumes that response variables are multivariate normal within each group and that the groups have similar covariance matrices (this property, called sphericity, extends the assumption of equal variance from univariate ANOVA to a multivariate context). To achieve normality, we log10-transformed the fold-change expression data to correct for right skews in the data for all 8 miRNAs. However, we were unable to meet the sphericity assumption (Mauchly’s test of sphericity on log10 fold-change: W = 0.0020, P value < .0001). As a nonparametric alternative, we applied a permutational MANOVA, 3 as implemented in the Adonis function of the vegan package in R. 49 Permutation repeatedly reshuffles the group labels that are associated with individual data-points and recalculates a test statistic. This procedure breaks any association between the explanatory and response variables in the permuted data and yields an approximate distribution for the test statistic under the null hypothesis of no difference between the groups.
Our second analysis used Bayesian logistic regression to model the odds of membership in the DLBCL group as a function of miRNA expression and identify a combination of biomarkers to use in lymphoma diagnosis. Specifically, we fitted all possible additive models with a combination of 1, 2, or 3 miRNAs (92 models in total), and then compared the models using Akaike’s information criterion (AIC). AIC is a measure of parsimony; it decreases with model fit and increases with the number of model parameters (smaller scores are better). We selected a Bayesian approach because several of the models in our candidate set perfectly discriminated DLBCL cases from controls in our data, and conventional methods cannot estimate regression parameters under these conditions. 18 To address this issue, Bayesian methods specify a prior probability distribution for the parameter values, which identifies the relative plausibility of different values before accounting for the information in the data. The data are then used to refine this distribution. We performed the analysis using the bayesglm function in the ARM package in R 32 and a “weak” prior recommended by Gelman and collaborators. 18 We report the results only for those models that correctly predicted DLBCL with >95% accuracy. Model goodness-of-fit was evaluated by comparing the Hosmer-Lemeshow χ2-statistic for the actual data to null distributions generated using data simulated from the fitted model. 19
Results
The average total RNA concentration measured by UV spectrophotometry was 229.9 ng/µl (±100 ng/μl), and the ratio of absorbance at 260 nm and 280 nm was 1.95 (±0.15). Results from the capillary electrophoresis are shown in Supplemental File S2. All samples tested had amplifiable miRNA by qPCR. RNU6B was chosen as a normalizer because it showed better stability regarding the level of expression across samples with a mean Ct = 20.3 for all normal samples (range = 19.1–21.7) and a mean Ct = 20.1 for all DLBCL samples (range = 19.2–21.4) across all qPCR plates. Mean Ct values for each gene tested are shown in Supplemental Table S3. Primer specificity was verified by analyzing dissociation curves. Mature miRNA PCR products showed single peaks from the specific amplification products with expected melting temperatures (74–76 °C) for mature miRNAs, according to the manufacturer’s instructions.
Permutational MANOVA showed significant separation in the multivariate locations of the DLBCL and control groups (F = 18.993, P = .0002, based on 5000 permutations). When analyzed individually, the miRNAs let-7a (P = .0004662, t = 3.8737, df = 33.883), let-7b (P = 1.15e-05, t = 5.1374, df = 33.878), let-7c (P = .0007556, t = 3.7169, df = 32.585), let-7e (P = 1.578e-08, t = 7.9491, df = 26.862), let-7f (P = 1.619e-09, t = 8.1916, df = 33.6), let-7g (P = 6.526e-06, t = 5.3669, df = 32.508), and miR-155 (P = .02739, t = 2.3295, df = 27.573) were downregulated in the DLBCL group compared to the control group. In contrast, miR-34a (P = .002373, t = −3.307, df = 31.339) was upregulated in the DLBCL group compared to the control group (Figs. 1, 2). miRNA levels in let-7 family were all strongly or moderately correlated with each other, and more weakly correlated with miR-155. In contrast, miR-34a showed a weak negative correlation with miR-155, but had no significant correlations with any of the let-7 miRNAs.
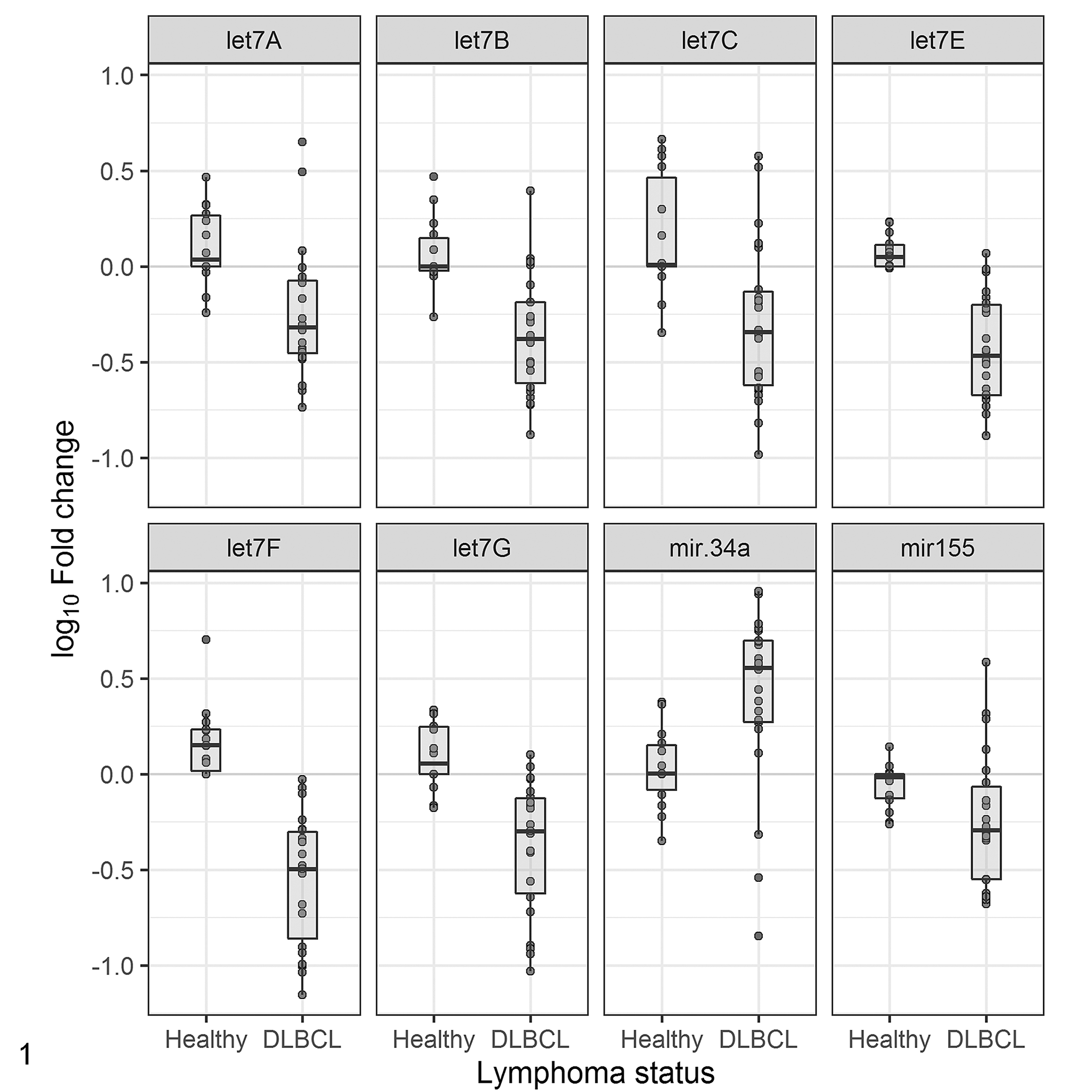
Boxplot comparing log10-transformed relative miRNA expression of let-7a, b, c, e, f, g, miR-155, and miR-34a between control nonneoplastic lymph node (N = 14) and diffuse large B-cell lymphoma (DLBCL; N = 22). Value >0 indicate upregulation relative to the control, and values <0 indicate downregulation based on Welch 2-sample t tests. Points indicate individual data values. Quartiles and medians are indicated by the boxes and the horizontal bar. MicroRNAs downregulated in DLBCL compared to controls included some let-7 family members (P ≤ .001) and miR-155 (P = .02739), whereas miR-34a was upregulated in DLBCL compared to controls (P = .002373).
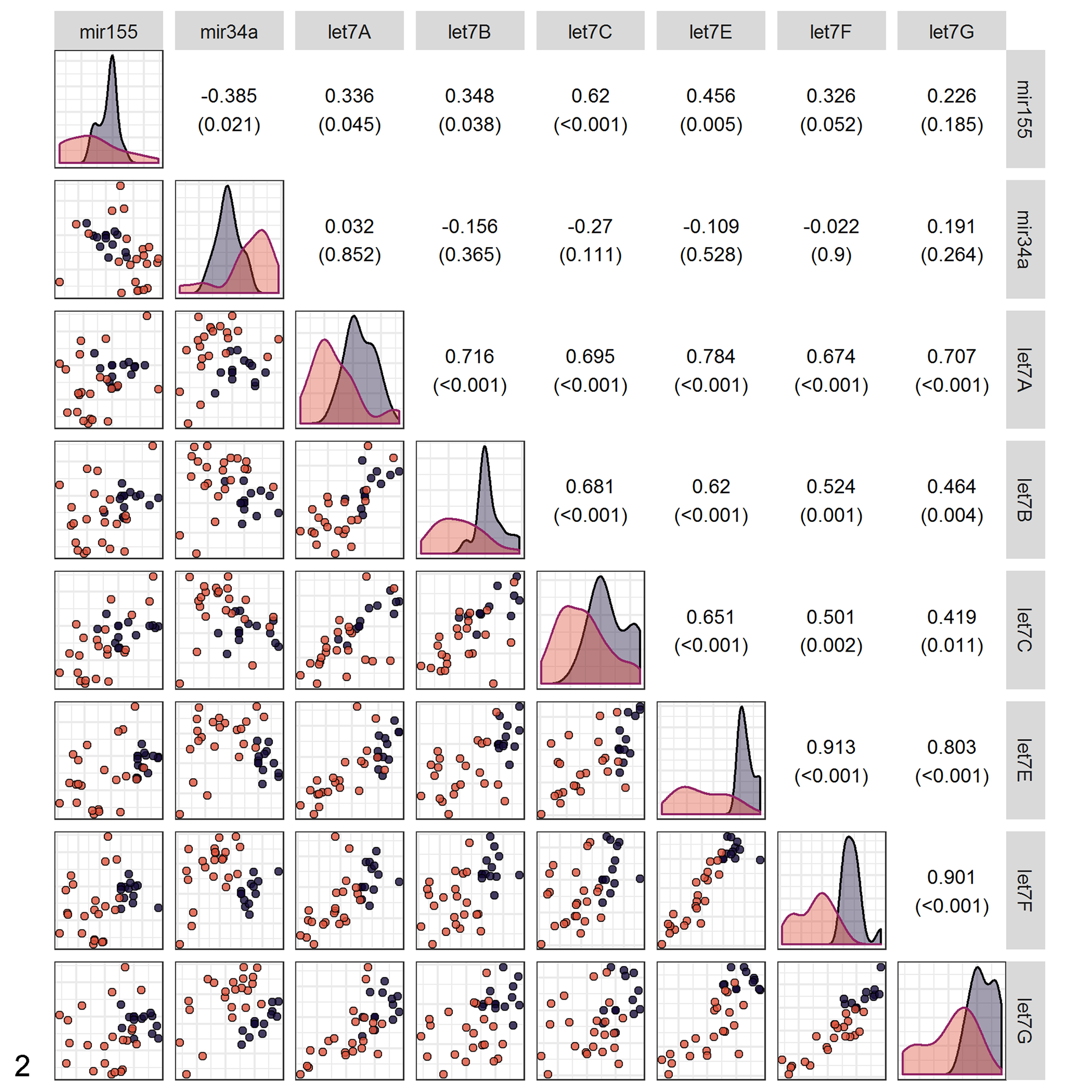
Pairwise relationships among the 8 differently expressed miRNAs in 36 dogs with diffuse large B-cell lymphoma (DLBCL) or nonneoplastic lymph nodes. Values in the upper right of the matrix give the Pearson correlation (and P value) for the corresponding scatterplot in the lower left of the matrix. Plots along the diagonal show kernel density estimates for the distribution of expression of each miRNA, broken out by lymphoma status. Colors show controls (dark purple) versus DLBCL samples (light orange). Note the clear separation of clusters in the plot of let-7f versus miR-34a.
Based on these results, the AIC-best model for discriminating DLBCL and control groups used a combination of miR-34a and let-7f (Table 1, Fig. 2). This was closely followed by the model with let-7b and let-7f, and then by let-7f alone. The remaining models possessed similar discriminatory power but were more complex (they all contained 3 markers, rather than 1 or 2 markers). All of the reported models provided a reasonable fit to the data (Table 1). In general, a goodness-of-fit test examines the null hypothesis that a particular model could have generated the observed data. In this case, the Hosmer-Lemeshow χ2 was used to check the assumptions in a logistic regression. The nonsignificant P values mean that any of these models could plausibly have generated the data demonstrated. Thus, there are no apparent glitches with the model fit that was used.
Combination of miRNA Markers According to the Akaike Information Criterion (AIC) Value Used to Select the Best Combination for the Diagnosis of DLBCL in Dogs. All Additive Models With 3 or Fewer miRNA Markers Were Considered. The Models That Correctly Predicted DLBCL >95% Are Reported†.
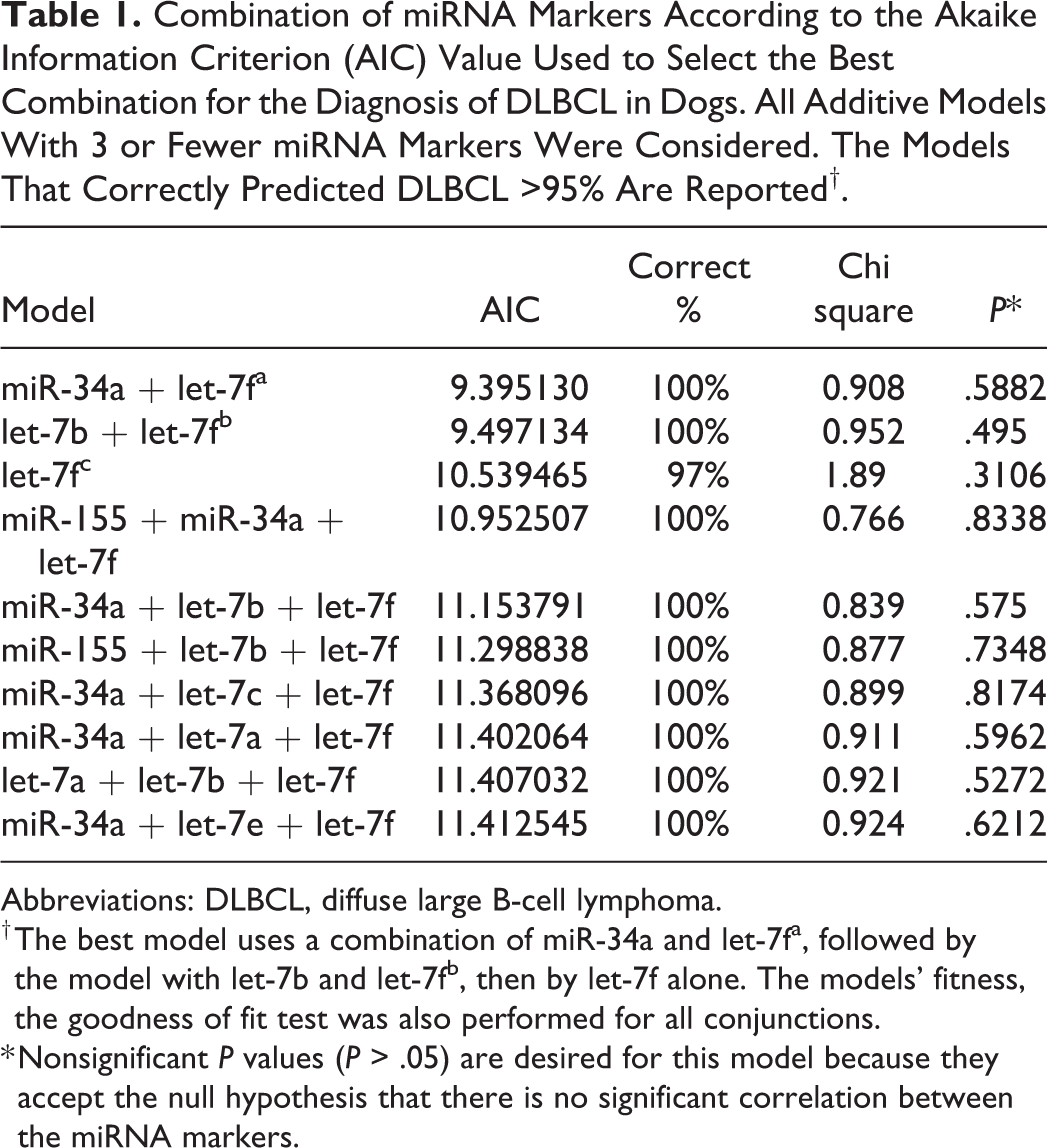
Abbreviations: DLBCL, diffuse large B-cell lymphoma.
† The best model uses a combination of miR-34a and let-7fa, followed by the model with let-7b and let-7fb, then by let-7f alone. The models’ fitness, the goodness of fit test was also performed for all conjunctions.
* Nonsignificant P values (P > .05) are desired for this model because they accept the null hypothesis that there is no significant correlation between the miRNA markers.
Discussion
Lymphoma is a complex type of cancer, making it challenging to achieve an accurate and rapid definitive diagnosis. Currently, the definitive diagnosis of lymphoma relies on the morphologic evaluation from a biopsy (fine needle aspirate or histopathology) of the affected lymph node or other affected organs in conjunction with immunophenotyping. The expression of CD79a or Pax-5 and absence of CD3 expression in neoplastic cells is commonly required for confirmation of DLBCL by IHC. PARR can be used to support the diagnosis of lymphoma based on clonality with approximately 94% specificity for lymphoid neoplasia and 75% sensitivity. 6 Flow cytometry phenotyping has a superior sensitivity than PARR; 45 however, fresh cells are not always available to run the test. 51 In general, the cost and availability of these procedures are limiting factors for definitive diagnosis with subtyping in some patients. Thus, molecular diagnostic tools that could differentiate between DLBCL and healthy lymphoid tissues are valuable. Furthermore, further investigations to accurately differentiate DLBCL miRNA profile from other lymphoma subtypes are highly desired.
Our study identified 8 miRNAs that were differentially expressed in DLBCL compared to nonneoplastic lymph node. Previous studies have shown that the expression of miRNAs extracted from FFPE tissue correlates well with fresh-frozen samples in lymph node/lymphoma samples 15,28,42,53 despite the degradation of longer RNAs, cross-linking, chemical modifications, and regardless of storage time. Both the lymph node controls and DLBCL FFPE samples were processed using the same reagents and protocols and over the same period (batches of controls and DLBCL samples were processed together) for reliability and consistency in all steps from preparation to data analysis. We were able to acquire extracted miRNA in high concentrations and with high quality based on UV spectrophotometry. Moreover, we were able to amplify miRNA from all samples tested.
The members of the let-7 family of miRNAs were discovered in Caenorhabditis elegans; these miRNAs are expressed during cell development at different stages. 36 The let-7 family is composed of tumor suppressor miRNAs that are conserved among different animal species from worms to humans. 33 Studies in human B-cell lymphoma have shown that the oncogenic transcription factor c-Myc suppresses the let-7 family by binding to the upstream conserved region of the let-7 gene. 11 The study herein reports that the members of the let-7 family expressed in dogs are significantly downregulated with variable significance, with the most significant being let-7f, followed by let-7e and let-7g (Fig. 1).
The miRNA miR-34a is widely expressed in human cells and is one of the direct transcriptional targets of the P53 gene. 35 P53 is a tumor suppressor protein, which modulates cell fate 34 and transcriptionally induces the expression of the pro-apoptotic miR-34a. 10 To some extent, miR-34a has a critical role in apoptosis mediated by P53 35 as well as an antiproliferative effect. 23,35 MiR-34a is downregulated in several human and animal cancers, including human cancers with p1chromosomal deletions, which is the region that encodes miR-34a in humans. 52 Surprisingly, we found that miR-34a is upregulated in canine DLBCL compared to nonneoplastic lymph node controls. Similar results were reported in a study on canine urothelial carcinoma when compared to nonneoplastic and inflamed bladder tissue, 48 and, more recently, in canine lymphoma. 13 These results suggest that the dog might have different mechanisms of miR-34a gene regulation.
miR-155 has a robust physiological role in hematopoiesis and inflammation, but also has a pathological role in tumorigenesis as it is considered an oncogenic miRNA. 16 miR-155 is upregulated in FFPE lymph node tissue from humans diagnosed with DLBCL 54 and is highly detected in human thymus and spleen. 30,44 Furthermore, the expression level of miR-155 is higher in activated B-cells than in germinal center B-cell of human DLBCL. 27 This aberrant expression of miR-155 seen in different types of lymphoma in humans as well as in canine splenic lymphoma, where the level of expression of miR-155 correlates with tumor behavior, 2 spiked our interest in examining its expression in canine DLBCL. Unlike many studies on human lymphoma, we found that miR-155 is downregulated in DLBCL in dogs. A recent study also showed the downregulation of miR-155 and demonstrated that the downregulation was associated with the outcome of canine lymphoma. 13 As with miR-34a, further investigation on the role of this miRNA in canine DLBCL is warranted.
Last, we were interested in finding the best combination of markers that could differentiate between DLBCL and nonneoplastic lymph node controls for all the cases while keeping the model as simple as possible with the fewest number of miRNA markers. The “AIC-best” model using let-7f and miR-34a fulfilled this goal, in part because the expression of these 2 miRNAs is uncorrelated (r = −0.021) and, thus, they provide separate pieces of information about a patient’s DLBCL status. However, the other models in Table 1 also provided good discrimination between DLBCL and control groups, and the miRNAs in these models may prove useful if additional markers are needed when comparing DLBCL to other types of lymphoma and reactive lymph nodes.
To the authors’ knowledge, this study is the first to approach the diagnosis of canine DLBCL using a combination of 1, 2, or 3 miRNA markers. Using the combination of miR-34a and let-7f or let-7b and let-7f, we achieved 100% differentiation between controls and the DLBCL dogs in this study, and we found that let-7f alone discriminated non-neoplastic and DLBCL in 97% of the cases.
We used RNU6B, a small noncoding RNA, as a normalizer for relative quantification of miRNA expression in our study. Although RNU6B is widely used for normalization of miRNAs, there is no consensus in the literature about its use, and many researchers consider RNU6B not suitable. 39 –41 Many researchers, however, agree on the fact that each experiment needs to define a normalizer that will not vary across all samples and conditions. 25 In our study, RNU6B showed acceptable variation across samples, despite the presence of disease. Moreover, in the absence of information regarding miRNA expression in dogs, the use of RNU6B prevented the possibility of using a miRNA normalizer that could be involved in the pathogenesis of disease.
The current study describes different miRNA-based biomarkers models that differentiate DLBCL and healthy lymphoid FFPE canine tissues. Additional studies are needed to confirm these findings and to determine the use of these models in other subtypes of lymphoma and in other sample types, including fresh tissue, fresh-frozen tissue, plasma/serum, whole blood, and lymph node aspirates to provide a more comprehensive profile of miRNA biomarkers for DLBCL.
Our results represent a step forward on the development of a novel, relatively rapid, sensitive, and accurate diagnostic test for DLBCL in dogs that could potentially use different sample types. Together with the clinical and morphologic alterations, multiple assays with multiple markers may be used to refine the diagnosis of lymphoma, providing the animal the best chance of survival. Future studies are needed to include other types of lymphoma and to evaluate the detection of these biomarkers in the blood to develop a minimally invasive test.
Supplemental Material
Supplemental Material, Combined_supplemental_materials-Elshafie_et_al - MicroRNA Biomarkers in Canine Diffuse Large B-Cell Lymphoma
Supplemental Material, Combined_supplemental_materials-Elshafie_et_al for MicroRNA Biomarkers in Canine Diffuse Large B-Cell Lymphoma by Nelly O. Elshafie, Naila C. do Nascimento, Nathanael I. Lichti, Andrea L. Kasinski, Michael O. Childress and Andrea P. dos Santos in Veterinary Pathology
Footnotes
Acknowledgements
We would like to thank Dr Jose Ramos-Vara for reviewing the histopathology slides. This work was supported by the Purdue Research Foundation (PRF) 2018-2019 of Purdue University.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Purdue Research Foundation (PRF) 2018-2019 of Purdue University.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
