Abstract
Schistosomus reflexus syndrome (SR) is a rare and lethal congenital malformation that has been reported in the olive ridley sea turtle (Lepidochelys olivacea) in Mexico. Although the etiology remains unclear, it is presumed to be genetic. Since embryonic development in sea turtles largely depends on environmental conditions, we investigated whether sea turtle total mercury content participates in the etiology of SR. Given that several toxins are known to affect both DNA methylation and/or mitochondrial DNA (mtDNA) copy number, we also probed for associations of these parameters to SR and mercury exposure. We measured the levels of each variable in malformed olive ridley sea turtle embryos (either with SR or other non-SR malformations) and embryos without malformations. Malformed embryos (with or without SR) showed higher mercury concentrations compared to normal embryos, while only embryos with SR showed higher levels of methylation compared to embryos without malformations and those with other malformations. Furthermore, we uncovered a positive correlation between mercury concentrations and DNA methylation in SR embryos. With respect to mtDNA copy number, no differences were detected across experimental groups. Because of sample size limitations, this study is an initial attempt to understand the association of environmental toxins (such as mercury) and epigenetic alterations (DNA methylation) in the etiology of SR in sea turtles.
Keywords
Schistosomus reflexus syndrome (SR) is a rare and lethal congenital malformation, characterized by exposure of the abdominal and thoracic viscera (schistosomus) due to a fissure of the ventral abdominal wall and spinal inversion (reflexus), with different levels of severity. 17,42 Other minor features include ankylosis of the extremities, in which extremities are positioned adjacent to the skull, and hypoplasia of the lung and diaphragm. 45
SR was thought to be exclusive to ruminants, occurring mainly in cattle 42,67 but occasionally in sheep and goats; 17,27,79 sporadic cases have been reported in other domestic and nondomestic mammals. 25,44,47,52,61 Recently, we reported SR in olive ridley (Lepidochelys olivacea) sea turtle embryos in a representative nesting beach from northwestern Mexico. 3 This species is considered vulnerable according to the Red List of Threatened Species of the IUCN. 1 Notably, of 20 257 embryos examined, 124 had congenital malformations, and 124 of these had SR (ie, 31% of malformations and 0.6% of all embryos examined). Those with SR had different degrees of severity, mostly short neck, anury, and different types of dysmelia. 3 Other reports of SR in reptiles include a case report in the bearded dragon (Pogona vitticeps) 34 and 2 studies in vipers that mention the presence of schistosoma 54,77 (although none of them were reported as SR).
It is suspected that SR originates before or during the differentiation of the primitive intraembryonic mesoderm. 10 Although its cause is not clear, both genetics 27,36,45 and consanguinity are known predisposing factors in ruminants. 12,27 In cats, SR has been associated with chromosomal aberrations (such as chromosomal breaks, nonhomologous pairing of chromosomes, and dicentric fragments), 62 as well as superfetation. 39 Although no specific genes have been directly associated with SR, mutant mice with knockout of the transforming growth factor β gene (Tgfb) or the AP-2 family of transcription factors show defects in closure of the ventral body wall, spinal malformation, and other anomalies. 58,76 It has also been proposed that anomalies similar to SR could involve exposure to environmental factors, suggesting a multifactorial etiology. 52 In fact, genetic and/or environmental causes can produce relatively similar malformation patterns in mammals. 41,47
A relevant environmental contaminant is mercury, a widely distributed chemical element of natural and anthropogenic origin. Mercury is found in organic and inorganic forms; the latter can be biotransformed by bacteria to the organic form, methylmercury, which is the fraction of total mercury transferred in the food chain. 8 Although mercury is normally present at low concentrations in aquatic systems, bioaccumulation leads to increased concentrations in the food chain. 80 Mercury concentrations in different sea turtle species depend on the feeding habits and feeding areas: the green sea turtle is predominantly herbivorous, the loggerhead sea turtle is omnivorous, and the olive ridley is omnivorous and opportunistic. 46,53 In the latter, maternal transfer of mercury has been reported in eggs, with higher concentrations in the yolk where lipophilic contaminants tend to concentrate. 63
In amphibians, maternal mercury transfer causes reduced reproductive success, as well as congenital malformations, including skeletal malformations. 6,81 In green sea turtles, mercury concentrations in the liver have been associated with different health problems, although an association with congenital malformations has not been reported. 83 In green and loggerhead sea turtles, high mercury concentrations have also been reported in the yolk, as well as lower concentrations in egg albumin and shell, with no apparent effects on reproductive success. 43,74
Thus, exposure to environmental toxicants during early development may cause congenital malformations. 56,85 Exposure to methylmercury has been associated with changes in epigenetic marks (DNA methylation and histone modifications) in mice and rats 9,60 that play important roles in transcriptional regulation, 7,16,33,48,51,86 genomic stability, 14 X-chromosome inactivation, 73 and genomic imprinting. 15,49,68 Furthermore, distinct developmental anomalies have been associated with alterations in DNA methylation. 92 For instance, mutations in the de novo methyltransferase DNMT3B are often found in the ICF syndrome (immunodeficiency, centromeric region instability, facial anomalies) that is characterized by hypomethylation of pericentromeric satellite repeats. 29,91 Likewise, mutant Dnmt3b–/– mouse embryos show multiple developmental defects, including growth impairment, rostral neural tube defects, and hypomethylation of minor satellite repeats. 59
Mitochondrial damage could also be related to mercury exposure, since mitochondrial DNA (mtDNA) is particularly susceptible to oxidative stress. 5,78 For example, genotoxic assays in Caenorhabditis elegans have shown that methylmercury causes a decrease in mtDNA copy number, disrupting mitochondrial end points in a compound-dependent fashion (organic or inorganic forms of mercury). 90
SR is a complex congenital syndrome with a high incidence (31%) 3 among malformed olive ridley sea turtle embryos in northwestern Mexico. Given the described effects of mercury exposure on congenital malformations, DNA methylation, and mtDNA copy number, we investigated whether these parameters differed in embryos with SR, embryos with other congenital malformations (non-SR), and embryos without congenital malformations.
Materials and Methods
Sample Collection
In a previous 7-month study conducted at El Verde Camacho nesting beach in Mexico (23° 44′ 22′′ N, 106° 58′ 27′′ W), we reported a 2% incidence of congenital malformations in olive ridley sea turtle embryos, including SR syndrome in 0.6% of the embryos, comprising 31% cases of congenital malformations. 2,3 From these embryos (which were in late developmental stages 29–31), we randomly selected 12 specimens with congenital malformations (5 of them had SR) and 12 with no externally apparent congenital malformations (normal embryos that died of unknown causes; Table 1). For DNA methylation and mtDNA copy number analyses, tissue samples (skin and underlying muscle) were taken from the back of the neck of each embryo and preserved in individual microcentrifuge tubes containing absolute ethanol at –20°C until use. To measure total mercury concentrations, the rest of each embryo was preserved individually in sealed plastic bags at –20°C until use. Since most embryos were in late developmental stages, the yolk sac was mostly reabsorbed, and therefore it was excluded from the analysis. All analyses were performed in individual embryos.
Levels of Total Mercury Concentrations, Percentage of 5-Methylcytosine (5-mC), and Mitochondrial DNA (mtDNA) Copy Number in Olive Ridley Sea Turtle Embryos.a

Abbreviations: FU, fluorescence units; Hg, mercury; NA, quantity of DNA for analysis was insufficient (not analyzed); SR, embryos with schistosomus reflexus (other malformations observed in these embryos are also indicated).
Total Mercury Concentrations
Homogenized tissue (1 g, wet mass) was digested with 5 ml of 50% HNO3 and later with 3 ml of H2O2 in a microwave digestion system (CEM MARSX CEM Corporation, USA) according to EPA method 3052. 19 Once the samples were digested, they were analyzed by atomic absorption spectrophotometry (Perkin Elmer 1100-B Perkin Elmer, Mexico). Mercury quantification was carried out using the hydride generator accessory (MHS-20 Perkin Elmer, Mexico); the limit of detection was 0.0002 μg/g. Quality control/quality assurance consisted of blanks, duplicates, and a certified reference material (DOLT-48 National Research Council, Canada). 55 Mean percent recovery was 91.3 (n = 5). Results are presented in μg/g wet mass.
Global DNA Methylation Analysis
Genomic DNA was extracted by the salting-out method 50 and quantified using SYBR Green I (DNA Genotek DNA Genotek Inc, Canada) and a spectrofluorometer (The Infinite M1000-Tecan Tecan Group Ltd, Switzerland). Duplicate measurements of global DNA methylation were performed with the MethylFlash Methylated DNA Quantification Kit (Epigentek Epigentek Group Inc, USA), using 160 ng of DNA following the instructions of the manufacturer.
Mitochondrial DNA (mtDNA) Copy Number
Analysis of mtDNA copy number was performed from same samples used for the methylation analysis using a quantitative (non-real-time) polymerase chain reaction (PCR) protocol. 70 Briefly, we amplified the coding region of the mitochondrial gene NADH-ubiquinone oxidoreductase chain 6 (ND6) that codes for a protein component in the electron transport chain. 84 Primers were designed using the publicly available genome of the green sea turtle Chelonia mydas (NC_000886.1; Table 2). To ensure that the PCR conditions were within the exponential phase, it was verified that a sample diluted at 1:2 resulted in a 50% reduction of the PCR product relative to the undiluted sample. PCR amplification profile was as follows: 94°C for 2 minutes; 24 cycles of 94°C for 30 seconds, 62°C for 30 seconds, and 72°C for 1 minute; and a final extension performed at 72°C for 5 minutes. The resulting PCR products were quantified in triplicate by spectrofluorometry (The Infinite M1000-Tecan; excitation wavelength: 485 nm, emission wavelength: 535 nm). Fluorescence values were used directly to compare relative mtDNA content between samples. 70
PCR Primers Used to Amplify the ND6 Coding Region of Mitochondrial DNA.
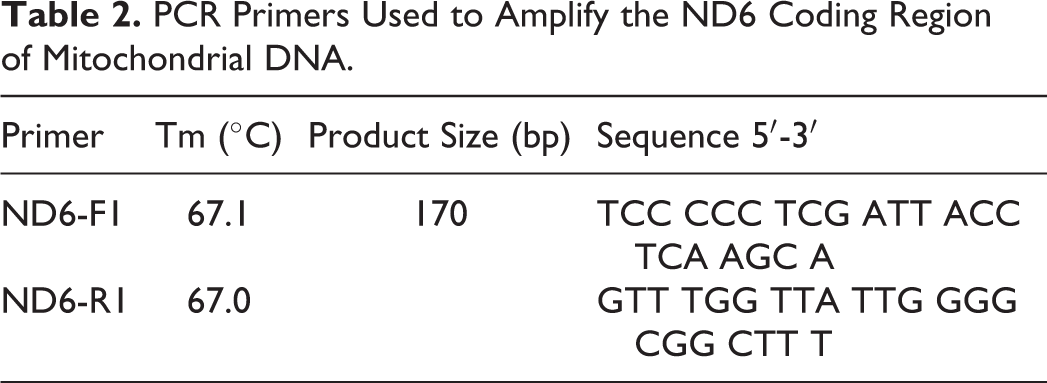

Statistical Analysis
Mercury concentrations, percentage of 5-methylcytosine (5-mC), and fluorescence of mtDNA between normal and malformed embryos, as well as between embryos with SR and non-SR malformations, were compared using the Student t test when the criteria of normality and homoscedasticity were met; otherwise, a Mann-Whitney U 1-way analysis of variance (ANOVA) was performed. In addition, a Spearman correlation analysis was used to assess correlations between mercury concentrations and DNA methylation. All statistical analyses were performed with the SigmaPlot v11 software (Systat, San Jose, CA).
Results
Total Mercury Concentrations
Concentrations of mercury in olive ridley sea turtle embryos are shown in Table 1 and Supplemental File S1. We observed that average mercury levels were about 2-fold higher in malformed relative to nonmalformed embryos (0.0289 and 0.0156 μg/g of mercury, respectively, P = .026; Fig. 1). However, mercury concentrations in embryos with SR did not differ significantly from those with other or no congenital malformations (P = .803 and P = .225, respectively; Figs. 2 and 3, respectively).
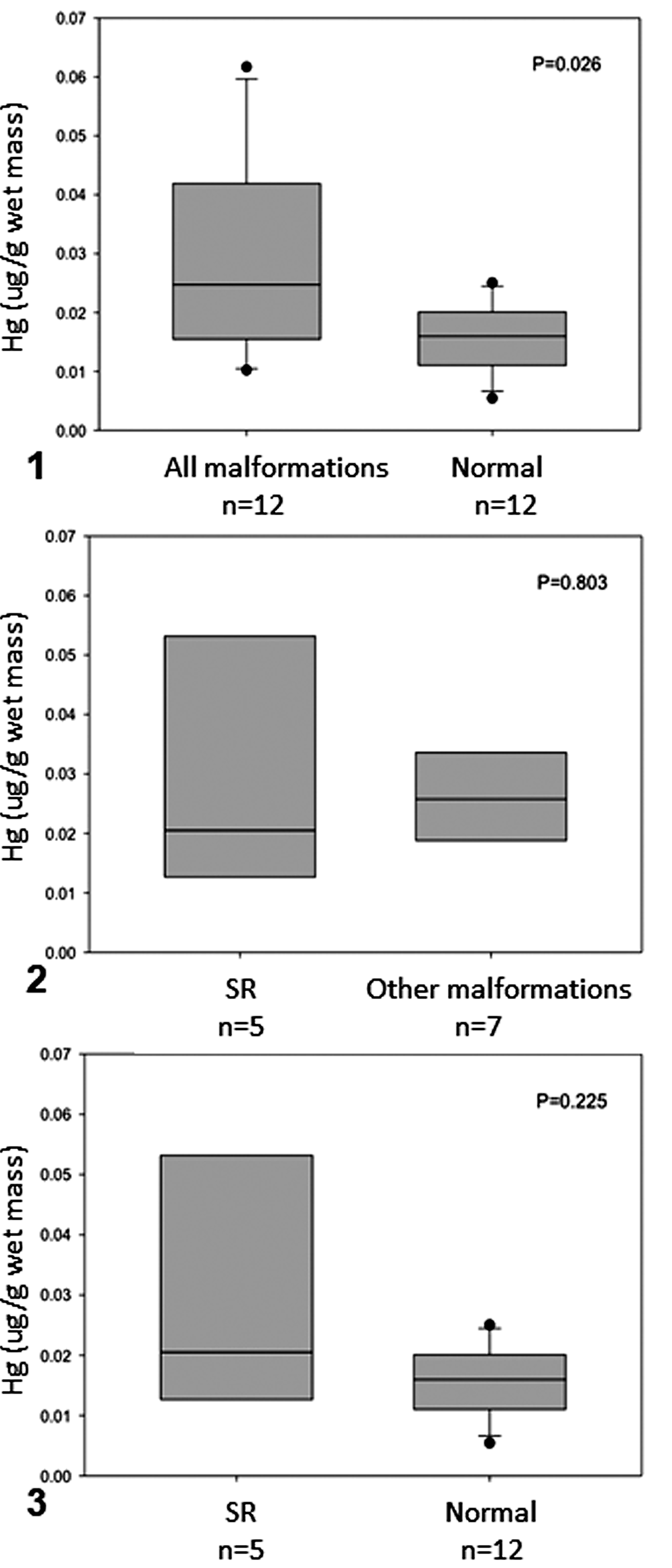
Mercury concentrations in olive ridley sea turtle embryos that had schistosomus reflexus (SR) malformation, had other malformations, or were normal. The boxes indicate the median and the 25th and 75th percentiles; error bars indicate 10th and 90th percentiles; data points indicate outliers. P < .05; n = sample size.
Genome-Wide DNA Methylation
The percentage of global DNA methylation (5-mC) in olive ridley sea turtle embryos is shown in Table 1 and Supplemental File S1. No significant differences were found in 5-mC between normal (0.106% ± 0.0606%) and malformed (0.111% ± 0.0891%) embryos (Fig. 4). However, 5-mC content was increased about 2-fold in SR embryos (0.187% ± 0.0859%) relative to those with other or no congenital malformations (P = .005 and P = .042, respectively; Figs. 5 and 6).
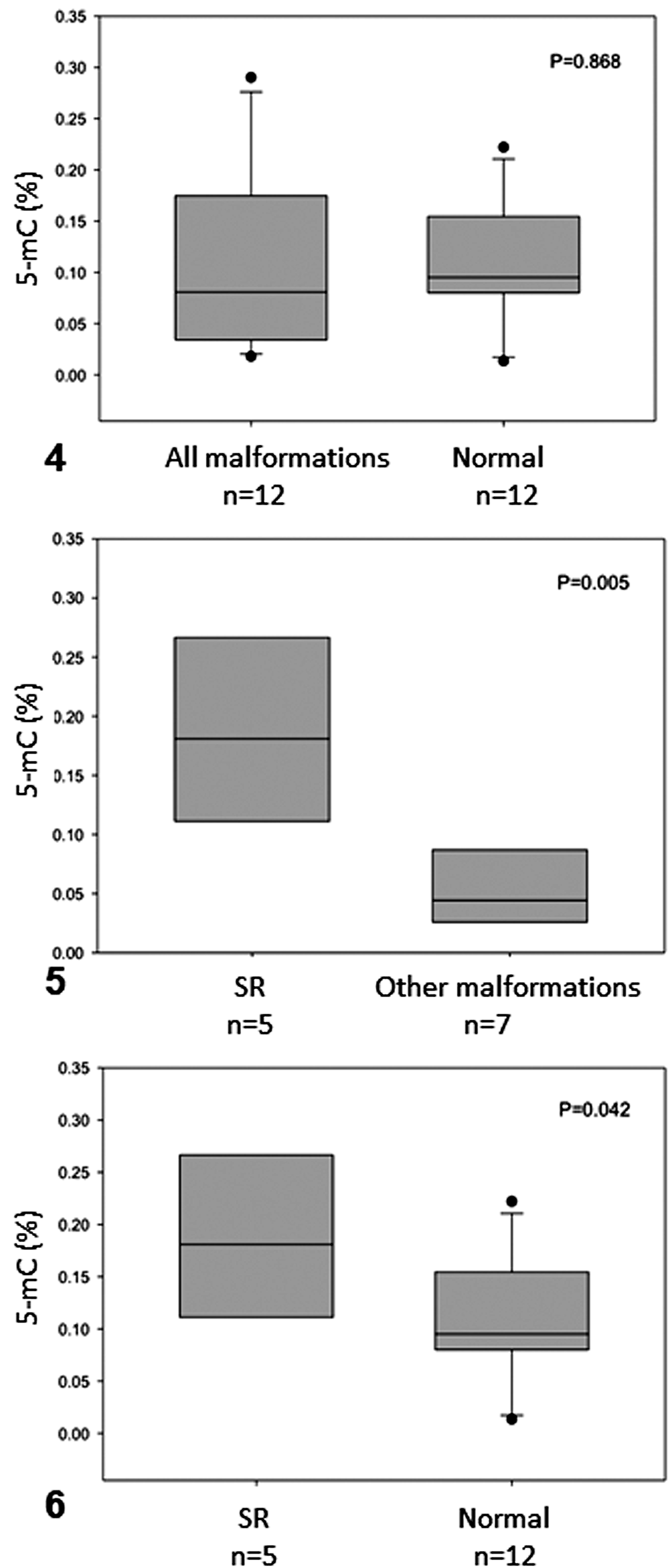
Percentage of 5-methylcytosine (5-mC) in olive ridley sea turtle embryos that had schistosomus reflexus (SR) malformation, had other malformations, or were normal. The boxes indicate the median and the 25th and 75th percentiles; error bars indicate 10th and 90th percentiles; data points indicate outliers. P < .05; n = sample size.
Correlations between mercury and DNA methylation content across experimental groups were tested (Figs. 7–11). Only SR embryos showed a positive and significant correlation between mercury concentrations and DNA methylation (rs = 1, P = .0167; Fig. 10).
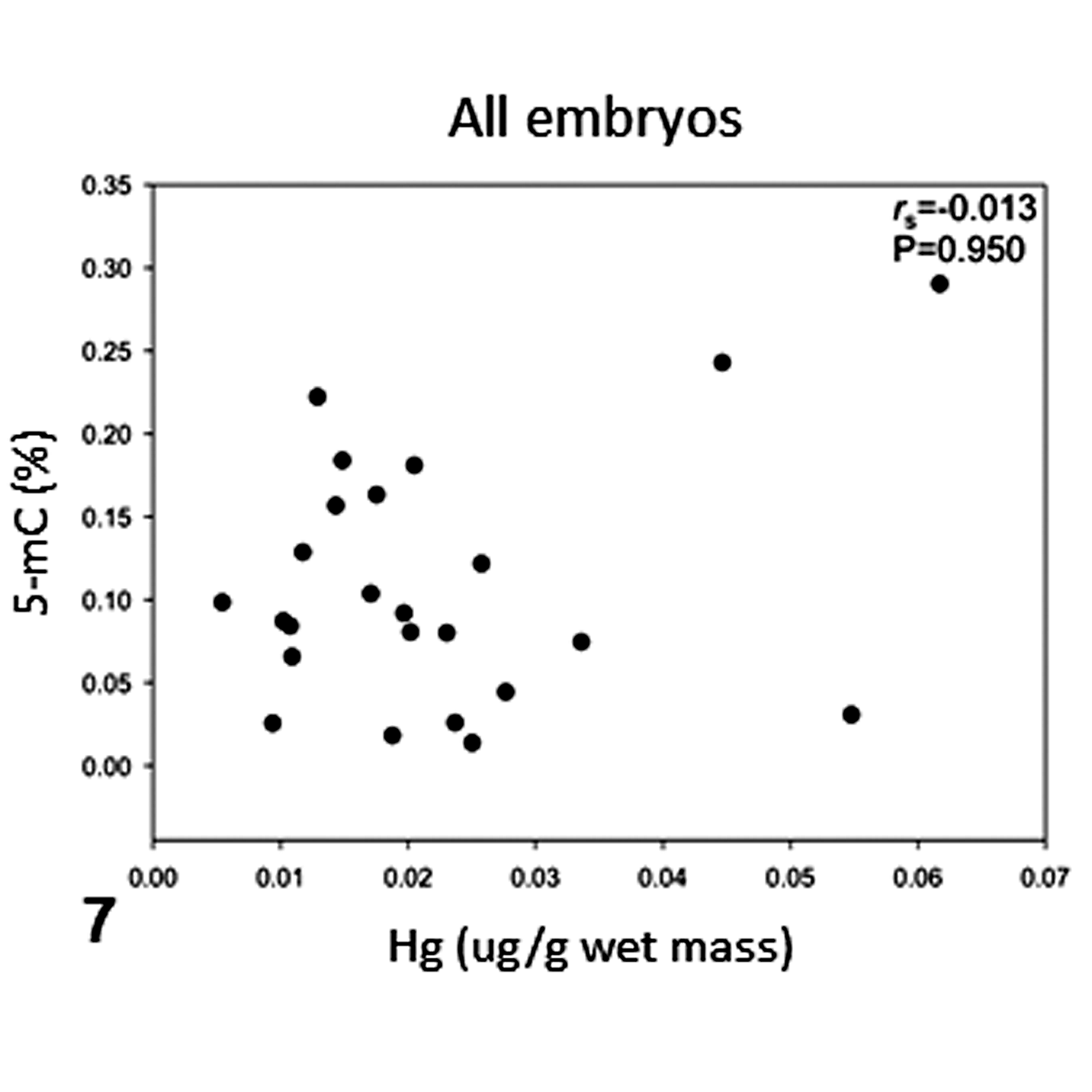
Correlation of mercury concentrations and percentage of 5-methylcytosine (5-mC) in all olive ridley sea turtle embryos analyzed. rs = Spearman correlation coefficient, P < .05.
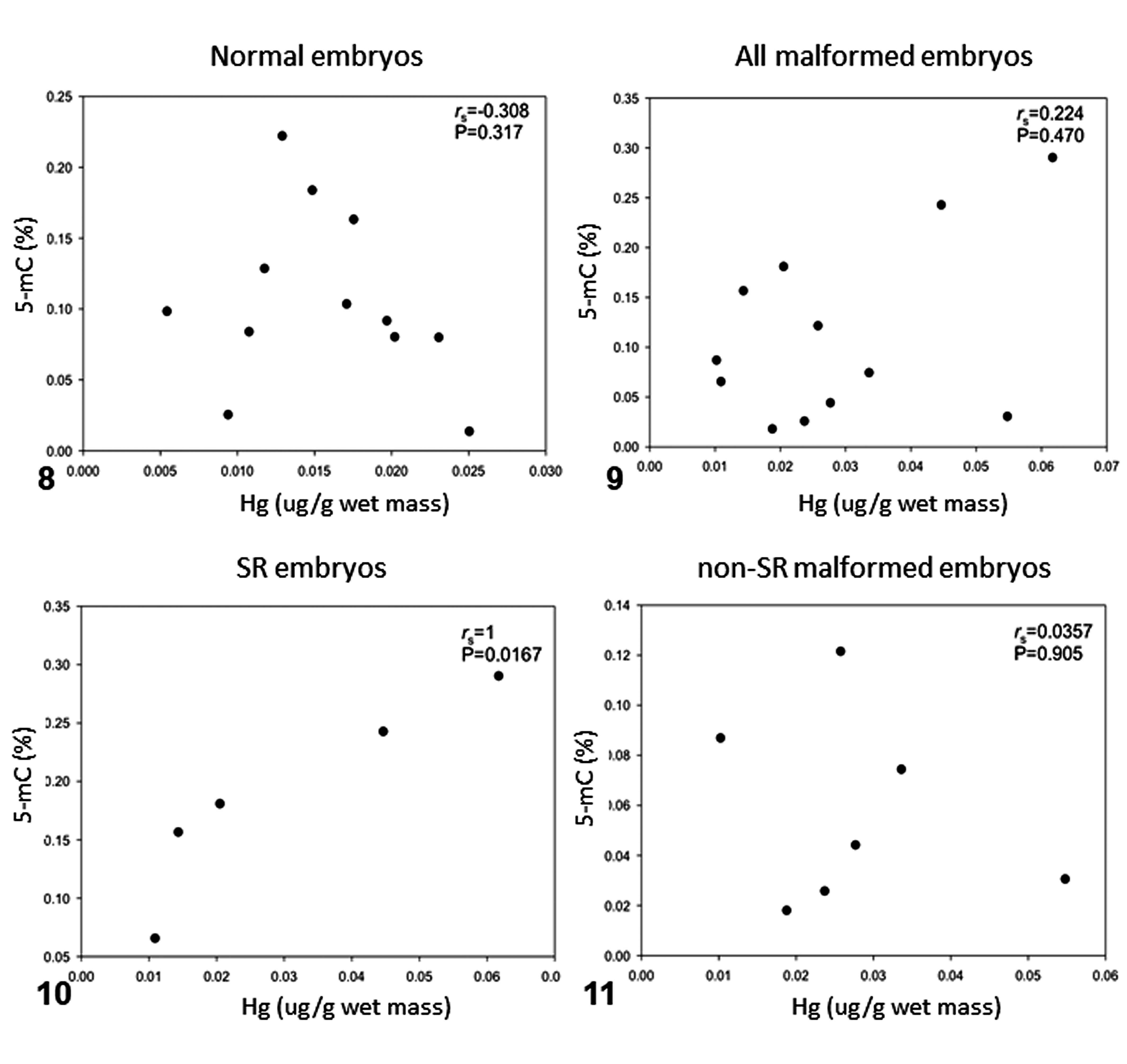
Correlation of mercury concentrations and percentage of 5-methylcytosine (5-mC) in olive ridley sea turtle embryos that had schistosomus reflexus (SR) malformation, had other malformations, or were normal. rs = Spearman correlation coefficient, P < .05.
Mitochondrial DNA (mtDNA) Copy Number
Mitochondrial DNA copy number in olive ridley sea turtle embryos is shown in Table 1 and Supplemental File S1. There were no significant differences in fluorescence levels, either between malformed (734.682 ± 471.347) and normal (739.250 ± 220.185) embryos or between SR (811.300 ± 481.827) and non-SR malformed embryos (670.833 ± 497.843) or normal embryos (Figs. 12, 13, and 14, respectively). However, malformed embryos presented higher mtDNA fluorescence variability—lower values in quartile 1 (Q1) and higher values in quartile 3 (Q3), very close to the maximum value compared to normal embryos (Fig. 12)—but these were not significantly different from the other groups.
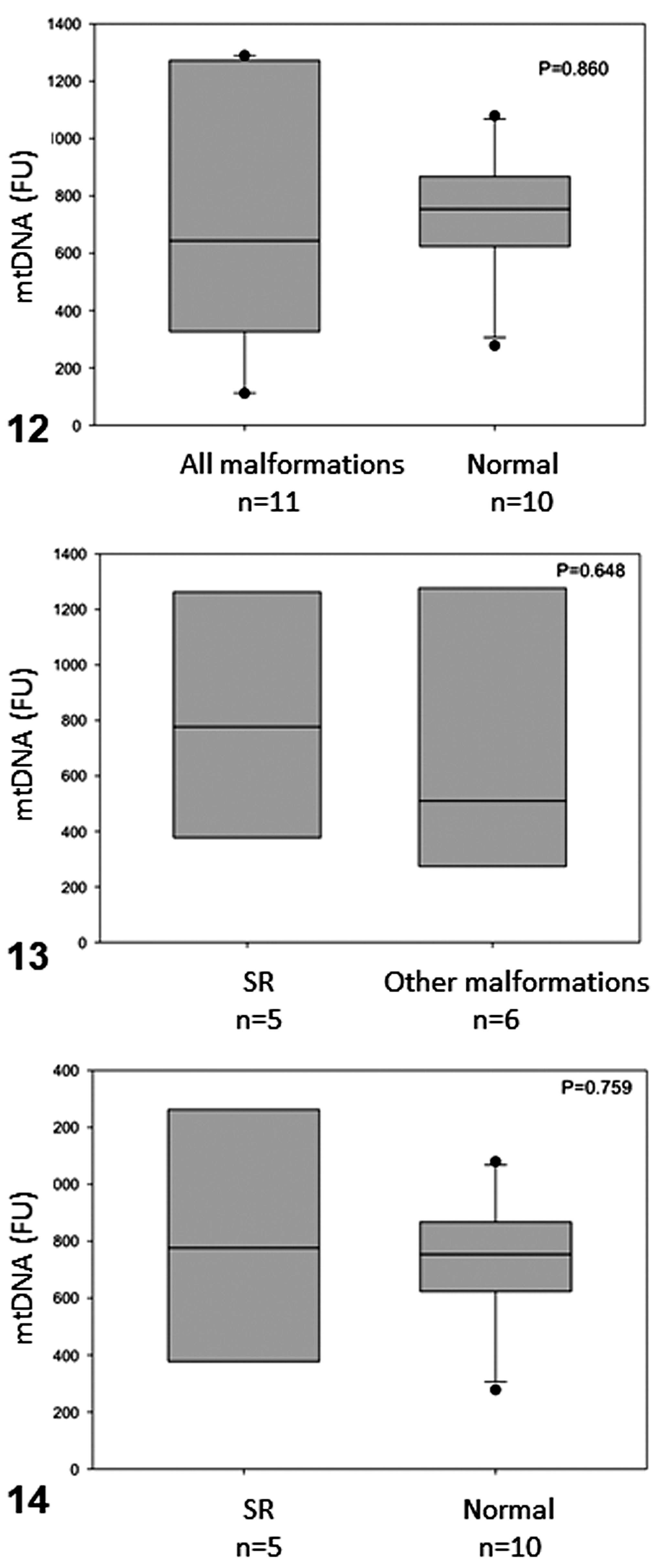
Box plots showing mtDNA fluorescence in olive ridley sea turtle embryos that had SR malformation, had other malformations, or were normal. The boxes indicate the median and the 25th and 75th percentiles; error bars indicate 10th and 90th percentiles; data points indicate outliers. P < .05; n = sample size.
Discussion
The results obtained in this study show that mercury concentrations were higher in malformed compared to normal embryos. Among malformed embryos, the highest mercury concentrations were detected in those with SR. Teratogenic effects of mercury have been reported in humans, as in Minamata disease caused by ingestion of marine food contaminated with methylmercury. This disease is characterized by neurological damage, including sensory alterations, ataxia, dysarthria, constriction of the visual field, and auditory alterations, 20,30 in addition to congenital malformations such as polydactyly, high palate, defective external acoustic meatus, microcornea, and protrusion of the coccyx. Likewise, exposure to methylmercury in fish, amphibians, birds, and other mammals causes skeletal malformations, gastroschisis, and optical, cardiovascular, limb, and skull malformations. 6,13,31,32,41,75,81,87 In sea turtles, the teratogenic effects of mercury exposure have not been documented, but mercury may contribute to health problems, 35 and maternal transfer has been reported. 26,63 Although feeding areas for olive ridley females seem to be more dispersed than for other sea turtle species, some reports point to the Gulf of California and the Gulf of Tehuantepec (in the Mexican Pacific) as potential feeding sites. 46 Mercury has been found in mollusks, crustaceans, fish, and birds from the Gulf of California 64,71,72 and in fish from the Gulf of Tehuantepec. 18 Thus, it is possible that olive ridley embryos were exposed to mercury by maternal transfer.
SR embryos showed the highest concentrations of mercury; however, when SR and non-SR malformed embryos were compared, no significant differences were detected in mercury concentrations. These results should be taken with caution due to the small sample size and the high variability in mercury content. First, the embryos used in this study were sampled from different nests, and nest-to-nest variations may contribute to the observed variability. 40 Second, mercury is not the only teratogenic compound present in the environment, and therefore the observed effect could result from synergy with other teratogenic compounds. 21 Nevertheless, the observed variability could be relevant in the biological context, since it has been reported that increased variability upon exposure could indicate environmental stress. 23
We observed higher levels of global DNA methylation in olive ridley sea turtle embryos with SR, compared to those with normal development or other congenital malformations. Therefore, it is plausible that global hypermethylation of DNA could be either a cause or a consequence of SR. In Drosophila melanogaster, DNA hypermethylation induced by overexpression of the de novo methyltransferase Dnmt3a is indeed associated with developmental defects due to reduced rates of cell cycle progression and uneven chromosome condensation. 88 Interestingly, SR turtle embryos also showed a positive correlation between mercury concentrations and global DNA methylation, although other studies of mercury exposure and DNA methylation in mammalian and nonmammalian vertebrates report a negative association. 11,57 Little is known of the mechanisms by which mercury exposure leads to alterations in DNA methylation profiles. 93
Hypomethylation of DNA has been related to genomic instability, aging, and cancer, 24,89 while DNA hypermethylation in placenta has been related to developmental defects. 37,69
Most studies of mammals exposed to mercury or methylmercury have focused on neurological tissues and report a hypomethylated status of DNA. 4,9,60,66 Nevertheless, in other groups such as birds and amphibians, methylmercury does not seem to affect DNA methylation. 4 In addition, methylmercury is able to induce hypermethylation of promoter regions, which may be gene, tissue, or cell specific, 28,38 indicating that the status of DNA methylation depends on the functional characteristics of a specific DNA region. This is important because several approaches are available to quantify global DNA methylation. Some of them are based on restriction enzymes sensitive to methylation (such as the luminometric methylation assay, LUMA); others are based on the analysis of Alu transposable elements or LINE-1 retrotransposable elements, which are dispersed in the human (or mammalian) genome; and other approaches use enzyme-linked immunosorbent assay (ELISA)–like principles depending on antibodies to capture and detect methylated DNA. Thus, the method of choice may provide different results regarding a global DNA methylation status, depending on the principle and specific characteristics of the samples. So, to compare the complexity of global DNA methylation data in different species and under different scenarios, the quantification principle should be the same.
Finally, mitochondrial function and dynamics are essential for normal development. 82 In particular, a low copy number of mtDNA is often predictive of mitochondrial genetic damage, 90 while a high copy number is indicative of exposure to stress. For instance, in mice, mtDNA copy number is increased by in utero exposure to secondhand cigarette smoke 22 and in humans following chronic exposure to polycyclic aromatic hydrocarbons. 65 In this study, mtDNA copy number did not differ significantly, neither between normal and malformed embryos nor between SR and other malformed or normal embryos. Nevertheless, a higher variability in mtDNA copy number was observed in malformed embryos (both SR and other malformations) compared to normal embryos. This could be due to cell-to-cell differences in mtDNA copy number within each abnormal embryo as a response to environmental or developmental factors, and further studies are necessary to understand such variability.
Conclusion
This study represents an initial attempt to understand the cause of the rare malformative syndrome SR in olive ridley sea turtle embryos, particularly the role of mercury and DNA methylation. Despite sample size limitations, the results from this study show that SR syndrome in olive ridley sea turtle embryos is associated with higher mercury concentrations in tissue and DNA methylation content, with no apparent association with mtDNA copy number. These findings suggest genetic-environment interactions in the development of SR in sea turtle embryos.
Supplemental Material
Supplemental Material, DS1_VET_10.1177_0300985819868649 - Mercury Concentration, DNA Methylation, and Mitochondrial DNA Damage in Olive Ridley Sea Turtle Embryos With Schistosomus Reflexus Syndrome
Supplemental Material, DS1_VET_10.1177_0300985819868649 for Mercury Concentration, DNA Methylation, and Mitochondrial DNA Damage in Olive Ridley Sea Turtle Embryos With Schistosomus Reflexus Syndrome by Rodolfo Martín-del-Campo, Annelisse Bárcenas-Ibarra, Gertrud Lund, Dalia Rodríguez-Ríos, Lenin Yong-Villalobos, Jaqueline García-Hernández and Alejandra García-Gasca in Veterinary Pathology
Footnotes
Acknowledgements
We thank all the people working at El Verde Camacho nesting beach, especially Daniel Ríos-Olmeda and Raquel Briseño-Dueñas. We also thank the Wildlife Department of the Ministry of the Environment in Mexico (DGVS-SEMARNAT) for capture permits and volunteer Rogelio Iván Lozano-Guzmán for assistance in collecting and cleaning the olive ridley nests. We are thankful to Rubí Hernández-Cornejo and Daniela Aguilera-Márquez for technical assistance and especially grateful to Isaías Rojas-Lleonart for his assistance in the identification and description of congenital malformations in the olive ridley sea turtle. We are also thankful to the National Council for Science and Technology (CONACYT) and the Research Center for Food and Development (CIAD) for the scholarship and financial support awarded to R.M.C. Finally, we are grateful to Miguel Betancourt-Lozano, Itzel Sifuentes-Romero, Raúl Llera-Herrera, Alberto Abreu-Grobois, and the anonymous reviewers for constructive comments to this article.
Author Contribution
Rodolfo Martín-del-Campo and Annelisse Bárcenas-Ibarra contributed equally to this study.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: National Council for Science and Technology (CONACYT) and the Research Center for Food and Development (CIAD) provided scholarship and financial support to R.M.C and A.B.I..
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
