Abstract

Key to the concept of neoplasia is the presence of a clonal cell population. Identification of such a clonal cell population (also referred to as clonality testing or clonality assessment) is dependent on the presence of markers that under normal conditions are highly polymorphic in the cell type of interest. For leukemias and lymphomas, collectively termed neoplastic lymphoid proliferations, such highly polymorphic markers exist in the form of the antigen receptor genes that have to rearrange in a cell-specific way to encode the immunoglobulin (IG) or T-cell receptor (TCR) molecule. In a reactive cell population the repertoire of rearranged IG/TCR genes is thus highly diverse, whereas in a neoplastic context the IG/TCR repertoire is to some or to a large extent identical; of note, these are like the 2 ends of a spectrum with variants in between (Fig. 1). Still, clonality testing of rearranged IG/TCR genes forms a useful tool in the diagnostic armamentarium of leukemias and lymphomas.
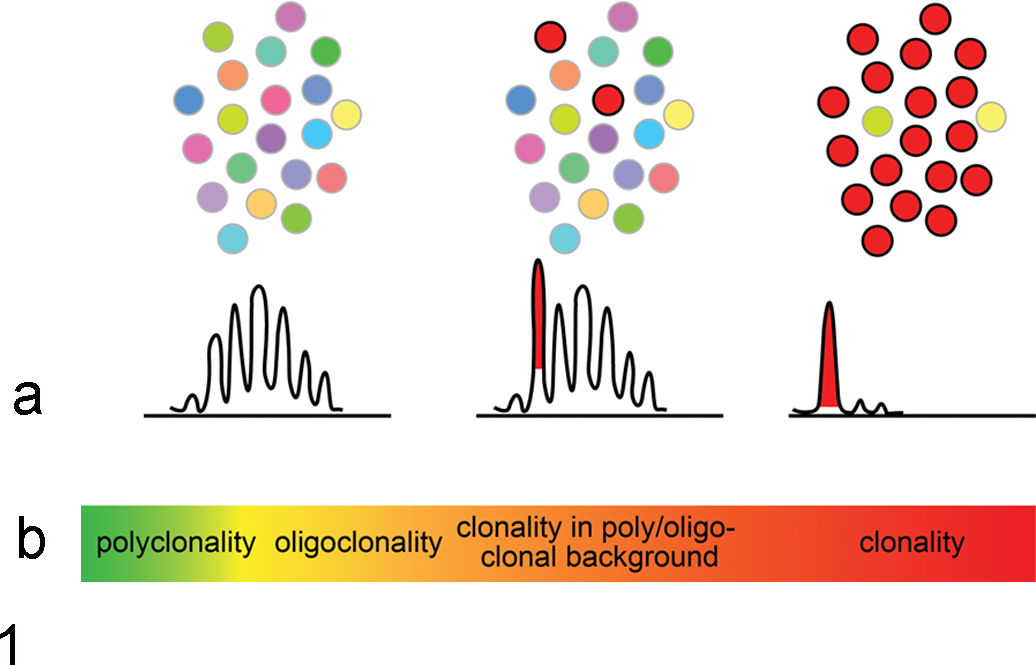
(a) Schematic representation of cell populations and the typical resulting IG/TCR GeneScan profiles reflecting polyclonality (left), clonality in a polyclonal background (middle), and clonality (right). (b) Profiles are part of a continuum ranging from truly polyclonal at 1 end to full-blown clonal at the opposite end.
In this issue Keller et al 1 review the state-of-the-art in clonality testing in veterinary medicine, highlighting the main aspects of all 3 phases of the clonality testing work flow, that is the preanalytical, analytical, and postanalytical phases. Next to that, the authors provide a rationale for harmonization of clonality assessment in veterinary medicine, building on what has been developed and achieved in human medicine for that matter. 2,3
Given the current differences between veterinary laboratories with respect to laboratory practices and interpretation guidelines, harmonization of terminology, protocols and interpretation is a good step toward improving the accuracy of clonality testing assays worldwide. The glossary of terms as provided by Keller et al 1 would help to avoid Babylonian misunderstandings for a clinical test that has been designated “molecular morphology” referring to the somewhat subjective description of profiles (see also Fig. 1). Furthermore, the detailed discussion of interpretation pitfalls in the review by Keller et al 1 is most useful to create awareness amongst pathologists and molecular biologists active in the veterinary field.
However, the veterinary field would strongly profit from making more steps, or better from making a big leap, by striving toward standardization, as has been done in human medicine. Standardization goes 1 step further than harmonization and implies much more attuned, evidence-based optimization to deliver an approach that is robust enough in a multicenter context. What would be the required actions and instruments for that purpose?
A first thing to do would be to create a (small) consortium that starts to directly compare existing IG/TCR assays on the very same samples in multiple veterinary laboratories. The best performing assays for the different species would be the winners and could become the gold standards, unless it is clear that no single assay has a clinical sensitivity that is acceptable. In that case, novel multiplex PCR assays should be considered with newly designed primers, taking advantage of additional information that comes from high-throughput sequencing efforts (both on topology and expressed repertoires). In this respect the already existing IMGT veterinary page (www.imgt.org/IMGTveterinary) could be further exploited as a central platform for submitting sequence data in order to better annotate genes, identify polymorphic variants, and denote pseudogenes.
Another essential step would be to design a quality assessment scheme for parallel testing of selected samples in multiple laboratories. Such a scheme would be instrumental to evaluate and improve performance of individual laboratories, especially when accompanied by some kind of feedback on the results through educational lectures and/or reports.
Finally, it might be worthwhile investing in additional targets to increase the rate of clonality detection. The fact that many of the species are predominantly IG lambda expressing, would make the somatic hyper mutation-insensitive IGK Kde rearrangements an interesting target to consider. The same holds for incomplete IGH D-J rearrangements. That being said, setting up new assays is not an easy task in the veterinary field, as every species would require its own primer set, whereas the lack of complete genome sequence information might hamper design of primers in particular cases. Even though currently the available sequence information might not be sufficient, it might be worth investing in this. The current developments in next generation sequencing (NGS) technologies might provide possibilities that could be exploited in this respect.
Another development that could potentially be of interest for the veterinary field as well, is capture-based NGS of IG/TR genes. 4 Proof of principle is out for the human field, where probes directed against regions of the IG and also TCR gene complexes have been shown to allow sequencing of rearranged IG/TCR genes as the basis for clonality testing, repertoire analysis and even translocation testing. 4,5
In summary, the clonality testing field in veterinary medicine is moving in the right direction by starting to harmonize assays, but given the current momentum the movement should not stop there. Rather, this first action should pave the way toward full standardization, which eventually will lift the average quality of clonality testing for different species in the veterinary field, which is essential to accurately distinguish lymphoma from inflammatory conditions in pets. Standardization should thus become the vet’s pet.
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
