Abstract
Enterospora nucleophila is a microsporidian responsible for an emaciative disease in gilthead sea bream (Sparus aurata). Its intranuclear development and the lack of in vitro and in vivo models hinder its research. This study investigated the associated lesions, its detection by quantitative polymerase chain reaction, and the cellular immune response of naturally infected fish. The intensity of infection in the intestine was correlated with stunted growth and reduced body condition. At the beginning of the outbreaks, infection prevalence was highest in intestine and stomach, and in subsequent months, the prevalence decreased in the intestine and increased in hematopoietic organs and stomach. In heavy infections, the intestine had histologic lesions of enterocyte hypercellularity and proliferation of rodlet cells. Infected enterocytes had E. nucleophila spores in the cytoplasm, and a pyknotic nucleus, karyorhexis or karyolysis. Lymphocytes were present at the base of the mucosa, and eosinophilic granule cells were located between the enterocytes. In intestinal submucosa, macrophage aggregates containing spores were surrounded by lymphocytes and granulocytes, with submucosal infiltration of granulocytes. Macrophage aggregates appeared to develop into granulomata with necrotic areas containing parasite remnants. Immunohistochemistry revealed mast cells as the main type of granulocyte involved. Abundant IgM+ and IgT+ cells were identified by in situ hybridization in the submucosa when intracytoplasmic stages were present. This study describes the lesions of E. nucleophila in gilthead sea bream, an important aquaculture species.
Microsporidia are highly specialized obligate intracellular (intracytoplasmic and/or intranuclear) eukaryotes that are closely related to the fungi. 18 They have a relatively simple development, consisting of 2 general stages: merogony and sporogony. Meronts multiply inside the infected host cell, eventually forming sporonts and then spores, which are ultimately released from the host and transmit the infection. There are almost 200 genera and over 1400 described microsporidian species, though the actual diversity is estimated to be much higher. The host range extends from protists and invertebrates (especially insects and other arthropods) to vertebrates, particularly fish and mammals including humans. 7 Diverse pathological effects of microsporidia have been described in healthy and immunocompromised humans, 9,19 in companion animals, and especially in livestock such as bees, silkworms, crabs, lobsters, and fish. 5,15,17,24,25,36,44 Over half of the known genera infect aquatic organisms and some are considered emerging pathogens in the food chain. 40 About 120 species of microsporidia have been described from fish, and several are involved in diseases of wild, ornamental, experimental models, and farmed stocks. Their impact on fish health is well acknowledged, either producing direct mortalities, diminishing catch value, reducing growth rates, increasing the chances of predation by other animals, or impairing the reproductive or the cardiorespiratory capacities. 21,23,40
Enterospora nucleophila is an emerging pathogen in gilthead sea bream (Sparus aurata) culture in the Mediterranean area. It is an intranuclear microsporidian species belonging to the family Enterocytozoonidae, which includes human and aquatic hosts. Since its first description in Spanish aquaculture facilities, 27 it has also been detected in Italian and Greek gilthead sea bream farms, and in different facilities including on-growing net cages and land-based nurseries. 1,6 Information related to other fish intranuclear microsporidians is scarce. Mortalities have been reported due to Nucleospora salmonis in Chinook and Atlantic salmon, 4,10 to Desmozoon lepeophtherii (syn. Paranucleospora theridion) in Atlantic salmon, 26,46 to Nucleospora cyclopteri in the cleaner fish lumpsucker (Cyclopterus lumpus), 12 and to an unidentified intranuclear microsporidium in grouper (Epinephelus spp.). 47
Studies on E. nucleophila are hindered by its very small size (spores are 1 × 1.5 μm), its intranuclear development, and the lack of in vitro and in vivo propagation models. Limited information about the pathology has been presented elsewhere. 1,27 In this work, we investigated the pathological effects of this infection, its location within the host, and the cellular immune response invoked in naturally infected gilthead sea bream from different disease outbreaks in Mediterranean farms.
Materials and Methods
Fish and Samplings
All experimental protocols involving fish were approved by the Ethics and Animal Welfare Committee of IATS, CSIC, and Generalitat Valenciana (Approval Number 151-2014). They were carried out in a registered installation facility (Code ES120330001055) in accordance with the principles published in the European Animal Directive (2010/63/EU) and Spanish laws (Royal Decree RD53/2013) for the protection of animals used in scientific experiments.
Three lots of gilthead sea bream (n = 300 for lot 1 and n = 200 each for lots 2 and 3) on-growing in commercial sea cages in the Western Mediterranean and displaying emaciation were transported alive to the Institute of Aquaculture Torre de la Sal (IATS-CSIC) facilities. On arrival, 200, 92, and 20 fish (from lots 1, 2, and 3, respectively) were sacrificed by overexposure to the anesthetic (MS-222, 0.1 g/L; Sigma). After biometry and necropsy, intestinal samples were taken for molecular diagnosis by quantitative polymerase chain reaction (qPCR), as well as formalin fixation for histopathology. The remaining 100, 108, and 180 fish of lots 1, 2, and 3, respectively, were kept in the open-flow facilities of IATS-CSIC and mortality was recorded for 152, 79, and 296 days (lots 1, 2, and 3, respectively). Lot 3 was also used to study the progression of the infection in different time points for 296 days post-initial detection (dpid). Fish were sampled at 44 (n = 10), 57 (n = 10), 92 (n = 10), 130 (n = 10), and 296 (n = 21) dpid, and their condition factor (calculated as 100 × body weight)/length3) and specific growth rate (calculated as 100 × ([ln final body weight] − [ln initial body weight])/days) were calculated. From the same fish, intestines were collected for molecular diagnosis by qPCR. At 92, 130, and 296 dpid, additional samples of stomach, gallbladder, head kidney, and spleen were collected for molecular diagnosis. During this time, fish were fed commercial pellets (Biomar) and disease signs and mortalities were recorded.
Simultaneously, another lot of 100 clinically healthy gilthead sea bream (no disease signs of any bacterial or parasitic disease) from the same fish farm, and qPCR-negative for E. nucleophila, were reared under the same conditions as the infected ones.
Diagnosis of Enterospora nucleophila Infection
qPCR
Oligonucleotide probes targeting the SSU rDNA gene of E. nucleophila were designed from an alignment of available Microsporidia SSU rDNA genotypes, including 209 Enterocytozoonidae. 1 Unique E. nucleophila segments, or segments with sufficient variation with respect to the closest taxa, were visually identified in the alignment. Choice of the best probes targeting these segments, according to specificity and thermodynamic properties, was aided by the software package OLIGO 7. 32 Two oligonucleotides were chosen as primers for a SYBR qPCR test: forward ENU-101F23: 5′ TGC GCT AAT TTG CTT TAT ACG AG 3′ and reverse ENU-151R22: 5′ ATT GCA CCT TAT TCT ACA CTC T3′. PCR reactions were assembled in 20 µl volume with 2 µl template DNA and 300 nM each primer in a SYBR commercial master-mix (PyroTaq Evagreen, CMB, Spain). The reactions were programmed in a thermal cycler with a 15-minute Taq activation step (95 °C) followed by 40 cycles of denaturation (95 °C, 15 seconds); annealing (50 °C, 30 seconds), and extension (72 °C, 30 seconds). Standards consisting of 10-fold dilutions with known amounts of a plasmid containing the target gene (covering 7 orders of magnitude) were run with each plate. Only data from reactions with standard curves within an efficiency range (E = 0.95–1.05), and R 2 > 0.99 were accepted. Samples with Ct < 38 were considered positive.
DNA was extracted from clinical samples using a robotic system (EpMotion 5070, Eppendorf, Hamburg, Germany) with dedicated DNA extraction kits (Nucleospin, Macherey-Nagel, Germany). The test samples consisted of small pieces of tissue (50–100 mg) or an aliquot (200 μl) of an intestine homogenate. For the homogenates, the tissue was placed in 5 ml tubes containing ice-cold saline and 3 g of mixed homogenization beads (2 mm, 3.2 mm, and 3.5 mm UFO shape), which were then processed in a laboratory blender (Bullet Blender, Next Advance, USA) with 3× 5-minute cycles at 4 °C. Blood samples were drawn with 0.5 M EDTA-coated syringes. Small aliquots (10 μl) were transferred to microtubes and directly diluted in lysis buffer and processed with a dedicated kit (Nucleospin Blood). Quantification and quality control of extracted DNA samples was monitored with a spectrophotometer (Nanodrop, Thermo Scientific, Spain).
Calcofluor-White Stain
Paraffin sections (4 µm thickness) were cut and mounted onto electrostatically charged glass slides (Superfrost-Plus; Thermo Fisher Scientific, Spain). Air-dried blood smears were also prepared. They were stained with 0.1% calcofluor-white M2R stain in ddH2O (double-distilled water) with a drop of 10% potassium hydroxide, for 1 minute. The slides were rinsed immediately with 30% ethanol. A light counterstaining was achieved with 0.1% Evans blue solution containing a few drops of glacial acetic acid for 1 minute. Slides were dehydrated, mounted in DPX and examined using an Olympus BX51 fluorescence microscope under UV excitation light.
We use the term “clinically diseased” fish for those in the enzootically affected farms that showed the typical disease signs previously described for E. nucleophila infection: anorexia, cachexia, swollen abdominal cavity, and transparent intestinal walls). 27 Those fish were generally positive by qPCR and histology. On the contrary, “non–clinically diseased” fish were those that did not show clinical signs but were positive by qPCR, and in most cases negative by histology.
Histology, Immunohistochemistry, and In Situ Hybridization
Formalin-fixed tissues were dehydrated in a graded ethanol series, embedded in paraffin or methacrylate resin (Technovit 7100, Kultzer, Wehrheim, Germany), sectioned at 1 µm for resin or 4 µm for paraffin) and stained with Giemsa. Immunohistochemistry was used to discriminate between mast cells and acidophilic granulocytes, among the eosinophilic granule cells. For such purpose, paraffin sections (4 µm thick) were mounted on super-frost plus slides and a rabbit-anti-histamine antibody (Sigma-Aldrich) was used to detect mast cells. 11
RNA-in situ hybridization (RNA-ISH) was used to detect transcripts of IgM and IgT. It was performed using the ViewRNA ISH Tissue 2-plex Assay kit (Affymetrix), which uses a proprietary DNA signal amplification technology allowing simultaneous detection of 2 target mRNAs in tissue sections with single copy sensitivity. The probe sets were designed using the sequences of gilthead sea bream IgM (Accession Numbers: JQ811851 and KX599199) and IgT (KX599200 and KX599201) heavy chains, in regions 20-903 and 2-901, respectively. Variable domains and regions not expressed both in the soluble and the membrane-bound forms were excluded, to allow detection of all transcripts regardless of the form. The ISH procedure was performed with 4-μm-thick tissue sections following the manufacturer’s instructions, using a 10-minute heat pretreatment at 90 to 95 °C and a 30-minute protease step at 40 °C. After probe hybridization and development with Fast Blue and Fast Red substrates, the slides were counterstained with Gill’s hematoxylin 25% for 3 minutes, washed, and mounted in aqueous mounting medium.
Statistics
Statistical analyses were performed with GraphPad Prism 8 (GraphPad Software, Inc, San Diego, CA). We searched for covariations between infection status and fish biometrical variables using nonparametric Spearman correlation analyses. Differences between groups were studied with appropriate tests according to the variable studied, as detailed in the Results section. Average Ct values from qPCR reactions are given as the estimated mean ± CI95%.
Results
Disease Signs
Growth Arrest and Emaciation
The specific growth rate of a naturally infected stock (100% prevalence at the arrival to the laboratory) was determined to be 0.38 during 296 days, about 20% of that of a healthy stock of gilthead sea bream of equivalent size held under the same diet and conditions for the same period (ie, 1.9). Using grouped data from all the fish identified as infected in this study, a statistically significant positive correlation between the condition factor and the infection intensity (as determined by the qPCR Ct values from intestinal samples) was found (Fig. 1), indicating an inverse relationship between infection intensity and condition factor. In a subsample of naturally infected fish from lot 3 maintained in the laboratory up to 296 dpid and sampled periodically, a clear decrease in the prevalence and intensity of infection and an improved condition factor with time was observed (Figs. 2, 3).
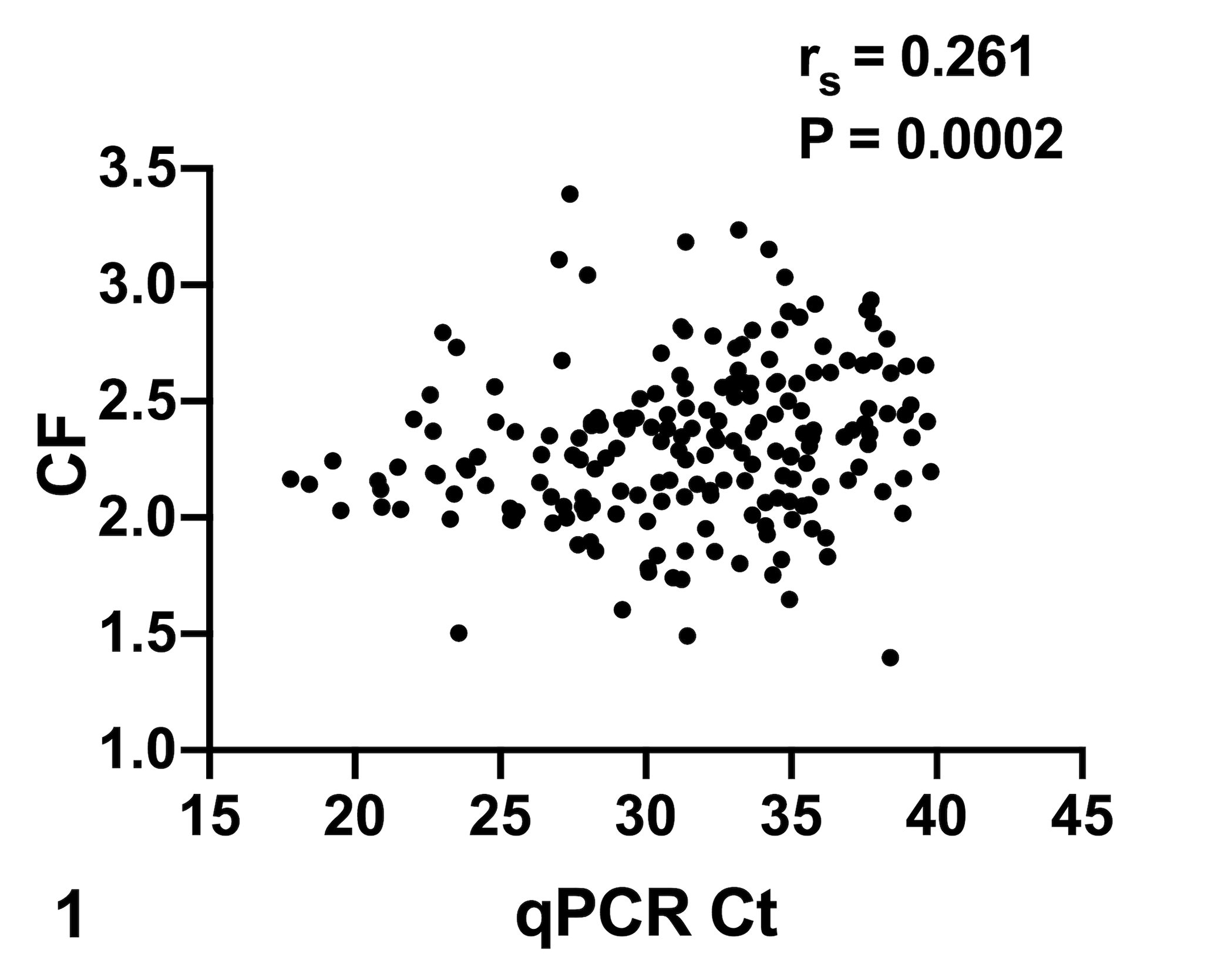
Correlation between intensity of infection by Enterospora nucleophila (qPCR Ct values) and fish body condition factor (CF). Higher intensity of infection (lower Ct) correlates with worse CF in fish obtained from different outbreaks (Lot 1: n = 61, Lot 2: n = 92, and Lot 3: n = 48). Each dot represents the data from a single infected fish. CI95% = [0.1236, 0.3893]. r s = Spearman correlation coefficient, P < .05.
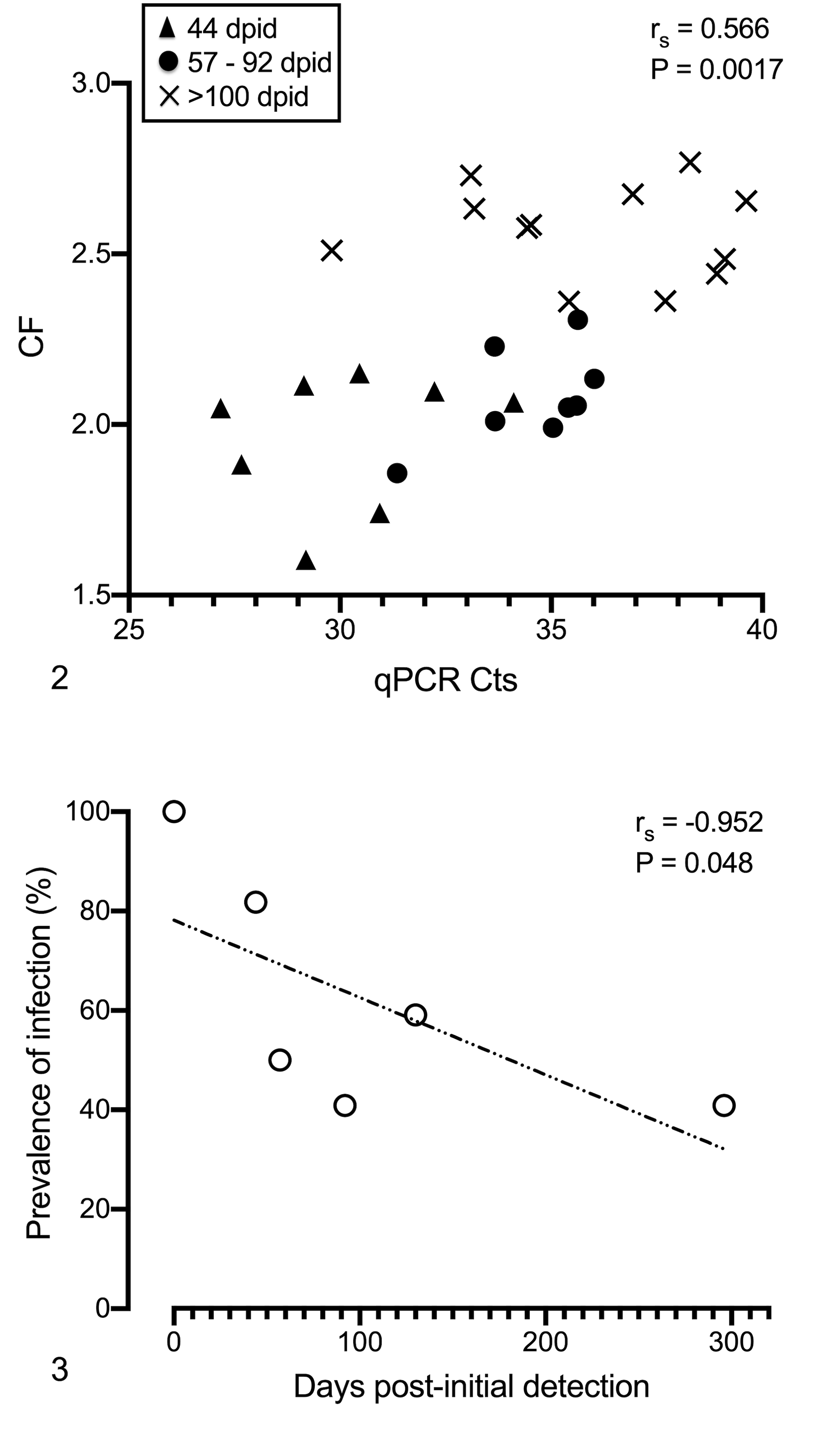
Enterospora nucleophila–infected fish from a stock with 100% prevalence and high intensity (lot 3) were maintained in the laboratory up to 296 days post-initial detection (dpid) and sampled periodically.
Mortality
In the 3 lots, mortality started on arrival of the fish to the IATS facilities and reached 50% or more in a minimum period of 82 days (Table 1).
Epidemiology of Natural Enterospora nucleophila Infection in 3 Lots of Gilthead Sea Bream. Diagnosis was done by qPCR except in *, which was by histology.
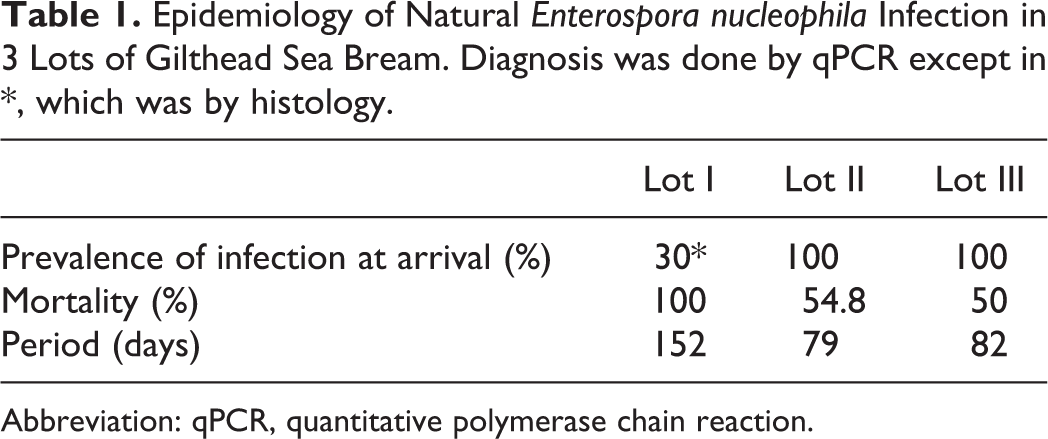

Abbreviation: qPCR, quantitative polymerase chain reaction.
Other Disease Signs
Most fish with high infection levels exhibited cachexia, emaciation, and lethargy. Cases of extreme cachexia were clearly visible from a dorsal view (Fig. 4). At necropsy, internal organs appeared pale, and the intestine was thinned and transparent and frequently contained clear or greenish fluid and white feces in the terminal portion (Fig. 5). Occasionally, ascites was present.
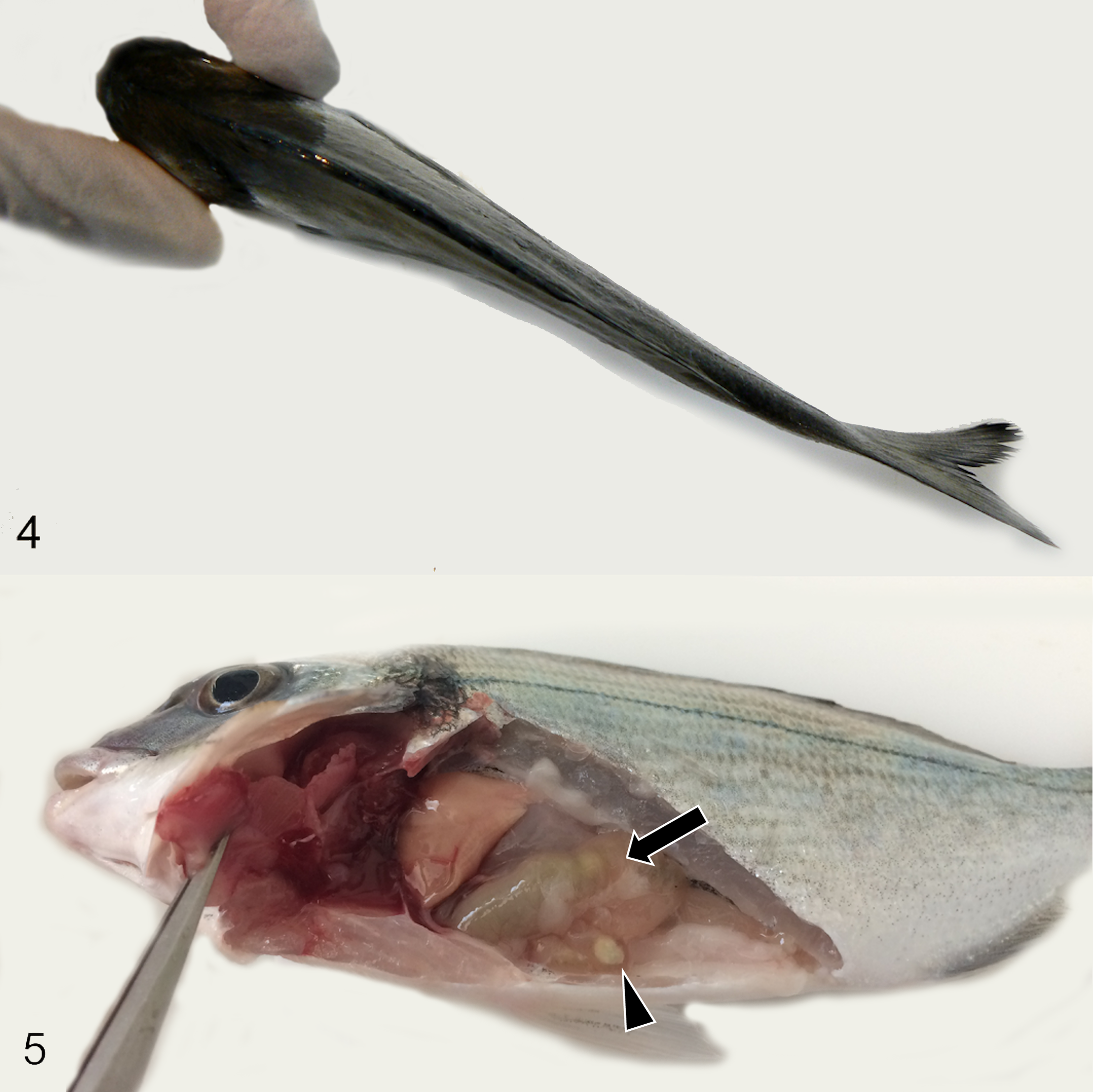

Enterospora nucleophila infection, gilthead sea bream.
Location of Enterospora nucleophila in the Host
Intestinal samples of a subset of fish (n = 20) from clinically diseased gilthead sea bream were selected at an enzootic farm were tested by qPCR for the E. nucleophila SSU rDNA gene. The data showed individual-animal differences in the distribution of E. nucleophila along the intestine. In general, the anterior intestine had the highest prevalence (75%), whereas posterior and middle intestine segments had lower prevalence (70% and 65%, respectively). Ct values obtained for anterior intestine segments were also, on average, lower, indicating higher intensity of infection (average Ct of 28.37, vs 29.82 for middle intestine, and 30.29 for anterior intestine), and this pattern was present in most fish presenting moderate to high infection levels. However, individual patterns varied widely, and only a whole-intestine homogenate could reveal the true prevalence of the stock (100%).
The presence of blood stages was confirmed by qPCR in 44% of a panel of samples from fish with a positive qPCR test for the intestine (n = 63). However, the average Ct values of blood samples were much higher (36.47 ± 0.49, vs 30.16 ± 1.19 at the intestine), and often close to the routine positive cutoff value of 38. Indeed, when taking Ct = 40 as a positive cutoff value, 100% of these blood samples were positive.
Enterospora nucleophila was detected by qPCR in organs other than intestine sampled over a 296 dpid period (n = 41, Lot 3; Fig. 6). At the time of initial diagnosis, this stock had 100% prevalence based on qPCR of the intestine. E. nucleophila infection tended to become more prevalent in the stomach and especially in the hemopoietic tissues (spleen and head kidney) later in the infection. In the last sampling point (296 dpid), 100% of the sampled fish (n = 21) were positive in spleen and head kidney and 95% in the stomach. In general, as observed with blood samples, samples from organs other than the intestine had relatively high Ct values. For gall bladder samples only 2/41 samples were positive Ct = 35.9 and 37.8 during the study period. At the last sampling (296 dpid), the infection intensity of hemopoietic tissues was similar or higher than those of the intestine and stomach (ie, head kidney samples mean Ct = 33.69 ± 0.652; spleen: Ct = 33.80 ± 0.258; stomach: Ct = 34.15 ± 1.03; intestine Ct = 34.38 ± 2.06).
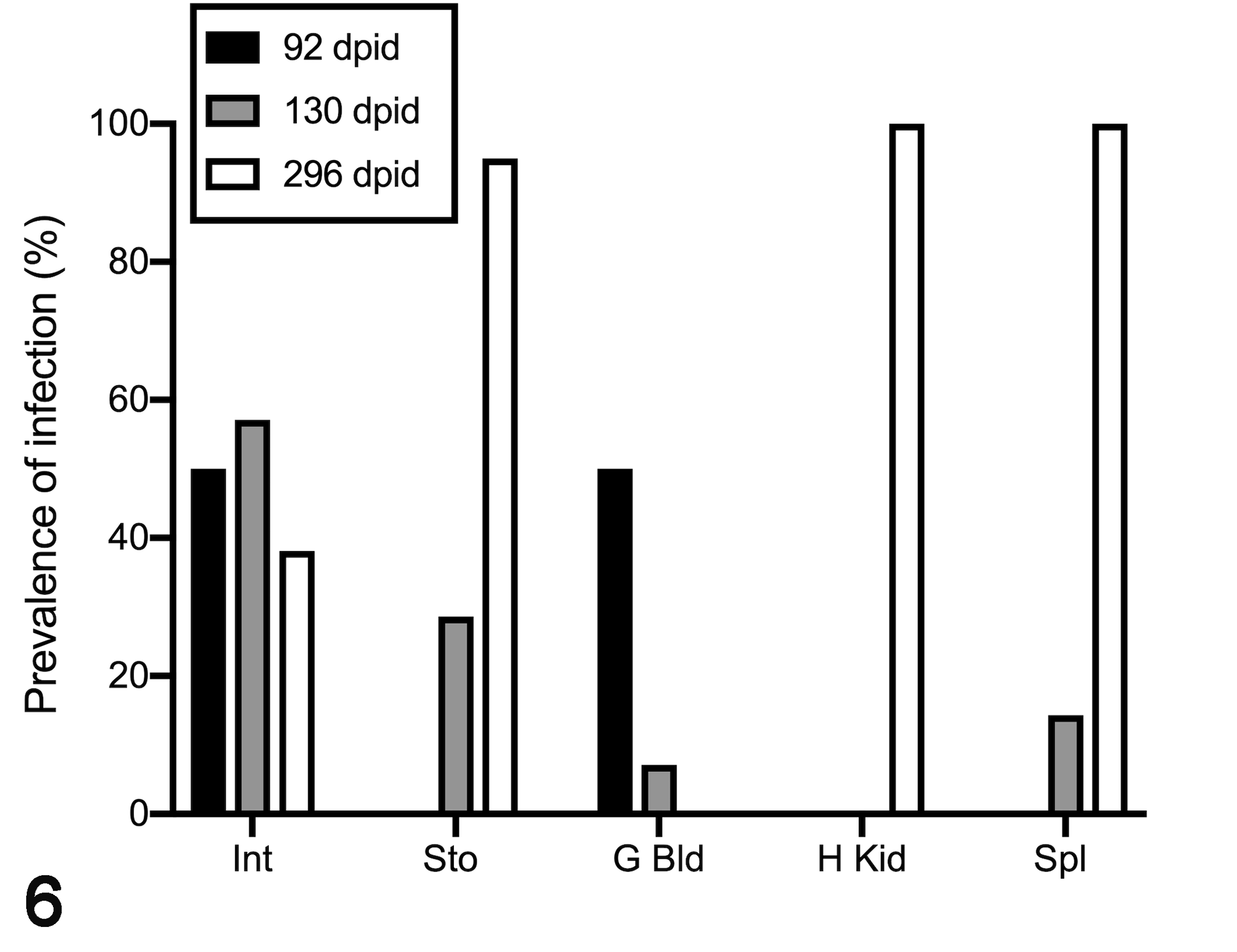
Prevalence of Enterospora nucleophila infection determined by qPCR in different organs from naturally infected fish maintained at the laboratory and sampled periodically up to 296 days post-initial detection (dpid) of infection in the lot of fish. Sto = stomach; Int = intestine; G bld = gall bladder; H Kid = head kidney; Spl = spleen.
In contrast to this increase in the prevalence of infection in stomach and hemopoietic organs, the prevalence in the intestine declined to 50% in the first 92 dpid, and to 38% at the last sampling (Fig. 6). All individuals with detectable infection in nondigestive organs also tested positive somewhere in the digestive tract in the first samplings up to 130 dpid. In contrast, at the last sampling point (296 dpid), detection in nondigestive organs did not always coincide with detection in the digestive tract.
Histopathology
Clinically Diseased Fish (Figs. 7–25)
This category included fish in which E. nucleophila was clearly detectable by histology (Giemsa and/or calcofluor-white-stained specimens), normally in large amounts and mainly in the intestinal mucosa, in addition to being positive by qPCR. During the initial period in which the outbreak was detected, the most obvious histologic lesion in the intestine was an increased number of enterocytes that had a large nucleus, and increase number of rodlet cells occupying the mucosa and disorganizing the normal appearance of the palisade (Figs. 7, 8). At a later stage, E. nucleophila was also present within the cytoplasm of rodlet cells, enterocytes, and macrophages, either in the mucosa or the submucosa (Figs. 9, 10, 12). The nucleus of infected cells was pyknotic, or had karyorrhexis or karyolysis (Figs. 29–31). Moreover, there was lymphocyte infiltration at the base of the mucosa and frequent eosinophilic granule cells between enterocytes. This profile was typical of infected fish sampled in the farms in winter. In these fish, host cells harboring cytoplasmic spores were frequently being shed to the gut lumen, with release of microsporidian spores from them (Fig. 11). Merogonial stages and spores were found in cells within blood vessels (Figs. 13–15).
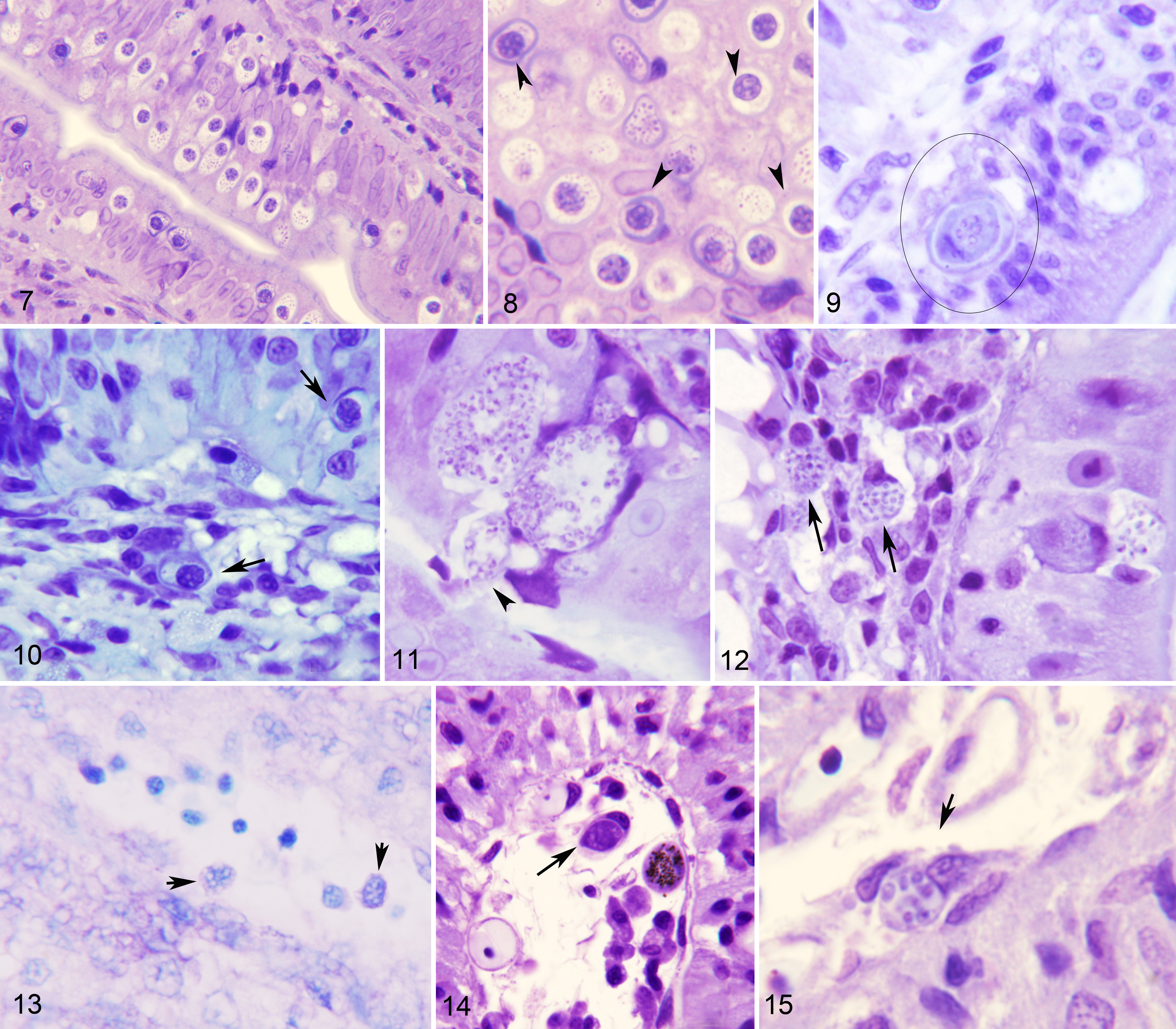
Severe Enterospora nucleophila infection, intestines, gilthead sea bream. Resin sections, Giemsa stain.
In the subsequent stage (several months after the initial outbreak), macrophage aggregates containing spores were seldom found in the mucosa, but were frequent in the submucosa, where they were often surrounded by an inner layer of lymphocytes and an outer layer of granulocytes (Figs. 16, 17), very rarely encapsulated by fibrocytes, and seldom with melanin deposits. At this point, the submucosa displayed inflammatory cellular infiltration with abundant granulocytes, and granulomata were sometimes present in the intestinal serosa. In a later stage of infection (some months post-initial detection), macrophage aggregates became larger, full of foamy macrophages with ingested spores, and sometimes harboring what appeared to be remnants of infected rodlet cells (Fig. 18). These aggregates eventually appeared to develop into granulomata with necrotic areas that were filled with remnants of E. nucleophila (Fig. 24). In parallel, epithelial cells showed cytoplasmic rarefaction (Fig. 19) or necrosis (Figs. 20, 21) and the mucosa contained abundant cellular debris (Fig. 25).
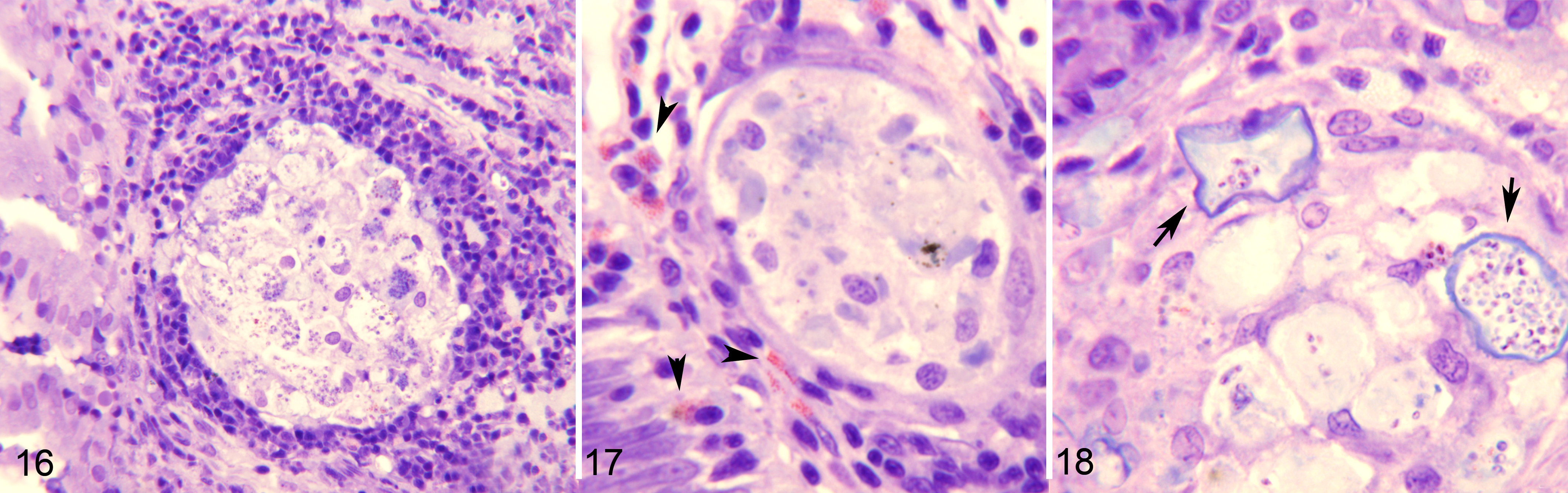
Severe Enterospora nucleophila infection, intestines, gilthead sea bream. Resin sections, Giemsa stain.
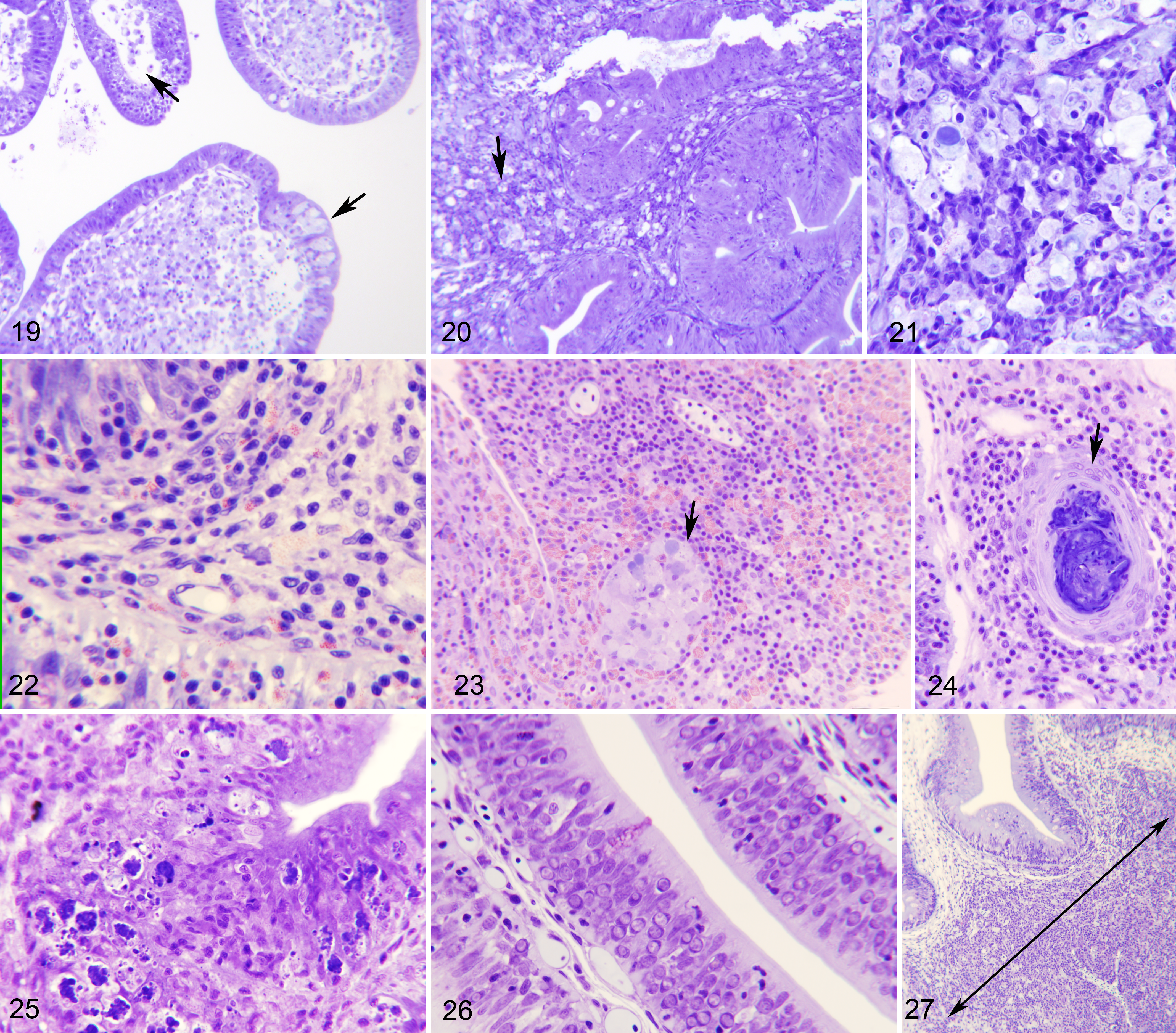
Advanced Enterospora nucleophila infection, intestines, gilthead sea bream. Resin sections, Giemsa stain.
Non–Clinically Diseased Fish (Figs. 26–32)
Fish that were positive by qPCR (albeit often with high Ct values) but with no E. nucleophila stages detected by conventional histology, nonetheless had histologic lesions in the intestine. As above, the lesions were characterized by increased numbers of epithelial cells that had large nuclei and an abnormal chromatin pattern (Fig. 26). Occasionally, merogonial stages were present within these nuclei (Figs. 29–31). Cells frequently appeared vacuolated or contained debris. In some cases, rodlet cells were abundant, especially in the posterior segment of the intestine. As for clinically diseased fish, infiltration of lymphocytes and eosinophilic granule cells and early macrophage aggregates (Figs. 27, 28, 32) were common. These observations corresponded to the late disease course and coincided with fish in which E. nucleophila was detected by qPCR in hematopoietic organs, such as spleen and head kidney, in which macrophages with E. nucleophila remnants accumulated.
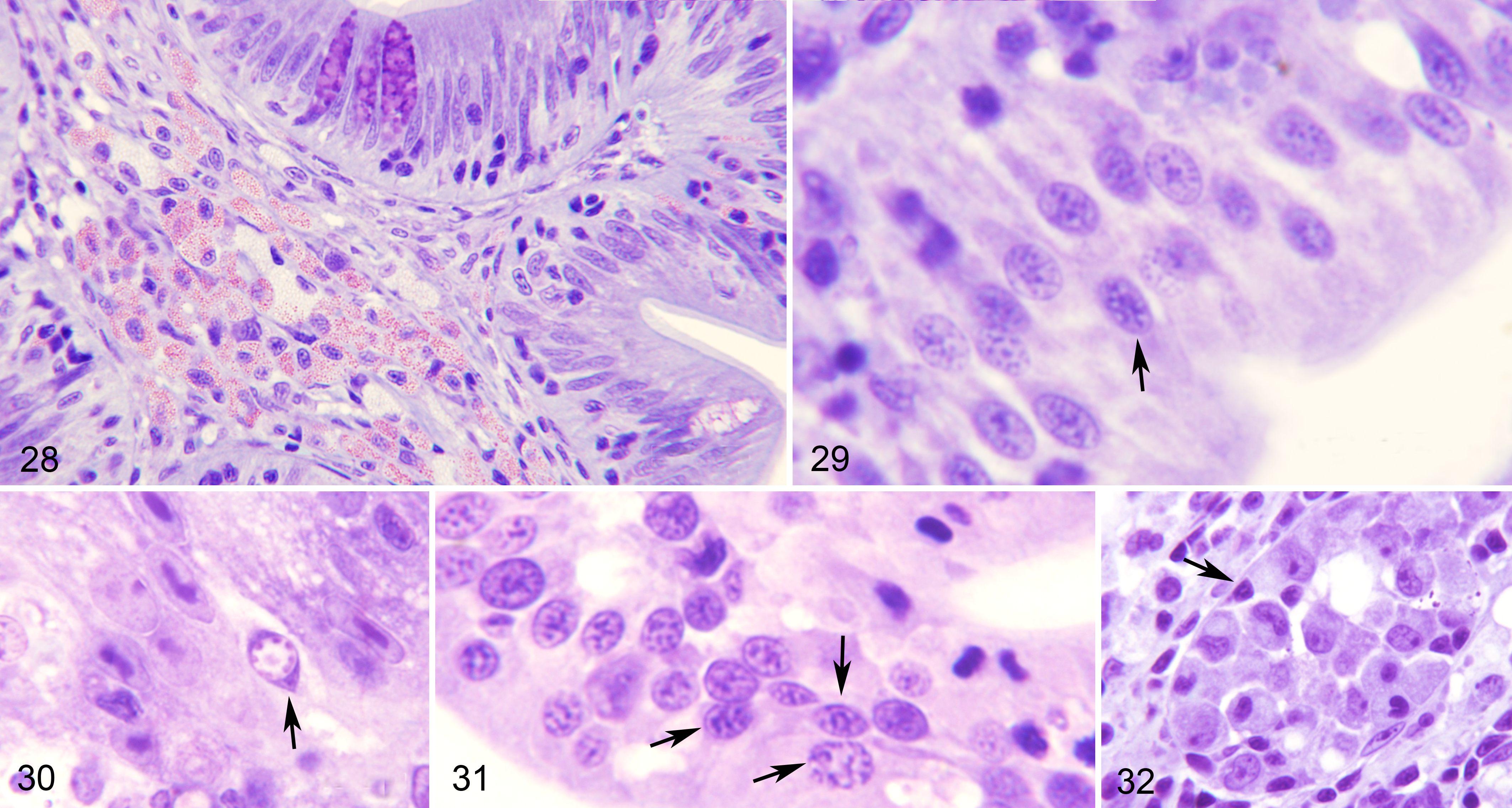
Enterospora nucleophila infection without disease signs, intestine, gilthead sea bream. Resin sections, Giemsa stain.
Cellular Immune Response
Immunohistochemistry showed that mast cells were the main type of granulocyte forming the cellular inflammatory infiltrate in the submucosa and mucosa. Very few acidophilic granulocytes were present, even in severe inflammatory infiltrates (Figs. 33, 34). RNA-ISH labelled IgM and IgT transcripts in cells in the submucosa when intracytoplasmic stages of E. nucleophila were present, and the labelled cells were sometimes in close proximity to the spores. IgT-expressing cells were particularly abundant in the anterior intestine and even infiltrated in the mucosa in contact with infected cells. IgT- and IgM-expressing cells were also abundant around macrophage aggregates in the submucosa (Figs. 35–39).
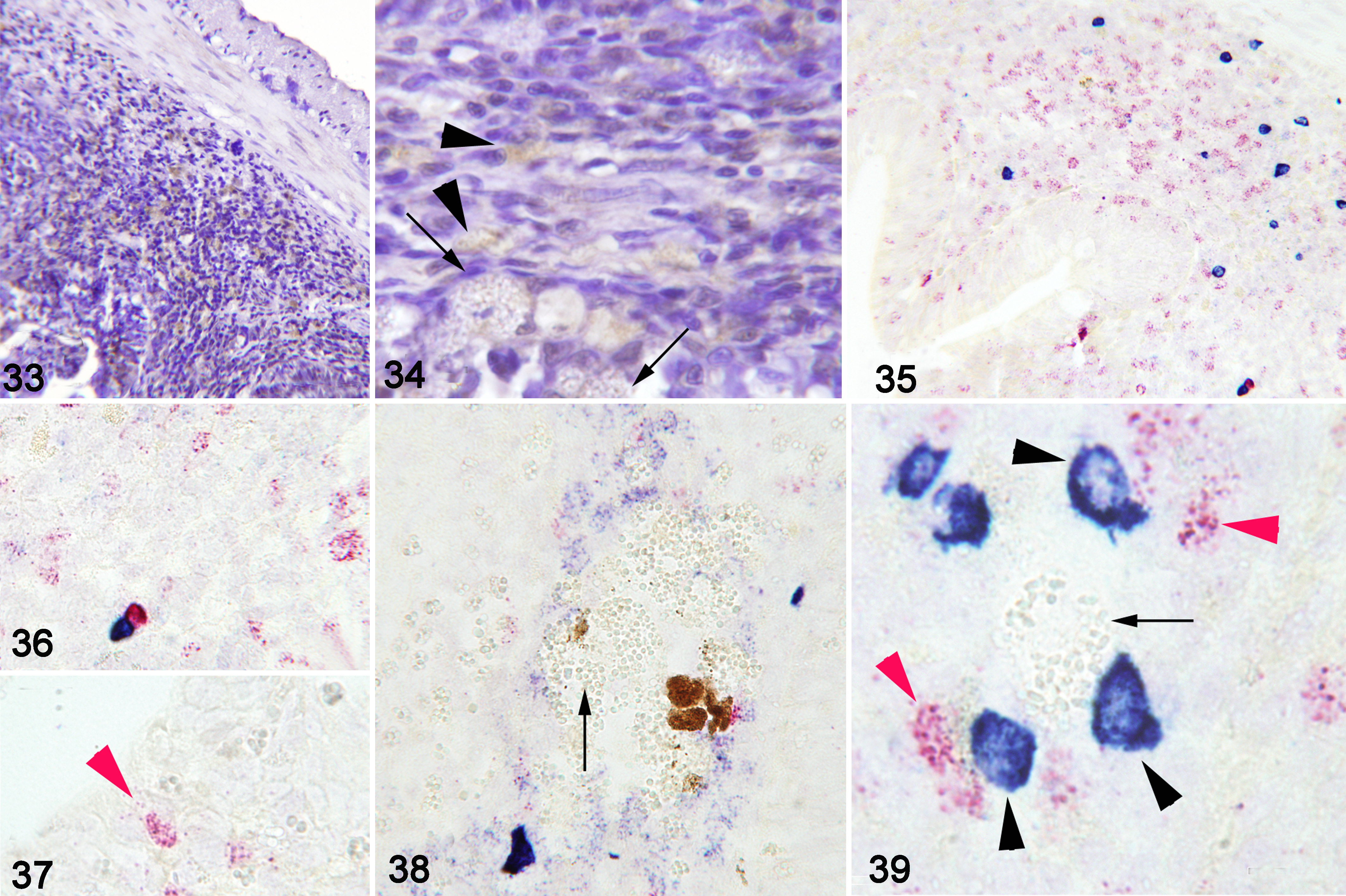
Severe Enterospora nucleophila infection, intestines, gilthead sea bream.
Discussion
Gilthead sea bream with severe E. nucleophila infection had arrested growth, emaciation, and thinned intestinal walls. These findings are similar to those recently described in juvenile groupers infected with a related intranuclear enteric microsporidian, including severe emaciation, anorexia, and intestinal edema with whitish content and a thin transparent wall. 47 D. lepeophtherii infection of Atlantic salmon was previously reported exclusively in the gills and skin, but more recent intestinal infections were associated with ascites, hyperemia, and often peritonitis, among other signs. 46 Whitish feces are also frequently seen in gilthead sea bream affected by the myxozoan Enteromyxum leei or the coccidian Cryptosporidium molnari. 2 Therefore, these signs are not specific for microsporidiosis but common to enteric microparasites affecting feed intake and disrupting intestinal functions such as digestion and absorption of nutrients and minerals and osmotic balance. In the present study, whitish feces from naturally infected gilthead sea bream were qPCR-positive for E. nucleophila (data not shown), suggesting that contamination of the water by feces is possible.
Stunted growth is a characteristic of this disease. 27 In the current study, the body weight of naturally infected fish was approximately half that of the unaffected stock at the farm. Indeed, an abnormally high coefficient of variation in the routine biometrical check-points appears as one of the presumptive indicators of this infection. In addition, a statistically significant relationship between infection intensity and body condition factor was found in the entire dataset of infected fish sampled during 3 different outbreaks. Stunted growth and poor body condition factor are nonspecific signs and they may also be caused by inadequate diets or feeding regimes, or by other diseases such as enteromyxosis. 35,37 Growth arrest is common in microsporidian infections, partly due to the induced anorexia. In this study, the specific growth rate in one lot of infected fish was only 1/6 that of normal fish. By comparison, the average weight of moribund grouper infected by an enteric microsporidian was 20% to 40% of that for uninfected fish, and D. lepeophtherii–infected Atlantic salmon weighed only 20% to 25% of normal growing fish. 46,47 In rainbow trout experimentally infected with Loma salmonae, the onset of significant growth rate suppression was associated with the appearance of gill lesions, 38 with xenoma onset, and accompanied by a reduction in feed intake. 30 Zebrafish (Danio rerio) experimentally infected with Pseudoloma neurophilia also had a reduced size. 31 Therefore, the economic losses due to E. nucleophila are potentially significant not only due to mortality (which was low to moderate in the sea cages), but more importantly to the severely stunted growth.
In vertebrate and invertebrate hosts, cell-mediated immunity confers protection against microsporidians and the key mechanisms are phagocytosis and encapsulation. 16,28,42 Most information on fish immune responses against microsporidia is based on xenoma-forming species (reviewed in Rodriguez-Tovar et al 33 ) and very little is known about intranuclear species. In the current study, the cellular immune response of gilthead sea bream was very mild or absent when E. nucleophila was intranuclear in the intestinal mucosa. In this circumstance, hypercellularity of the mucosa (detected at low magnification) was the main histologic lesion. The cellular immune response apparently started when E. nucleophila was located in the cytoplasm of the host cells and/or when it passed into the submucosa. At that point, macrophages and mast cells predominated, as shown by histology and immunohistochemistry, whereas acidophilic granulocytes (equivalent to mammalian neutrophils) 8 seemed to be depleted, similar to that reported in E. leei infections. 11 This picture is similar to other fish microsporidia in which the inflammatory response is triggered when xenomas rupture, leading to severe, persistent, and extensive cellular proliferation, like in Loma salmonae gill infections in salmonids. 3 Conversely, in other microsporidians, such as Spraguea in the Japanese anglerfish, even heavy infections of the nervous system had no associated lesions. 13 There were high number of IgT-expressing cells in the intestine, particularly the anterior segment, and these cells were distributed across the epithelium and in close proximity to the microsporidia. The abundance of these cells would be in line with the mucosal role of IgT described for teleosts, 48 and could suggest a role of this immunoglobulin in the local response to E. nucleophila.
Foamy macrophages are laden with lipid generated from the massive uptake of modified low-density lipoproteins and the intracytoplasmic accumulation of cholesteryl esters. Foam cells can also be generated as a consequence of infection by persistent pathogens such as Mycobacterium, Chlamydia, and Toxoplasma in humans and other mammals. 43 These foamy macrophages have been described in other microsporidioses, such as in AIDS patients with Encephalitozoon intestinalis infection in the lamina propria. 22 Foamy macrophages containing numerous intracytoplasmic microsporidia were also observed in granulomatous infiltrates of parrots. 14 In fish, macrophages with foamy cytoplasm have been described ingesting spores of a microsporidian parasite causing liquefaction of the skeletal muscle of the Norway pout, Trisopterus esmarkii, 29 and in infiltrations in gut, spleen, kidney, heart, liver, gill and dermis of D. lepeophtherii–infected Atlantic salmon. 46 The foamy macrophages that harbor microsporidial stages seem resistant to degradation and it is thought that when the immune system is unable to eliminate substances perceived as foreign, it produces a granuloma encapsulated by fibroblasts, attempting to wall off the non–self-material. The granuloma is therefore typical of fish in a late stage of this disease. The role of macrophages in resistance and clearing of fish microsporidian spores has been described, but some parasites are capable of blocking the intracellular killing mechanisms of phagocytes, 33 thus avoiding destruction by immune cells. In fact, it has been suggested that P. theridion–infected macrophages could be responsible for propagating the parasite within the host. 41 The current study confirms the dual role of macrophages. On one hand, the observation of different viable parasite stages within macrophages in the intestinal mucosa and submucosa, and in blood vessels, suggests these cells may act as “Trojan horses.” On the other hand, as the observation of parasite debris within macrophage aggregates suggest that they may combat the parasite in the last stages of the infection, perhaps acting in antigen presentation and attracting other leukocytes. This latter role is also in line with the accumulation of macrophages as well as higher detection of the parasite by qPCR in spleen and head kidney during the late stage of infection. The survival of microsporidians within macrophages has already been described for other species, as they have evolved mechanisms that allow them to resist destruction within macrophages. For example, phagolysosome formation is prevented by Glugea hertwigi, 45 and Enterocytozoon cuniculi is capable of surviving despite the host’s active immune response. Latent asymptomatic infections may persist as long as multiplication of the microsporidian and the immune response are in equilibrium. 42 In the subset of fish from a naturally infected stock sampled periodically along 296 days in this study, a progressive partial recovery from the infection was noted. Although the data are likely biased by the mortality observed in the stock, a clear decrease in the prevalence from the 100% initial value was still obvious. In addition, infected fish in this stock presented, on average, a much lower intensity and a recovered body condition factor by the end of the follow-up period. Nonetheless, the presence of fish harboring the parasite at 10 months after the outbreak, even if in small numbers, is remarkable and can have obvious epidemiological implications. It is currently unknown if these patterns would correspond to a full recovery stage or to a relapsing infection. 1
Besides the severe disease signs, E. nucleophila can cause extensive histopathological changes in acute cases, even when present at lower levels (only detectable by qPCR) in fish that do not show disease signs. Similarly, PCR showed the presence of N. cyclopteri in most organs of apparently healthy lumpfish. 12 The existence of nonsymptomatic carriers has relevant consequences for the spread of the disease, and early detection of E. nucleophila in non-diseased fish may be useful to prevent further dissemination of infection within farmed stocks, or to avoid stressful handling of the fish that could trigger clinical disease.
Enterospora nucleophila was detected in blood, but its pathological significance is difficult to ascertain. Blood samples were PCR-positive for E. nucleophila in only 44% of clinically infected fish from the farms. This finding suggests hematogenous dissemination of E. nucleophila within the host. This opens the possibility of cryptic or dormant E. nucleophila life stages in other organs as described for Loma salmonae. 34,39 However, in-depth examination of diseased fish including ISH was not conclusive. 1 Experimental infections and time-lapse sequential samplings may be needed to properly determine the presence of E. nucleophila life stages in blood and extraintestinal organs.
Several members of the Enterocytozonidae family have a remarkable impact on aquatic hosts. Prestocked ponds of cultured prawns in Thailand had a high prevalence of Enterocytozoon hepatopenaei, with the resultant economic losses due to slowed growth and early harvest estimated at $32 000 per hectare in every culture cycle. 20 There are no economic estimates on the impact of E. nucleophila in the culture of gilthead sea bream.
Footnotes
Acknowledgements
The authors acknowledge the collaboration of anonymous fish farming companies allowing access to the animals during the disease outbreaks. We thank J. Monfort and L. Rodríguez for the technical assistance on histological processing.
Declaration of Conflicting Interests
The authors declared no competing interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work has been carried out with financial support from the European Union and the Spanish MINECO under grant projects ParaFishControl (H2020-634429) and AGL2013-R-48560-C2-2-R, respectively. APS was contracted under ParaFishControl project, RDP was contracted under the PTA-Program from the Spanish Ministry of Science, Innovation and Universities (PTA2018-015315-I), and MCP under CSIC PIE Project No. 201740E013. NHA was the recipient of a fellowship from the Egyptian Cultural Office in Spain. This publication reflects only the authors’ view, and the European Commission cannot be held responsible for any use which may be made of the information contained herein.
