Abstract
Scrapie is a naturally occurring transmissible spongiform encephalopathy of sheep and goats. There are different strains of sheep scrapie that are associated with unique molecular, transmission, and phenotype characteristics. However, in the United States, very little is known about the potential presence of scrapie strains. Scrapie strain and PRNP genotype could both affect susceptibility, potential for transmission, incubation period (IP), and control measures required for eliminating scrapie from a flock. The investigators evaluated 2 US scrapie isolates, No. 13-7 and x124, after intranasal inoculation to compare clinical signs, IPs, spongiform lesions, and patterns of PrPSc deposition in sheep with scrapie-susceptible PRNP genotypes (QQ171). After inoculation with x124, susceptibility and IP were associated with valine at codon 136 (V136) of the prion protein: VV136 sheep had short IPs (6.9 months), those in AV136 sheep were 11.9 months, and AA136 sheep did not develop scrapie. All No. 13-7 inoculated sheep developed scrapie, with IPs of 20.1 months for AA136 sheep, 22.8 months for AV136 sheep, and 26.7 months for VV136 sheep. Patterns of immunoreactivity in the brain were influenced by inoculum isolate and host genotype. Differences in PrPSc profiles versus isolate were most striking when examining brains from sheep with the VV136 genotype. Inoculation into C57BL/6 mice resulted in markedly different attack rates (90.5% for x124 and 5.9% for No. 13-7). Taken together, these data demonstrate that No. 13-7 and x124 represent 2 distinct strains of scrapie with different IPs, genotype susceptibilities, and PrPSc deposition profiles.
Keywords
Scrapie is a naturally occurring transmissible spongiform encephalopathy (TSE) of sheep and goats. Other TSEs include bovine spongiform encephalopathy in cattle, chronic wasting disease in cervids, and Creutzfeldt-Jakob disease in humans. The central event in the pathogenesis of TSEs is the conversion of the host-encoded cellular prion protein (PrPC) into its misfolded pathogenic form (PrPSc), which leads to neurodegeneration and, eventually, death.
Strain traits arise from differences in the structure of the misfolded protein. 36 In the natural host, these strains may be differentiated by incubation period (IP), clinical presentation, 35 patterns of PrPSc immunoreactivity or microscopic lesions in the brain, 14 and the physiochemical properties of the PrPSc itself. 43 Inoculation of naturally occurring scrapie isolates into inbred mouse strains has resulted in the identification of multiple strains of scrapie on the basis of IPs and lesion profiles. 6,10
Susceptibility to scrapie is influenced by amino acid polymorphisms in the prion protein. In sheep, codons 136, 154, and 171 influence susceptibility to classical scrapie with 136 valine (V), 154 arginine (R), and 171 glutamine (Q) (VRQ) being associated with increased susceptibility, whereas 136 alanine (A), 11,25 154 histidine (H), 28 and 171 arginine (R) (AHR) 12,34 are associated with resistance to natural scrapie. Codon 171 appears to have the most discernible influence; sheep with QQ171 are susceptible and those with RR171 are resistant.
Very little is known about the number and diversity of scrapie strains present in the United States. In this report we describe the clinical and pathologic findings in sheep experimentally challenged via the intranasal route with 2 US scrapie isolates. A previous study demonstrated that these isolates have genotype-dependent differences in IP after inoculation by the intracranial route. 23 The purpose of this study was to compare clinical signs, IPs, spongiform lesions, and patterns of PrPSc deposition in sheep with scrapie-susceptible prion protein gene (PRNP) genotypes to determine if inoculation with these scrapie isolates resulted in phenotypes that warrant consideration as separate scrapie strains.
Materials and Methods
Ethics Statement
All animal experiments described were reviewed and approved by the National Animal Disease Center’s Institutional Animal Care and Use Committee (protocol nos. 3893 [sheep study] and 2414 [mouse bioassay study]) and were carried out in strict accordance with the Guide for the Care and Use of Laboratory Animals (Institute of Laboratory Animal Resources, National Academy of Sciences, Washington, DC) and the Guide for the Care and Use of Agricultural Animals in Research and Teaching (Federation of Animal Science Societies, Champaign, IL).
Sources of Sheep Scrapie Inocula
The No. 13-7 inoculum was derived from a sheep receiving a fourth intracranial serial passage through ARQ/ARQ Suffolk sheep. 19 The original inoculum No. 13-7 was prepared from a pool of 13 brains from scrapie-affected ARQ/ARQ sheep (all were positive by immunohistochemistry) from 7 source flocks. 22 The tissue was ground in a mechanical grinder, gentamicin was added at a concentration of 100 μg/ml, and the final concentration of 10% weight per volume was made with phosphate-buffered saline.
The x124 inoculum was prepared from the brain of a Suffolk sheep (ear tag no. 3004, genotype VRQ/ARR) from the University of Idaho scrapie flock that was challenged with the x124 scrapie brain pool inoculum. 43 The original x124 inoculum was prepared from a pool of 7 scrapie-affected sheep brains from 5 flocks obtained from the University of Idaho Caine Veterinary Teaching Center, as described previously. 8,23 All donor sheep were PrPSc positive by immunohistochemistry and western blot testing of the brain stem medulla at the level of the obex. The PRNP genotypes at codons 136, 154, and 171 of the original donor sheep were as follows: n = 5 AARRQQ, n = 1 AVRRQQ, and n = 1 VVRRQQ.
Animal Procedures
At 4.5 months of age, recipient sheep from a scrapie-free flock were intranasally (n = 12) or intracranially (n = 4) inoculated with 1 ml of 10% suspension of either the No. 13-7 or x124 inocula. 21 Inoculated sheep had polymorphisms at codon 136 (see Table 1) but were homozygous at positions 112 M, 127 H, 137 M, 141 L, 151 R, 154 R, 157 M, 171Q, 176 N, 180 H, 189Q, 195 T, 196 T, 211 R, 220Q and 223 R. Briefly, the inoculum was drawn into a syringe and, without the aid of a needle, squirted into the right nostril while the head was elevated slightly. Inoculated animals were housed in a biosafety level 2 containment facility for the initial 2 weeks postinoculation (PI) and later were moved to outside pens at the National Animal Disease Center. The sheep were fed pelleted growth and maintenance rations that contained no ruminant protein, and clean water was available ad libitum. Control sheep were kept with the scrapie-free flock at the National Animal Disease Center. Both groups were fed pelleted growth ration and alfalfa hay. Animals were observed daily for the development of clinical signs and euthanized on the development of clinical signs consistent with scrapie or at the end of the experiment (30 months PI).
Animal Data and Immunohistochemistry Results for Non–Central Nervous System Tissues.
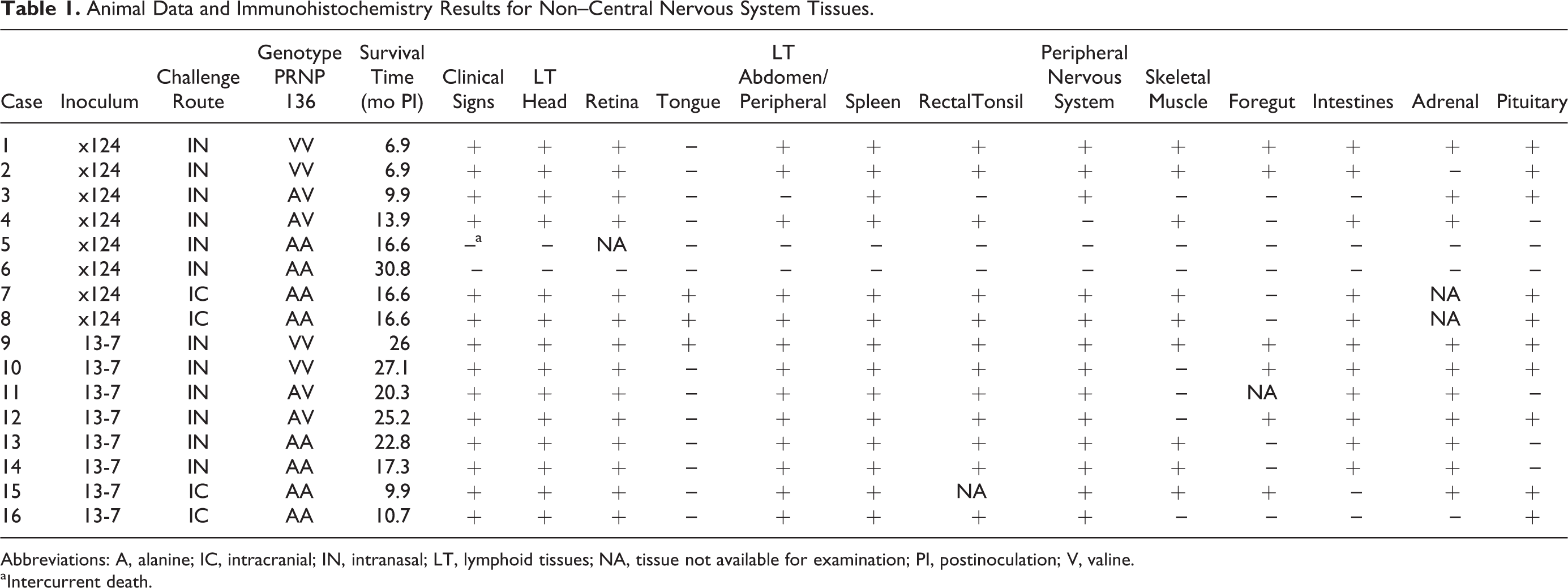

Abbreviations: A, alanine; IC, intracranial; IN, intranasal; LT, lymphoid tissues; NA, tissue not available for examination; PI, postinoculation; V, valine.
aIntercurrent death.
Sample Collection and Processing
Rectal mucosal biopsies 13 were collected from all sheep at 5 weeks PI and then at 3.5, 5, 6.5, 8, and 14 months PI. A detailed necropsy was carried out on each animal, and 2 sets of tissue samples were collected. One set of tissues included representative sections of lymphoid tissues (palatine tonsil, pharyngeal tonsil, and 3 rd eyelid), lymph nodes (LNs) (retropharyngeal, mesenteric, renal, prescapular, and popliteal), nasal planum, tongue, esophagus, reticulum, rumen, omasum, abomasum, intestines (jejunum, ileum, and cecum), liver, spleen, kidney, urinary bladder, pancreas, adrenal gland, thymus gland, thyroid gland, pituitary gland, turbinates, trachea, lung, skin, striated muscles (heart, tongue, diaphragm, masseter, biceps femoris, and psoas major), eye (including retina), nerves (optic, sciatic, and vagus), trigeminal ganglion, spinal cord (cervical, thoracic, and lumbar), and brain (cerebral cortex, cerebellum, midbrain including superior colliculus, and brain stem including obex). These tissues were fixed in 10% buffered formalin, embedded in paraffin wax, sectioned at 4 μm, and stained with hematoxylin and eosin for light microscopy. The second set of tissues was frozen.
All paraffin-embedded tissues were stained by an automated immunohistochemical method for detection of PrPSc as described previously 16 with a cocktail of 2 monoclonal antibodies, F89/160.1.5 33 and F99/97.6.1, 39 each used at a concentration of 5 μg/ml.
The frozen portion of brain was used for immunodetection of PrPres by a commercial western blot (Prionics Check western blot; Prionics AG, Schlieren, Switzerland) as described previously 16 and developed with monoclonal antibody P4 (Rida mAb P4; R-Biopharm AG, Darmstadt, Germany), which targets amino acids 89 to 104 of the ovine prion protein sequence 24 at a 1:10,000 dilution (0.1 μg/ml).
PrPSc Short Profiles
PrPSc short profiles that can be used to compare and differentiate TSE strains in sheep by quantifying the morphologic type and distribution of PrPSc deposits were recorded as described previously. 30 Briefly, the morphology of PrPSc deposits in the brain depends on the subcellular location of PrPSc and the cell type affected. 14,15 Intracellular PrPSc deposit types include intraneuronal, intramicroglial, and intra-astrocytic. Extracellular accumulations include stellate, subpial, perivascular, perivacuolar, particulate/coalescing, perineuronal, linear, nonvascular plaques, and vascular plaques. PrPSc deposits may be associated with the gray matter neuropil (linear, perineuronal, particulate/coalescing), neurons (intraneuronal), microglial cells (intramicroglial), astrocytes (intra-astrocytic), and astrocyte processes that form the glial limitans (subpial, subependymal, perivascular), or glial cells of uncertain origin (stellate, perivacuolar). Plaques are fibrillar, radiate, relatively large accumulations of PrPSc that are termed vascular plaques when they are unequivocally distributed around blood vessels and nonvascular plaques when they are not. 14 For each of 11 neuroanatomic areas in the corpus striatum (basal nuclei and frontal cortex), the magnitude of PrPSc accumulation associated with each of the above 12 morphologic labeling types was scored on a scale of 0 to 3 (see Gonzalez et al 15 for pictorial examples of each score). The resulting PrPSc profiles were compared between animals in this study.
Mouse Bioassay
To further characterize the No. 13-7 and x124 isolates, they were inoculated into C57BL/6 mice after isoflurane anesthesia as previously described. 38 Mice (n = 23 for x124, n = 21 for No. 13-7) were concurrently inoculated by the intracranial (20 μl) and intraperitoneal (100 μl) routes using the same 10% brain homogenate as used in sheep. Mice were monitored daily for the development of clinical signs by animal care staff members. When signs suggestive of prion disease such as ataxia, listing or rolling gate, pelvic limb paresis, lethargy, or poor grooming with urine-stained fur were recognizable by observation, animals were humanely euthanized and their brains prepared for analysis by western blot for the identification of PrPSc as described above except using monoclonal antibody 6H4 (Prionics AG) at a 1:10,000 dilution (0.1 μg/ml).
IPs are expressed as days PI. All animals that died of intercurrent disease or without the development of clinical signs 2 standard deviations or less prior to the average incubation time were included in the calculation of attack rate. Using these criteria to calculate attack rate, 21 mice were included in the x124 inoculation group, and 17 mice were used for the No. 13-7 group. Survival analysis was done using GraphPad Prism 6 for Mac OSX (GraphPad Software, San Diego, CA).
Results
All sheep challenged with isolate No. 13-7 developed clinical disease. Mean IPs for AA136 sheep (20.1 months) were shorter than for AV136 (22.8 months) and VV136 (26.7 months) sheep (see Table 1). Of the sheep challenged with x124 inoculum, 4 of 6 developed disease during the project. The mean IP for VV136 sheep (n = 2) was 6.9 months and for AV136 sheep (n = 2) was 11.9 months. IPs of sheep challenged with x124 were significantly shorter than those challenged with No. 13-7 (Welch 2-sample t test, P < 0.01). Neither sheep of the AA136 genotype inoculated with isolate x124 developed clinical signs consistent with scrapie: 1 AA136 sheep was euthanized at 16.6 months PI because of intercurrent disease, and a second did not develop clinical disease and was euthanized at the end of the project (30 months PI). Clinical signs observed in both inoculated groups included reluctance to stand, hindlimb weakness, and stumbling. Clinical signs consistent with pruritus (eg, rubbing and nibbling at the hind legs and flanks and loss of wool over the hipbones) were observed only in VV136 or AV136 sheep challenged intranasally with x124.
Preclinical PrPSc accumulation, assessed using rectal mucosal (RAMALT) biopsy, was first detected at the 6.5 months PI in a VV136 sheep inoculated intranasally with x124. This sheep was euthanized because of severe clinical disease 2 weeks later. None of the remaining animals (including intracranially inoculated AA136 positive control animals) were positive at the 8-month PI sampling. At the 14-month PI sampling, 5 of 6 No. 13-7 intranasally inoculated sheep were positive, as were both x124 intracranially inoculated sheep of the AA136 genotype. RAMALT samples collected at necropsy were strongly positive for all No. 13-7 challenged animals and VV136 x124 animals. One AV136 sheep challenged with x124 was negative, and the other was weakly positive. Both AA136 sheep challenged intranasally with x124 were negative in all RAMALT samples collected.
Widespread PrPSc accumulation in non–central nervous system tissues was observed in all challenged sheep with the exception of the 2 AA136 x124 intranasally inoculated sheep, which were negative on all tissues examined. The esophagus, diaphragm, liver, lung, trachea, turbinate, nasal planum, thymus, thyroid, and urinary bladder were negative in all sheep. A summary of the results for other tissues can be found in Table 1.
In positive lymphoid tissues of the head (palatine tonsil, pharyngeal tonsil, 3 rd eyelid, and retropharyngeal LN), abdominal LNs (mesenteric LN, renal LN), gut-associated lymphoid tissue (GALT) (jejunum, ileum, caecum, RAMALT), peripheral LNs (prescapular LN, popliteal LN), and spleen, the PrPSc immunolabeling pattern consisted of fine to coarse granules of PrPSc in tingible body macrophages with or without reticular labeling of follicular dendritic cells. 41
Immunolabeling of the enteric nervous system (ENS) manifested as fine granules of PrPSc within the cytoplasm of neurons and glial cells and, occasionally, deposition of PrPSc on the cell membrane of enteric neurons of the myenteric (Auerbach’s) plexus and submucosal (Meissner’s) plexus. 42 In foregut tissues, ENS labeling was the only type of immunolabeling seen: the reticulum was positive in 1 sheep (No. 13-7 intracranial AA136); the rumen was positive in 1 sheep (No. 13-7 intranasal VV136), and this animal was also positive in the abomasum; and the omasum was positive in 4 sheep (n = 2 x124 intranasal VV136, n = 1 No. 13-7 intranasal AV136, n = 1 No. 13-7 intranasal AV136).
In the intestines (jejunum, ileum, cecum), immunolabeling of GALT and ENS was variably present. Both GALT and ENS immunolabeling were observed concurrently in all (n = 6) No. 13-7 intranasally inoculated sheep and x124 intranasally inoculated VV136 sheep. The ENS was negative in x124 intranasally inoculated AA136 sheep and No. 13-7 intracranially inoculated AA136 sheep. In x124 intranasally inoculated AV136 sheep, the ENS was positive in 1 of 2 animals, but insufficient tissue was available to assess GALT. In x124 intracranially inoculated AA136 sheep, the ENS was positive (n = 2) and GALT positive (n = 1) or negative (n = 1).
Positive immunolabeling in the peripheral nervous system (optic nerve, sciatic nerve, vagus nerve, trigeminal ganglion) presented as punctate deposits in ganglion cells and/or satellite cells. 17 The plexiform layers of the retina contained confluent accumulations, and multifocal punctate immunolabeling was seen in the photoreceptor, nuclear, and ganglion cell layers. The location and morphology of PrPSc immunolabeling in the tongue was consistent with accumulation in the sensory nerve fibers in the core of the lingual papillae. 32
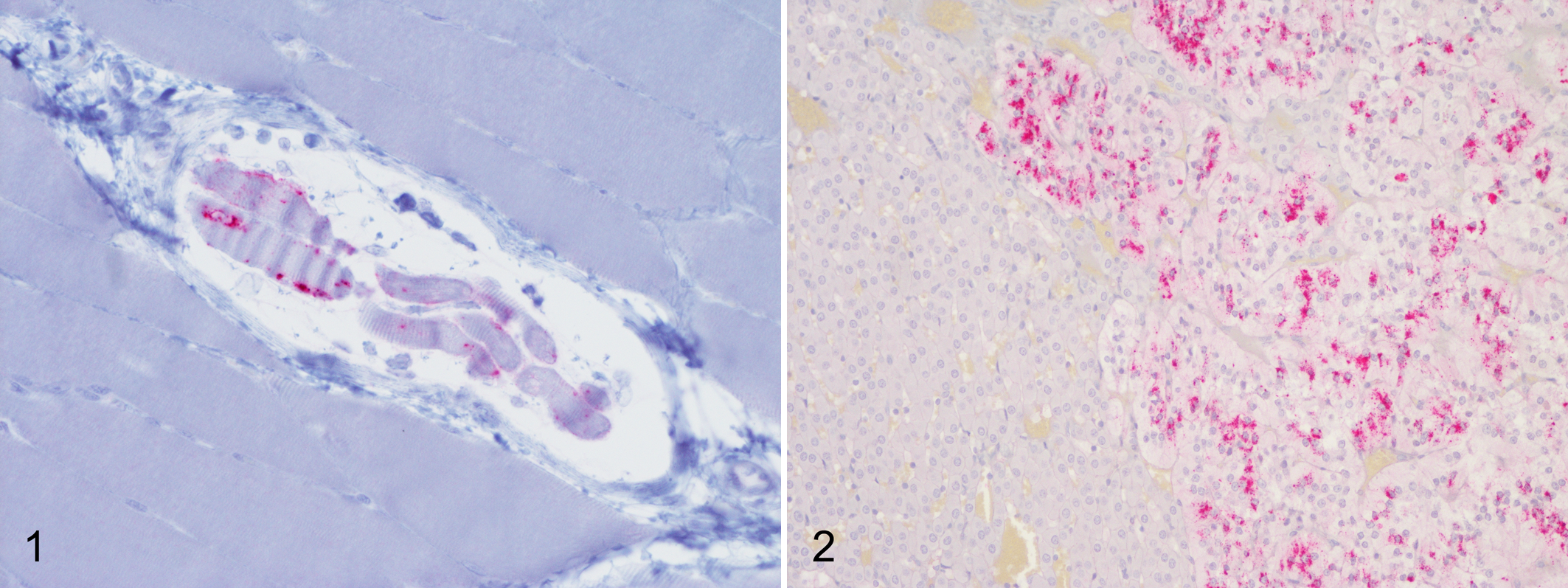
In skeletal muscle (biceps femoris, masseter, psoas major), immunoreactivity was observed as punctate deposits in cells of the neuromuscular spindles (Figure 1). 1 In the adrenal gland, granular immunolabeling was observed in the chromaffin cells of the adrenal medulla (Figure 2). 29 In the pituitary, granular immunolabeling was present in the pars intermedia and nervosa. 37
Genotype appeared to have an effect on the PrPSc profile in No. 13-7 inoculated animals (Figure 3). PrPSc profiles for AA136 and AV136 sheep were similar. In contrast, PrPSc profiles for VV136 sheep featured relatively prominent and widespread intracellular immunolabeling types, in particular intra-astrocytic and intramicroglial labeling, and relatively less glial membrane associated labeling types stellate, subpial, perivascular, and perivacuolar (Figures 4–7).
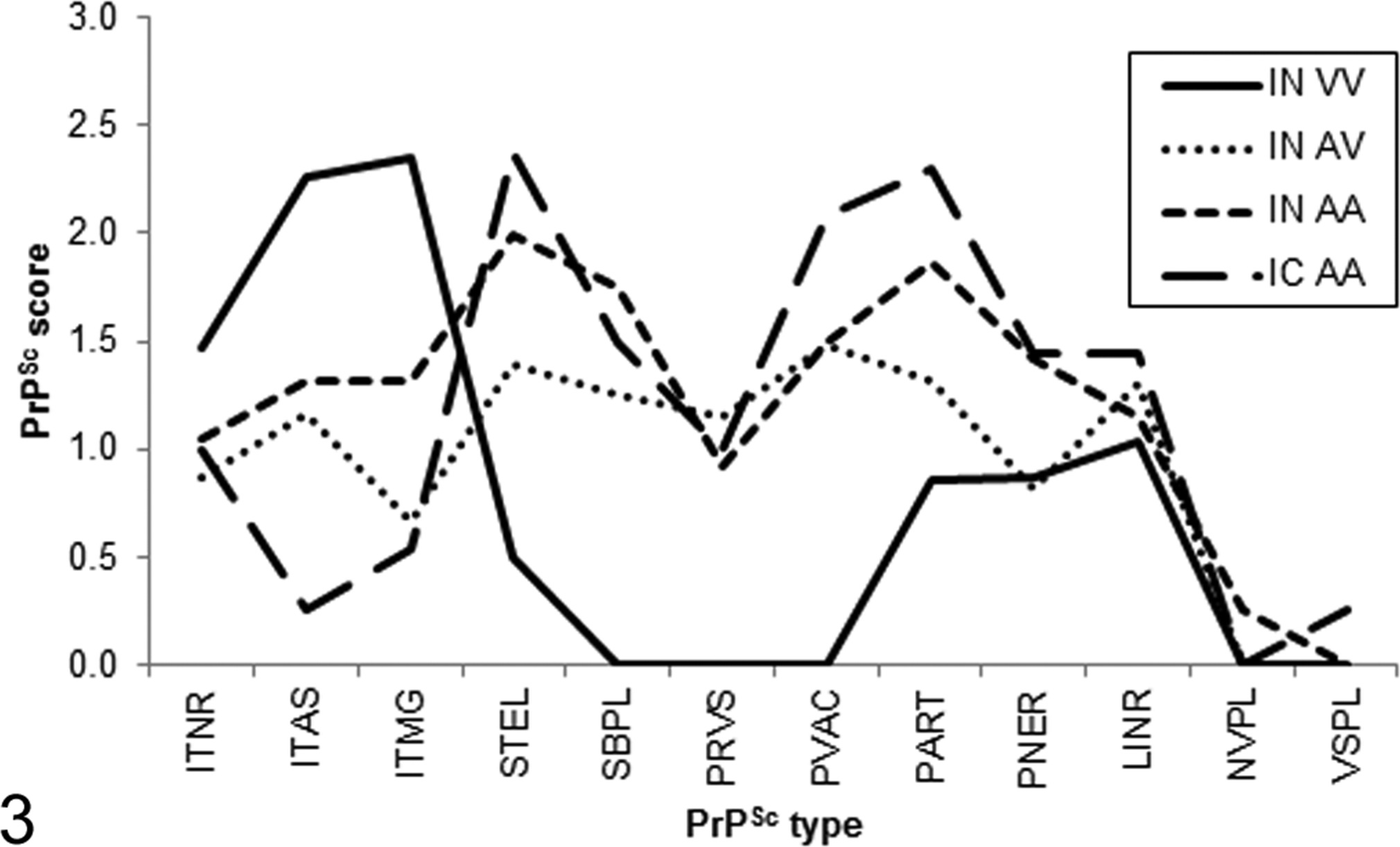
PrPSc profiles for sheep challenged with the No. 13-7 inoculum. IC, intracranial challenge; IN, intranasal challenge. Genotype at codon 136: A, alanine; V, valine. PrPSc types: ITNR, intraneuronal; ITAS, intra-astrocytic; ITMG, intramicroglial; STEL, stellate; SBPL, subpial; PRVS, perivascular; PVAC, perivacuolar; PART, particulate/coalescing; PNER, perineuronal; LINR, linear; NVPL, nonvascular plaques; VSPL, vascular plaques.
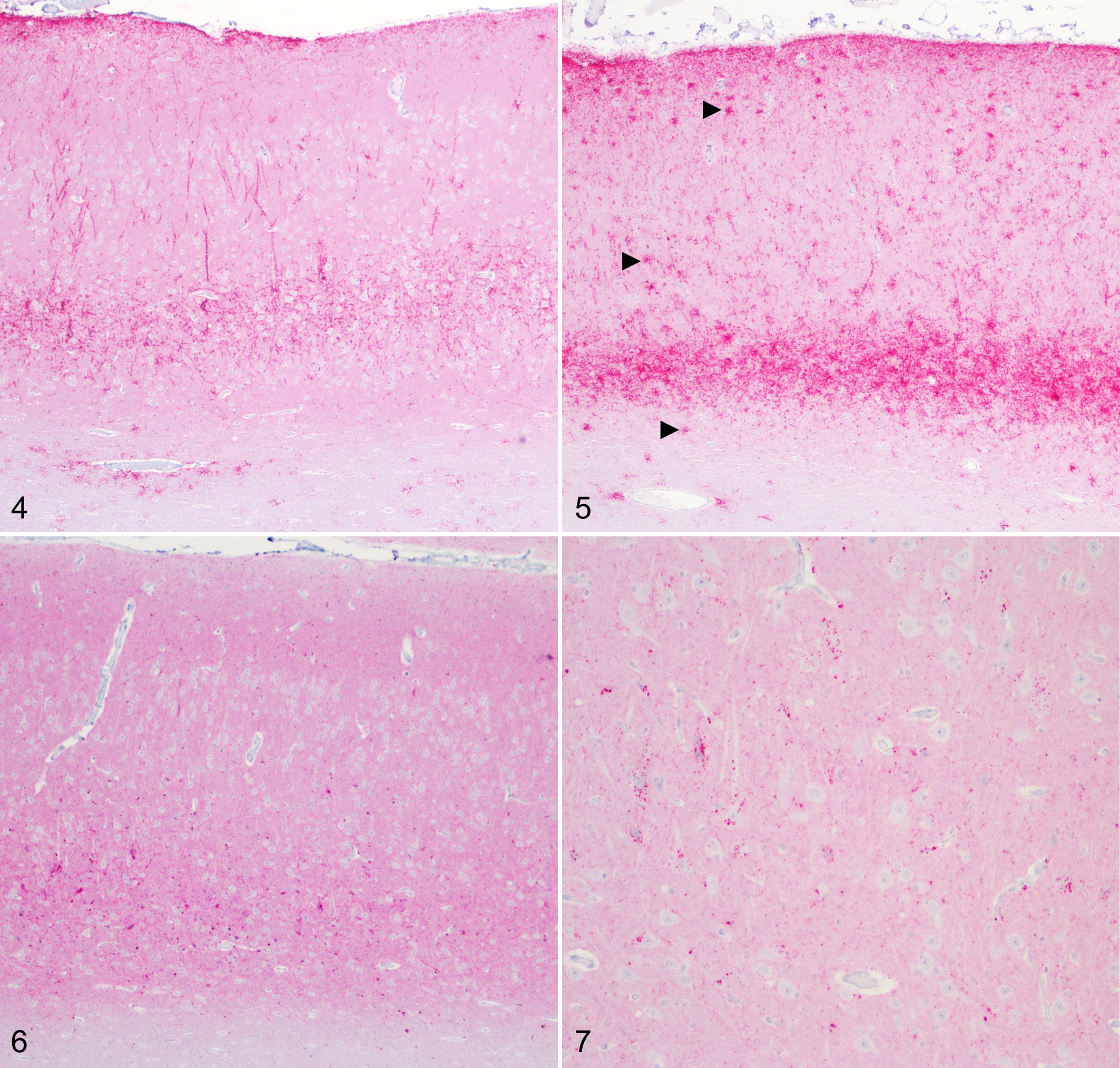
Scrapie, brain, sheep. Immunohistochemistry for PrPSc, monoclonal antibody F99/97.6.1.
PrPSc profiles for AA136 sheep challenged with No. 13-7 via the intranasal or intracranial route were broadly similar. However, a small reduction in the magnitude of intra-astrocytic and intramicroglial labeling types was observed in intracranially challenged sheep (Figure 3).
PrPSc was not detected in the brains or other tissues of the 2 AA136 sheep inoculated intranasally with x124. PrPSc profiles for VV136 and AV136 sheep challenged with this isolate were broadly similar, although AV sheep had relatively less intra-astrocytic and intramicroglial immunolabeling. Intracranially challenged sheep had less subpial labeling (Figure 8).
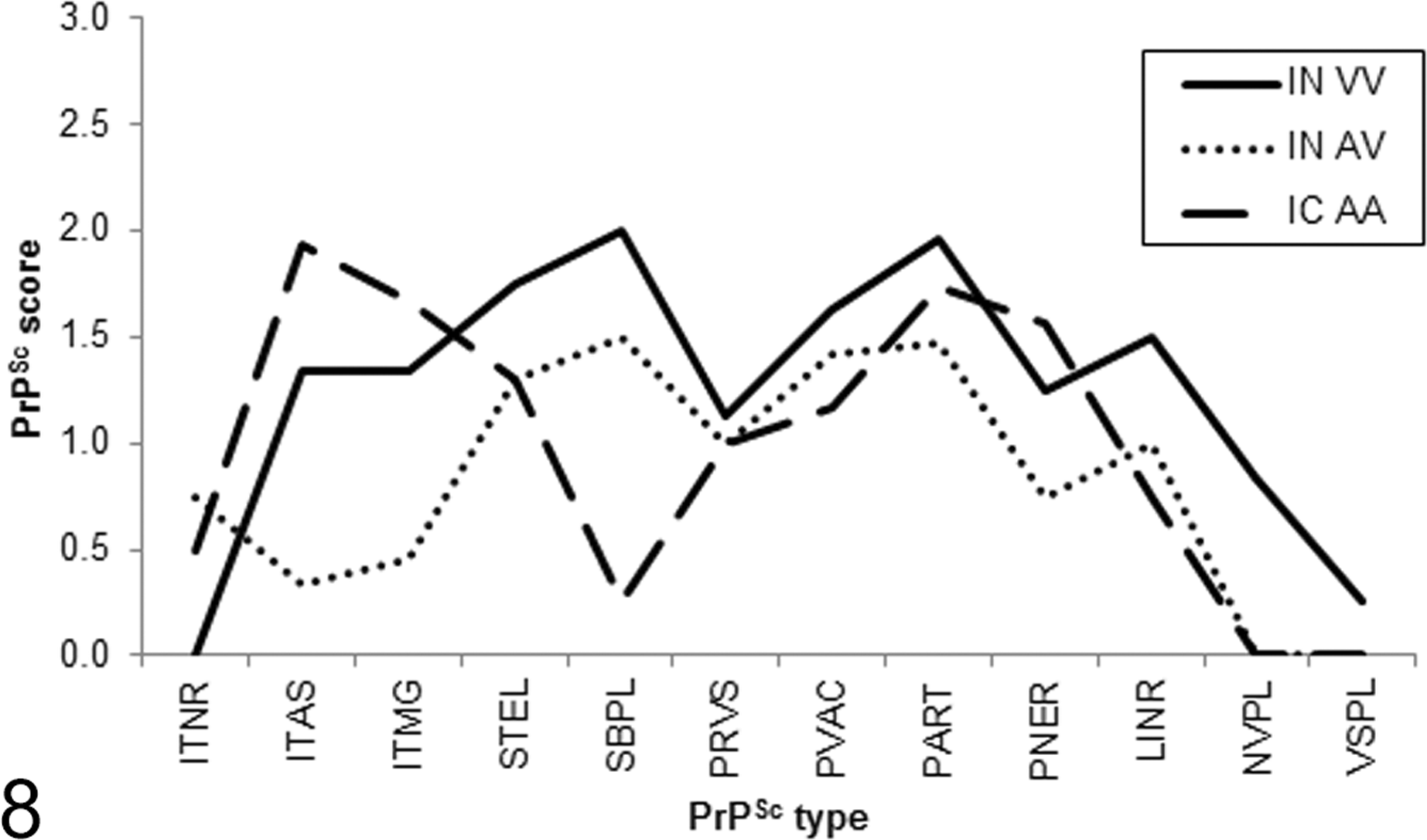
PrPSc profiles for sheep challenged with the x124 inoculum. IC, intracranial challenge; IN, intranasal challenge. Genotype at codon 136: A, alanine; V, valine. PrPSc types: ITNR, intraneuronal; ITAS, intra-astrocytic; ITMG, intramicroglial; STEL, stellate; SBPL, subpial; PRVS, perivascular; PVAC, perivacuolar; PART, particulate/coalescing; PNER, perineuronal; LINR, linear; NVPL, nonvascular plaques; VSPL, vascular plaques.
Inoculum appeared to have an effect on PrPSc profiles for VV136 and intracranially challenged AA136 sheep. When considering VV136 sheep, those inoculated with No. 13-7 had relatively more intracellular labeling (intraneuronal, intra-astrocytic, intramicroglial), less glial membrane associated labeling (stellate, subpial, perivascular, perivacuolar), and less neuropil labeling (particulate) than those inoculated with x124 (Figure 9). In contrast, PrPSc profiles for AV136 sheep were similar for both inocula (Figure 10). A comparison could not be made for intranasally challenged AA136 sheep, because x124 challenged sheep did not accumulate PrPSc in the brain (Figure 11). However, for intracranially challenged AA136 sheep, sheep inoculated with x124 had more intraglial labeling (intra-astrocytic, intramicroglial) and less glial membrane labeling (stellate, subpial, perivacuolar) than sheep challenged with No. 13-7 (Figure 12).
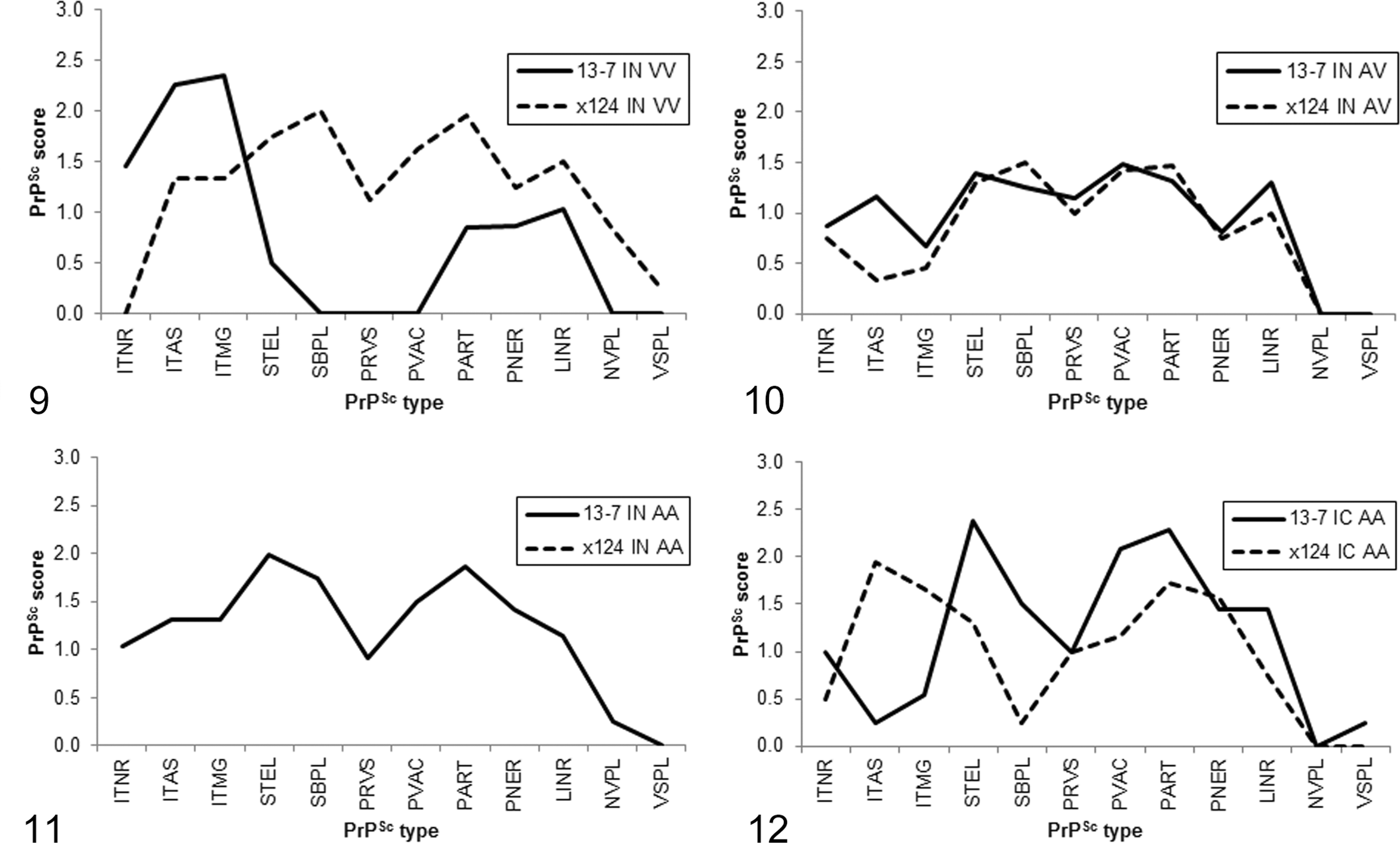
PrPSc profiles for VV136 sheep (Figure 9) and AV136 sheep (Figure 10) challenged intranasally and for AA136 sheep challenged intranasally (Figure 11) or intracranially (Figure 12). IC, intracranial challenge; IN, intranasal challenge. PrPSc types: ITNR, intraneuronal; ITAS, intra-astrocytic; ITMG, intramicroglial; STEL, stellate; SBPL, subpial; PRVS, perivascular; PVAC, perivacuolar; PART, particulate/coalescing; PNER, perineuronal; LINR, linear; NVPL, nonvascular plaques; VSPL, vascular plaques.
Molecular profiles obtained from western blot of brain homogenates from all sheep were similar regardless of inoculum used or genotype of sheep (Figure 13).
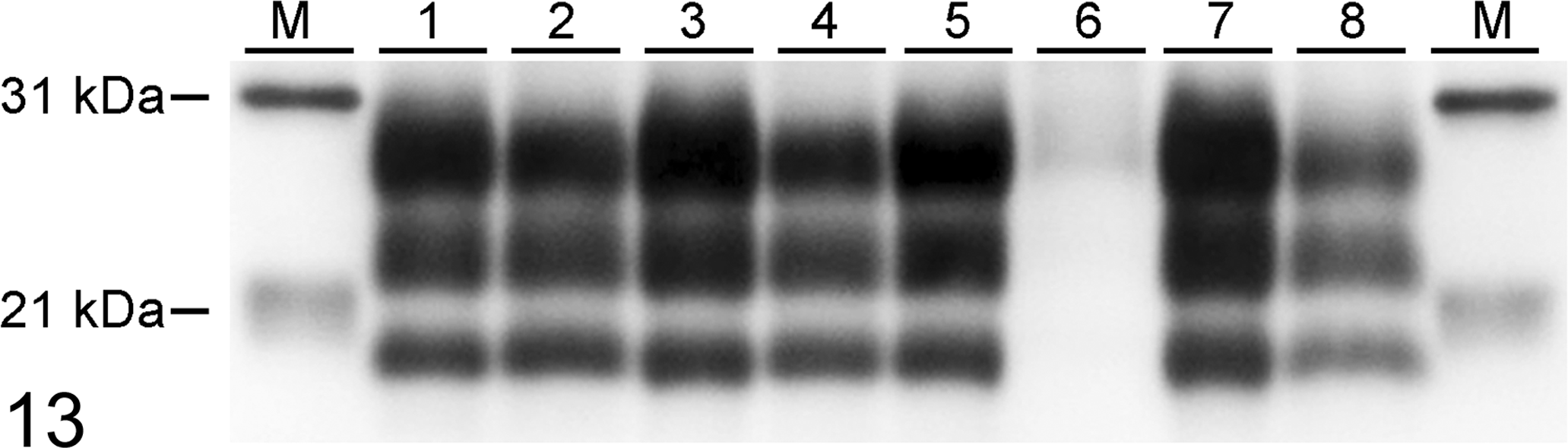
Western immunoblot of brain stem samples from scrapie-inoculated sheep developed using monoclonal antibody P4 to detect PrPres Lane key: M, molecular mass marker. 1 and 2, VV136 sheep challenged intranasally with No. 13-7 (lane 1, case #10) or x124 (2, case #2). 3 and 4, AV136 sheep challenged intranasally with No. 13-7 (3, case #11) or x124 (4, case #3). 5 and 6, AA136 sheep challenged intranasally with No. 13-7 (5, case #13) or x124 (6, case #5). 7 and 8, AA136 sheep challenged intracranially with No. 13-7 (7, case #15) or x124 (8, case #8).
The No. 13-7 and x124 inocula had significantly different effects in C57BL/6 mice (P < .0001). The attack rate for the x124 inoculum was 90.5% (19 of 21 mice), whereas the attack rate for No. 13-7 was 5.9% (1 of 17 mice). The mean incubation time for the x124 inoculum was 557 days, whereas the incubation time for the single positive animal inoculated with No. 13-7 inoculum was 907 days (Figure 14).
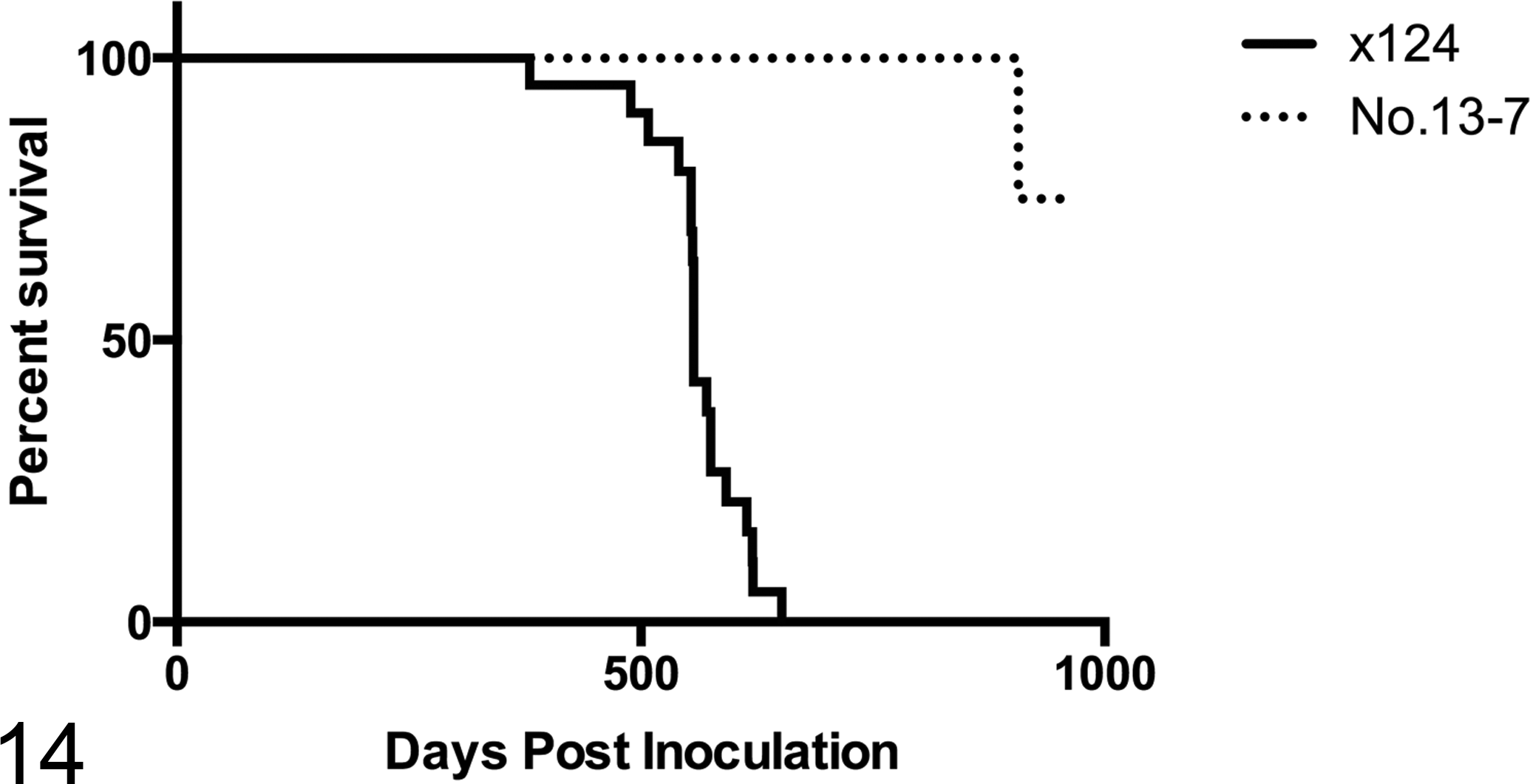
Survival time of C57BL/6 mice inoculated with x124 (n = 23) and No. 13-7 (n = 21) sheep scrapie isolates. Mice inoculated with x124 (solid line) developed scrapie, with a mean incubation of 557 days and an attack rate of 90.5%. Only a single mouse inoculated with No. 13-7 (dashed line) developed scrapie, with an incubation period of 907 days.
Discussion
This study provides further characterization of the phenotype of the No. 13-7 and x124 US scrapie isolates in sheep after intranasal challenge. This work presents evidence for 2 distinct scrapie strains in US sheep on the basis of differences in clinical signs, genotypes of susceptible sheep, IPs, and lesion profiles in genotype-matched experimental groups of sheep and attack rates and incubation times on mouse bioassay.
There were differences in clinical signs demonstrated by sheep receiving the different inocula in the present study. In sheep inoculated intranasally with x124 inoculum, clinical signs of pruritus (eg, itching, biting, and wool loss) were observed, but not in those challenged intranasally with No. 13-7. Lack of obvious pruritus or loss of wool was previously reported for the No. 13-7 inoculum after oral and intracranial challenge. 19,20 Interestingly, intracranial challenge with x124 did not lead to pruritus, 23 consistent with the results reported in the present study. Mild pruritus was observed rarely following oral challenge with x124. 8
All sheep used in this study were homozygous Q171 and considered to be scrapie susceptible. However, the amino acid at codon 136 played a major role in IP differences between sheep inoculated with either the x124 or No. 13-7 inoculum. IPs in sheep inoculated with x124 were markedly shorter than those in sheep challenged with No. 13-7. These observations are consistent with previous results. 8,23 Interestingly, PrPSc was not detected in sheep of the AA136 genotype challenged intranasally with x124. In contrast, AA136 sheep inoculated intracranially with x124 accumulated PrPSc in their lymphoid tissue and developed clinical disease, but with prolonged IPs relative to sheep carrying the V136 codon. The susceptibility of AA136 sheep to intracranial 23 but not intranasal challenge suggests that initial replication of PrPSc may be inefficient in the periphery of sheep with the AA136 genotype or that perhaps a higher dose of inoculum would be required to infect sheep of this genotype. The shorter IP observed with the x124 inoculum correlates with a lower stability of PrPSc as determined by unfolding in guanidine hydrochloride. 43
A similar trend in the IPs produced by the x124 and No. 13-7 inocula was observed on mouse bioassay. IPs for mice challenged with the x124 isolate were significantly shorter than for mice challenged with the No. 13-7 isolate. In addition, the attack rate of the x124 isolate (90.5%) was significantly higher than that of the No. 13-7 isolate (5.9%). Therefore mouse bioassay results further support the designation of x124 and No. 13-7 as distinct strains of the sheep scrapie agent.
In previous studies, the No. 13-7 inoculum has been used to challenge sheep by the intracranial, 18 –20 intralingual, 18 oral, 20 intranasal, 21 peritoneal, 21 and conjunctival 21 routes, and the x124 inoculum was shown to result in rapid incubation times in sheep with the V136 codon after oral or intracranial inoculation. 8,23 In the present study, IPs after intranasal inoculation with the x124 inoculum were similar to those previously reported for oral challenge 8 but were longer than after intracranial challenge. The mean IP for x124 intranasally challenged AV136 sheep (11.9 months PI) reported in the present study is similar to that reported for AV136 sheep challenged orally with the x124 inoculum at birth (10.2 months PI) or 9 months of age (12.6 months PI). 8 This suggests that passage of x124 in AV136 sheep does not markedly alter its phenotype, at least with regard to IP in AV136 sheep. With regard to the x124 inoculum used here that is derived from a single animal challenged with the x124 isolate, 43 it is difficult to form a direct comparison with the results of previous studies in which the inoculum was derived from a pool of brains from scrapie affected sheep of different genotypes. 8,23
With the exception of AA136 sheep inoculated with x124 (which did not develop scrapie), there was widespread distribution of PrPSc immunoreactivity in non–central nervous system tissues in all sheep. For x124 inoculated animals, PrPSc was detected in the tongue only in the 2 AA136 sheep inoculated intracranially. The location and morphology of PrPSc immunolabeling in these animals was consistent with accumulation in the sensory nerve fibers in the core of the lingual papillae, 32 as reported previously for 7 of 10 naturally infected ARQ/ARQ sheep in Italy. 9 Of the animals challenged with No. 13-7, PrPSc was only observed in the tongue of 1 intranasally challenged VV136 sheep. The absence of PrPSc in the tongue from ARQ/ARQ sheep is consistent with a previous study in which PrPSc was not detected in the tongue from ARQ/ARQ sheep inoculated intralingually (n = 3) or intracranially (n = 5) with the No. 13-7 inoculum. 18 Anterograde spread of prions from the brain to the tongue along tongue-associated cranial nerves, and retrograde spread from the tongue to the brain, has been reported for hamsters inoculated with transmissible mink encephalopathy. 2,3 Detection of PrPSc in tongues from both intranasally and intracranially challenged animals in the present study suggests that similar mechanisms exist in sheep.
Profiling of PrPSc immunoreactivity patterns further supports differences between these 2 scrapie strains. Among No. 13-7 inoculated sheep, the PrPSc profile of VV136 sheep was clearly different from other codon 136 genotypes in this group (AV, AA). VV136 sheep inoculated with No. 13-7 had relatively more intraneuronal labeling, less glial membrane–associated labeling, and less neuropil labeling than sheep inoculated with x124. Similar to sheep inoculated with No. 13-7, VV136 sheep inoculated with x124 had more intra-astrocytic and intramicroglial labeling than AV136 sheep. However, in contrast to the No. 13-7 challenged sheep, the amounts of glial membrane and neuropil labeling were similar in x124 intranasally challenged VV136 and AV136 sheep. Because the AA136 sheep inoculated with x124 did not develop scrapie, it was not possible to obtain a profile for these sheep.
Different routes of inoculation have previously been shown to result in differences in the magnitude of immunolabeling in sheep, although the overall shape of the PrPSc profiles curve is usually conserved. 15 Given that the overall shape of the PrPSc profile curves for AA136 sheep challenged intracranially or intranasally with No. 13-7 are similar for all labeling types except intra-astrocytic and intramicroglial, it is likely that the apparent reduction in magnitude of these 2 labeling types only in intracranially challenged sheep is the result of magnification of individual variation effects by small group sizes. Repeating the challenge with larger group sizes could help resolve this issue.
PrPSc profiles for the No. 13-7 and x124 inocula were similar in intranasally challenged AV136 sheep but distinctly different in intranasally challenged VV136 sheep and intracranially challenged AA136 sheep. These observations are likely to be the result of agent strain versus host genotype interactions.
The No. 13-7 inoculum has been passaged in AA136 sheep, while the x124 inoculum was prepared from the brain of a VRQ/ARR (codons 136, 154, and 171) sheep that had been challenged with a mixed-genotype brain pool. In QR171 sheep with scrapie, more than 90% of the PrPSc is composed of the Q171 allelotype, as the ARR allelotype has low conversion efficiency for PrPSc. 26 In sheep, the V136 codon does not occur on the same allele as the R171 codon. Therefore, the PrPSc in the No. 13-7 inoculum was ARQ and that of x124 was VRQ.
Conversion efficiencies in cell culture and in vitro assays can be used to infer interactions between PrPSc and PrPC proteins from sheep of different genotypes. Conversion is more likely to occur 7,40 and to be more efficient 4,5,27 with a homozygous inoculum-substrate combination (eg, inoculum from sheep with the VRQ [PrPSc-VRQ] allele to susceptible VRQ [PrPC-VRQ] sheep) than with a heterozygous combination (eg, VRQ inoculum to ARQ sheep). Using in vitro conversion assays, inocula from 6 VRQ/VRQ sheep naturally infected with scrapie were not able to amplify in a substrate derived from the brains of scrapie-free ARQ/ARQ sheep, while 3 of 6 ARQ/ARQ isolates were able to amplify in either ARQ and VRQ substrates. 40 The x124 inoculum causes disease in intracranially challenged, but not intranasally challenged, AA136 sheep. This suggests that although the x124 inoculum is able to convert PrPC-ARQ, it does so with a low conversion efficiency but that this barrier can be overcome by inoculation via the intracranial route.
With regard to VRQ/ARQ heterozygous sheep, cell free conversion assays have shown that PrPSc-VRQ/ARQ converts PrPC-VRQ at least 3 times as efficiently as to PrPC-ARQ. 4 In vivo, purified PrPC from scrapie-affected sheep was found to contain equivalent amounts of PrPC-VRQ and PrPC-ARQ, while PrPSc from the same sheep contained 75% PrPSc-VRQ and only 25% PrPSc-ARQ. 31 This preferential accumulation of PrPSc-VRQ, presumably regardless of inoculum genotype, may explain why PrPSc profiles for No. 13-7 and x124 are similar in 136AV sheep. The longer IPs observed in No. 13-7 intranasally challenged 136AV sheep compared with x124 challenged 136AV sheep could be due to less efficient conversion of PrPC-VRQ/ARQ by PrPSc-ARQ than by PrPSc-VRQ.
Conclusions
The results of the present study, based on differences between these 2 inocula with regards to clinical signs, genotype susceptibilities, IPs, and PrPSc profiles observed in sheep, and on IPs and attack rates in wild-type mice support the fact there are at least 2 strains of scrapie present in the US national flock. Different strains may require additional consideration when addressing potential transmission within a flock or control measures applied on a national basis.
Footnotes
Dr Smith’s current address is Department of Veterinary Pathology, College of Veterinary Medicine, Iowa State University, Ames, IA
Dr Richt’s current address is Kansas State University, College of Veterinary Medicine, Manhattan, KS
Acknowledgements
We thank Martha Church, Kevin Hassall, Joe Lesan, Leisa Mandell, and Trudy Tatum for providing technical support to this project. Mention of trade names or commercial products in this article is solely for the purpose of providing specific information and does not imply recommendation or endorsement by the Department of Agriculture. The Department of Agriculture is an equal-opportunity provider and employer.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This research was funded in its entirety by congressionally appropriated funds to the United States Department of Agriculture, Agricultural Research Service. The funders of the work did not influence study design, data collection and analysis, decision to publish, or preparation of the manuscript.
